Submitted:
13 February 2026
Posted:
14 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Cells and Viruses
2.2. Construction of OAV-j F Gene Expression Cassette
2.3. In Vitro Expression of the Recombinant F Protein
2.3.1. Indirect Immunofluorescent Assay
2.3.2. Western Immunoblotting
2.4. Scaling-up and Purification of pDNA-F
2.5. Preparation of Nanoparticle-Based DNA Vaccine Formulations
2.5.1. Preparation of Chitosan Nanoparticles (Cs NPs) – Based pDNA-F Vaccine
2.5.2. Preparation of PLGA Nanoparticles (PLGA NPs) - Based pDNA-F Vaccine
2.5.3. Preparation of PAMAM Dendrimers–Based pDNA-F Vaccine
2.6. Physicochemical Characterization of Nanoparticles
2.6.1. Hydrodynamic Size, Polydispersity Index, and Zeta Potential Analysis
2.6.2. Scanning Electron Microscopy (SEM)
2.6.3. Fourier Transform Infrared (FTIR) Spectroscopy
2.6.4. Entrapment Efficiency (EE%) and Loading Capacity (LC%)
2.7. In Vitro DNA Release Study
2.8. In Vivo Evaluation of DNA Vaccine Formulations
2.8.1. Experimental Animal Groups and Vaccination Protocol
2.8.2. Serological Evaluation of Humoral Immunity
2.8.3. Evaluation of Protective Efficacy
2.9. Statistical Analysis
3. Results
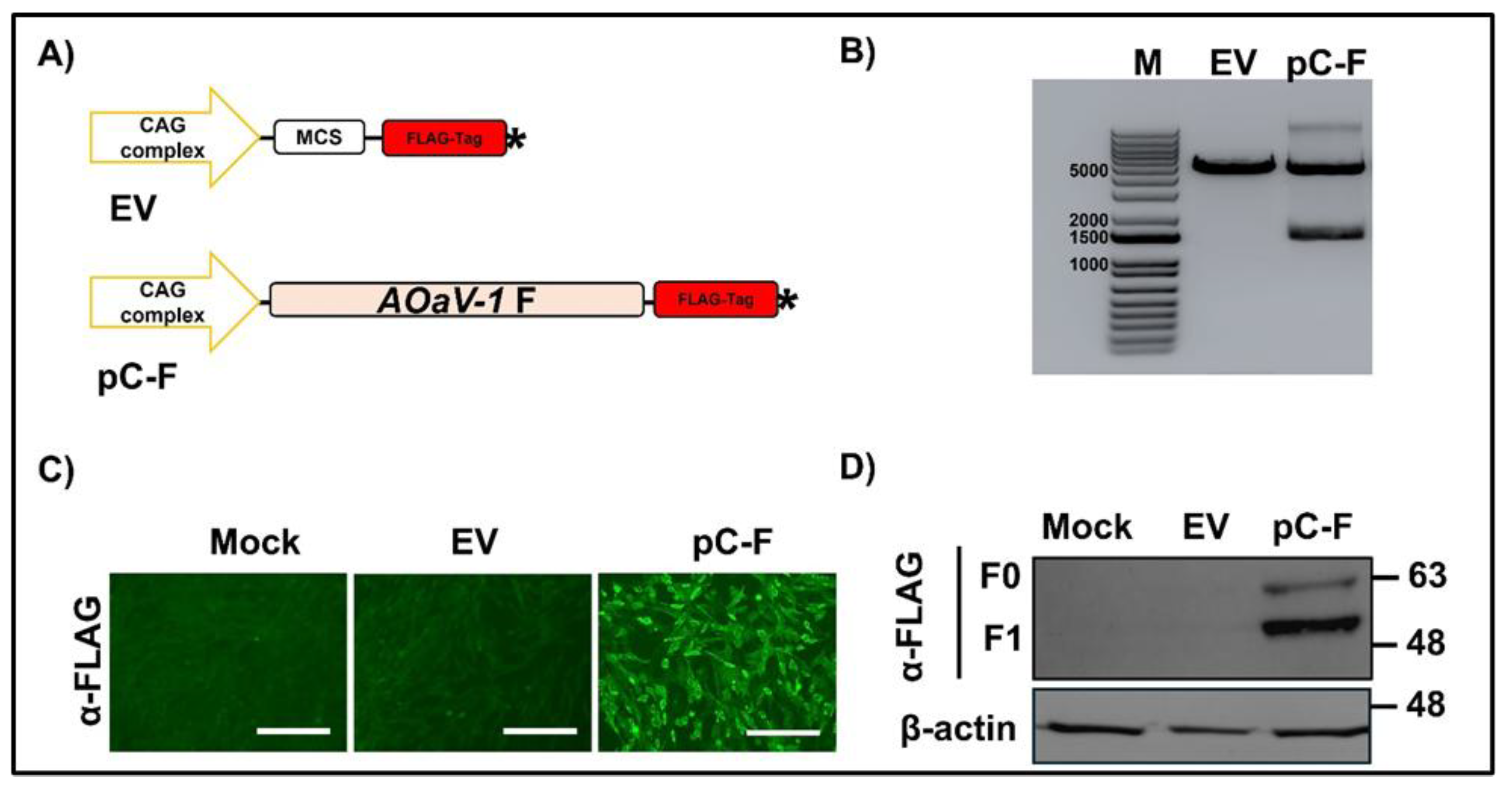
3.1. Generation of pDNA-F and Characterization of the Recombinant F Protein
3.2. Physicochemical Characterization of the pDNA-F–Loaded Nanoparticles
3.2.1. Hydrodynamic Size, Polydispersity Index (PDI), and Zeta Potential
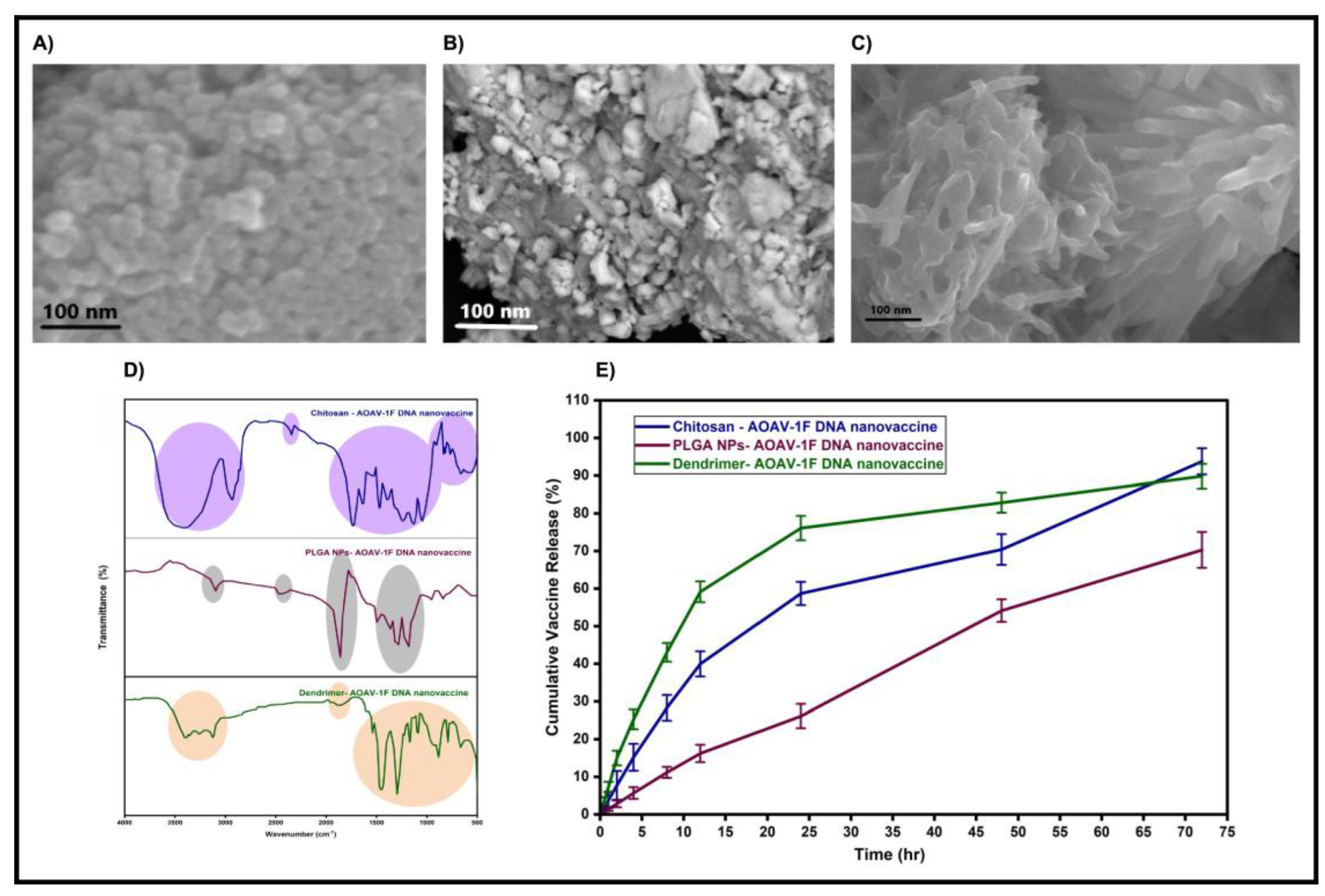
3.2.2. Scanning Electron Microscopy (SEM)
3.2.3. Fourier Transform Infrared (FTIR) Analysis
3.2.4. Entrapment Efficiency (EE%) and Loading Capacity (LC%)
3.2.5. In Vitro Release of pDNA-F DNA from Nanoparticles
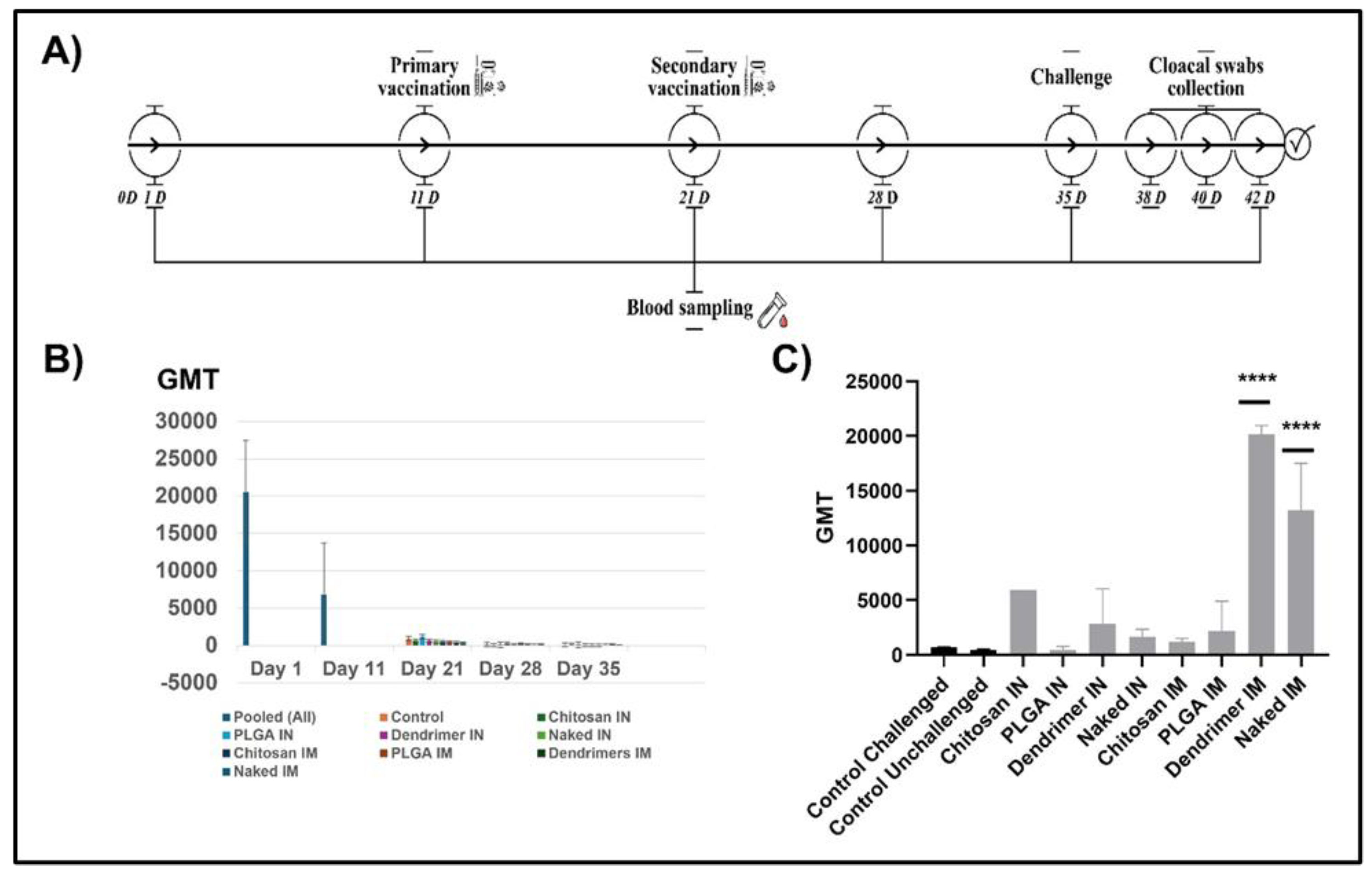
3.3. Robust Humoral Immune Responses Elicited by PAMAM-Dendrimers-pDNA-F
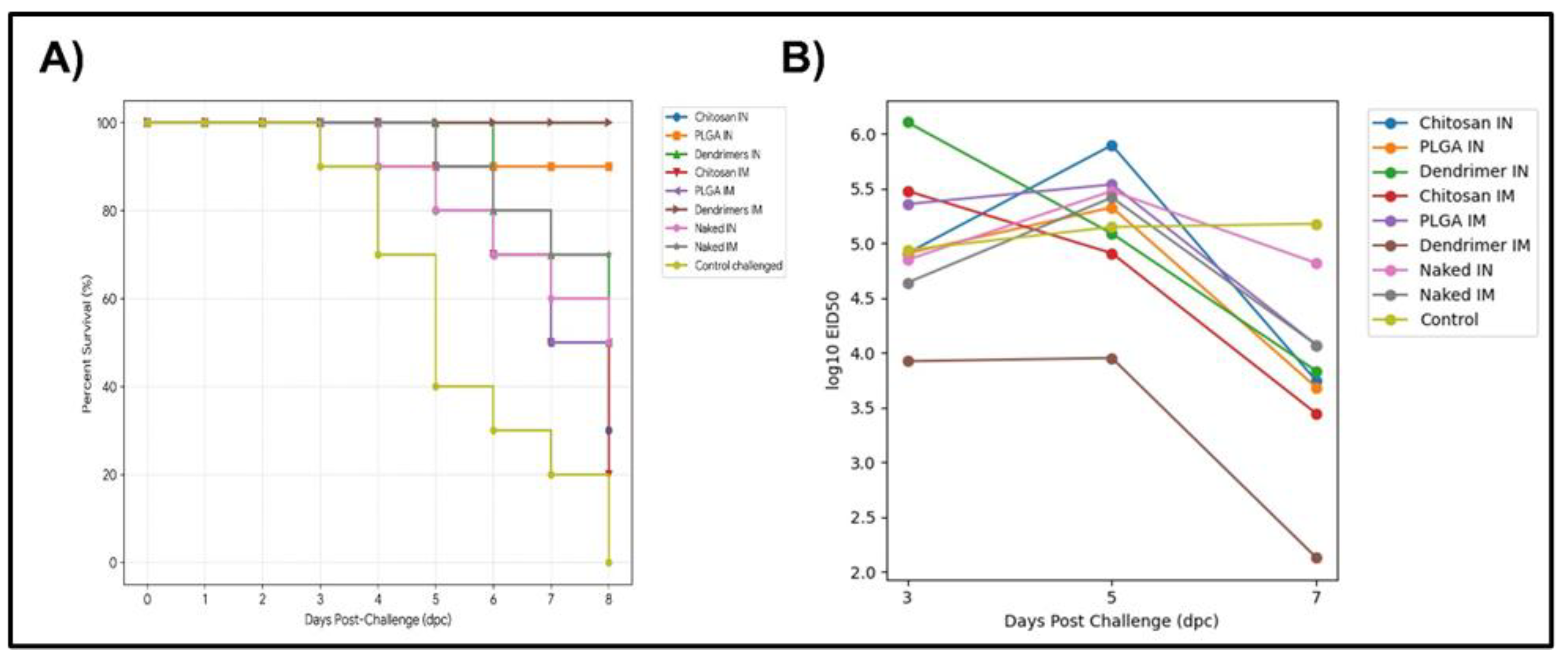
3.4. PAMAM-Dendrimers-pDNA-F Exhibits Potent Protective Efficacy Against Virulent Challenge
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ND | Newcastle disease |
| OAV-j | Orthoavulavirus javaense |
| F | Fusion |
| pDNA | Plasmid DNA |
| NPs | Nanoparticles |
| Cs | Chitosan |
| PLGA | Poly (lactic-co-glycolic acid) |
| PAMAM | Poly(amidoamine) |
| DLS | Dynamic light scattering |
| PDI | Polydispersity index |
| SEM | Scanning Electron Microscopy |
| FTIR | Fourier-transform infrared spectroscopy |
| IFA | Indirect immunofluorescence assay |
| WB | Western blot |
| ELISA | Enzyme-Linked Immunosorbent Assay |
| GMT | Geometric mean titer |
