Submitted:
06 February 2026
Posted:
09 February 2026
You are already at the latest version
Abstract
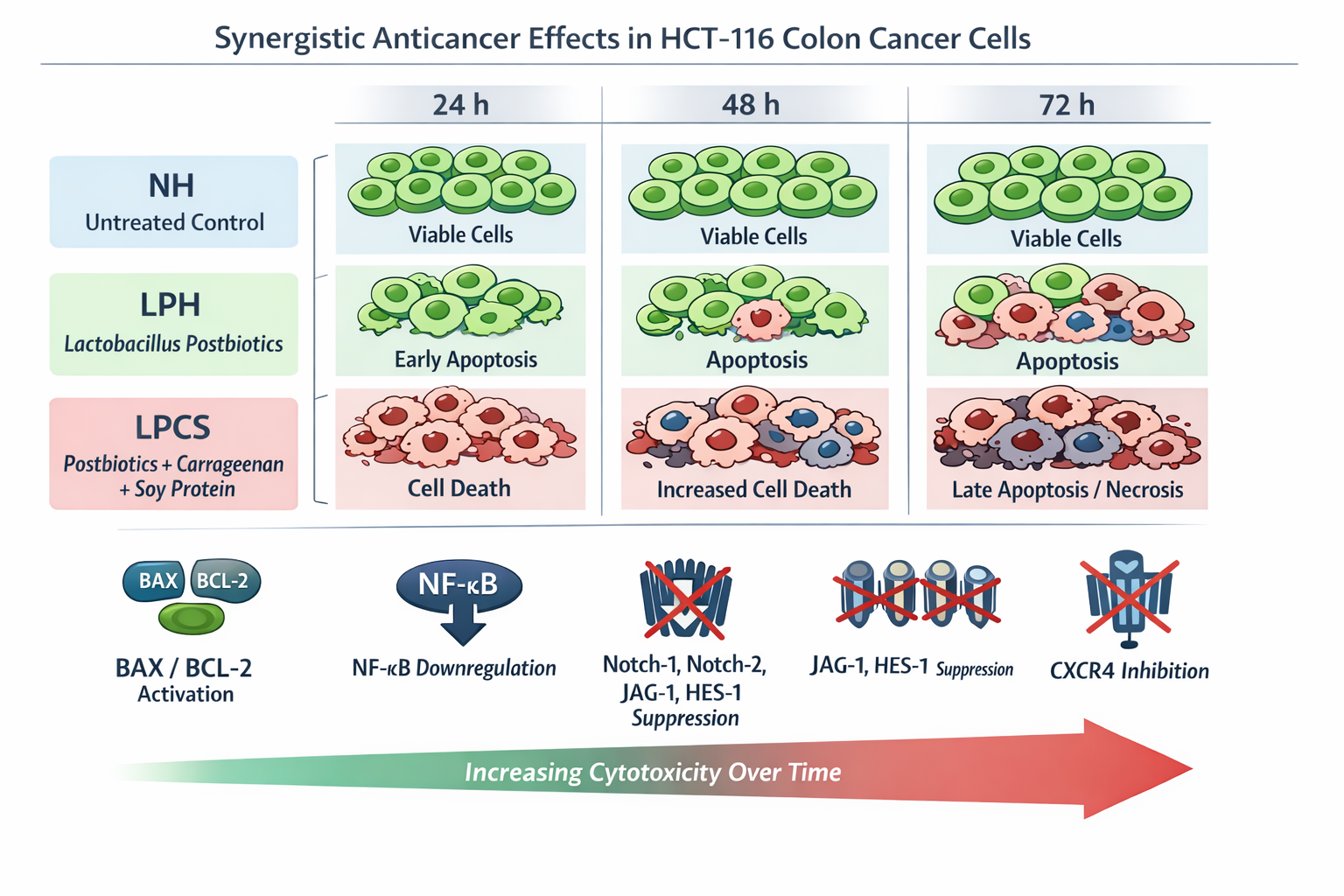
Background: Colorectal cancer remains a major cause of cancer-related mortality, highlighting the need for effective and low-toxicity therapeutic approaches. This study examined the time-dependent anticancer effects of Lactobacillus postbiotics alone and in combination with a carrageenan/soy protein blend in HCT-116 colorectal cancer cells. Methods: Cells were divided into three groups: untreated control (NH), postbiotics alone (LPH), and the combined formulation (LPCS). Cell viability, morphological alterations, apoptotic stages, gene expression, and temporal gene–gene correlations were analyzed at 24, 48, and 72 h. Results: The combined formulation induced the strongest and most sustained cytotoxic effects, with maximal late apoptosis and necrosis observed at 72 h. Postbiotics alone mainly triggered early apoptosis at 24 h, which diminished over time. Dynamic, time-dependent modulation of apoptotic regulators (BAX and