Submitted:
09 February 2026
Posted:
10 February 2026
You are already at the latest version
Abstract
Keywords:
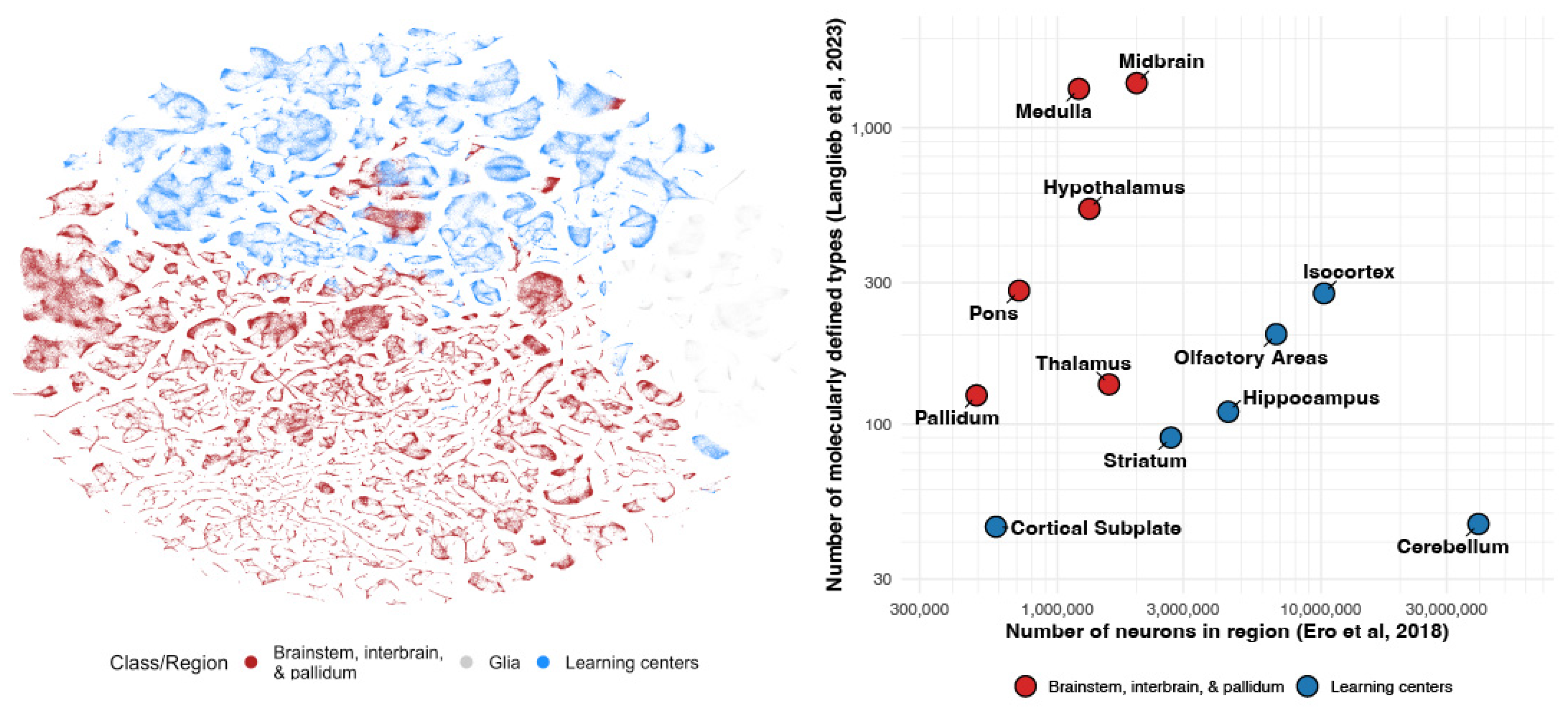
1. High Cellular Complexity in Deep Brain Structures
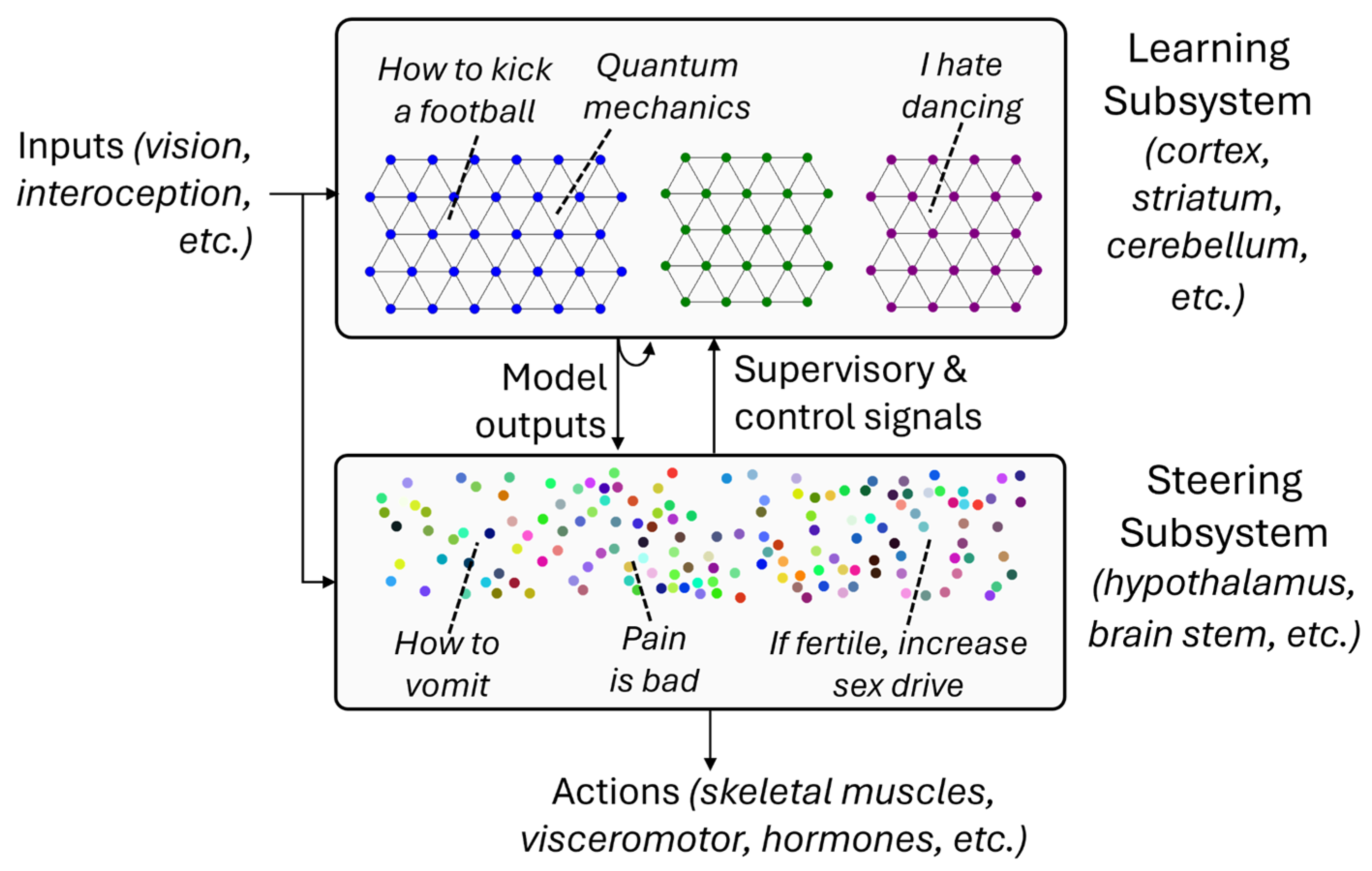
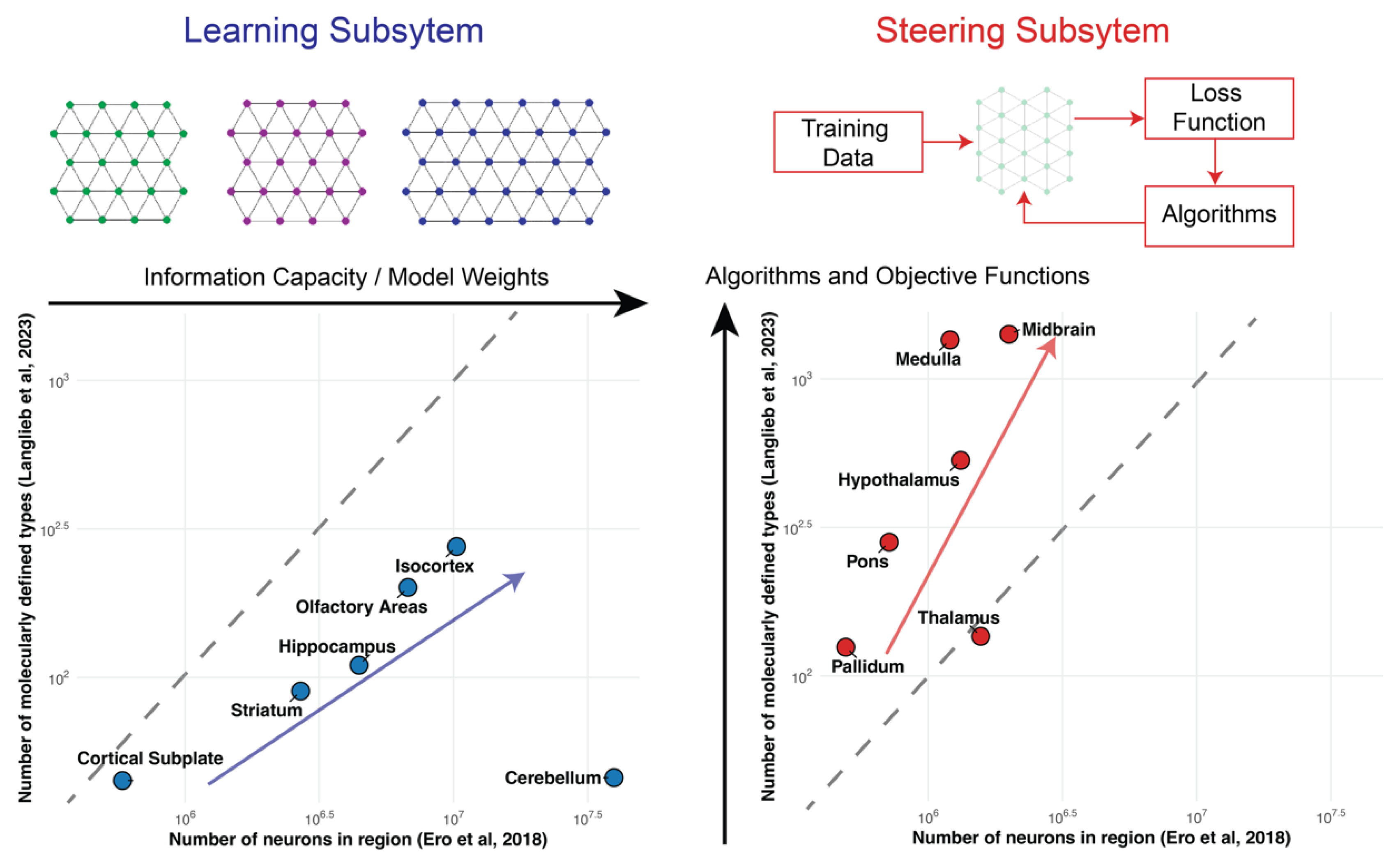
2. The Learning Subsystem: Scaling via Replication
3. The Steering Subsystem: Scaling via Diversification
The Steering Subsystem as a Biological Reward Function
4. Comparisons with Existing Frameworks
5. Maladaptation: When Fixed Drives Meet Changing Environments
6. Architectural Vulnerabilities: Lessons for Neuropsychiatric and Neurodegenerative Disease
7. Conclusion: Lessons of the Steering Subsystem for Neuroscience and AI Design
Author Contributions
Acknowledgments
Competing Interests
References
- Ramón y Cajal, S. (1909). Histologie du système nerveux de l’homme & des vertébrés (Maloine).
- Masland, R.H. The fundamental plan of the retina. Nat. Neurosci. 2001, 4, 877–886. [Google Scholar] [CrossRef]
- Masland, R.H.; Sanes, J.R. Retinal ganglion cell types: Current states and lessons for the brain. Annu. Rev. Neurosci. in press. 2015. [Google Scholar]
- Kim, I.J.; Zhang, Y.; Yamagata, M.; Meister, M.; Sanes, J.R. Molecular identification of a retinal cell type that responds to upward motion. Nature 2008, 452, 478–482. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Kim, I.-J.; Sanes, J.R.; Meister, M. The most numerous ganglion cell type of the mouse retina is a selective feature detector. Proc. Natl. Acad. Sci. USA 2012, 109, E2391–E2398. [Google Scholar] [CrossRef]
- Münch, T.A.; da Silveira, R.A.; Siegert, S.; Viney, T.J.; Awatramani, G.B.; Roska, B. Approach sensitivity in the retina processed by a multifunctional neural circuit. Nat. Neurosci. 2009, 12, 1308–1316. [Google Scholar] [CrossRef] [PubMed]
- Masland, R.H. Neuronal cell types. Curr. Biol. 2004, 14, R497–R500. [Google Scholar] [CrossRef]
- Kawaguchi, Y.; Kubota, Y. GABAergic cell subtypes and their synaptic connections in rat frontal cortex. Cereb. Cortex 1997, 7, 476–486. [Google Scholar] [CrossRef]