| RT-qPCR | Reverse Transcription quantitative Polymerase Chain Reaction |
| Ct | Cycle threshold |
| EID₅₀ | 50% Egg Infectious Dose |
| Dpc | Day post challenge |
References
- Vlaicu, PA; Untea, AE; Oancea, AG. Sustainable Poultry Feeding Strategies for Achieving Zero Hunger and Enhancing Food Quality. Agriculture 2024, 14(10), 1811. [Google Scholar] [CrossRef]
- ALEXANDER, DJ. Newcastle disease and other avian paramyxovirus. Revue Scientifique et Technique de l’OIE 2000, 19(2), 443–62. [Google Scholar] [CrossRef] [PubMed]
- Cattoli, G; Susta, L; Terregino, C; Brown, C. Newcastle disease. Journal of Veterinary Diagnostic Investigation 2011, 23(4), 637–56. [Google Scholar] [CrossRef] [PubMed]
- Abd Elfatah, KS; Elabasy, MA; El-khyate, F; Elmahallawy, EK; Mosad, SM; El-Gohary, FA; et al. Molecular Characterization of Velogenic Newcastle Disease Virus (Sub-Genotype VII.1.1) from Wild Birds, with Assessment of Its Pathogenicity in Susceptible Chickens. Animals 2021, 11(2), 505. [Google Scholar] [CrossRef]
- Ul-Rahman, A; Ishaq, HM; Raza, MA; Shabbir, MZ. Zoonotic potential of Newcastle disease virus: Old and novel perspectives related to public health. Rev Med Virol 2022, 32(1). [Google Scholar] [CrossRef]
- Yusoff, K; Tan, WS. Newcastle disease virus: Macromolecules and opportunities. Avian Pathology 2001, 30(5), 439–55. [Google Scholar] [CrossRef]
- Kolakofsky, D; Roux, L; Garcin, D; Ruigrok, RWH. Paramyxovirus mRNA editing, the ‘rule of six’ and error catastrophe: a hypothesis. Journal of General Virology 2005, 86(7), 1869–77. [Google Scholar] [CrossRef]
- Chen, L; Gorman, JJ; McKimm-Breschkin, J; Lawrence, LJ; Tulloch, PA; Smith, BJ; et al. The Structure of the Fusion Glycoprotein of Newcastle Disease Virus Suggests a Novel Paradigm for the Molecular Mechanism of Membrane Fusion. Structure 2001, 9(3), 255–66. [Google Scholar] [CrossRef]
- Pantua, H; McGinnes, LW; Leszyk, J; Morrison, TG. Characterization of an Alternate Form of Newcastle Disease Virus Fusion Protein. J Virol. 2005, 79(18), 11660–70. [Google Scholar] [CrossRef]
- Toyoda, T; Gotoh, B; Sakaguchi, T; Kida, H; Nagai, Y. Identification of amino acids relevant to three antigenic determinants on the fusion protein of Newcastle disease virus that are involved in fusion inhibition and neutralization. J Virol. 1988, 62(11), 4427–30. [Google Scholar] [CrossRef]
