Submitted:
11 February 2026
Posted:
12 February 2026
You are already at the latest version
Abstract

Keywords:
1. Introduction
2. Methods
2.1. Data acquisition and database construction
2.2. Hidden Markov model construction and proteome mining
2.3. Multiple sequence alignment and phylogenetic reconstruction
2.4. Physicochemical profiling and structural fingerprinting
2.5. Motif conservation profiling
2.6. Deep structural validation via protein language modeling
2.7. Quantitative alignment visualization
2.8. Candidate refinement and systematic nomenclature
2.9. Transcriptomic data integration and reciprocal homology mapping
2.10. Integrated phylogenomic visualization and statistical profiling
3. Results
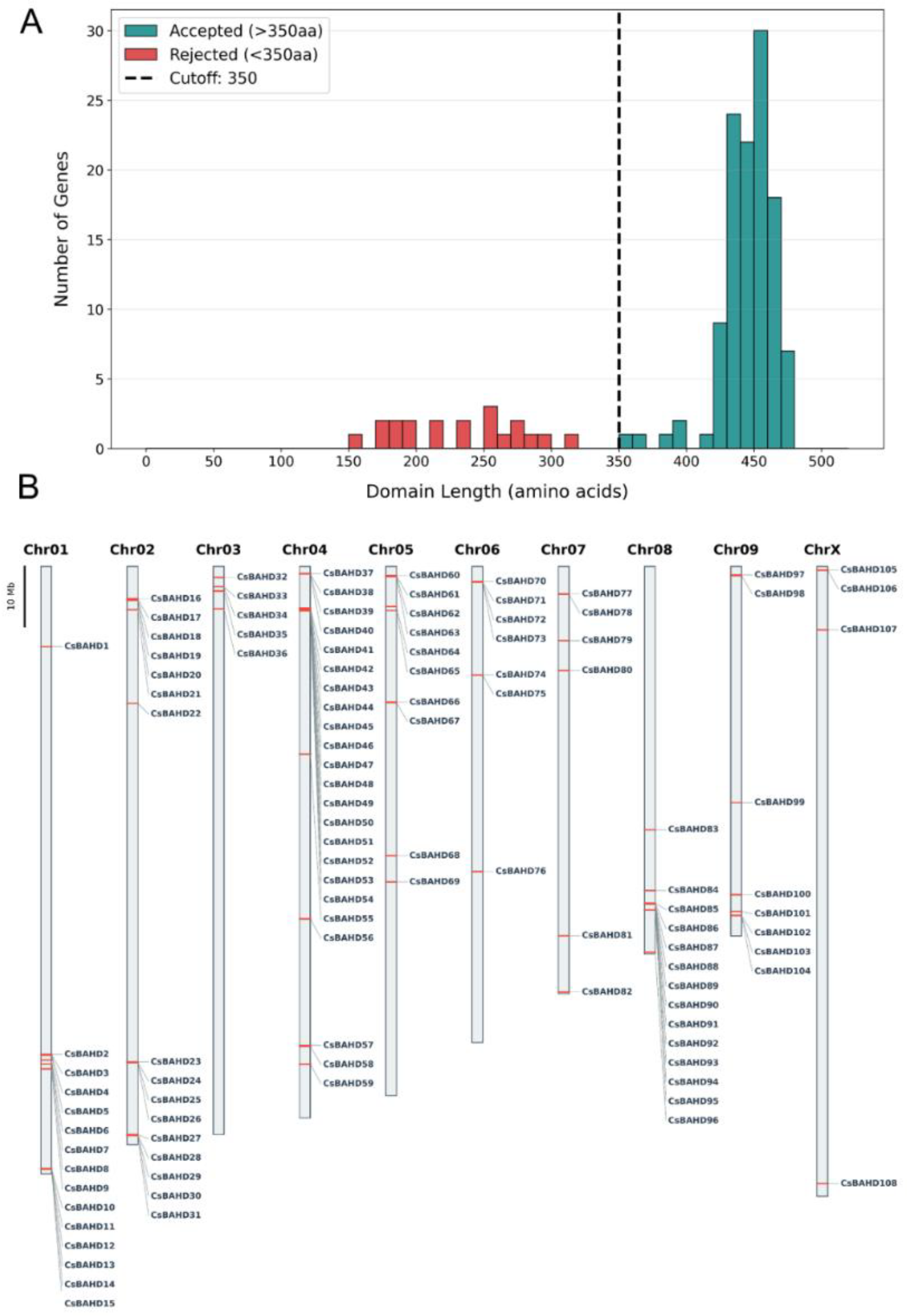
3.1. Genome-wide identification and chromosomal distribution
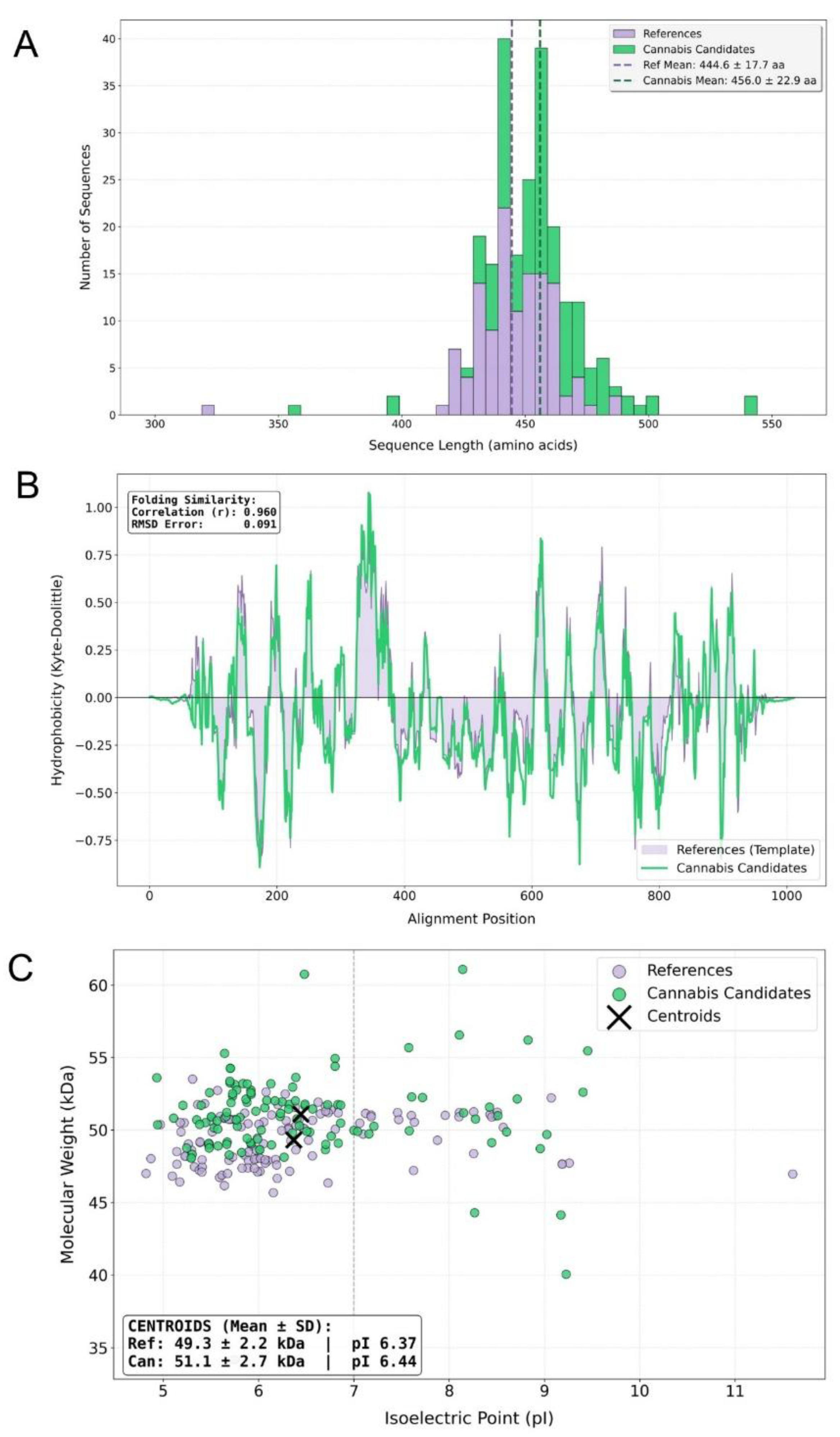
3.2. Structural conservation and physicochemical landscape
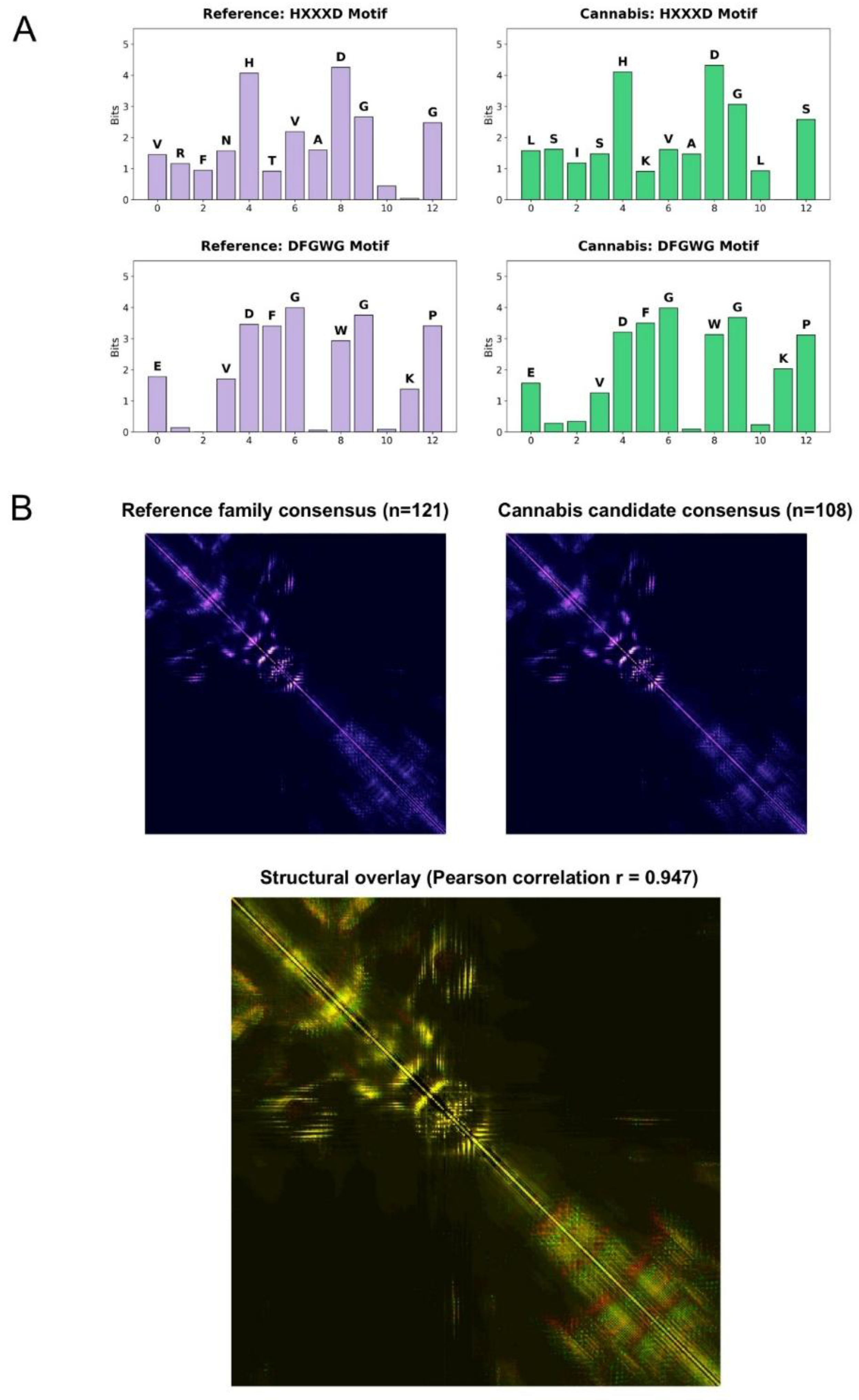
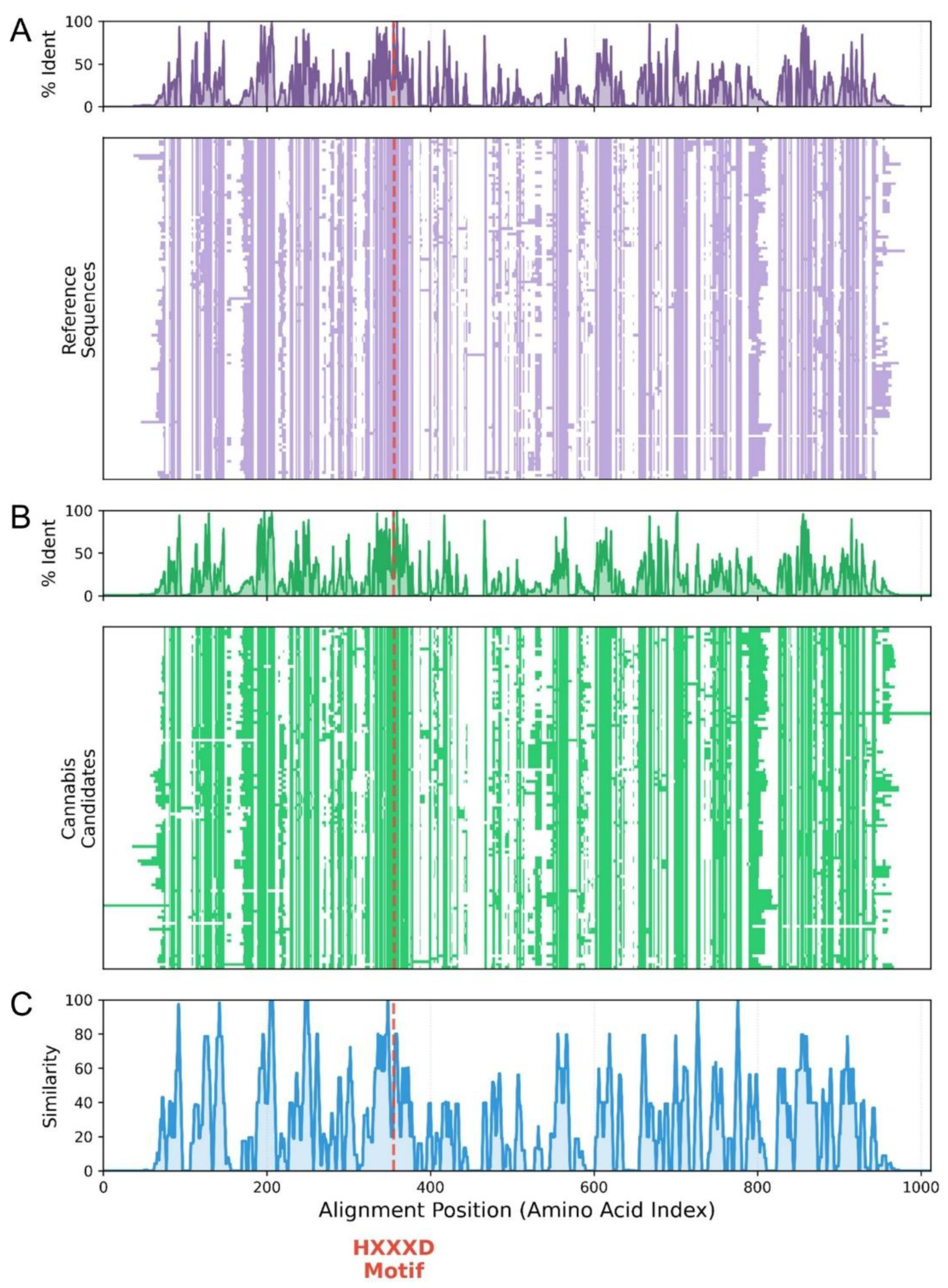
3.3. Conservation of catalytic motifs and deep structural modeling
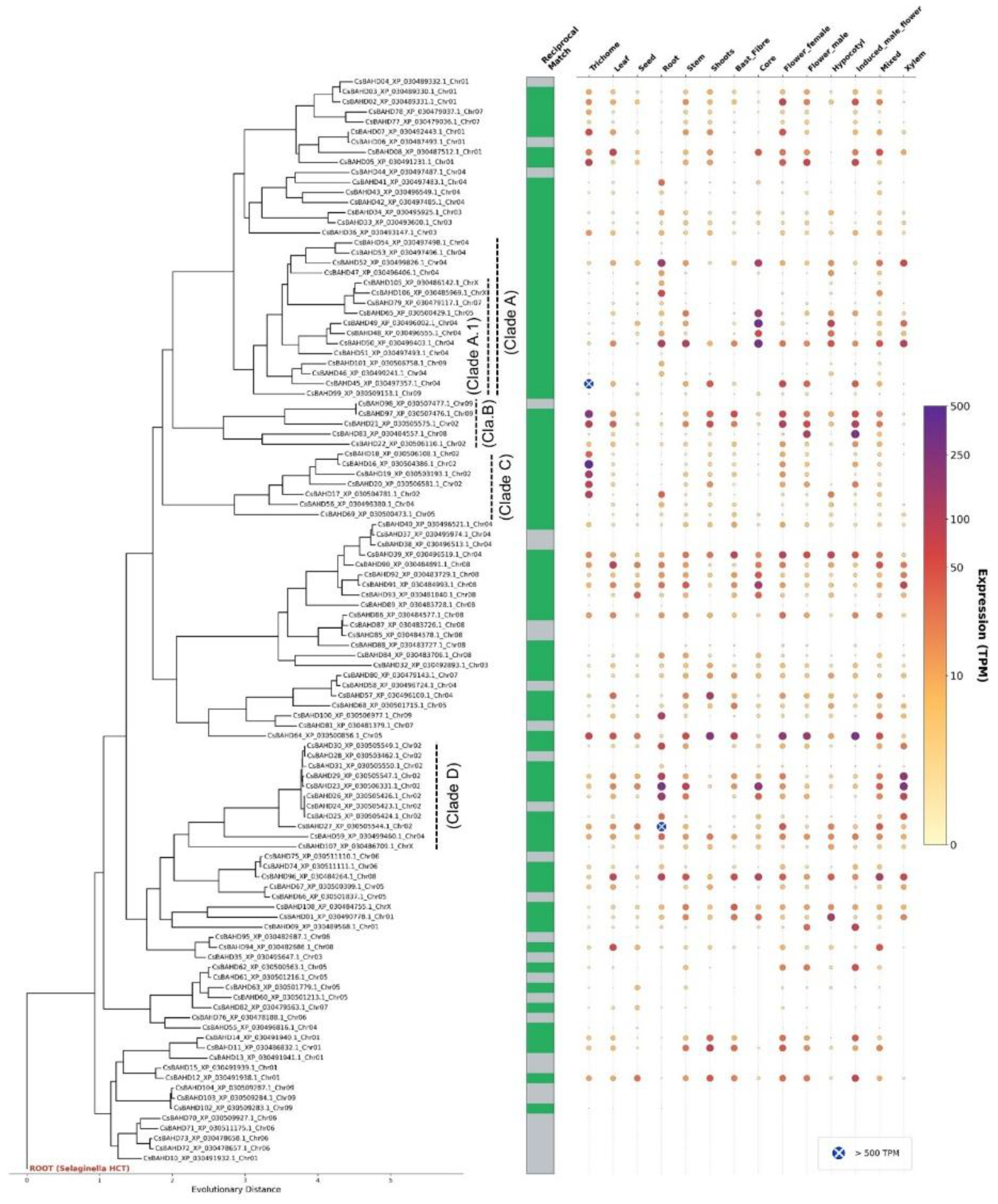
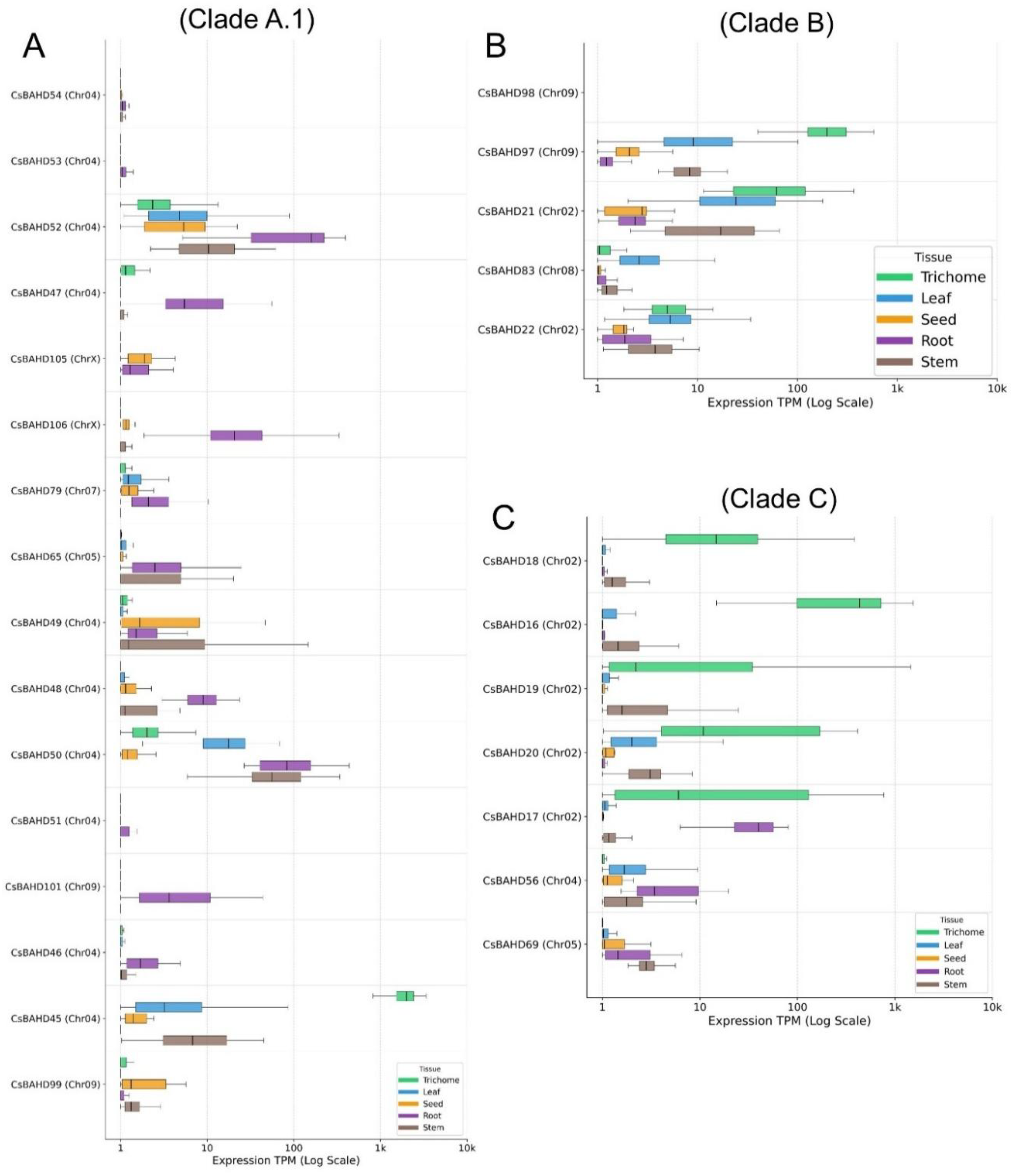
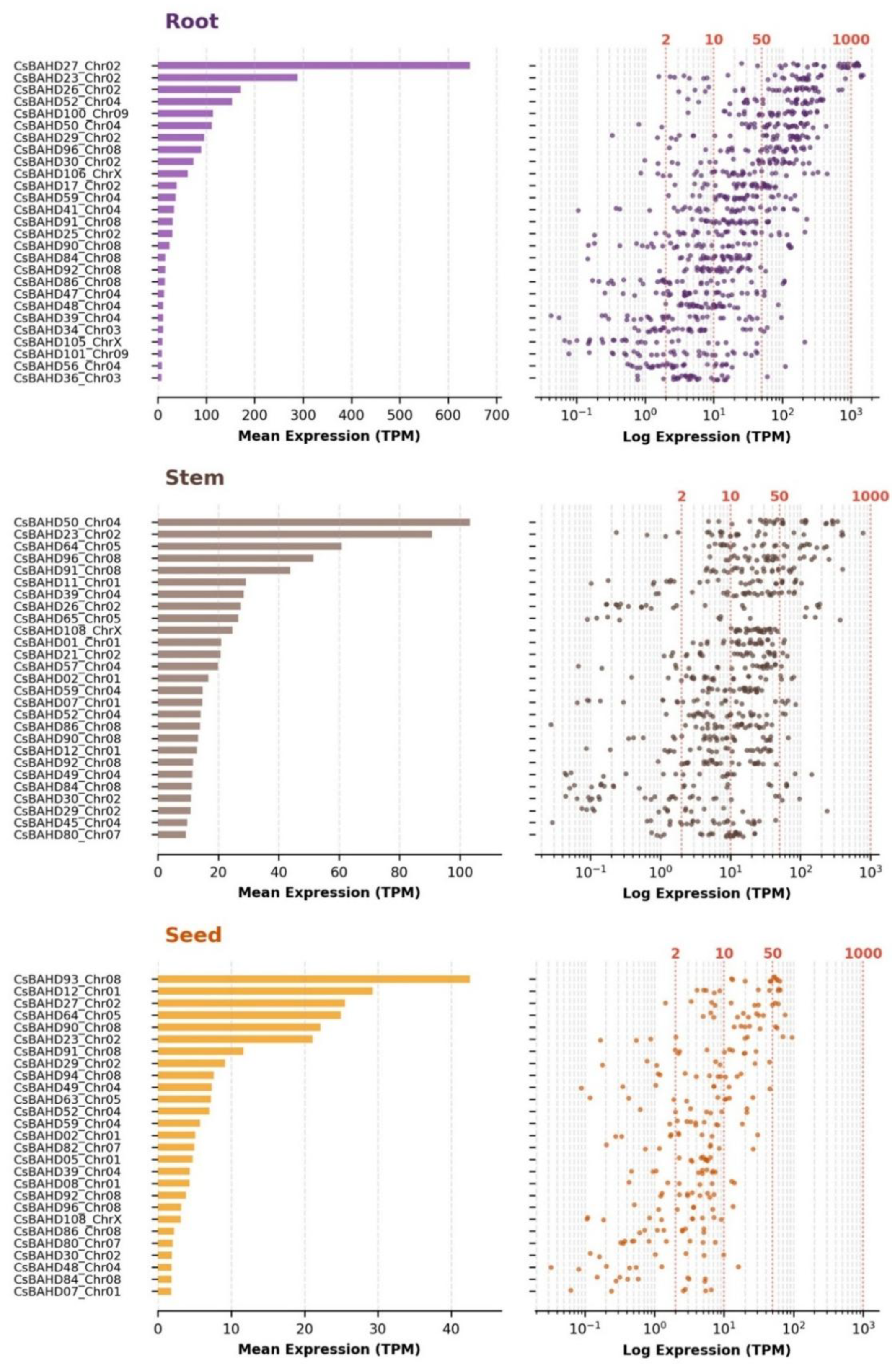
3.4. Phylotranscriptomic landscape and chemotypic functionalization
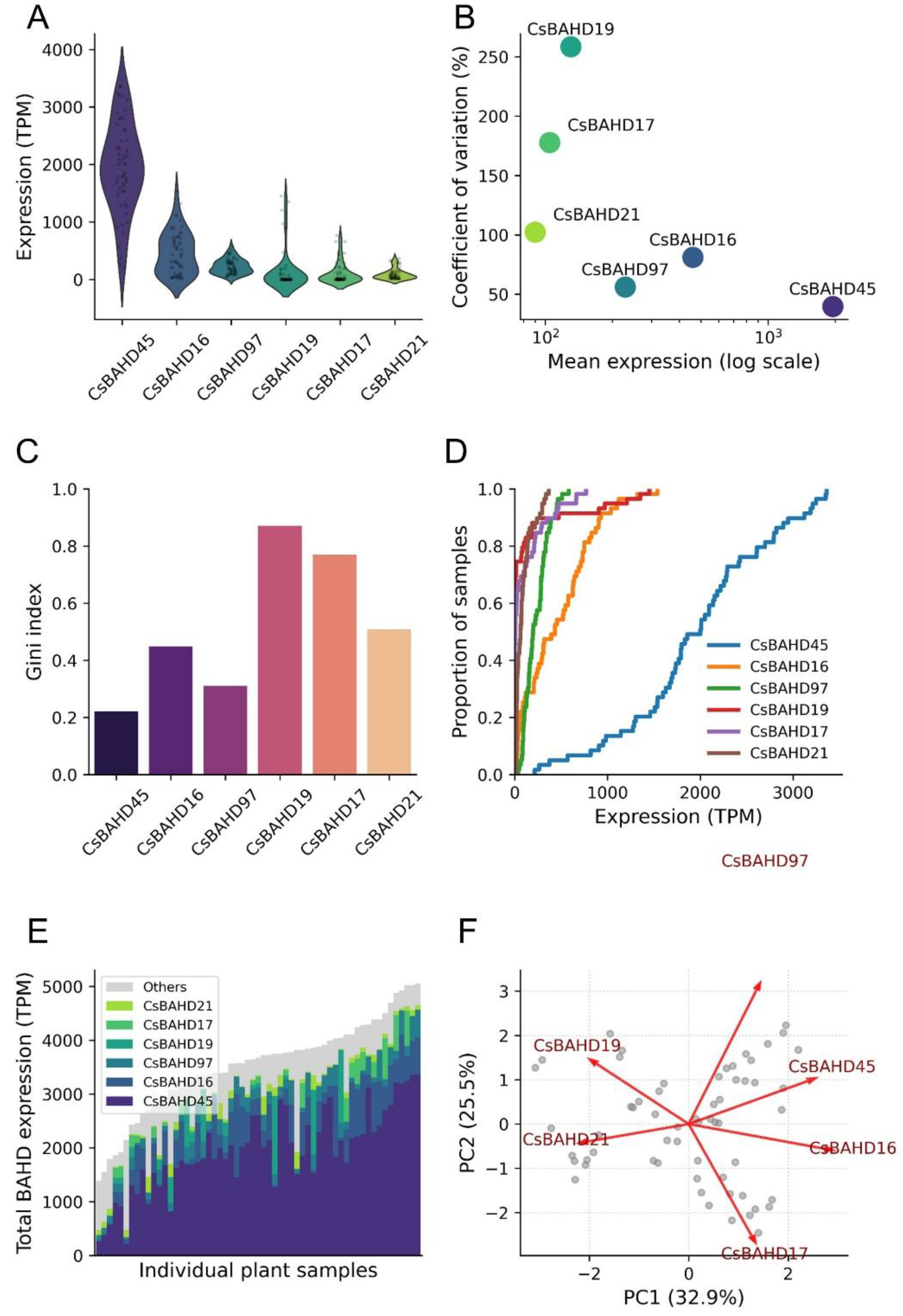
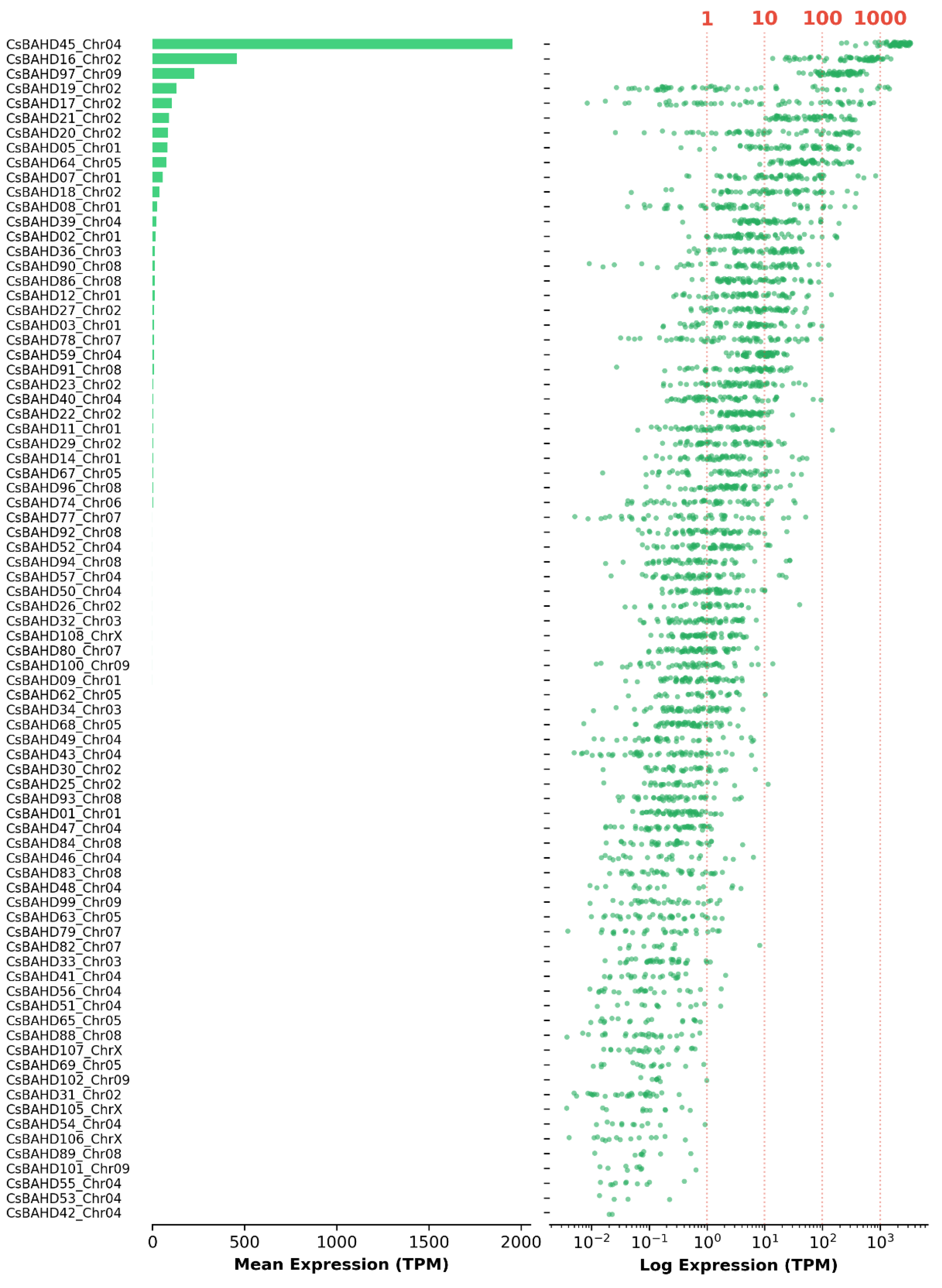
3.5. Stochastic regulation drives chemotypic diversity