- Gupta, A.; Wang, Y.; Markram, H. Organizing principles for a diversity of GABAergic interneurons and synapses in the neocortex. Science 2000, 287, 273–278. [Google Scholar] [CrossRef]
- Petilla Interneuron Nomenclature Group; Ascoli, G.A.; Alonso-Nanclares, L.; Anderson, S.A.; Barrionuevo, G.; Benavides-Piccione, R.; Burkhalter, A.; Buzsaki, G.; Cauli, B.; Defelipe, J.; et al. Petilla terminology: Nomenclature of features of GABAergic interneurons of the cerebral cortex. Nat. Rev. Neurosci. 2008, 9, 557–568. [Google Scholar] [CrossRef]
- Dong, H.W.; Petrovich, G.D.; Swanson, L.W. Topography of projections from amygdala to bed nuclei of the stria terminalis. Brain Res. Brain Res. Rev. 2001, 38, 192–246. [Google Scholar] [CrossRef]
- Dong, H.-W.; Swanson, L.W. Organization of axonal projections from the anterolateral area of the bed nuclei of the stria terminalis. J. Comp. Neurol. 2004, 468, 277–298. [Google Scholar] [CrossRef]
- Swanson, L.W. Cerebral hemisphere regulation of motivated behavior11Published on the World Wide Web on 2 November 2000. Brain Res. 2000, 886, 113–164. [Google Scholar] [CrossRef]
- Zeng, H.; Sanes, J.R. Neuronal cell-type classification: Challenges, opportunities and the path forward. Nat. Rev. Neurosci. 2017, 18, 530–546. [Google Scholar] [CrossRef]
- Nelson, S.B.; Sugino, K.; Hempel, C.M. The problem of neuronal cell types: A physiological genomics approach. Trends Neurosci. 2006, 29, 339–345. [Google Scholar] [CrossRef] [PubMed]
- Zeisel, A.; Muñoz-Manchado, A.B.; Codeluppi, S.; Lönnerberg, P.; La Manno, G.; Juréus, A.; Marques, S.; Munguba, H.; He, L.; Betsholtz, C.; et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 2015, 347, 1138–1142. [Google Scholar] [CrossRef]
- Macosko, E.Z.; Basu, A.; Satija, R.; Nemesh, J.; Shekhar, K.; Goldman, M.; Tirosh, I.; Bialas, A.R.; Kamitaki, N.; Martersteck, E.M.; et al. Highly Parallel Genome-wide Expression Profiling of Individual Cells Using Nanoliter Droplets. Cell 2015, 161, 1202–1214. [Google Scholar] [CrossRef]