- Shi, B; Yang, G; Xiao, Y; Qian, K; Shao, H; Xu, M; et al. Long-Term Protection against Virulent Newcastle Disease Virus (NDV) in Chickens Immunized with a Single Dose of Recombinant Turkey Herpesvirus Expressing NDV F Protein. Vaccines (Basel) 2024, 12(6). [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations. Newcastle disease and its control in village chickens; Rome, 2002. [Google Scholar]
- Dortmans, JCFM; Peeters, BPH; Koch, G. Newcastle disease virus outbreaks: Vaccine mismatch or inadequate application? Vet Microbiol 2012, 160(1–2), 17–22. [Google Scholar] [CrossRef]
- Dortmans, JCFM; Venema-Kemper, S; Peeters, BPH; Koch, G. Field vaccinated chickens with low antibody titres show equally insufficient protection against matching and non-matching genotypes of virulent Newcastle disease virus. Vet Microbiol 2014, 172(1–2), 100–7. [Google Scholar] [CrossRef] [PubMed]
- Liu, H; de Almeida, RS; Gil, P; Majó, N; Nofrarías, M; Briand, FX; et al. Can genotype mismatch really affect the level of protection conferred by Newcastle disease vaccines against heterologous virulent strains? Vaccine 2018, 36(27), 3917–25. [Google Scholar] [CrossRef] [PubMed]
- Miller, PJ; King, DJ; Afonso, CL; Suarez, DL. Antigenic differences among Newcastle disease virus strains of different genotypes used in vaccine formulation affect viral shedding after a virulent challenge. Vaccine 2007, 25(41), 7238–46. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z; He, X; Deng, J; Hu, J; Liu, X. Current situation and future direction of Newcastle disease vaccines; Veterinary research. NLM (Medline), 2022; Vol. 53, p. 99. [Google Scholar]
- Bolhassani, A; Yazdi, SR. DNA immunization as an efficient strategy for vaccination. Avicenna J Med Biotechnol. 2009, 1(2), 71–88. [Google Scholar]
- Gurunathan, S; Klinman, DM; Seder, RA. DNA Vaccines: Immunology, Application, and Optimization. Annu Rev Immunol 2000, 18(1), 927–74. [Google Scholar] [CrossRef]
- Zhou, ZP; Chen, ZR; Bandara, RA; Duan, R; Cao, H; Liu, J; et al. DNA-Based Vaccines: Advances, Applications, and Future Prospects. Genes Dis 2026, 102025. [Google Scholar] [CrossRef]
- Neeli, P; Chai, D; Roy, D; Prajapati, S; Bonam, SR. DNA Vaccines in the Post-mRNA Era: Engineering, Applications, and Emerging Innovations. Int J Mol Sci. 2025, 26(17), 8716. [Google Scholar] [CrossRef]
- Kutzler, MA; Weiner, DB. DNA vaccines: ready for prime time? Nat Rev Genet 2008, 9(10), 776–88. [Google Scholar] [CrossRef]
- Li, J; Wu, G; Huang, Z; Hu, J; Tie, X; Wu, H; et al. Advances and prospects of respiratory mucosal vaccines: mechanisms, technologies, and clinical applications. NPJ Vaccines 2025, 10(1), 230. [Google Scholar] [CrossRef] [PubMed]
