Submitted:
12 February 2026
Posted:
12 February 2026
You are already at the latest version
Abstract
Horizontal Gene Transfer (HGT) is a hallmark of the evolution of parasitic plants, facilitated by the haustorial connection. While mitochondrial HGT is widespread, the extent of nuclear HGT and the long-term retention of foreign genetic material in holoparasitic lineages remain poorly understood. This study explores the genomic architecture of Prosopanche americana (Hydnoraceae), a non-photosynthetic holoparasite currently specialized on Fabaceae. Through a comparative phylogenomic approach integrating draft mitochondrial genomes (mtDNA) and nuclear transcriptomes of P. americana, we identified a multi-layered landscape of foreign DNA. The mtDNA of P. americana contains 18 foreign regions (>500 bp) primarily derived from Solanales, Malvales, and Fabales. Notably, 13 of these regions are shared with P. panguanensis, indicating they were acquired in their common ancestor before speciation and ecological shift. In the nuclear genome, we identified 305 horizontally acquired transcripts (101 orthogroups) with high confidence. Functional analysis revealed an enrichment of foreign genes involved in metabolic pathways and plastid functions (e.g., photosystems and thylakoids) exclusively derived from the ancestral host order Solanales. Our results demonstrate that the genome of P. americana acts as a “molecular fossil,” preserving evidence of past