- Klein, A.M.; Mazutis, L.; Akartuna, I.; Tallapragada, N.; Veres, A.; Li, V.; Peshkin, L.; Weitz, D.A.; Kirschner, M.W. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 2015, 161, 1187–1201. [Google Scholar] [CrossRef]
- Habib, N.; Avraham-Davidi, I.; Basu, A.; Burks, T.; Shekhar, K.; Hofree, M.; Choudhury, S.R.; Aguet, F.; Gelfand, E.; Ardlie, K.; et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 2017, 14, 955–958. [Google Scholar] [CrossRef]
- Chen, K.H.; Boettiger, A.N.; Moffitt, J.R.; Wang, S.; Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 2015, 348, aaa6090. [Google Scholar] [CrossRef] [PubMed]
- Moffitt, J.R.; Bambah-Mukku, D.; Eichhorn, S.W.; Vaughn, E.; Shekhar, K.; Perez, J.D.; Rubinstein, N.D.; Hao, J.; Regev, A.; Dulac, C.; et al. Molecular, spatial, and functional single-cell profiling of the hypothalamic preoptic region. Science 2018, 362, eaau5324. [Google Scholar] [CrossRef] [PubMed]
- Rodriques, S.G.; Stickels, R.R.; Goeva, A.; Martin, C.A.; Murray, E.; Vanderburg, C.R.; Welch, J.; Chen, L.M.; Chen, F.; Macosko, E.Z. Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution. Science 2019, 363, 1463–1467. [Google Scholar] [CrossRef]
- Ståhl, P.L.; Salmén, F.; Vickovic, S.; Lundmark, A.; Navarro, J.F.; Magnusson, J.; Giacomello, S.; Asp, M.; Westholm, J.O.; Huss, M.; et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 2016, 353, 78–82. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; He, Y.; Zhou, Y.; Huang, J.; Maher, K.; Wang, B.; Tang, Z.; Luo, S.; Tan, P.; Wu, M.; et al. Spatial atlas of the mouse central nervous system at molecular resolution. Nature 2023, 622, 552–561. [Google Scholar] [CrossRef] [PubMed]
- Langlieb, J.; Sachdev, N.S.; Balderrama, K.S.; Nadaf, N.M.; Raj, M.; Murray, E.; Webber, J.T.; Vanderburg, C.; Gazestani, V.; Tward, D.; et al. The molecular cytoarchitecture of the adult mouse brain. Nature 2023, 624, 333–342. [Google Scholar] [CrossRef]