BCL2), inflammatory signaling (NF- κB), and survival-related pathways (Notch-1, Notch-2, JAG-1, HES-1, and CXCR4) were observed. Combination treatment led to early stress responses followed by suppression of survival signaling and terminal disruption of apoptotic balance. Conclusion: These findings demonstrate that combining Lactobacillus postbiotics with carrageenan and soy protein enhances both the potency and durability of anticancer effects, supporting the development of multi-component postbiotic strategies for colorectal cancer.
Keywords:
1. Introduction
2. Results
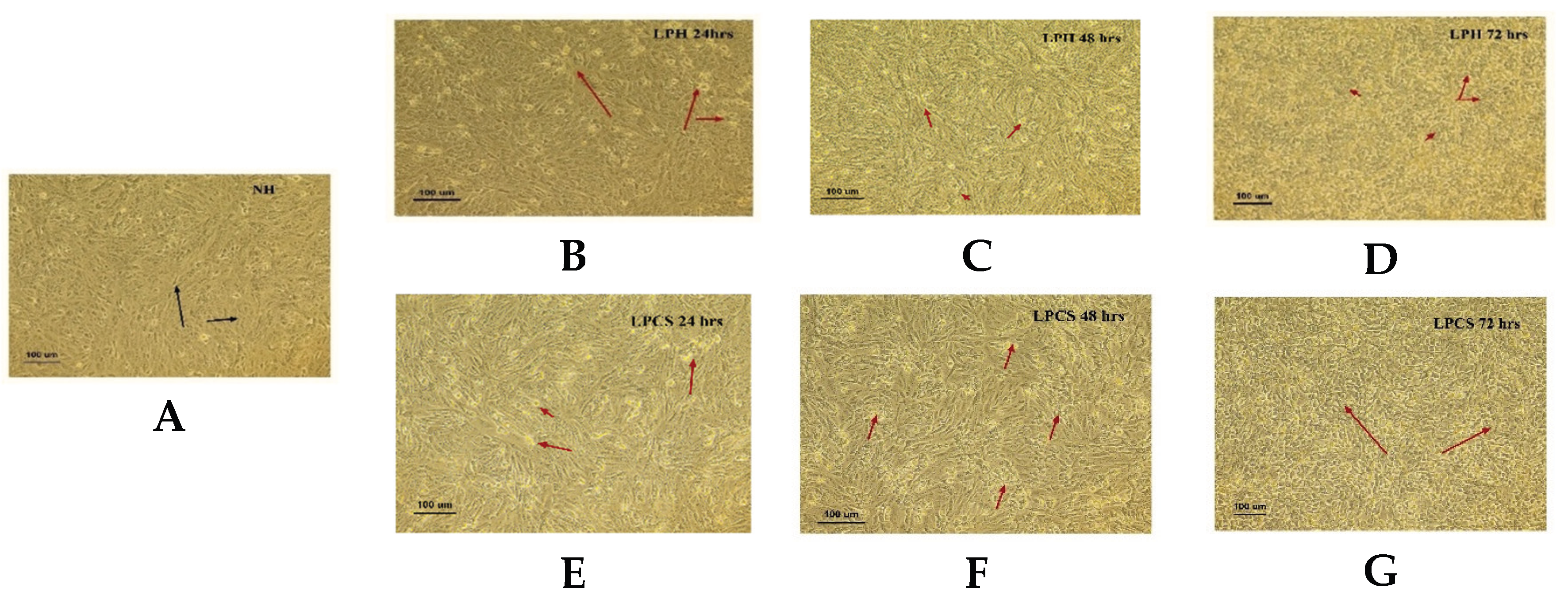
2.1. Morphological Evidence of Apoptosis in HCT Cells Following Postbiotic and Combination Treatments
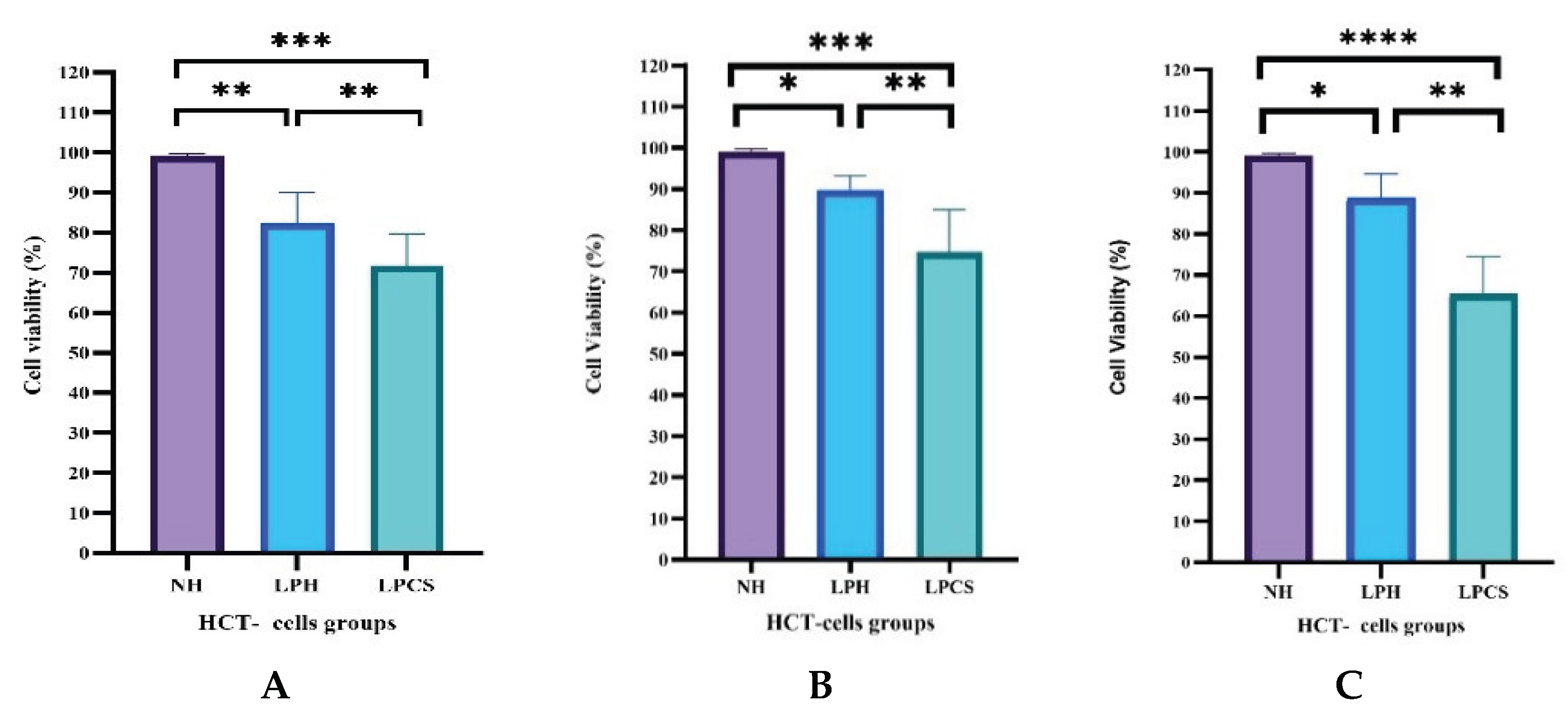
2.2. Time-Dependent Effects of Lactobacillus Postbiotics and Combination Therapy on HCT Cell Viability
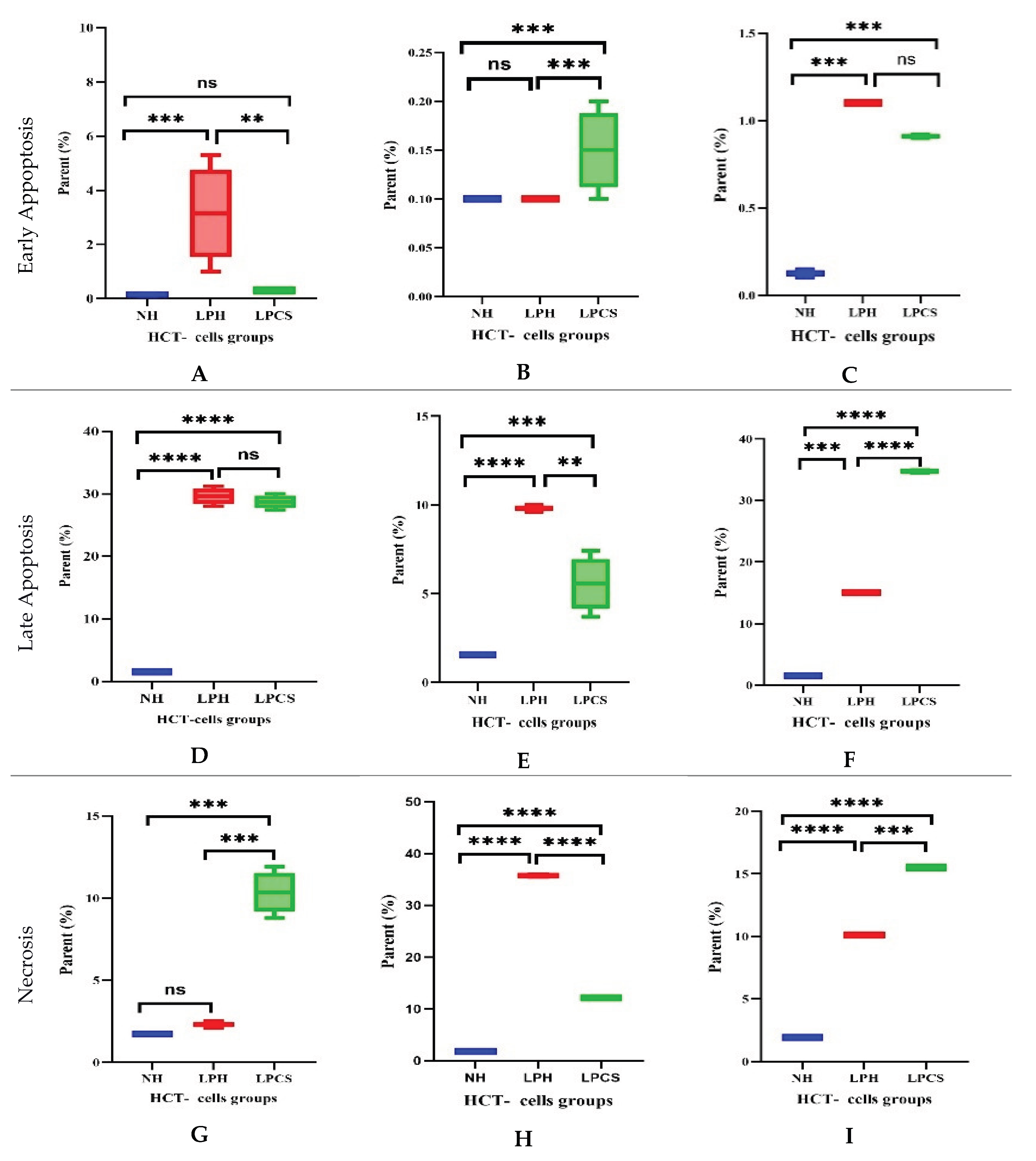
2.3. Time-Dependent Progression of Apoptosis and Necrosis in HCT Cells
2.4. Time-Dependent Apoptotic Gene Expression Responses in HCT Cells
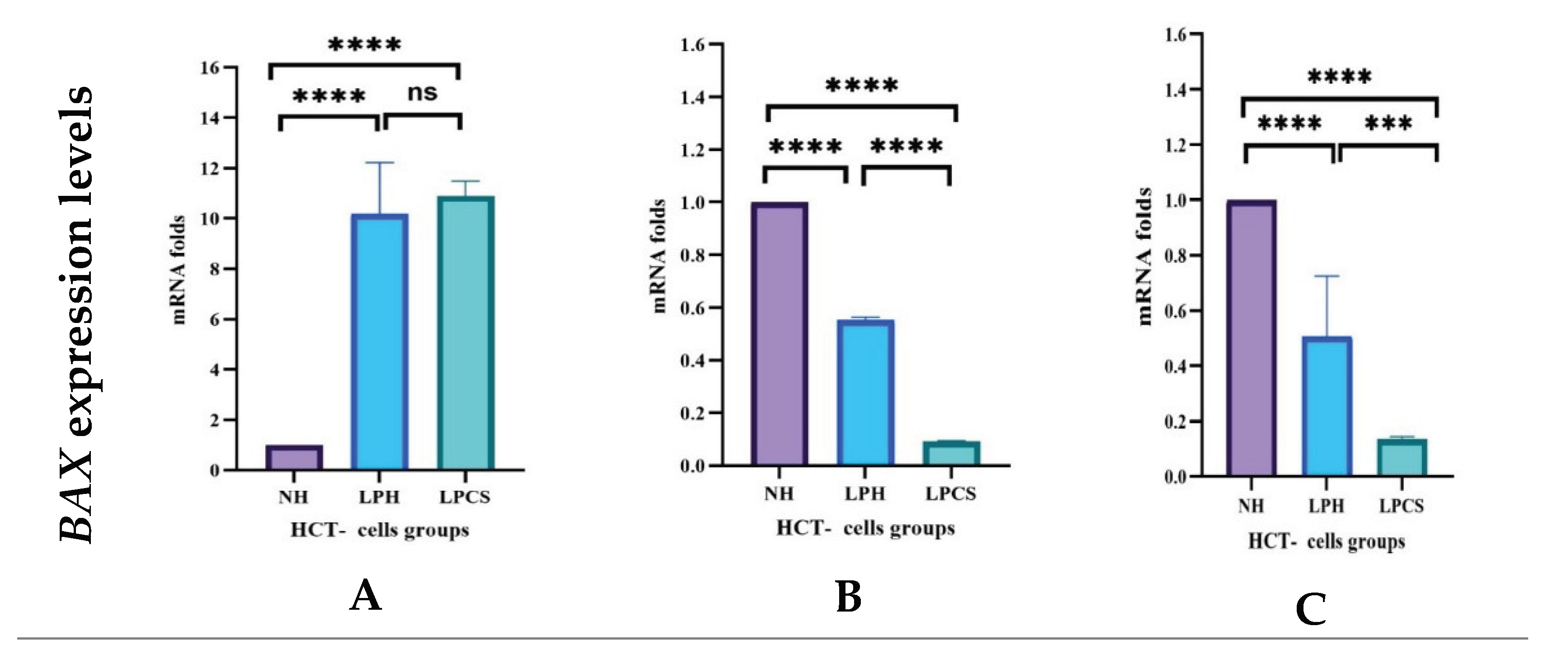