ecological interactions with diverse host lineages. The disparity in HGT footprints between the current host (Fabaceae) and ancestral hosts suggests a period of high genomic plasticity followed by host specialization, providing new insights into the timing and dynamics of horizontal gene flow in holoparasitic Piperales
Keywords:
1. Introduction
2. Materials and Methods
2.1. Mitochondrial Genome Assemblies
2.2. Identification of High-Confidence Mitochondrial HGT Candidate Intergenic Regions
2.3. Sequencing and Assembly of the Prosopanche americana Transcriptome
2.4. Orthogroup Inference and Identification of Foreign Nuclear Genes
- Similarity Filter (BLASTP): We selected OGs where at least one Prosopanche sequence had its best hit (BLASTP, e-value < 1e-5) against a species from the host families.
- Phylogenetic Informativeness: OGs with a minimum of three taxa were retained to ensure robust topological inference.
2.5. Phylogenetic Reconstruction of Nuclear Foreign Candidate Genes
2.6. Nuclear HGT Event Detection and Curation
2.7. Functional Annotation of Foreign Nuclear Genes
3. Results
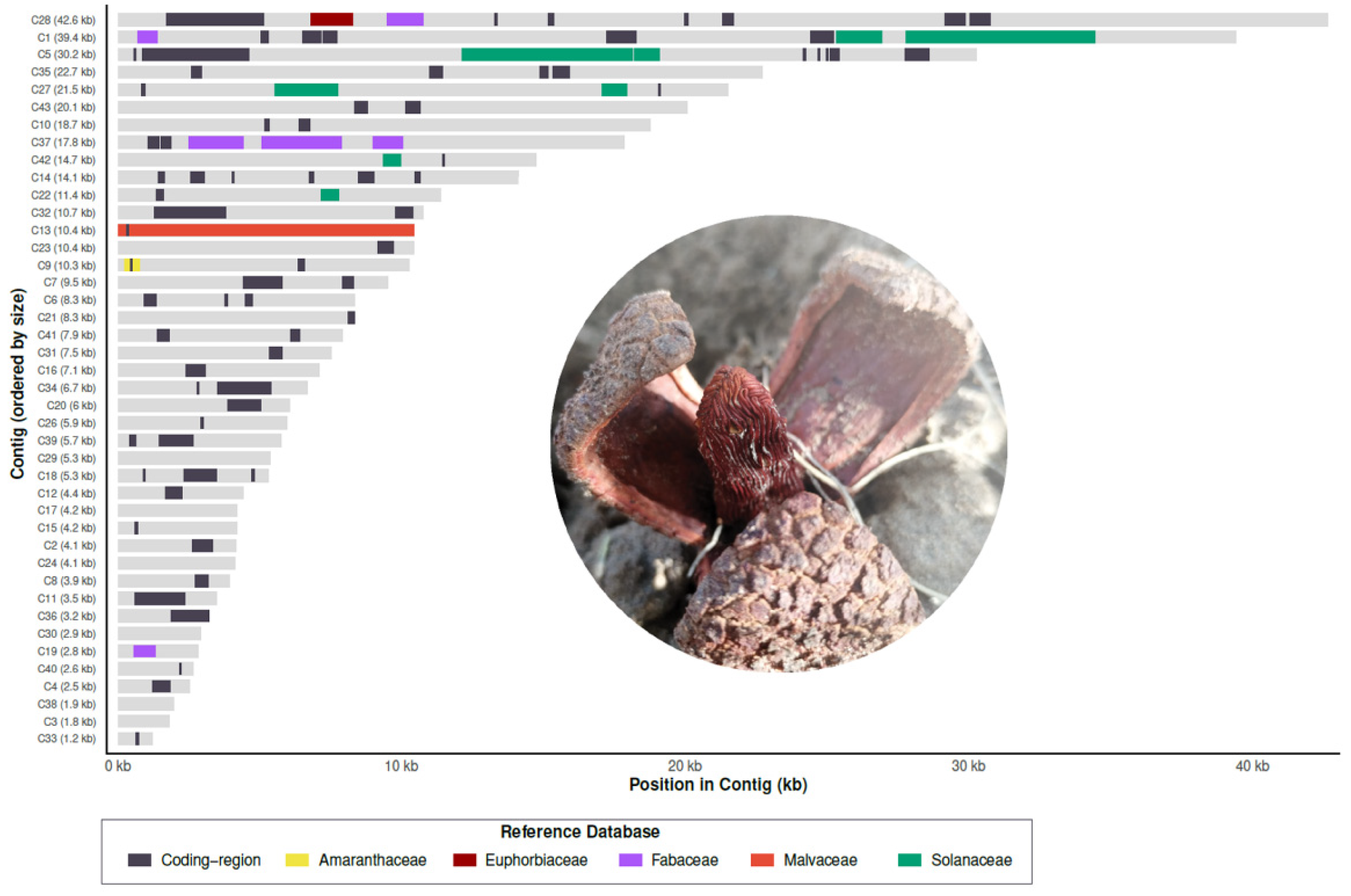
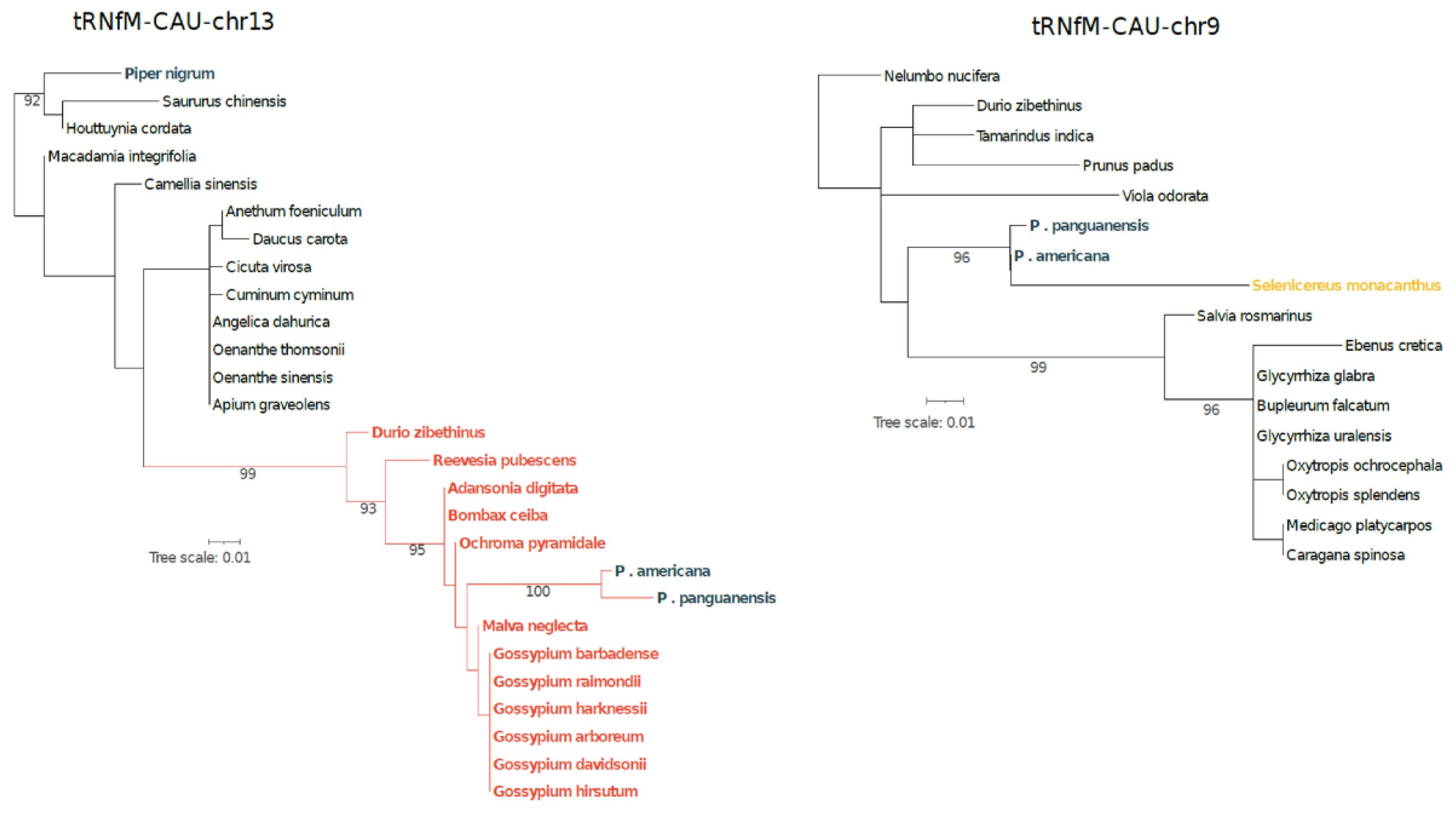
3.1. Characterization and Origin of Mitochondrial HGTs in Prosopanche americana
3.2. HGT Impact on Mitochondrial Coding Regions
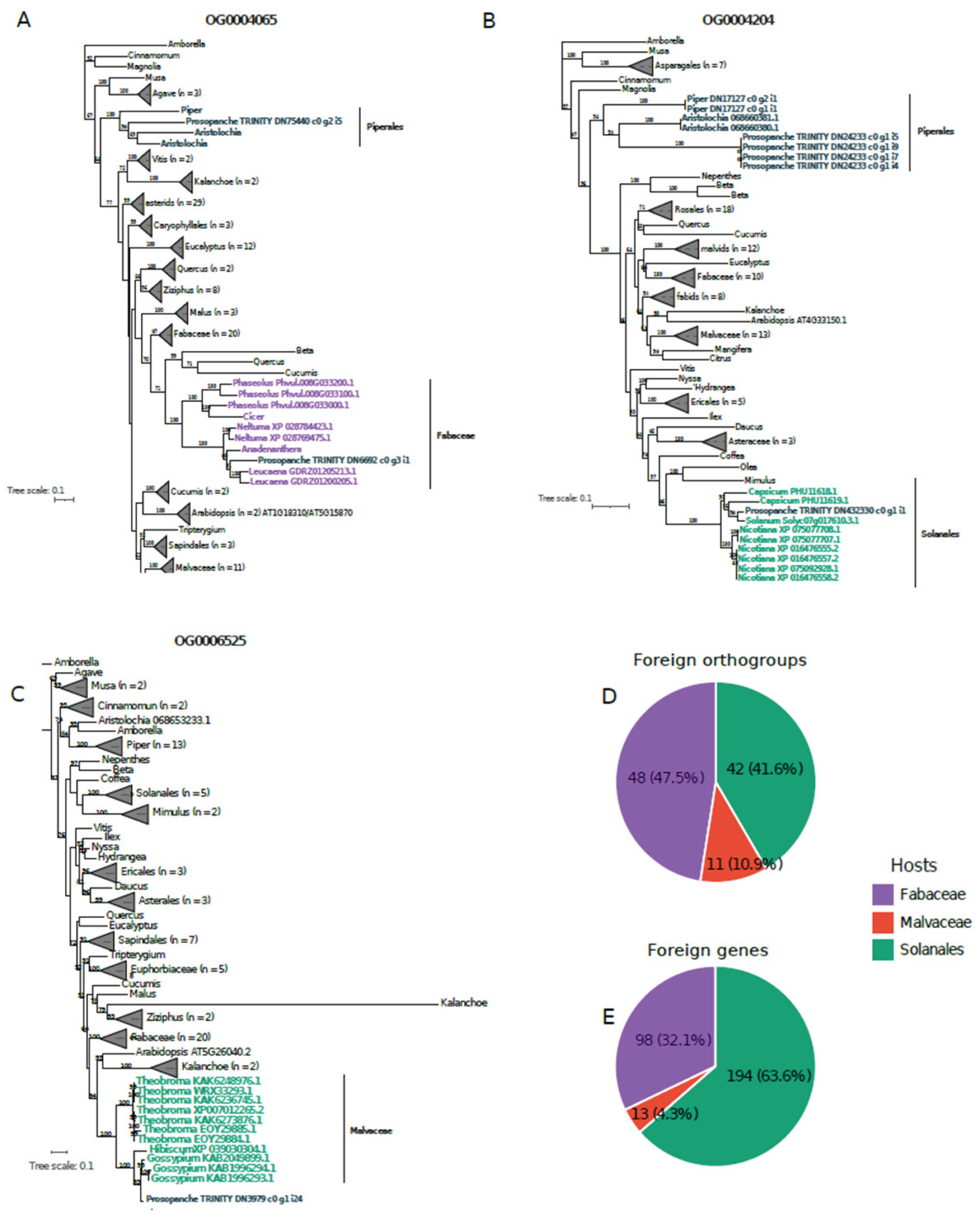
3.3. HGT from Different Hosts in the Nuclear Genome of Prosopanche americana
3.4. Functional Analyses of Foreign Nuclear Transcripts