- Li, Y; Jin, L; Chen, T. The Effects of Secretory IgA in the Mucosal Immune System. Biomed Res Int. 2020, 2020(1). [Google Scholar] [CrossRef] [PubMed]
- Lu, B; Lim, JM; Yu, B; Song, S; Neeli, P; Sobhani, N; et al. The next-generation DNA vaccine platforms and delivery systems: advances, challenges and prospects. Front Immunol 2024, 15. [Google Scholar] [CrossRef] [PubMed]
- Lim, M; Badruddoza, AZM; Firdous, J; Azad, M; Mannan, A; Al-Hilal, TA; et al. Engineered Nanodelivery Systems to Improve DNA Vaccine Technologies. Pharmaceutics 2020, 12(1), 30. [Google Scholar] [CrossRef]
- Tang, J; Cai, L; Xu, C; Sun, S; Liu, Y; Rosenecker, J; et al. Nanotechnologies in Delivery of DNA and mRNA Vaccines to the Nasal and Pulmonary Mucosa. Nanomaterials 2022, 12(2), 226. [Google Scholar] [CrossRef]
- Bolhassani, A; Javanzad, S; Saleh, T; Hashemi, M; Aghasadeghi, MR; Sadat, SM. Polymeric nanoparticles: potent vectors for vaccine delivery targeting cancer and infectious diseases. Hum Vaccin Immunother. 2014, 10(2), 321–32. [Google Scholar] [CrossRef]
- Shi, Y; Zhu, L; Qiao, Z; Zhai, Y; Di, J; Wang, S; et al. Polymeric particle-based antigen delivery system: From immunological engineering to clinical translation. Int J Pharm X 2025, 10, 100466. [Google Scholar] [CrossRef]
- Kisakova, LA; Apartsin, EK; Nizolenko, LF; Karpenko, LI. Dendrimer-Mediated Delivery of DNA and RNA Vaccines. Pharmaceutics 2023, 15(4), 1106. [Google Scholar] [CrossRef]
- REED, LJ; MUENCH, H. A SIMPLE METHOD OF ESTIMATING FIFTY PER CENT ENDPOINTS12. Am J Epidemiol 1938, 27(3), 493–7. [Google Scholar] [CrossRef]
- Dimitrov, KM; Taylor, TL; Marcano, VC; Williams-Coplin, D; Olivier, TL; Yu, Q; et al. Novel Recombinant Newcastle Disease Virus-Based In Ovo Vaccines Bypass Maternal Immunity to Provide Full Protection from Early Virulent Challenge. Vaccines (Basel) 2021, 9(10), 1189. [Google Scholar] [CrossRef]
- Hitoshi, N; Ken-ichi, Y; Jun-ichi, M. Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene [Internet] 1991 Dec 15 [cited 2026 Jan 23, 108(2), 193–9. Available online: https://pubmed.ncbi.nlm.nih.gov/1660837/. [CrossRef] [PubMed]
- Abdel-Moneim, A; El-Shahawy, A; Yousef, AI; Abd El-Twab, SM; Elden, ZE; Taha, M. Novel polydatin-loaded chitosan nanoparticles for safe and efficient type 2 diabetes therapy: In silico, in vitro and in vivo approaches. Int J Biol Macromol 2020, 154, 1496–504. [Google Scholar] [CrossRef] [PubMed]