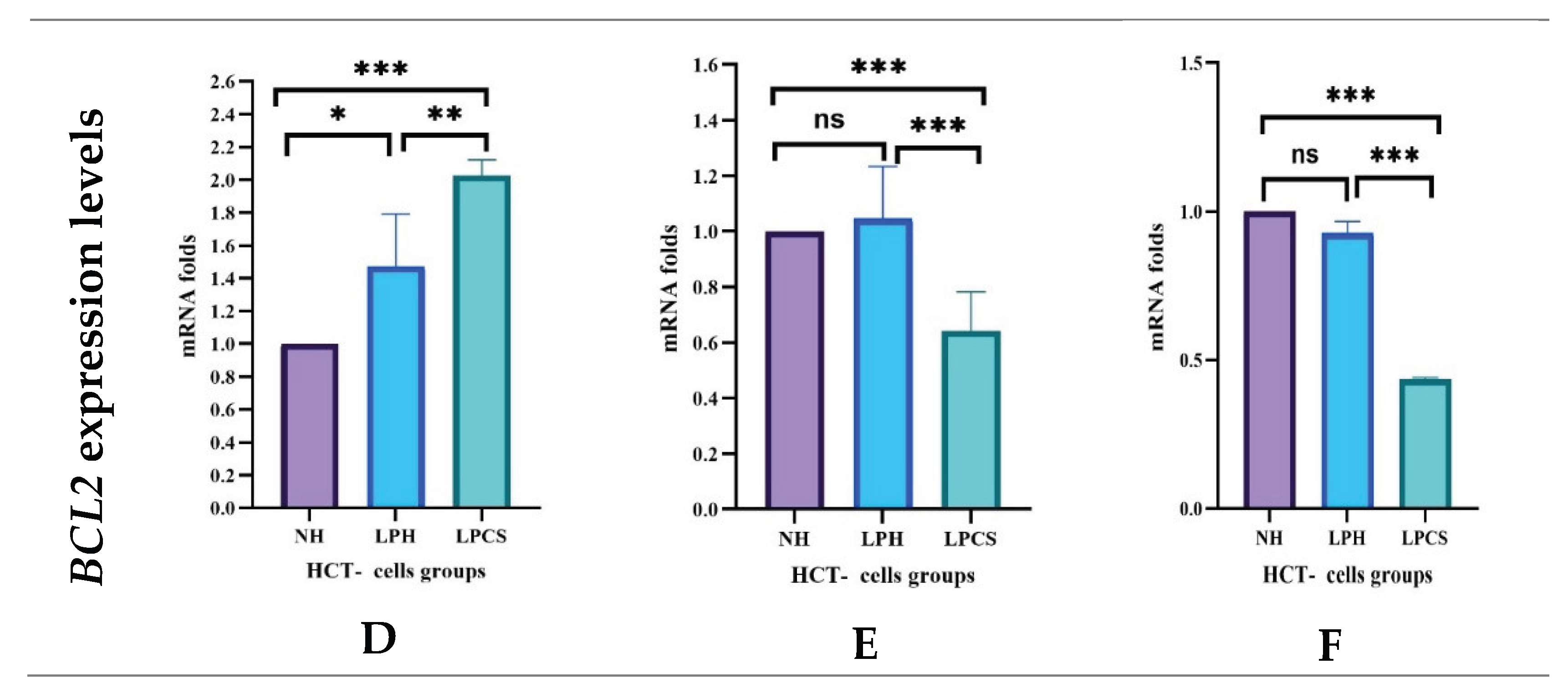
2.4.1. Levels of Apoptotic Regulatory Genes
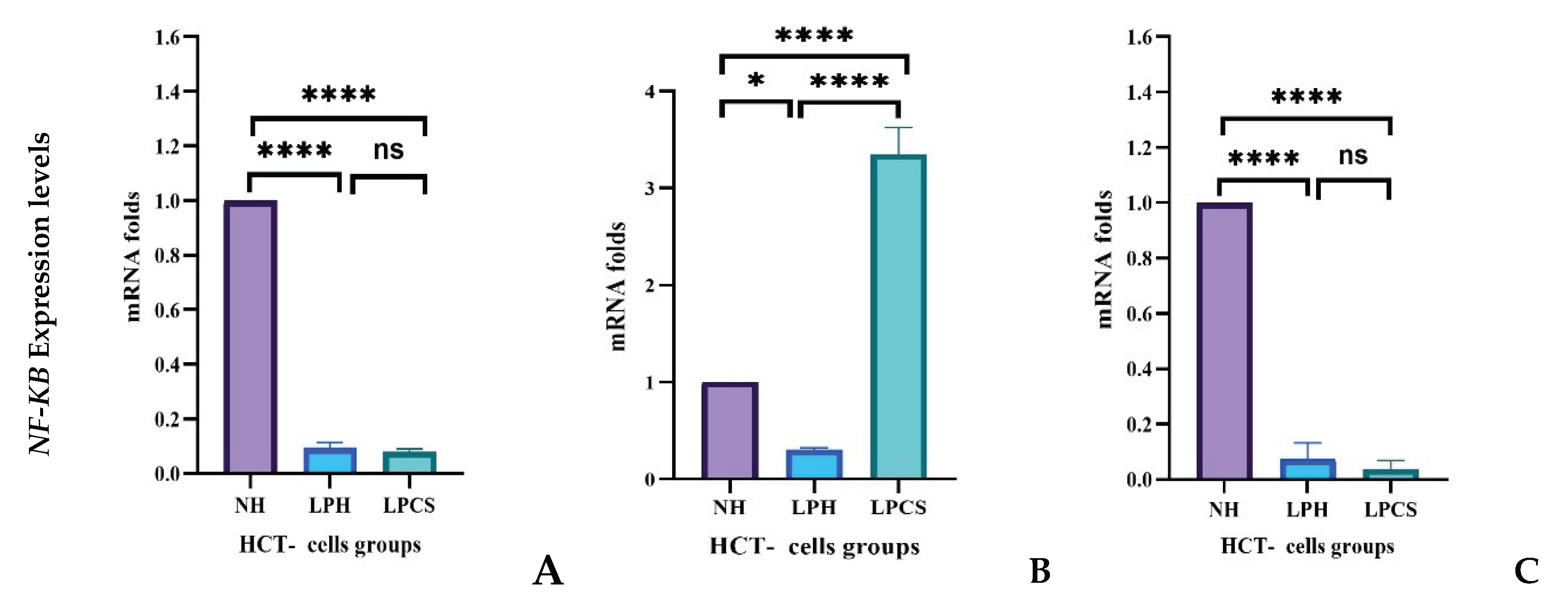
2.4.2. Levels of Inflammatory Gene Expression
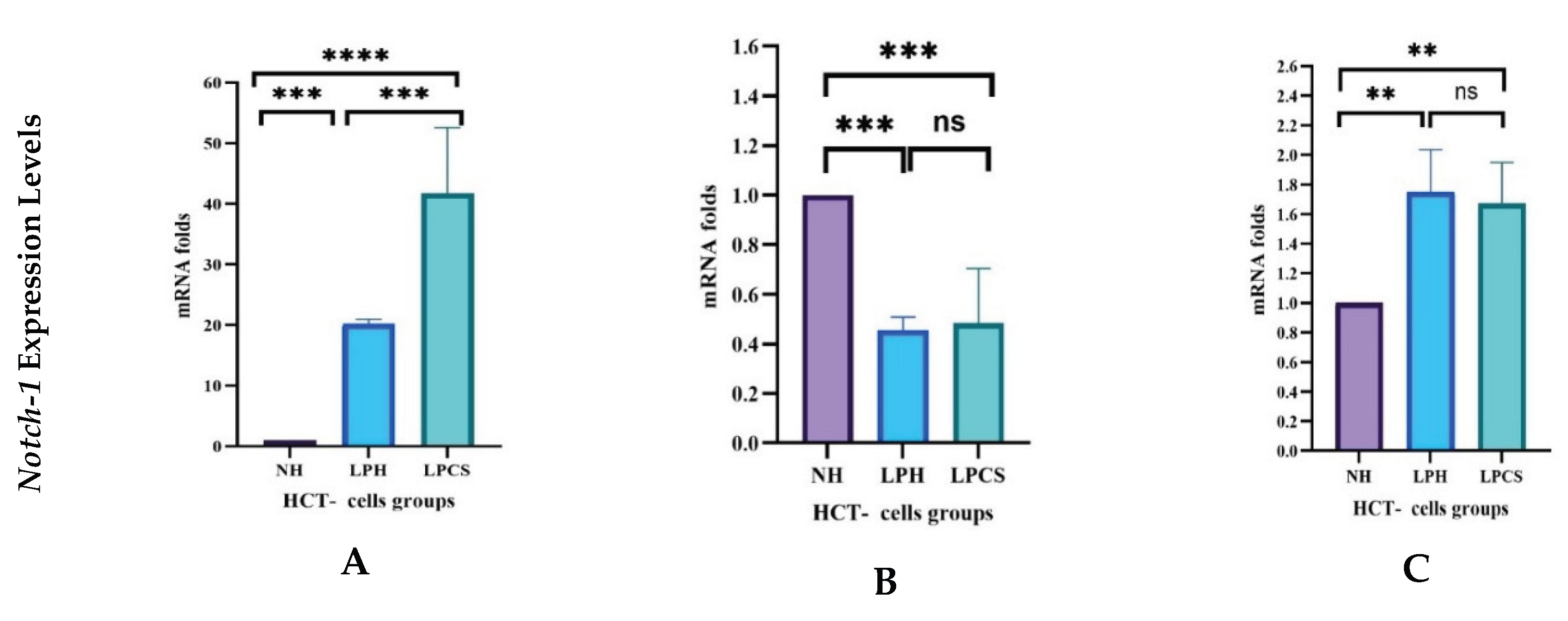
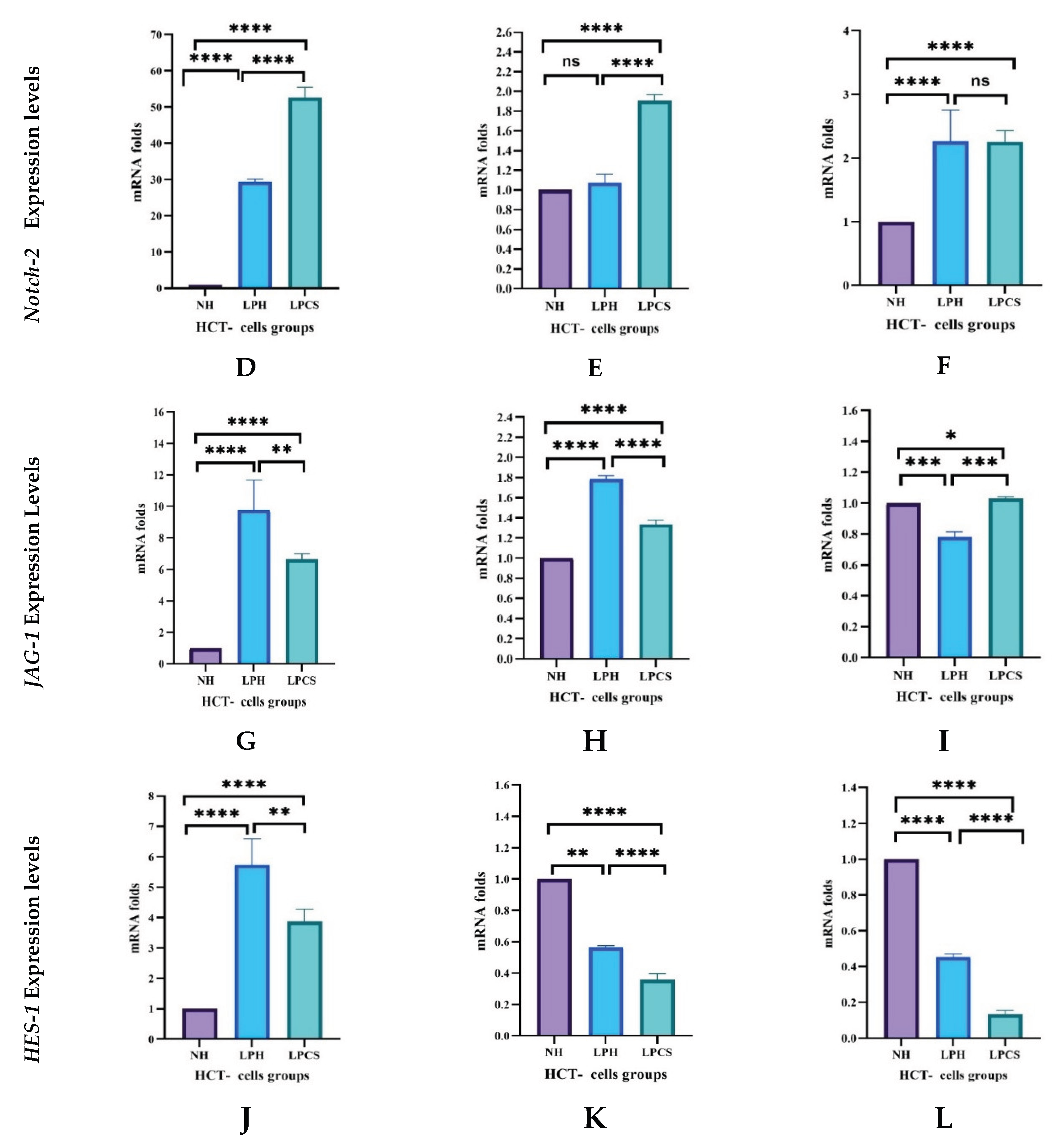
2.4.3. Levels of Notch Pathway Gene Expressions
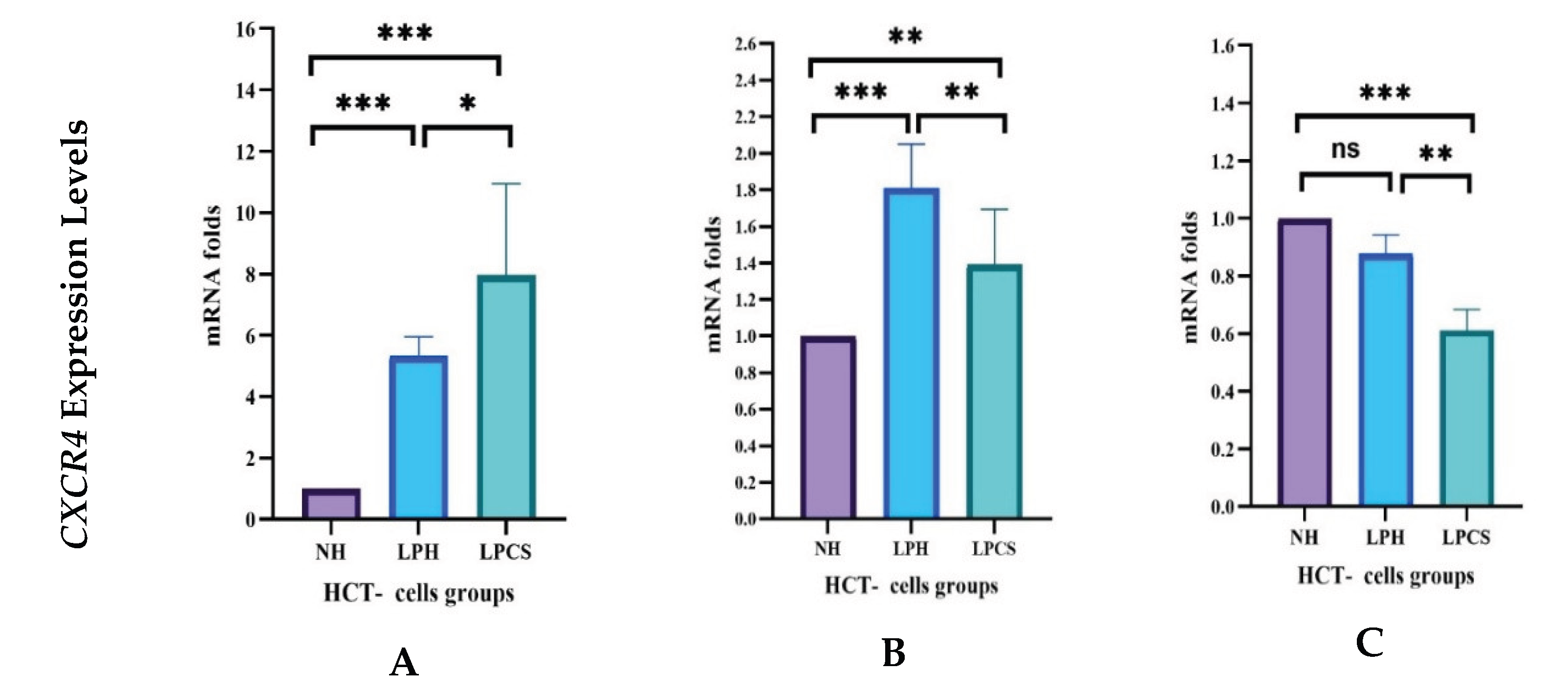
2.4.4. Levels of CXCR4 Receptor Gene Expression
2.5. Detailed Analysis of Gene Expression Correlations Post-Treatment
2.5.1. Temporal Analysis of Gene Regulatory Networks in Lactobacillus, Carrageenan, and Soy Protein Combination Treatment
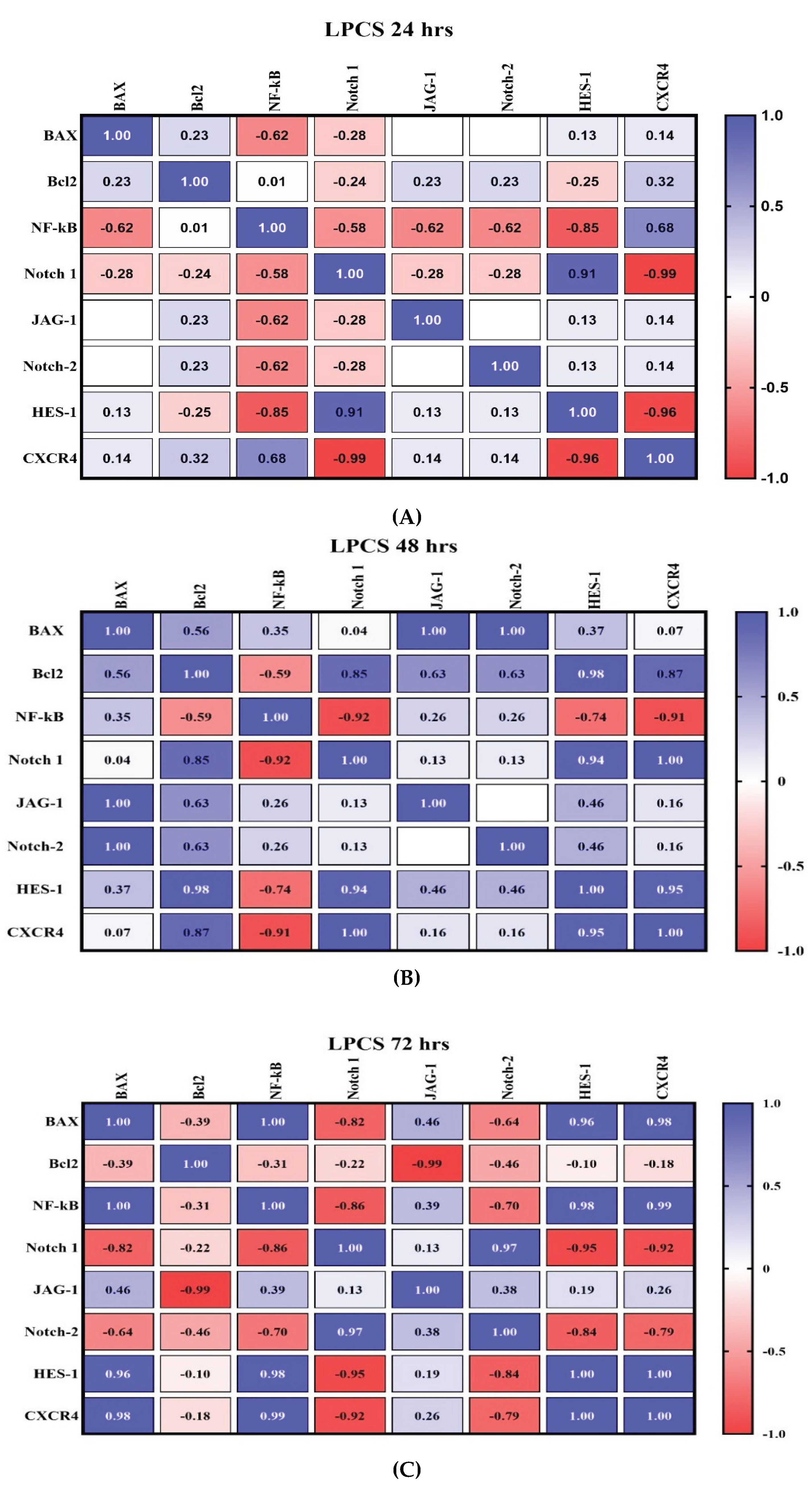
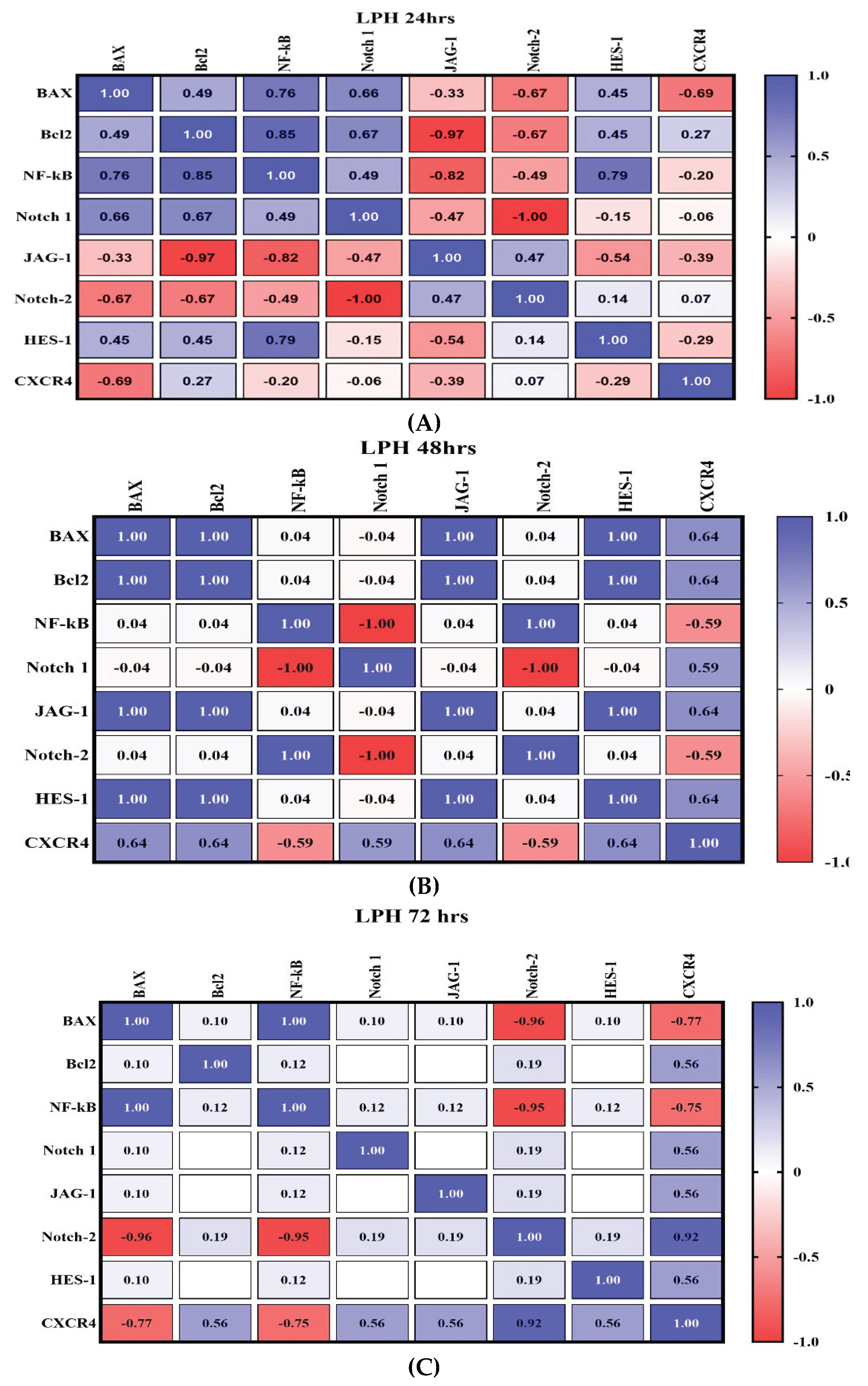
2.5.2. Temporal Analysis of Gene Regulatory Networks in Lactobacillus Postbiotic-Treated Cells
3. Discussion
4. Materials and Methods
4.1. Preparation of Lactobacillus acidophilus Postbiotics and Treatment Formulations
4.2. HCT-116 Cell Maintenance, Treatment, and Sample Harvesting
4.3. Evaluation of Cytotoxicity and Apoptosis in HCT-116 Cells
4.3.1. MTT Cell Viability Assay
4.3.2. Flow Cytometric Analysis of Apoptosis