3.6. Subterranean and reproductive specialization: The functional differentiation of vegetative modules
4. Discussion
4.1. Genomic landscape and telomeric plasticity for accelerated precision breeding
4.2. Functional specialization in vegetative and reproductive sinks: lignification, rhizosphere defense, and seed maturation as evolutionary scaffolds
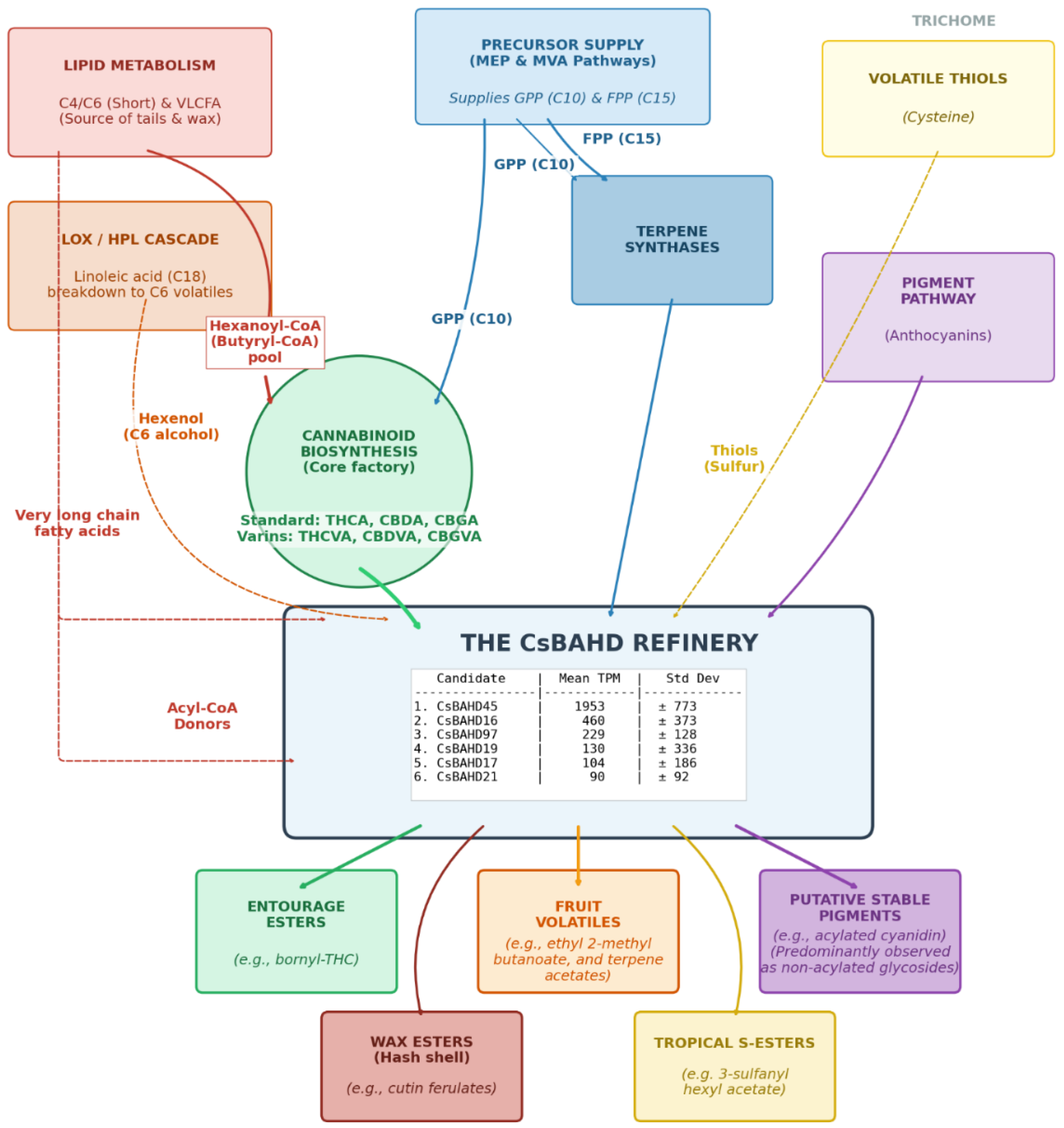
4.3. Transcriptional architecture of the glandular trichome: biosynthesis of fruit esters and the modulation of terpene profiles
4.3.1. Biosynthesis of short-chain fruit esters for floral, sweet and exotic aroma
4.3.2. Acetylation of the terpene pool to create soft delicate floral notes
4.3.3. Structural modeling and physicochemical sequestration of volatile terpenes
4.3.4. The evolutionary enigma of cannabinoid biosynthesis
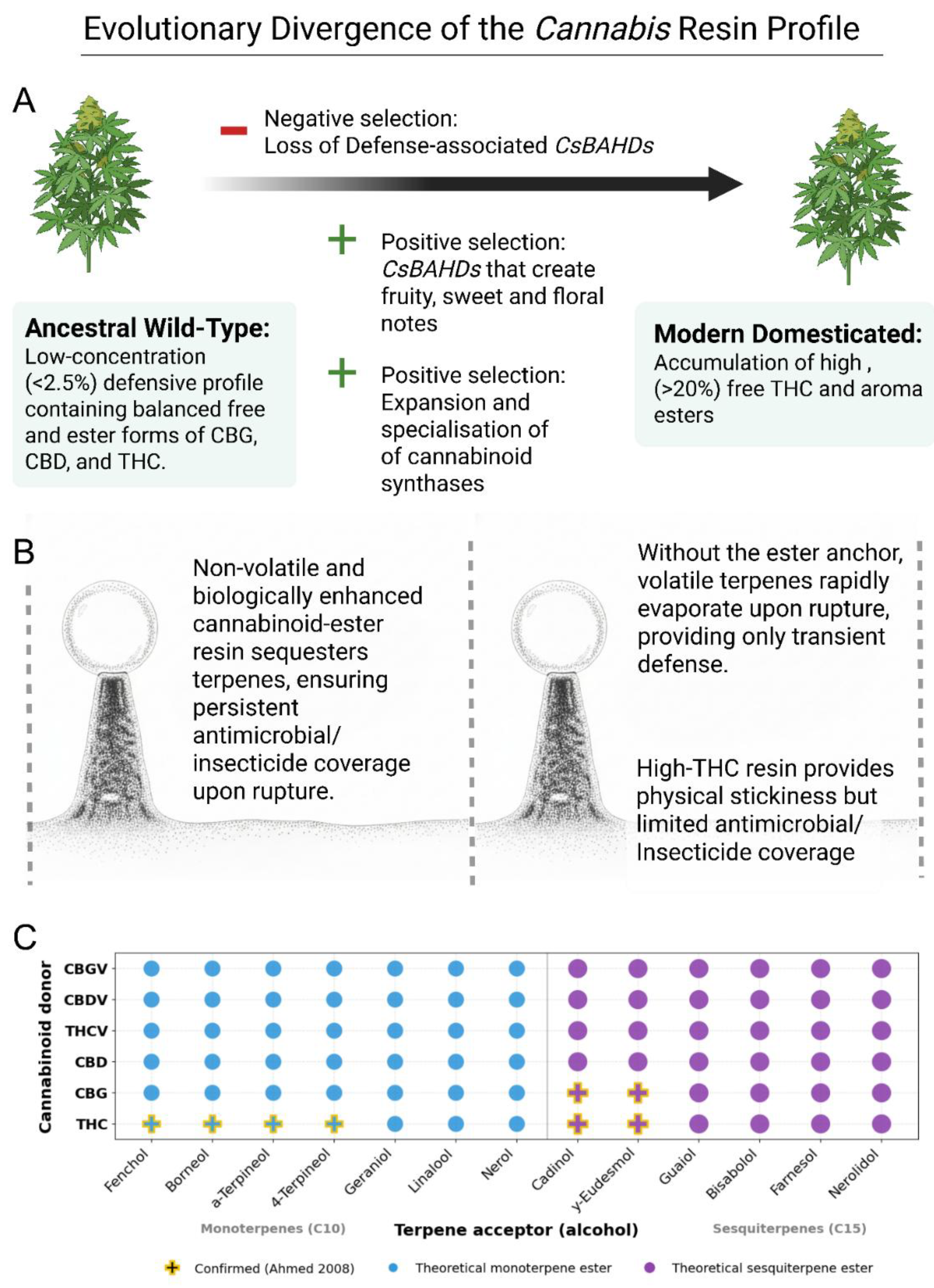
4.3.5. Physicochemical sequestration of volatile terpenes: esters as a mechanism for contact toxicity
4.3.6. Chemotypic divergence trough domestication and the uncoupling of ancestral defense pathways
Funding
Conflicts of Interest
Abbreviations
| 4CL | 4-coumarate:CoA ligase |
| AAT | alcohol acyltransferase |
| ATP | adenosine triphosphate |
| BLAST | basic local alignment search tool |
| CBD | cannabidiol |
| CBDA | cannabidiolic acid |
| CBDV | cannabidivarin |
| CBG | cannabigerol |
| CBGA | cannabigerolic acid |
| CBGV | cannabigerovarin |
| CoA | coenzyme A |
| CNV | copy number variation |