- Akerele, G; Ramadan, N; Renu, S; Renukaradhya, GJ; Shanmugasundaram, R; Selvaraj, RK. In vitro characterization and immunogenicity of chitosan nanoparticles loaded with native and inactivated extracellular proteins from a field strain of Clostridium perfringens associated with necrotic enteritis. Vet Immunol Immunopathol 2020, 224, 110059. [Google Scholar] [CrossRef] [PubMed]
- Salvador, A; Sandgren, KJ; Liang, F; Thompson, EA; Koup, RA; Pedraz, JL; et al. Design and evaluation of surface and adjuvant modified PLGA microspheres for uptake by dendritic cells to improve vaccine responses. Int J Pharm. 2015, 496(2), 371–81. [Google Scholar] [CrossRef]
- Adediran, E; Arte, T; Pasupuleti, D; Vijayanand, S; Singh, R; Patel, P; et al. Delivery of PLGA-Loaded Influenza Vaccine Microparticles Using Dissolving Microneedles Induces a Robust Immune Response. Pharmaceutics 2025, 17(4), 510. [Google Scholar] [CrossRef]
- Azab, AA; Yehia, N; Makhareta, M; Samir, M; Shoukry, A; Elhalem Mohamed, AA; et al. Evaluation of inactivated avian influenza virus and Newcastle disease virus bivalent vaccination program against newly circulated H5N8 and NDV strains. Poult Sci. 2023, 102(10), 102952. [Google Scholar] [CrossRef]
- Ganar, K; Das, M; Sinha, S; Kumar, S. Newcastle disease virus: Current status and our understanding. Virus Res 2014, 184, 71–81. [Google Scholar] [CrossRef]
- Miller, PJ; Afonso, CL. Newcastle Disease Virus. In Encyclopedia of Life Sciences; Wiley, 2011. [Google Scholar]
- Ferraro, B; Morrow, MP; Hutnick, NA; Shin, TH; Lucke, CE; Weiner, DB. Clinical Applications of DNA Vaccines: Current Progress. Clinical Infectious Diseases 2011, 53(3), 296–302. [Google Scholar] [CrossRef]
- Shi, R; Liu, X; Wang, Y; Pan, M; Wang, S; Shi, L; et al. Long-term stability and immunogenicity of lipid nanoparticle COVID-19 mRNA vaccine is affected by particle size. Hum Vaccin Immunother 2024, 20(1). [Google Scholar] [CrossRef]
- Hassett, KJ; Higgins, J; Woods, A; Levy, B; Xia, Y; Hsiao, CJ; et al. Impact of lipid nanoparticle size on mRNA vaccine immunogenicity. Journal of Controlled Release 2021, 335, 237–46. [Google Scholar] [CrossRef]
- Saade, F; Petrovsky, N. Technologies for enhanced efficacy of DNA vaccines. Expert Rev Vaccines 2012, 11(2), 189–209. [Google Scholar] [CrossRef] [PubMed]
- Hashemi, P; Osanloo, M; Farjadfar, A; Nasiri-Ghiri, M; Zarenezhad, E; Mahmoodi, S. A multi-epitope protein vaccine encapsulated in alginate nanoparticles as a candidate vaccine against Shigella sonnei. Sci Rep. 2024, 14(1), 22484. [Google Scholar] [CrossRef] [PubMed]