4. Discussion
4.1. Molecular Fossils of an Ancestral Generalist in the mtDNA and in the Nucleus of Prosopanche
4.2. Comparative HGT Dynamics and Functional Landscape
4.3. Functional Bias in Nuclear Retained Sequences
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Cai, Liming. Rethinking Convergence in Plant Parasitism through the Lens of Molecular and Population Genetic Processes. American Journal of Botany 2023, 110(5), e16174. [Google Scholar] [CrossRef] [PubMed]
- Cai, Liming; Arnold, Brian J.; Xi, Zhenxiang; et al. Deeply Altered Genome Architecture in the Endoparasitic Flowering Plant Sapria Himalayana Griff. (Rafflesiaceae). Current Biology 2021, 31(5), 1002–1011.e9. [Google Scholar] [CrossRef] [PubMed]
- Cantalapiedra, Carlos P; Hernández-Plaza, Ana; Letunic, Ivica; Bork, Peer; Huerta-Cepas, Jaime. eggNOG-Mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Molecular Biology and Evolution 2021, 38(12), 5825–29. [Google Scholar] [CrossRef] [PubMed]
- Chen, Xiaoli; Fang, Dongming; Wu, Chenyu; et al. Comparative Plastome Analysis of Root- and Stem-Feeding Parasites of Santalales Untangle the Footprints of Feeding Mode and Lifestyle Transitions. Genome Biology and Evolution 2020, 12(1), 3663–76. [Google Scholar] [CrossRef] [PubMed]
- Chen, Xiaoli; Fang, Dongming; Xu, Yuxing; et al. Balanophora Genomes Display Massively Convergent Evolution with Other Extreme Holoparasites and Provide Novel Insights into Parasite–Host Interactions. Nature Plants 2023, 9(10), 1627–42. [Google Scholar] [CrossRef]