4.4. Quantitative Gene Expression Analysis
4.5. Statistical Analysis and Data Visualization
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Primer Name | Primer sequence | Primer length |
| Notch-1 Forward | GAC ATC ACG GAT CAT ATG GA | 20 |
| Notch-1 Reverse | CTC GCA TTG ACC ATT CAA AC | 20 |
| Notch-2 Forward | GAT GCC ACC TGA ACA ACT GC | 20 |
| Notch-2 _Reverse | TGA CAA CAG CAA CAG CAA GG | 20 |
| Jadded-1_Forward | AGC GAC CTG TGT GGA TGA G | 19 |
| Jadded-1_Reverse | GGC TGG AGA CTG GAA GAC C | 19 |
| HES-1 Forward | CCA GTT TGC TTT CCT CAT TCC | 21 |
| HES-1 Reverse | TCT TCT CTC CCA GTA TTC AAG TTCC | 25 |
| BAX Forward | CCT GTG CAC CAA GGT GCC GGA ACT | 24 |
| BAX_Reverse | CCA CCC TGG TCT TGG ATC CAG CCC | 24 |
| BCL-2 Forward | TTG TGG CCT TCT TTG AGT TCG GTG | 24 |
| BCL-2_Reverse | GGT GCC GGT TCA GGT ACT CAG TCA | 24 |
| NF-kB Forward | ATC CCA TCT TTG ACA ATC GTG C | 22 |
| NF-kB Reverse | CTG GTC CCG TGA AAT ACA CCT C | 22 |
| CXCR-4 Forward | TTC TAC CCC AAT GAC TTG TG | 20 |
| CXCR-4 Reverse | ATG TAG TAA GGC AGC CAA CA | 20 |