- Yao, Z.; van Velthoven, C.T.J.; Kunst, M.; Zhang, M.; McMillen, D.; Lee, C.; Jung, W.; Goldy, J.; Abdelhak, A.; Aitken, M.; et al. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 2023, 624, 317–332. [Google Scholar] [CrossRef]
- Zhang, M.; Pan, X.; Jung, W.; Halpern, A.R.; Eichhorn, S.W.; Lei, Z.; Cohen, L.; Smith, K.A.; Tasic, B.; Yao, Z.; et al. Molecularly defined and spatially resolved cell atlas of the whole mouse brain. Nature 2023, 624, 343–354. [Google Scholar] [CrossRef]
- Erö, C.; Gewaltig, M.-O.; Keller, D.; Markram, H. A cell atlas for the mouse brain. Front. Neuroinform 2018, 12, 84. [Google Scholar] [CrossRef]
- Campbell, J.N.; Macosko, E.Z.; Fenselau, H.; Pers, T.H.; Lyubetskaya, A.; Tenen, D.; Goldman, M.; Verstegen, A.M.J.; Resch, J.M.; McCarroll, S.A.; et al. A molecular census of arcuate hypothalamus and median eminence cell types. Nat. Neurosci. 2017, 20, 484–496. [Google Scholar] [CrossRef]
- Chen, R.; Wu, X.; Jiang, L.; Zhang, Y. Single-Cell RNA-Seq Reveals Hypothalamic Cell Diversity. Cell Rep. 2017, 18, 3227–3241. [Google Scholar] [CrossRef] [PubMed]
- Byrnes, S.J. Intro to brain-like-AGI safety. 2025. [Google Scholar] [CrossRef]
- Marblestone, A.H.; Wayne, G.; Kording, K.P. Toward an integration of deep learning and neuroscience. Front. Comput. Neurosci. 2016, 10, 94. [Google Scholar] [CrossRef]
- Kaplan, J.; McCandlish, S.; Henighan, T.; Brown, T.B.; Chess, B.; Child, R.; Gray, S.; Radford, A.; Wu, J.; Amodei, D. Scaling laws for neural language models. arXiv 2020, arXiv:2001.08361. [Google Scholar] [CrossRef]
- Mora-Bermúdez, F.; Badsha, F.; Kanton, S.; Camp, J.G.; Vernot, B.; Köhler, K.; Voigt, B.; Okita, K.; Maricic, T.; He, Z.; et al. Differences and similarities between human and chimpanzee neural progenitors during cerebral cortex development. eLife 2016, 5, e18683. [Google Scholar] [CrossRef]
- Herculano-Houzel, S. The remarkable, yet not extraordinary, human brain as a scaled-up primate brain and its associated cost. Proc. Natl. Acad. Sci. USA 2012, 109, 10661–10668. [Google Scholar] [CrossRef]