- Cocucci, Alfredo. Estudios en el genero Prosopanche (Hydnoraceae); Sec. 2. (Cordoba), 1965. [Google Scholar]
- Cocucci, ALfredo. Estudios en el genero Prosopanche II . Organización de la Flor 1975, 8 (December), 7–15. [Google Scholar]
- Cocucci, Alfredo. Estudios En El Genero Prosopanche III, Embriologia; Sec. 9. (Cordoba), 1976. [Google Scholar]
- Cusimano, Natalie; Renner, Susanne S. Sequential Horizontal Gene Transfers from Different Hosts in a Widespread Eurasian Parasitic Plant, Cynomorium Coccineum. American Journal of Botany 2019, 106(5), 679–89. [Google Scholar] [CrossRef]
- Davis, Charles C.; Wurdack, Kenneth J. Host-to-Parasite Gene Transfer in Flowering Plants: Phylogenetic Evidence from Malpighiales. Science 2004, 305(5684), 676–78. [Google Scholar] [CrossRef]
- Davis, Charles C; Xi, Zhenxiang. Horizontal Gene Transfer in Parasitic Plants. Current Opinion in Plant Biology 2015, 26 (August), 14–19. [Google Scholar] [CrossRef]
- Emms, David M.; Kelly, Steven. OrthoFinder: Phylogenetic Orthology Inference for Comparative Genomics. Genome Biology 2019, 20(1), 238. [Google Scholar] [CrossRef]
- Feng, Yanlei; Wicke, Susann. Systemic Organellar Genome Reconfiguration along the Parasitic Continuum in the Broomrape Family (Orobanchaceae). Plant and Cell Physiology 2025, pcaf131. [Google Scholar] [CrossRef]