| GAPDH Forward | GCA CCG TCA AGG CTG AGA AC | 20 |
| GAPDH Reverse | TGG TGA AGA CGC CAG TGG A | 19 |
References
- Zou, J.-Y.; Chen, Q.-L.; Luo, X.-C.; Damdinjav, D.; Abdelmohsen, U. R.; Li, H.-Y.; Battulga, T.; Chen, H.-B.; Wang, Y.-Q.; Zhang, J.-Y., Natural products reverse cancer multidrug resistance. Frontiers in Pharmacology 2024, Volume 15 - 2024.
- Garg, P.; Malhotra, J.; Kulkarni, P.; Horne, D.; Salgia, R.; Singhal, S. S., Emerging Therapeutic Strategies to Overcome Drug Resistance in Cancer Cells. Cancers 2024, 16, (13), 2478.
- Nan, Y.; Su, H.; Zhou, B.; Liu, S., The function of natural compounds in important anticancer mechanisms. Front Oncol 2022, 12, 1049888.
- Qian, S.; Wei, Z.; Yang, W.; Huang, J.; Yang, Y.; Wang, J., The role of BCL-2 family proteins in regulating apoptosis and cancer therapy. Front Oncol 2022, 12, 985363.
- Mustafa, M.; Ahmad, R.; Tantry, I. Q.; Ahmad, W.; Siddiqui, S.; Alam, M.; Abbas, K.; Moinuddin; Hassan, M. I.; Habib, S.; Islam, S., Apoptosis: A Comprehensive Overview of Signaling Pathways, Morphological Changes, and Physiological Significance and Therapeutic Implications. Cells 2024, 13, (22), 1838.