| CV | coefficient of variation |
| DAT | deacetylvindoline 4-O-acetyltransferase |
| ECDF | empirical cumulative distribution function |
| ESM | evolutionary scale modeling |
| FPP | farnesyl pyrophosphate |
| GPP | geranyl pyrophosphate |
| HCBT | anthranilate N-hydroxycinnamoyl/benzoyltransferase |
| HCT | hydroxycinnamoyl-CoA:shikimate/quinate hydroxycinnamoyltransferase |
| HMM | hidden Markov model |
| HPL | hydroperoxide lyase |
| KDE | kernel density estimate |
| LG | Le-Gascuel |
| LOX | lipoxygenase |
| MD | molecular dynamics |
| MEP | methylerythritol phosphate |
| MVA | mevalonate |
| MW | molecular weight |
| PAV | presence/absence variation |
| PC | principal component |
| PCA | principal component analysis |
| pI | isoelectric point |
| RBH | reciprocal best hit |
| RMSD | root mean square deviation |
| RNA-seq | RNA sequencing |
| TDA | tandem duplication array |
| THC | tetrahydrocannabinol |
| THCA | tetrahydrocannabinolic acid |
| THCV | tetrahydrocannabivarin |
| TPM | transcripts per million |
| UV-B | ultraviolet B |
| VOC | volatile organic compound |
| VSC | volatile sulfur compound |
References
- T.K.L. Tran, T. Avellaneda, A. André, E. Gillich, M. Steinhaus, D.Á. Carrera, L. Katsir, I. Chetschik, The Plant of Many Scents: Unraveling the Odorant Composition of Selected CBD Hemp Cultivars, J. Agric. Food Chem. 73 (2025) 24314–24325. [CrossRef]