- Jadaa, N; Aldor, N; Jenik, K; Oberhoffner, S; Butkowsky, C; Tran, A; et al. Evaluating Cationic Nanoparticles as Carriers of Antiviral Nucleic Acids; 2024; pp. 309–20. [Google Scholar]
- Nafee, N; Taetz, S; Schneider, M; Schaefer, UF; Lehr, CM. Chitosan-coated PLGA nanoparticles for DNA/RNA delivery: effect of the formulation parameters on complexation and transfection of antisense oligonucleotides. Nanomedicine 2007, 3(3), 173–83. [Google Scholar] [CrossRef] [PubMed]
- Braun, CS; Vetro, JA; Tomalia, DA; Koe, GS; Koe, JG; Russell Middaugh, C. Structure/Function Relationships of Polyamidoamine/DNA Dendrimers as Gene Delivery Vehicles. J Pharm Sci. 2005, 94(2), 423–36. [Google Scholar] [CrossRef]
- Amir Kalvanagh, P; Ebtekara, M; Kokhaei, P; Soleimanjahi, H. Preparation and Characterization of PLGA Nanoparticles Containing Plasmid DNA Encoding Human IFN-lambda-1/IL-29. Iran J Pharm Res. 2019, 18(1), 156–67. [Google Scholar]
- Danhier, F; Ansorena, E; Silva, JM; Coco, R; Le Breton, A; Préat, V. PLGA-based nanoparticles: An overview of biomedical applications. Journal of Controlled Release 2012, 161(2), 505–22. [Google Scholar] [CrossRef]
- WAECKERLEMEN, Y; GROETTRUP, M. PLGA microspheres for improved antigen delivery to dendritic cells as cellular vaccines. Adv Drug Deliv Rev. 2005, 57(3), 475–82. [Google Scholar] [CrossRef]
- Demento, SL; Cui, W; Criscione, JM; Stern, E; Tulipan, J; Kaech, SM; et al. Role of sustained antigen release from nanoparticle vaccines in shaping the T cell memory phenotype. Biomaterials 2012, 33(19), 4957–64. [Google Scholar] [CrossRef]
- Abedi-Gaballu, F; Dehghan, G; Ghaffari, M; Yekta, R; Abbaspour-Ravasjani, S; Baradaran, B; et al. PAMAM dendrimers as efficient drug and gene delivery nanosystems for cancer therapy. Appl Mater Today 2018, 12, 177–90. [Google Scholar] [CrossRef]
- Siepmann, J; Peppas, NA. Higuchi equation: Derivation, applications, use and misuse. Int J Pharm. 2011, 418(1), 6–12. [Google Scholar] [CrossRef]
- Upputuri, RTP; Mandal, AKA. Mathematical Modeling and Release Kinetics of Green Tea Polyphenols Released from Casein Nanoparticles. Iran J Pharm Res. 2019, 18(3), 1137–46. [Google Scholar] [PubMed]
- Guo, Z; Wang, H; Yang, T; Wang, X; Lu, D; Li, Y; et al. Priming with a DNA vaccine and boosting with an inactivated vaccine enhance the immune response against infectious bronchitis virus. J Virol Methods 2010, 167(1), 84–9. [Google Scholar] [CrossRef] [PubMed]