- Garcia, Laura E.; Edera, Alejandro A.; Palmer, Jeffrey D.; Sato, Hector; Sanchez-Puerta, M. Virginia. Horizontal Gene Transfers Dominate the Functional Mitochondrial Gene Space of a Holoparasitic Plant. New Phytologist 2021, 229(3), 1701–14. [Google Scholar] [CrossRef] [PubMed]
- Gatica-Soria, Leonardo Martin; Roulet, M. Emilia; Tulle, Walter D.; Sato, Hector A.; Barrandeguy, M. Eugenia; Sanchez-Puerta, M. Virginia. Highly Variable Mitochondrial Chromosome Content in a Holoparasitic Plant Due to Recurrent Gains of Foreign Circular DNA. Physiologia Plantarum 2025, 177(2). [Google Scholar] [CrossRef] [PubMed]
- Góralski, Grzegorz; Denysenko-Bennett, Magdalena; Burda, Anna; Staszecka-Moskal, Natalia; Kwolek, Dagmara. Achievements in Horizontal Gene Transfer Studies in Parasitic Plants. Acta Biologica Cracoviensia s. Botanica 2022, 17–28. [Google Scholar] [CrossRef]
- Grabherr, Manfred G; Haas, Brian J; Yassour, Moran; et al. Full-Length Transcriptome Assembly from RNA-Seq Data without a Reference Genome. Nature Biotechnology 2011, 29(7), 644–52. [Google Scholar] [CrossRef]
- Hatt, Sebastian A; Grace, Olwen M; Zuntini, Alexandre R; Cameron, Duncan D; Thorogood, Chris J. Parasitic Plants Show Striking Convergence in Host Preference across Angiosperm Lineages. Annals of Botany 2025, 135(6), 1135–46. [Google Scholar] [CrossRef]
- Hatt, Sebastian A; Thorogood, Chris J; Bolin, Jay F; Musselman, Lytton J; Cameron, Duncan D; Grace, Olwen M. A Taxonomic Revision of the Genus Hydnora (Hydnoraceae) 2022.