- Florio, M.; Albert, M.; Taverna, E.; Namba, T.; Brandl, H.; Lewitus, E.; Haffner, C.; Sykes, A.; Wong, F.K.; Peters, J.; et al. Human-specific gene ARHGAP11B promotes basal progenitor amplification and neocortex expansion. Science 2015, 347, 1465–1470. [Google Scholar] [CrossRef]
- Hansen, D.V.; Lui, J.H.; Parker, P.R.L.; Kriegstein, A.R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 2010, 464, 554–561. [Google Scholar] [CrossRef] [PubMed]
- Nowakowski, T.J.; Bhaduri, A.; Pollen, A.A.; Alvarado, B.; Mostajo-Radji, M.A.; Di Lullo, E.; Haeussler, M.; Sandoval-Espinosa, C.; Liu, S.J.; Velmeshev, D.; et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 2017, 358, 1318–1323. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, M.T.; Sandoval, K.; Chen, C.P.; Mostajo-Radji, M.A.; Seeley, W.W.; Nowakowski, T.J.; Ye, C.J.; Paredes, M.F.; Pollen, A.A. The development and evolution of inhibitory neurons in primate cerebrum. Nature 2022, 603, 871–877. [Google Scholar] [CrossRef]
- Nascimento, M.A.; Biagiotti, S.; Herranz-Pérez, V.; Santiago, S.; Bueno, R.; Ye, C.J.; Abel, T.J.; Zhang, Z.; Rubio-Moll, J.S.; Kriegstein, A.R.; et al. Protracted neuronal recruitment in the temporal lobes of young children. Nature 2024, 626, 1056–1065. [Google Scholar] [CrossRef] [PubMed]
- Jean, A. Brain stem control of swallowing: Neuronal network and cellular mechanisms. Physiol. Rev. 2001, 81, 929–969. [Google Scholar] [CrossRef]
- Kamath, T.; Abdulraouf, A.; Burris, S.J.; Langlieb, J.; Gazestani, V.; Nadaf, N.M.; Balderrama, K.; Vanderburg, C.; Macosko, E.Z. Single-cell genomic profiling of human dopamine neurons identifies a population that selectively degenerates in Parkinson’s disease. Nat. Neurosci. 2022, 25, 588–595. [Google Scholar] [CrossRef]
- Silver, D.; Hubert, T.; Schrittwieser, J.; Antonoglou, I.; Lai, M.; Guez, A.; Lanctot, M.; Sifre, L.; Kumaran, D.; Graepel, T.; et al. Mastering chess and shogi by self-play with a general reinforcement learning algorithm. arXiv 2017, arXiv:1712.01815. [Google Scholar] [CrossRef]
- Oka, Y.; Ye, M.; Zuker, C.S. Thirst driving and suppressing signals encoded by distinct neural populations in the brain. Nature 2015, 520, 349–352. [Google Scholar] [CrossRef]