- Shan, S; Fenwick, S; Ellis, T; Poinern, E; Edwards, J; Le, X; et al. Evaluation of different chemical adjuvants on an avian influenza H6 DNA vaccine in chickens. Avian Pathology 2016, 45(6), 649–56. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K; Rong, G; Hao, Y; Yu, L; Kang, H; Wang, X; et al. IgA response and protection following nasal vaccination of chickens with Newcastle disease virus DNA vaccine nanoencapsulated with Ag@SiO2 hollow nanoparticles. Sci Rep. 2016, 6(1), 25720. [Google Scholar] [CrossRef]
- Zhao, K; Zhang, Y; Wang, X; Shi, C; Wang, X; Wang, X; et al. Chitosan-coated poly(lactic-co-glycolic) acid nanoparticles as an efficient delivery system for Newcastle disease virus DNA vaccine. Int J Nanomedicine 2014, 4609. [Google Scholar] [CrossRef]
- Ensign, LM; Cone, R; Hanes, J. Oral drug delivery with polymeric nanoparticles: The gastrointestinal mucus barriers. Adv Drug Deliv Rev. 2012, 64(6), 557–70. [Google Scholar] [CrossRef]
- Kim, D; Kim, S; Na, DH. Dendrimer nanoplatforms for oral drug delivery applications. J Pharm Investig. 2025, 55(2), 169–89. [Google Scholar] [CrossRef]
- Hussain, Z; Khan, S; Imran, M; Sohail, M; Shah, SWA; de Matas, M. PEGylation: a promising strategy to overcome challenges to cancer-targeted nanomedicines: a review of challenges to clinical transition and promising resolution. Drug Deliv Transl Res. 2019, 9(3), 721–34. [Google Scholar] [CrossRef]
- Lai, SK; Wang, YY; Hanes, J. Mucus-penetrating nanoparticles for drug and gene delivery to mucosal tissues. Adv Drug Deliv Rev. 2009, 61(2), 158–71. [Google Scholar] [CrossRef]
- Fröhlich, E. The role of surface charge in cellular uptake and cytotoxicity of medical nanoparticles. Int J Nanomedicine 2012, 5577. [Google Scholar] [CrossRef]
- Hobson, P. Mucosal immunization with DNA vaccines. Methods 2003, 31(3), 217–24. [Google Scholar] [CrossRef]
- Kapczynski, DR; Afonso, CL; Miller, PJ. Immune responses of poultry to Newcastle disease virus. Dev Comp Immunol. 2013, 41(3), 447–53. [Google Scholar] [CrossRef]
- He, C; Hu, Y; Yin, L; Tang, C; Yin, C. Effects of particle size and surface charge on cellular uptake and biodistribution of polymeric nanoparticles. Biomaterials 2010, 31(13), 3657–66. [Google Scholar] [CrossRef]
- Mao, S; Sun, W; Kissel, T. Chitosan-based formulations for delivery of DNA and siRNA. Adv Drug Deliv Rev. 2010, 62(1), 12–27. [Google Scholar] [CrossRef]
- Huo, S; Zuo, Y; Li, N; Li, X; Zhang, Y; Wang, L; et al. Chicken IL-7 as a potent adjuvant enhances IBDV VP2 DNA vaccine immunogenicity and protective efficacy. Vet Microbiol 2016, 193, 145–55. [Google Scholar] [CrossRef]