- S. Rice, J.A. Koziel, Characterizing the Smell of Marijuana by Odor Impact of Volatile Compounds: An Application of Simultaneous Chemical and Sensory Analysis, PLOS ONE 10 (2015) e0144160. [CrossRef]
- I.W.H. Oswald, T.R. Paryani, M.E. Sosa, M.A. Ojeda, M.R. Altenbernd, J.J. Grandy, N.S. Shafer, K. Ngo, J.R. Peat, B.G. Melshenker, I. Skelly, K.A. Koby, M.F.Z. Page, T.J. Martin, Minor, Nonterpenoid Volatile Compounds Drive the Aroma Differences of Exotic Cannabis, ACS Omega 8 (2023) 39203–39216. [CrossRef]
- I.W.H. Oswald, M.A. Ojeda, R.J. Pobanz, K.A. Koby, A.J. Buchanan, J. Del Rosso, M.A. Guzman, T.J. Martin, Identification of a New Family of Prenylated Volatile Sulfur Compounds in Cannabis Revealed by Comprehensive Two-Dimensional Gas Chromatography, ACS Omega 6 (2021) 31667–31676. [CrossRef]
- K.P. Kaminski, J. Hoeng, K. Lach-Falcone, F. Goffman, W.K. Schlage, D. Latino, Exploring Aroma and Flavor Diversity in Cannabis sativa L.—A Review of Scientific Developments and Applications, Molecules 30 (2025) 2784. [CrossRef]
- T.R. Paryani, M.E. Sosa, M.F.Z. Page, T.J. Martin, M.V. Hearvy, M.A. Ojeda, K.A. Koby, J.J. Grandy, B.G. Melshenker, I. Skelly, I.W.H. Oswald, Nonterpenoid Chemical Diversity of Cannabis Phenotypes Predicts Differentiated Aroma Characteristics, ACS Omega 9 (2024) 28806–28815. [CrossRef]
- P. Janta, S. Vimolmangkang, Chemical profiling and clustering of various dried cannabis flowers revealed by volatilomics and chemometric processing, J. Cannabis Res. 6 (2024) 41. [CrossRef]