- Chang, R.B.; Strochlic, D.E.; Williams, E.K.; Umans, B.D.; Liberles, S.D. Vagal sensory neuron subtypes that differentially control breathing. Cell 2015, 161, 622–633. [Google Scholar] [CrossRef]
- Dong, X.; Dong, X. Peripheral and central mechanisms of itch. Neuron 2018, 98, 482–494. [Google Scholar] [CrossRef]
- Zhang, C.; Kaye, J.A.; Cai, Z.; Wang, Y.; Prescott, S.L.; Liberles, S.D. Area Postrema Cell Types that Mediate Nausea-Associated Behaviors. Neuron 2021, 109, 461–472.e5. [Google Scholar] [CrossRef]
- Matthews, G.A.; Nieh, E.H.; Vander Weele, C.M.; Halbert, S.A.; Pradhan, R.V.; Yosafat, A.S.; Glober, G.F.; Izadmehr, E.M.; Thomas, R.E.; Lacy, G.D.; et al. Dorsal Raphe Dopamine Neurons Represent the Experience of Social Isolation. Cell 2016, 164, 617–631. [Google Scholar] [CrossRef] [PubMed]
- Bejjani, B.P.; Damier, P.; Arnulf, I.; Thivard, L.; Bonnet, A.M.; Dormont, D.; Cornu, P.; Pidoux, B.; Samson, Y.; Agid, Y. Transient acute depression induced by high-frequency deep-brain stimulation. N. Engl. J. Med. 1999, 340, 1476–1480. [Google Scholar] [CrossRef] [PubMed]
- Parvizi, J.; Veit, M.J.; Barbosa, D.A.N.; Kucyi, A.; Perry, C.; Parker, J.J.; Shivacharan, R.S.; Chen, F.; Yih, J.; Gross, J.J.; et al. Complex negative emotions induced by electrical stimulation of the human hypothalamus. Brain Stimul. 2022, 15, 615–623. [Google Scholar] [CrossRef]
- Amstutz, D.; Michelis, J.P.; Debove, I.; Maradan-Gachet, M.E.; Lachenmayer, M.L.; Muellner, J.; Schwegler, K.; Krack, P. Reckless generosity, Parkinson’s disease and dopamine: A case series and literature review. Mov. Disord. Clin. Pract. 2021, 8, 469–473. [Google Scholar] [CrossRef] [PubMed]
- Hamani, C.; McAndrews, M.P.; Cohn, M.; Oh, M.; Zumsteg, D.; Shapiro, C.M.; Wennberg, R.A.; Lozano, A.M. Memory enhancement induced by hypothalamic/fornix deep brain stimulation. Ann. Neurol. 2008, 63, 119–123. [Google Scholar] [CrossRef]
- Koestler, A. (1989). The ghost in the machine (Arkana).