- Deb, R; Dey, S; Madhan Mohan, C; Gaikwad, S; Kamble, N; Khulape, SA; et al. Development and evaluation of a Salmonella typhimurium flagellin based chimeric DNA vaccine against infectious bursal disease of poultry. Res Vet Sci. 2015, 102, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K; Han, J; Zhang, Y; Wei, L; Yu, S; Wang, X; et al. Enhancing Mucosal Immune Response of Newcastle Disease Virus DNA Vaccine Using N -2-Hydroxypropyl Trimethylammonium Chloride Chitosan and N, O -Carboxymethyl Chitosan Nanoparticles as Delivery Carrier. Mol Pharm. 2018, 15(1), 226–37. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K; Li, W; Huang, T; Luo, X; Chen, G; Zhang, Y; et al. Preparation and Efficacy of Newcastle Disease Virus DNA Vaccine Encapsulated in PLGA Nanoparticles. PLoS One 2013, 8(12), e82648. [Google Scholar] [CrossRef] [PubMed]
| Gr. | Formulation administered | Carrier | Route | Challenge | Survival rate |
|---|---|---|---|---|---|
| 1 | Non-vaccinated-challenged control | — | — | + | 0% |
| 2 | Non-vaccinated-unchallenged control | — | — | - | 100% |
| 3 | Cs-NPs-pDNA-F | Chitosan | Intranasal | + | 30% |
| 4 | PLGA-NPs-pDNA-F | PLGA | Intranasal | + | 90% |
| 5 | PAMAM-Dendrimers-pDNA-F | PAMAM-Dendrimers | Intranasal | + | 50% |
| 6 | Naked pDNA-F | None | Intranasal | + | 50% |
| 7 | Cs-NPs-pDNA-F | Chitosan | Intramuscular | + | 20% |
| 8 | PLGA-NPs-pDNA-F | PLGA | Intramuscular | + | 50% |
| 9 | PAMAM-Dendrimers-pDNA-F | PAMAM-Dendrimers | Intramuscular | + | 100% |
| 10 | Naked pDNA-F | None | Intramuscular | + | 70% |
| Formulation | Hydrodynamic Size (nm) | Polydispersity Index (PDI) | Zeta Potential (mV) |
|---|---|---|---|
| Cs-NPs-pDNA-F | 210 ± 15.5 | 0.21 ± 0.04 | +32.5 ± 2.8 |
| PLGA-NPs-pDNA-F | 255 ± 20.1 | 0.24 ± 0.05 | -21.8 ± 1.9 |
| PAMAM-Dendrimers-pDNA-F | 120 ± 10.2 | 0.16 ± 0.03 | +45.3 ± 3.5 |
| Release mechanism | Cs-NPs-pDNA-F (R²) | PLGA-NPs-pDNA-F (R²) | PAMAM-Dendrimers-pDNA-F (R²) | Equations | Model |
|---|---|---|---|---|---|
| Constant release rate | 0.9034 | 0.9911 | 0.7443 | Mt=k0t+b | Zero order |
|
Concentration- dependent release |
0.9558 | 0.9679 | 0.9466 | ln(M∞−Mt)=ln(M∞)−k1t | First order |
|
Diffusion-controlled ‘ release |
0.9842 | 0.9560 | 0.9226 | Mt=kHt1/2+bMt=kHt1/2+b | Higuchi |
|
Polymer/transport controlled (early phase) |
0.9914 | 0.9869 | 0.9945 | ln(Mt/M∞)=lnkKP+nlnt (≤60%) | Korsmeyer–Peppas |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