- D. Xu, Z. Wang, W. Zhuang, T. Wang, Y. Xie, Family characteristics, phylogenetic reconstruction, and potential applications of the plant BAHD acyltransferase family, Front. Plant Sci. 14 (2023). [CrossRef]
- W. Zhang, J. Li, Y. Dong, Y. Huang, Y. Qi, H. Bai, H. Li, L. Shi, Genome-wide identification and expression of BAHD acyltransferase gene family shed novel insights into the regulation of linalyl acetate and lavandulyl acetate in lavender, J. Plant Physiol. 292 (2024) 154143. [CrossRef]
- Sharafi, H. Hashemi Sohi, A. Mousavi, P. Azadi, B. Dehsara, B. Hosseini Khalifani, Enhanced morphinan alkaloid production in hairy root cultures of Papaver bracteatum by over-expression of salutaridinol 7-o-acetyltransferase gene via Agrobacterium rhizogenes mediated transformation, World J. Microbiol. Biotechnol. 29 (2013) 2125–2131. [CrossRef]
- St-Pierre, P. Laflamme, A.-M. Alarco, V. D, E. Luca, The terminal O-acetyltransferase involved in vindoline biosynthesis defines a new class of proteins responsible for coenzyme A-dependent acyl transfer, Plant J. 14 (1998) 703–713. [CrossRef]
- K. Barbosa-Xavier, F. Pedrosa-Silva, F. Almeida-Silva, T.M. Venancio, Cannabis Expression Atlas: a comprehensive resource for integrative analysis of Cannabis sativa L. gene expression, Physiol. Plant. 176 (2024) e70010. [CrossRef]