- Wong, R. S., Apoptosis in cancer: from pathogenesis to treatment. J Exp Clin Cancer Res 2011, 30, (1), 87.
- Haynes, J.; Manogaran, P., Mechanisms and Strategies to Overcome Drug Resistance in Colorectal Cancer. International Journal of Molecular Sciences 2025, 26, (5), 1988.
- Sadati, S.; Khalaji, A.; Bonyad, A.; Khoshdooz, S.; Hosseini Kolbadi, K. S.; Bahrami, A.; Moeinfar, M. S.; Morshedi, M.; Ghamsaraian, A.; Eterafi, M.; Eshraghi, R.; Khaksary Mahabady, M.; Mirzaei, H., NF-κB and apoptosis: colorectal cancer progression and novel strategies for treatment. Eur J Med Res 2025, 30, (1), 616.
- Nishida, A.; Andoh, A., The Role of Inflammation in Cancer: Mechanisms of Tumor Initiation, Progression, and Metastasis. Cells 2025, 14, (7), 488.
- Bakrim, S.; Fessikh, M. E.; Elhrech, H.; Omari, N. E.; Amanullah, M.; Ming, L. C.; Moshawih, S.; Bouyahya, A., Targeting inflammation in cancer therapy: from mechanistic insights to emerging therapeutic approaches. J Transl Med 2025, 23, (1), 588.