- MacLean, P. A Triune Concept of the Brain and Behaviour: Hincks Memorial Lectures; Boag, T.J., Campbell, D., Eds.; University of Toronto Press: Toronto, ON, Canada, 1973. [Google Scholar]
- Hawkins, J.; Blakeslee, S. (2004). On intelligence: How a new understanding of the brain will lead to the creation of truly intelligent machines (Times Books).
- Cesario, J.; Johnson, D.J.; Eisthen, H.L. Your brain is not an onion with a tiny reptile inside. Curr. Dir. Psychol. Sci. 2020, 29, 255–260. [Google Scholar] [CrossRef]
- Naulé, L.; Maione, L.; Kaiser, U.B. Puberty, A sensitive window of hypothalamic development and plasticity. Endocrinology 2021, 162, bqaa209. [Google Scholar] [CrossRef]
- Pinto, S.; Roseberry, A.G.; Liu, H.; Diano, S.; Shanabrough, M.; Cai, X.; Friedman, J.M.; Horvath, T.L. Rapid rewiring of arcuate nucleus feeding circuits by leptin. Science 2004, 304, 110–115. [Google Scholar] [CrossRef]
- Stagkourakis, S.; Spigolon, G.; Liu, G.; Anderson, D.J. Experience-dependent plasticity in an innate social behavior is mediated by hypothalamic LTP. Proc. Natl. Acad. Sci. USA 2020, 117, 25789–25799. [Google Scholar] [CrossRef]
- Post, E.; Forchhammer, M.C. Climate change reduces reproductive success of an Arctic herbivore through trophic mismatch. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 2369–2375. [Google Scholar] [CrossRef]
- Williams, G.C.; Nesse, R.M. The dawn of Darwinian medicine. Q. Rev. Biol. 1991, 66, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Pennisi, E. Evolution. Darwinian medicine’s drawn-out dawn. Science 2011, 334, 1486–1487. [Google Scholar] [CrossRef] [PubMed]
- Stevens, A.; Price, J. Evolutionary Psychiatry; Routledge: London, UK, 2021. [Google Scholar]
- Faulty Reward Functions in the Wild. Available online: https://www.google.com/url?q=https://openai.com/index/faulty-reward-functions/&sa=D&source=docs&ust=1765851859210892&usg=AOvVaw2ZiaDdn7hBDDLY6bmNRRAK.
- Mullins, N.; Forstner, A.J.; O’Connell, K.S.; Coombes, B.; Coleman, J.R.I.; Qiao, Z.; Als, T.D.; Bigdeli, T.B.; Børte, S.; Bryois, J.; et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat. Genet. 2021, 53, 817–829. [Google Scholar] [CrossRef] [PubMed]
- Trubetskoy, V.; Pardiñas, A.F.; Qi, T.; Panagiotaropoulou, G.; Awasthi, S.; Bigdeli, T.B.; Bryois, J.; Chen, C.-Y.; Dennison, C.A.; Hall, L.S.; et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 2022, 604, 502–508. [Google Scholar] [CrossRef]
- Skene, N.G.; Bryois, J.; Bakken, T.E.; Breen, G.; Crowley, J.J.; Gaspar, H.A.; Giusti-Rodriguez, P.; Hodge, R.D.; Miller, J.A.; Muñoz-Manchado, A.B.; et al. Genetic identification of brain cell types underlying schizophrenia. Nat. Genet. 2018, 50, 825–833. [Google Scholar] [CrossRef]
- Hendershot, C.S.; Bremmer, M.P.; Paladino, M.B.; Kostantinis, G.; Gilmore, T.A.; Sullivan, N.R.; Tow, A.C.; Dermody, S.S.; Prince, M.A.; Jordan, R.; et al. Once-weekly semaglutide in adults with alcohol use disorder: A randomized clinical trial. JAMA Psychiatry 2025, 82, 395–405. [Google Scholar] [CrossRef]
- Klausen, M.K.; Thomsen, M.; Wortwein, G.; Fink-Jensen, A. The role of glucagon-like peptide 1 (GLP-1) in addictive disorders. Br. J. Pharmacol. 2022, 179, 625–641. [Google Scholar] [CrossRef]
- Tuesta, L.M.; Chen, Z.; Duncan, A.; Fowler, C.D.; Ishikawa, M.; Lee, B.R.; Liu, X.-A.; Lu, Q.; Cameron, M.; Hayes, M.R.; et al. GLP-1 acts on habenular avoidance circuits to control nicotine intake. Nat. Neurosci. 2017, 20, 708–716. [Google Scholar] [CrossRef]