- M. Shalit, I. Guterman, H. Volpin, E. Bar, T. Tamari, N. Menda, Z. Adam, D. Zamir, A. Vainstein, D. Weiss, E. Pichersky, E. Lewinsohn, Volatile Ester Formation in Roses. Identification of an Acetyl-Coenzyme A. Geraniol/Citronellol Acetyltransferase in Developing Rose Petals, Plant Physiol. 131 (2003) 1868–1876. [CrossRef]
- C. Liu, X. Qiao, Q. Li, W. Zeng, S. Wei, X. Wang, Y. Chen, X. Wu, J. Wu, H. Yin, S. Zhang, Genome-wide comparative analysis of the BAHD superfamily in seven Rosaceae species and expression analysis in pear (Pyrus bretschneideri), BMC Plant Biol. 20 (2020) 14. [CrossRef]
- S. Yadav, W. Maurya, R. Kumari, P. Rangan, A.B. Gaikwad, Genome-wide identification and characterization of BAHD acyltransferases in black pepper (Piper nigrum L.) reveals their important role in piperine biosynthesis, Sci. Rep. 15 (2025) 39415. [CrossRef]
- R.C. Lynch, L.K. Padgitt-Cobb, A.R. Garfinkel, B.J. Knaus, N.T. Hartwick, N. Allsing, A. Aylward, P.C. Bentz, S.B. Carey, A. Mamerto, J.K. Kitony, K. Colt, E.R. Murray, T. Duong, H.I. Chen, A. Trippe, A. Harkess, S. Crawford, K. Vining, T.P. Michael, Domesticated cannabinoid synthases amid a wild mosaic cannabis pangenome, Nature 643 (2025) 1001–1010. [CrossRef]
- Molina, Y. Li-Beisson, F. Beisson, J.B. Ohlrogge, M. Pollard, Identification of an Arabidopsis Feruloyl-Coenzyme A Transferase Required for Suberin Synthesis, Plant Physiol. 151 (2009) 1317–1328. [CrossRef]