- Jin, Jian-Jun; Yu, Wen-Bin; Yang, Jun-Bo; et al. GetOrganelle: A Fast and Versatile Toolkit for Accurate de Novo Assembly of Organelle Genomes. Genome Biology 2020, 21(1), 241. [Google Scholar] [CrossRef]
- Larsson, Anders. AliView: A Fast and Lightweight Alignment Viewer and Editor for Large Datasets. Bioinformatics 2014, 30(22), 3276–78. [Google Scholar] [CrossRef]
- Letunic, Ivica; Bork, Peer. Interactive Tree of Life (iTOL) v6: Recent Updates to the Phylogenetic Tree Display and Annotation Tool. Nucleic Acids Research 2024, 52(W1), W78–82. [Google Scholar] [CrossRef]
- Martel, Carlos; Fernandez-Hilario, Robin; TELLO, JUAN; Arteaga, Robert; Gerlach, Günter. Prosopanche Panguanensis (Aristolochiaceae), a New Species from Central Peru. Phytotaxa 2018, 364 (August), 241. [Google Scholar] [CrossRef]
- Mower, Jeffrey P; Stefanović, Saša; Hao, Weilong; et al. Horizontal Acquisition of Multiple Mitochondrial Genes from a Parasitic Plant Followed by Gene Conversion with Host Mitochondrial Genes. BMC Biology 2010, 8(1), 150. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, Lam-Tung; Schmidt, Heiko A.; Von Haeseler, Arndt; Minh, Bui Quang. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Molecular Biology and Evolution 2015, 32(1), 268–74. [Google Scholar] [CrossRef] [PubMed]
- Nickrent, Daniel L. Parasitic Angiosperms: How Often and How Many? TAXON 2020, 69(1), 5–27. [Google Scholar] [CrossRef]
- Phanstiel, Douglas H.; Boyle, Alan P.; Araya, Carlos L.; Snyder, Michael P. Sushi.R: Flexible, Quantitative and Integrative Genomic Visualizations for Publication-Quality Multi-Panel Figures. Bioinformatics 2014, 30(19), 2808–10. [Google Scholar] [CrossRef]
- Potter, Simon C; Luciani, Aurélien; Eddy, Sean R; Park, Youngmi; Lopez, Rodrigo; Finn, Robert D. HMMER Web Server: 2018 Update. Nucleic Acids Research 2018, 46(W1), W200–204. [Google Scholar] [CrossRef]
- Richardson, A. O.; Palmer, J. D. Horizontal Gene Transfer in Plants. Journal of Experimental Botany 2006, 58(1), 1–9. [Google Scholar] [CrossRef]
- Roulet, M. Emilia; Garcia, Laura E.; Gandini, Carolina L.; Sato, Hector; Ponce, Gabriela; Sanchez-Puerta, M. Virginia. Multichromosomal Structure and Foreign Tracts in the Ombrophytum Subterraneum (Balanophoraceae) Mitochondrial Genome. Plant Molecular Biology 2020, 103(6), 623–38. [Google Scholar] [CrossRef]
- Sanchez-Puerta, M Virginia; Ceriotti, Luis F; Gatica-Soria, Leonardo M; Roulet, M Emilia; Garcia, Laura E; Sato, Hector A. Invited Review Beyond Parasitic Convergence: Unravelling the Evolution of the Organellar Genomes in Holoparasites. Annals of Botany 2023, 132(5), 909–28. [Google Scholar] [CrossRef]
- Sanchez-Puerta, M. Virginia; Edera, Alejandro; Gandini, Carolina L.; et al. Genome-Scale Transfer of Mitochondrial DNA from Legume Hosts to the Holoparasite Lophophytum Mirabile (Balanophoraceae). Molecular Phylogenetics and Evolution 2019, 132 (March), 243–50. [Google Scholar] [CrossRef]
- Sanchez-Puerta, M. Virginia; García, Laura E.; Wohlfeiler, Josefina; Ceriotti, Luis F. Unparalleled Replacement of Native Mitochondrial Genes by Foreign Homologs in a Holoparasitic Plant. The New Phytologist (England) 2017, 214(1), 376–87. [Google Scholar] [CrossRef] [PubMed]