- Hernandez, N.S.; Weir, V.R.; Ragnini, K.; Merkel, R.; Zhang, Y.; Mace, K.; Rich, M.T.; Christopher Pierce, R.; Schmidt, H.D. GLP-1 receptor signaling in the laterodorsal tegmental nucleus attenuates cocaine seeking by activating GABAergic circuits that project to the VTA. Mol. Psychiatry 2021, 26, 4394–4408. [Google Scholar] [CrossRef] [PubMed]
- Vallöf, D.; Vestlund, J.; Jerlhag, E. Glucagon-like peptide-1 receptors within the nucleus of the solitary tract regulate alcohol-mediated behaviors in rodents. Neuropharmacology 2019, 149, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.D.; Mietlicki-Baase, E.G.; Ige, K.Y.; Maurer, J.J.; Reiner, D.J.; Zimmer, D.J.; Van Nest, D.S.; Guercio, L.A.; Wimmer, M.E.; Olivos, D.R.; et al. Glucagon-like peptide-1 receptor activation in the ventral tegmental area decreases the reinforcing efficacy of cocaine. Neuropsychopharmacology 2016, 41, 1917–1928. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, N.S.; O’Donovan, B.; Ortinski, P.I.; Schmidt, H.D. Activation of glucagon-like peptide-1 receptors in the nucleus accumbens attenuates cocaine seeking in rats. Addict. Biol. 2019, 24, 170–181. [Google Scholar] [CrossRef]
- Allingbjerg, M.-L.; Hansen, S.N.; Secher, A.; Thomsen, M. Glucagon-like peptide-1 receptors in nucleus accumbens, ventral hippocampus, and lateral septum reduce alcohol reinforcement in mice. Exp. Clin. Psychopharmacol. 2023, 31, 612–620. [Google Scholar] [CrossRef]
- La Joie, R.; Perrotin, A.; de La Sayette, V.; Egret, S.; Doeuvre, L.; Belliard, S.; Eustache, F.; Desgranges, B.; Chételat, G. Hippocampal subfield volumetry in mild cognitive impairment, Alzheimer’s disease and semantic dementia. NeuroImage Clin. 2013, 3, 155–162. [Google Scholar] [CrossRef]
- Greffard, S.; Verny, M.; Bonnet, A.-M.; Beinis, J.-Y.; Gallinari, C.; Meaume, S.; Piette, F.; Hauw, J.-J.; Duyckaerts, C. Motor score of the Unified Parkinson Disease Rating Scale as a good predictor of Lewy body-associated neuronal loss in the substantia nigra. Arch. Neurol. 2006, 63, 584–588. [Google Scholar] [CrossRef] [PubMed]
- Braak, H.; Thal, D.R.; Ghebremedhin, E.; Del Tredici, K. Stages of the pathologic process in Alzheimer disease: Age categories from 1 to 100 years. J. Neuropathol. Exp. Neurol. 2011, 70, 960–969. [Google Scholar] [CrossRef] [PubMed]
- Christiano, P.; Leike, J.; Brown, T.B.; Martic, M.; Legg, S.; Amodei, D. Deep reinforcement learning from human preferences. arXiv 2017, arXiv:1706.03741. [Google Scholar]
- Ouyang, L.; Wu, J.; Jiang, X.; Almeida, D.; Wainwright, C.L.; Mishkin, P.; Zhang, C.; Agarwal, S.; Slama, K.; Ray, A.; et al. Training language models to follow instructions with human feedback. arXiv 2022, arXiv:2203.02155. [Google Scholar] [CrossRef]
- Bai, Y.; Jones, A.; Ndousse, K.; Askell, A.; Chen, A.; DasSarma, N.; Drain, D.; Fort, S.; Ganguli, D.; Henighan, T.; et al. Training a helpful and harmless assistant with reinforcement learning from human feedback. arXiv 2022, arXiv:2204.05862. [Google Scholar] [CrossRef]
- Amodei, D.; Olah, C.; Steinhardt, J.; Christiano, P.; Schulman, J.; Mané, D. Concrete Problems in AI Safety. arXiv 2016, arXiv:1606.06565. [Google Scholar] [CrossRef]
- Leike, J.; Krueger, D.; Everitt, T.; Martic, M.; Maini, V.; Legg, S. Scalable agent alignment via reward modeling: A research direction. arXiv 2018, arXiv:1811.07871. [Google Scholar] [CrossRef]
- Kenton, Z.; Everitt, T.; Weidinger, L.; Gabriel, I.; Mikulik, V.; Irving, G. Alignment of language agents. arXiv 2021, arXiv:2103.14659. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