- L. de Vries, H.A. MacKay, R.A. Smith, Y. Mottiar, S.D. Karlen, F. Unda, E. Muirragui, C. Bingman, K. Vander Meulen, E.T. Beebe, B.G. Fox, J. Ralph, S.D. Mansfield, pHBMT1, a BAHD-family monolignol acyltransferase, mediates lignin acylation in poplar, Plant Physiol. 188 (2021) 1014–1027. [CrossRef]
- L.P.Y. Lam, Y. Tobimatsu, S. Suzuki, T. Tanaka, S. Yamamoto, Y. Takeda-Kimura, Y. Osakabe, K. Osakabe, J. Ralph, L.E. Bartley, T. Umezawa, Disruption of p-coumaroyl-CoA:monolignol transferases in rice drastically alters lignin composition, Plant Physiol. 194 (2024) 832–848. [CrossRef]
- R.A. Smith, E.T. Beebe, C.A. Bingman, K. Vander Meulen, A. Eugene, A.J. Steiner, S.D. Karlen, J. Ralph, B.G. Fox, Identification and characterization of a set of monocot BAHD monolignol transferases, Plant Physiol. 189 (2022) 37–48. [CrossRef]
- J. Luo, C. Fuell, A. Parr, L. Hill, P. Bailey, K. Elliott, S.A. Fairhurst, C. Martin, A.J. Michael, A Novel Polyamine Acyltransferase Responsible for the Accumulation of Spermidine Conjugates in Arabidopsis Seed, Plant Cell 21 (2009) 318–333. [CrossRef]
- J. Beekwilder, M. Alvarez-Huerta, E. Neef, F.W.A. Verstappen, H.J. Bouwmeester, A. Aharoni, Functional Characterization of Enzymes Forming Volatile Esters from Strawberry and Banana, Plant Physiol. 135 (2004) 1865–1878. [CrossRef]
- J. Gertsch, M. Leonti, S. Raduner, I. Racz, J.-Z. Chen, X.-Q. Xie, K.-H. Altmann, M. Karsak, A. Zimmer, Beta-caryophyllene is a dietary cannabinoid, Proc. Natl. Acad. Sci. U. S. A. 105 (2008) 9099–9104. [CrossRef]