- Sun, J.; Chen, Y.; Xu, Z.; Wang, W.; Li, P., Notch signaling in the tumor immune microenvironment of colorectal cancer: mechanisms and therapeutic opportunities. J Transl Med 2025, 23, (1), 315.
- Iluta, S.; Nistor, M.; Buruiana, S.; Dima, D., Notch and Hedgehog Signaling Unveiled: Crosstalk, Roles, and Breakthroughs in Cancer Stem Cell Research. Life 2025, 15, (2), 228.
- Shi, Q.; Xue, C.; Zeng, Y.; Yuan, X.; Chu, Q.; Jiang, S.; Wang, J.; Zhang, Y.; Zhu, D.; Li, L., Notch signaling pathway in cancer: from mechanistic insights to targeted therapies. Signal Transduct Target Ther 2024, 9, (1), 128.
- Kumar, V.; Vashishta, M.; Kong, L.; Wu, X.; Lu, J. J.; Guha, C.; Dwarakanath, B. S., The Role of Notch, Hedgehog, and Wnt Signaling Pathways in the Resistance of Tumors to Anticancer Therapies. Front Cell Dev Biol 2021, 9, 650772.
- Li, Q.; Geng, S.; Luo, H.; Wang, W.; Mo, Y. Q.; Luo, Q.; Wang, L.; Song, G. B.; Sheng, J. P.; Xu, B., Signaling pathways involved in colorectal cancer: pathogenesis and targeted therapy. Signal Transduct Target Ther 2024, 9, (1), 266.
- Magryś, A.; Pawlik, M., Postbiotic Fractions of Probiotics Lactobacillus plantarum 299v and Lactobacillus rhamnosus GG Show Immune-Modulating Effects. Cells 2023, 12, (21).
- D’Amore, T.; Zolfanelli, C.; Lauciello, V.; Di Ciancia, A.; Vagliasindi, A.; Smaoui, S.; Varzakas, T., Using Postbiotics from Functional Foods for Managing Colorectal Cancer: Mechanisms, Sources, Therapeutic Potential, and Clinical Perspectives. Microorganisms 2025, 13, (6), 1335.
- Guo, J.; Zhao, Y.; Guo, W.; Sun, Y.; Zhang, W.; Zhao, Q.; Zhang, Y.; Jiang, Y., Effects of Lactobacillus paracei JY062 Postbiotic on Intestinal Barrier, Immunity, and Gut Microbiota. Nutrients 2025, 17, (7), 1272.
- Szydłowska, A.; Sionek, B., Probiotics and Postbiotics as the Functional Food Components Affecting the Immune Response. Microorganisms 2022, 11, (1).
- Balendra, V.; Rosenfeld, R.; Amoroso, C.; Castagnone, C.; Rossino, M. G.; Garrone, O.; Ghidini, M., Postbiotics as Adjuvant Therapy in Cancer Care. Nutrients 2024, 16, (15).
- Krylova, N. V.; Kravchenko, A. O.; Iunikhina, O. V.; Pott, A. B.; Likhatskaya, G. N.; Volod’ko, A. V.; Zaporozhets, T. S.; Shchelkanov, M. Y.; Yermak, I. M., Influence of the Structural Features of Carrageenans from Red Algae of the Far Eastern Seas on Their Antiviral Properties. Marine Drugs 2022, 20, (1), 60.
- Komisarska, P.; Pinyosinwat, A.; Saleem, M.; Szczuko, M., Carrageenan as a Potential Factor of Inflammatory Bowel Diseases. Nutrients 2024, 16, (9).
- Silva-Pinto, P. A.; de Pontes, J. T. C.; Aguilar-Morón, B.; Canales, C. S. C.; Pavan, F. R.; Roque-Borda, C. A., Phytochemical insights into flavonoids in cancer: Mechanisms, therapeutic potential, and the case of quercetin. Heliyon 2025, 11, (4), e42682.
- Hadidi, M.; Liñán-Atero, R.; Tarahi, M.; Christodoulou, M. C.; Aghababaei, F., The Potential Health Benefits of Gallic Acid: Therapeutic and Food Applications. Antioxidants (Basel) 2024, 13, (8).
- Poschner, S.; Maier-Salamon, A.; Zehl, M.; Wackerlig, J.; Dobusch, D.; Pachmann, B.; Sterlini, K. L.; Jäger, W., The Impacts of Genistein and Daidzein on Estrogen Conjugations in Human Breast Cancer Cells: A Targeted Metabolomics Approach. Frontiers in Pharmacology 2017, Volume 8 - 2017.
- Fan, Y.; Wang, M.; Li, Z.; Jiang, H.; Shi, J.; Shi, X.; Liu, S.; Zhao, J.; Kong, L.; Zhang, W.; Ma, L., Intake of Soy, Soy Isoflavones and Soy Protein and Risk of Cancer Incidence and Mortality. Front Nutr 2022, 9, 847421.
- Basson, A. R.; Ahmed, S.; Almutairi, R.; Seo, B.; Cominelli, F., Regulation of Intestinal Inflammation by Soybean and Soy-Derived Compounds. Foods 2021, 10, (4), 774.
- Koch, D. T.; Yu, H.; Beirith, I.; Schirren, M.; Drefs, M.; Liu, Y.; Knoblauch, M.; Koliogiannis, D.; Sheng, W.; De Toni, E. N.; Bazhin, A. V.; Renz, B. W.; Guba, M. O.; Werner, J.; Ilmer, M., Tigecycline causes loss of cell viability mediated by mitochondrial OXPHOS and RAC1 in hepatocellular carcinoma cells. J Transl Med 2023, 21, (1), 876.
- Vitale, I.; Pietrocola, F.; Guilbaud, E.; Aaronson, S. A.; Abrams, J. M.; Adam, D.; Agostini, M.; et al., Apoptotic cell death in disease: Current understanding of the NCCD 2023. Cell Death Differ 2023, 30, (5), 1097-1154.
- Chu, X.; Tian, W.; Ning, J.; Xiao, G.; Zhou, Y.; Wang, Z.; Zhai, Z.; Tanzhu, G.; Yang, J.; Zhou, R., Cancer stem cells: advances in knowledge and implications for cancer therapy. Signal Transduct Target Ther 2024, 9, (1), 170.
- Hijová, E., Postbiotics as Metabolites and Their Biotherapeutic Potential. International Journal of Molecular Sciences 2024, 25, (10), 5441.
- Gomez, X.; Sanon, S.; Zambrano, K.; Asquel, S.; Bassantes, M.; Morales, J. E.; Otanez, G.; Pomaquero, C.; Villarroel, S.; Zurita, A.; Calvache, C.; Celi, K.; Contreras, T.; Corrales, D.; Naciph, M. B.; Pena, J.; Caicedo, A., Key points for the development of antioxidant cocktails to prevent cellular stress and damage caused by reactive oxygen species (ROS) during manned space missions. NPJ Microgravity 2021, 7, (1), 35.
- Caliman-Sturdza, O. A.; Vaz, J. A.; Lupaescu, A. V.; Lobiuc, A.; Bran, C.; Gheorghita, R. E., Antioxidant and Anti-Inflammatory Activities of Probiotic Strains. International Journal of Molecular Sciences 2026, 27, (2), 1079.
- Zong, Y.; Li, H.; Liao, P.; Chen, L.; Pan, Y.; Zheng, Y.; Zhang, C.; Liu, D.; Zheng, M.; Gao, J., Mitochondrial dysfunction: mechanisms and advances in therapy. Signal Transduct Target Ther 2024, 9, (1), 124.
- White, S. A.; Zhang, L. S.; Pasula, D. J.; Yang, Y. H. C.; Luciani, D. S., Bax and Bak jointly control survival and dampen the early unfolded protein response in pancreatic beta-cells under glucolipotoxic stress. Sci Rep 2020, 10, (1), 10986.
- Palabiyik, A. A., The role of Bcl-2 in controlling the transition between autophagy and apoptosis (Review). Mol Med Rep 2025, 32, (1).
- Asoudeh-Fard, A.; Yeylagh-Beygi, M.; Asoudeh-Fard, M.; Abbasi, M.; Bagher Nazari, M.; Gholami, A.; Parsaei, A., Bifidobacterium enhances the antitumor efficacy of carboplatin in glioblastoma cells: targeting apoptotic and cell cycle regulatory pathways via Caspase, AKT/PTEN, and P53/P21 signaling. Cancer Cell Int 2025, 26, (1), 12.