- Sequeira, Andrea; Rocamundi, Nicolás; Ferrer, M.; Baranzelli, Matias; Marvaldi, Adriana. Unveiling the History of a Peculiar Weevil-Plant Interaction in South America: A Phylogeographic Approach to Hydnorobius Hydnorae (Belidae) Associated with Prosopanche Americana (Aristolochiaceae). Diversity 2018, 10(2), 33. [Google Scholar] [CrossRef]
- Steenwyk, Jacob L.; Buida, Thomas J.; Li, Yuanning; Shen, Xing-Xing; Rokas, Antonis. ClipKIT: A Multiple Sequence Alignment Trimming Software for Accurate Phylogenomic Inference. PLOS Biology 2020, 18(12), e3001007. [Google Scholar] [CrossRef] [PubMed]
- Sun, Guiling; Xu, Yuxing; Liu, Hui; et al. Large-Scale Gene Losses Underlie the Genome Evolution of Parasitic Plant Cuscuta Australis. Nature Communications 2018, 9(1), 2683. [Google Scholar] [CrossRef]
- Svetlikova, Petra; Su, Huei-Jiun; Suetsugu, Kenji; Husnik, Filip. Phylogenomics Clarifies Balanophora Evolution, Metabolic Retention in Reduced Plastids, and the Origins of Obligate Agamospermy. New Phytologist 2025, nph.70761. [Google Scholar] [CrossRef]
- Teixeira-Costa, Luiza. A Living Bridge between Two Enemies: Haustorium Structure and Evolution across Parasitic Flowering Plants. Brazilian Journal of Botany 2021, 44(1), 165–78. [Google Scholar] [CrossRef]
- Vogel, Alexander; Schwacke, Rainer; Denton, Alisandra K.; et al. Footprints of Parasitism in the Genome of the Parasitic Flowering Plant Cuscuta Campestris. Nature Communications 2018, 9(1), 2515. [Google Scholar] [CrossRef]
- Waterhouse, Robert M.; Seppey, Mathieu; Simao, Felipe A.; et al. BUSCO Applications from Quality Assessments to Gene Prediction and Phylogenomics. Molecular Biology and Evolution 2018, 35(3), 543–48. [Google Scholar] [CrossRef]
- Wick, Ryan R.; Schultz, Mark B.; Zobel, Justin; Holt, Kathryn E. Bandage: Interactive Visualization of de Novo Genome Assemblies. Bioinformatics 2015, 31(20), 3350–52. [Google Scholar] [CrossRef]
- Xi, Zhenxiang; Wang, Yuguo; Bradley, Robert K.; et al. Massive Mitochondrial Gene Transfer in a Parasitic Flowering Plant Clade. PLoS Genetics 2013, 9(2), e1003265. [Google Scholar] [CrossRef]
- Yang, Zhenzhen; Wafula, Eric K.; Kim, Gunjune; et al. Convergent Horizontal Gene Transfer and Cross-Talk of Mobile Nucleic Acids in Parasitic Plants. Nature Plants 2019, 5(9), 991–1001. [Google Scholar] [CrossRef] [PubMed]
- Yang, Zhenzhen; Zhang, Yeting; Wafula, Eric K.; et al. Horizontal Gene Transfer Is More Frequent with Increased Heterotrophy and Contributes to Parasite Adaptation. Proceedings of the National Academy of Sciences 2016, 113(45). [Google Scholar] [CrossRef]
- Yu, Runxian; Chen, Xudong; Long, Lingjie; et al. De Novo Assembly and Comparative Analyses of Mitochondrial Genomes in Piperales. Genome Biology and Evolution 2023, 15(3), evad041. [Google Scholar] [CrossRef]
- Yu, Runxian; Sun, Chenyu; Zhong, Yan; et al. The Minicircular and Extremely Heteroplasmic Mitogenome of the Holoparasitic Plant Rhopalocnemis Phalloides. Current Biology 2022, 32(2), 470–479.e5. [Google Scholar] [CrossRef]
- Zeng, Ying; Yang, Tao. RNA Isolation from Highly Viscous Samples Rich in Polyphenols and Polysaccharides. Plant Molecular Biology Reporter 2002, 20(4), 417–417. [Google Scholar] [CrossRef]
- Zou, Rong; Huang, Jian; Xie, Hong; et al. A Chromosome-Level Genome Assembly of Cistanche Deserticola Provides Insights into Its Evolution and Molecular Mechanisms of Parasitism. Plant Communications 2026, 7(1), 101581. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