- S.A. Ahmed, S.A. Ross, D. Slade, M.M. Radwan, F. Zulfiqar, R.R. Matsumoto, Y.-T. Xu, E. Viard, R.C. Speth, V.T. Karamyan, M.A. ElSohly, Cannabinoid ester constituents from high-potency Cannabis sativa, J. Nat. Prod. 71 (2008) 536–542. [CrossRef]
- Levsh, Y.-C. Chiang, C.F. Tung, J.P. Noel, Y. Wang, J.-K. Weng, Dynamic Conformational States Dictate Selectivity toward the Native Substrate in a Substrate-Permissive Acyltransferase, Biochemistry 55 (2016) 6314–6326. [CrossRef]
- D.W. Pate, Possible role of ultraviolet radiation in evolution ofCannabis chemotypes, Econ. Bot. 37 (1983) 396–405. [CrossRef]
- J. Lydon, A.H. Teramura, C.B. Coffman, UV-B RADIATION EFFECTS ON PHOTOSYNTHESIS, GROWTH and CANNABINOID PRODUCTION OF TWO Cannabis sativa CHEMOTYPES, Photochem. Photobiol. 46 (1987) 201–206. [CrossRef]
- D.W. Pate, Chemical ecology of Cannabis, (1994). https://www.druglibrary.net/olsen/HEMP/IHA/iha01201.html (accessed February 9, 2026).
- V. Mediavilla, S. Steinemann, Essential oil of Cannabis sativa L. strains, (n.d.). https://www.druglibrary.net/olsen/HEMP/IHA/jiha4208.html (accessed February 9, 2026).
- C. Villard, I. Baser, A.C. van de Peppel, K. Cankar, M.E. Schranz, R. van Velzen, Resurrected Ancestral Cannabis Enzymes Unveil the Origin and Functional Evolution of Cannabinoid Synthases, Plant Biotechnol. J. n/a (n.d.). [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