- Park, S. H., Recent Research on the Role of Lactobacilli Probiotics in Cancer Management. Nutrients 2026, 18, (2).
- Guo, Q.; Jin, Y.; Chen, X.; Ye, X.; Shen, X.; Lin, M.; Zeng, C.; Zhou, T.; Zhang, J., NF-κB in biology and targeted therapy: new insights and translational implications. Signal Transduct Target Ther 2024, 9, (1), 53.
- Szrejder, M.; Piwkowska, A., Gut Microbiome-Derived Short-Chain Fatty Acids in Glomerular Protection and Modulation of Chronic Kidney Disease Progression. Nutrients 2025, 17, (17), 2904.
- Ito, Y.; Kikuchi, E.; Tanaka, N.; Kosaka, T.; Suzuki, E.; Mizuno, R.; Shinojima, T.; Miyajima, A.; Umezawa, K.; Oya, M., Down-regulation of NF kappa B activation is an effective therapeutic modality in acquired platinum-resistant bladder cancer. BMC Cancer 2015, 15, 324.
- Gallenstein, N.; Tichy, L.; Weigand, M. A.; Schenz, J., Notch Signaling in Acute Inflammation and Sepsis. International Journal of Molecular Sciences 2023, 24, (4), 3458.
- Luan, W.; Wright, A. L.; Brown-Wright, H.; Le, S.; San Gil, R.; Madrid San Martin, L.; Ling, K.; Jafar-Nejad, P.; Rigo, F.; Walker, A. K., Early activation of cellular stress and death pathways caused by cytoplasmic TDP-43 in the rNLS8 mouse model of ALS and FTD. Mol Psychiatry 2023, 28, (6), 2445-2461.
- Edelmann, J., NOTCH1 Signalling: A key pathway for the development of high-risk chronic lymphocytic leukaemia. Frontiers in Oncology 2022, Volume 12 - 2022.
- Chen, C.; Du, Y.; Nie, R.; Wang, S.; Wang, H.; Li, P., Notch signaling in cancers: mechanism and potential therapy. Frontiers in Cell and Developmental Biology 2025, Volume 13 - 2025.
- Dai, Y.; Wilson, G.; Huang, B.; Peng, M.; Teng, G.; Zhang, D.; Zhang, R.; Ebert, M. P.; Chen, J.; Wong, B. C.; Chan, K. W.; George, J.; Qiao, L., Silencing of Jagged1 inhibits cell growth and invasion in colorectal cancer. Cell Death Dis 2014, 5, (4), e1170.
- He, J.; Li, L.; Lv, L.; Chen, X.; Ge, M.; Ren, Y.; Tang, X.; Liu, P.; Gao, W., JAG1 is correlated to suppressive immune microenvironment and predicts immunotherapy resistance in lung adenocarcinoma. Frontiers in Oncology 2023, Volume 13 - 2023.
- Zhang, Q.; Wang, H. Y.; Nayak, A.; Nunez-Cruz, S.; Slupianek, A.; Liu, X.; Basappa, J.; Fan, J. S.; Chekol, S.; Nejati, R.; Bogusz, A. M.; Turner, S. D.; Swaminathan, K.; Wasik, M. A., Induction of Transcriptional Inhibitor HES1 and the Related Repression of Tumor-Suppressor TXNIP Are Important Components of Cell-Transformation Program Imposed by Oncogenic Kinase NPM-ALK. Am J Pathol 2022, 192, (8), 1186-1198.
- Asefa, Z.; Belay, A.; Welelaw, E.; Haile, M., Postbiotics and their biotherapeutic potential for chronic disease and their feature perspective: a review. Frontiers in Microbiomes 2025, Volume 4 - 2025.
- Giambra, V.; Caldarelli, M.; Franza, L.; Rio, P.; Bruno, G.; di Iasio, S.; Mastrogiovanni, A.; Gasbarrini, A.; Gambassi, G.; Cianci, R., The Role of Notch Signaling and Gut Microbiota in Autoinflammatory Diseases: Mechanisms and Future Views. Biomedicines 2025, 13, (4), 768.
- Schioppa, T.; Uranchimeg, B.; Saccani, A.; Biswas, S. K.; Doni, A.; Rapisarda, A.; Bernasconi, S.; Saccani, S.; Nebuloni, M.; Vago, L.; Mantovani, A.; Melillo, G.; Sica, A., Regulation of the chemokine receptor CXCR4 by hypoxia. J Exp Med 2003, 198, (9), 1391-402.
- Fleszar, M. G.; Zawadzki, M.; Fortuna, P.; Bednarz-Misa, I.; Krauze, I.; Maciejewska, K.; Klekowski, J.; Chabowski, M.; Witkiewicz, W.; Krzystek-Korpacka, M., Robot-Assisted Colorectal Cancer Surgery Mitigates Early Postoperative Immunosuppression and Angiogenesis. International Journal of Molecular Sciences 2025, 26, (20), 10041.
- Artusa, V.; Calabrone, L.; Mortara, L.; Peri, F.; Bruno, A., Microbiota-Derived Natural Products Targeting Cancer Stem Cells: Inside the Gut Pharma Factory. International Journal of Molecular Sciences 2023, 24, (5), 4997.
- Smaldone, G.; Di Matteo, F.; Castelluccio, R.; Napolitano, V.; Miranda, M. R.; Manfra, M.; Campiglia, P.; Vestuto, V., Targeting the CXCR4/CXCL12 Axis in Cancer Therapy: Analysis of Recent Advances in the Development of Potential Anticancer Agents. Molecules 2025, 30, (6), 1380.
- Li, M.; Chen, F.; Clifton, N.; Sullivan, D. M.; Dalton, W. S.; Gabrilovich, D. I.; Nefedova, Y., Combined inhibition of Notch signaling and Bcl-2/Bcl-xL results in synergistic antimyeloma effect. Mol Cancer Ther 2010, 9, (12), 3200-9.
- Shin, H. C.; Seo, J.; Kang, B. W.; Moon, J. H.; Chae, Y. S.; Lee, S. J.; Lee, Y. J.; Han, S.; Seo, S. K.; Kim, J. G.; Sohn, S. K.; Park, T. I., Clinical significance of nuclear factor κB and chemokine receptor CXCR4 expression in patients with diffuse large B-cell lymphoma who received rituximab-based therapy. Korean J Intern Med 2014, 29, (6), 785-92.
- Hasnan, J.; Yusof, M. I.; Damitri, T. D.; Faridah, A. R.; Adenan, A. S.; Norbaini, T. H., Relationship between apoptotic markers (Bax and Bcl-2) and biochemical markers in type 2 diabetes mellitus. Singapore Med J 2010, 51, (1), 50-5.
- Naseri, M. H.; Mahdavi, M.; Davoodi, J.; Tackallou, S. H.; Goudarzvand, M.; Neishabouri, S. H., Upregulation of Bax and downregulation of Bcl2 during 3-NC mediated apoptosis in human cancer cells. Cancer Cell Int 2015, 15, 55.
- Gurunathan, S.; Thangaraj, P.; Kim, J. H., Postbiotics: Functional Food Materials and Therapeutic Agents for Cancer, Diabetes, and Inflammatory Diseases. Foods 2023, 13, (1).
- Xia, H.-H.; Huang, J.-Z., Tryptophan metabolism at the crossroads of the neuro-immuno-microbial axis: implications for precision medicine in chronic diseases. Frontiers in Cellular and Infection Microbiology 2026, Volume 15 - 2025.
- Yuliana, T.; Tyano, F. N.; Harlina, P. W.; Cahyana, Y.; Marta, H.; Krama, A., Characterizing Probiotic Lactic Acid Bacteria from Buffalo Milk Fermentation (Dadih) for Beef Biopreservation. Applied Sciences 2023, 13, (24), 13089.
- Livak, K. J.; Schmittgen, T. D., Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, (4), 402-8.

| Comparative Item | Untreated Control group | Lactobacillus postbiotics group | Lactobacillus postbiotics, κ-Carrageenan, and soy protein group |
|---|---|---|---|
| Number of HCT cells in each flask | 4 x 106 | ||
| Treatment concentration | Untreated | 50 mg postbiotic /10 ml growth media | 50 mg postbiotic + 25 mg carrageenan + 5 mg soy protein/10 ml growth media |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





