Submitted:
05 February 2026
Posted:
06 February 2026
You are already at the latest version
Abstract
Keywords:
Introduction
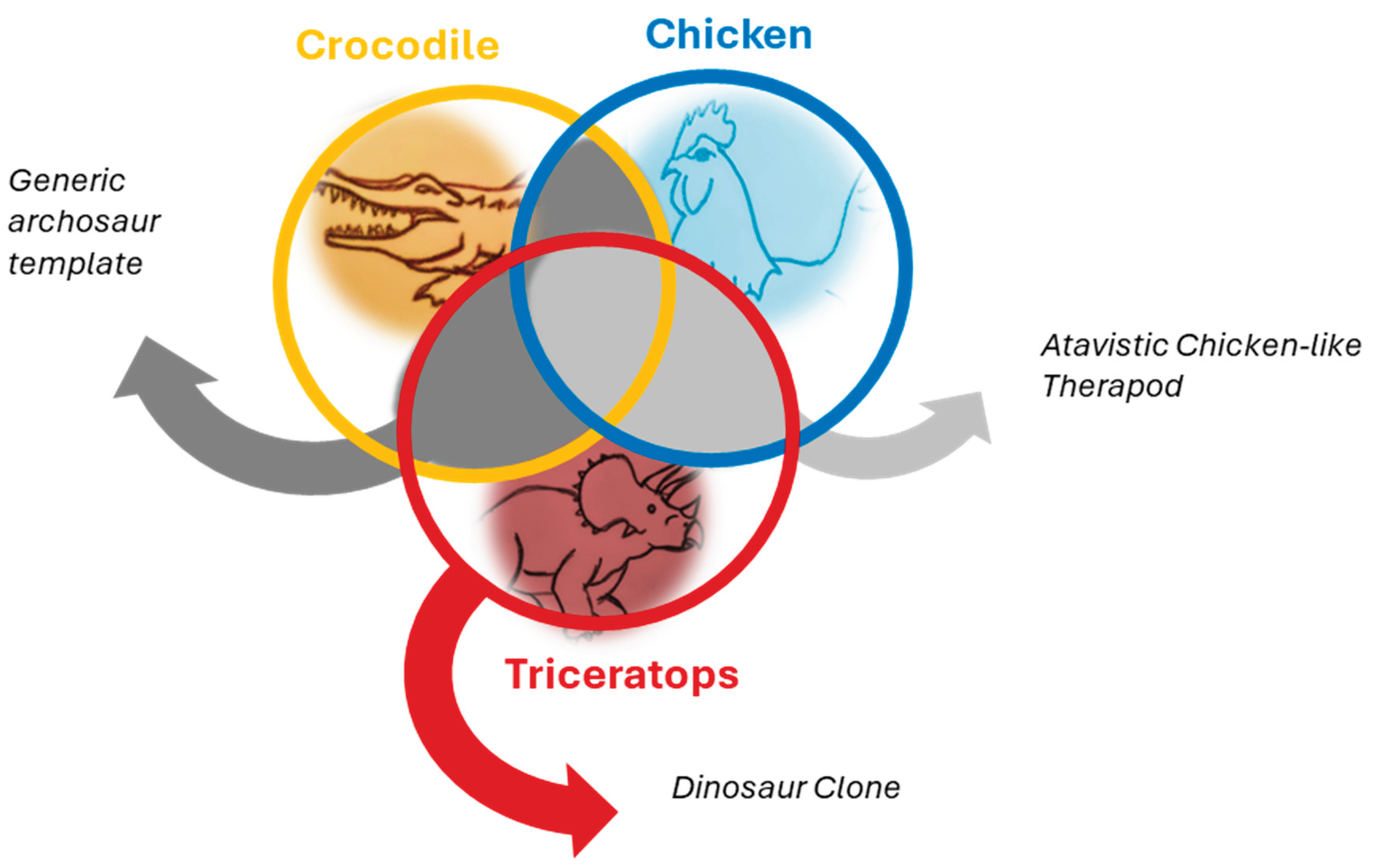
The Genetic Shrapnel Approach
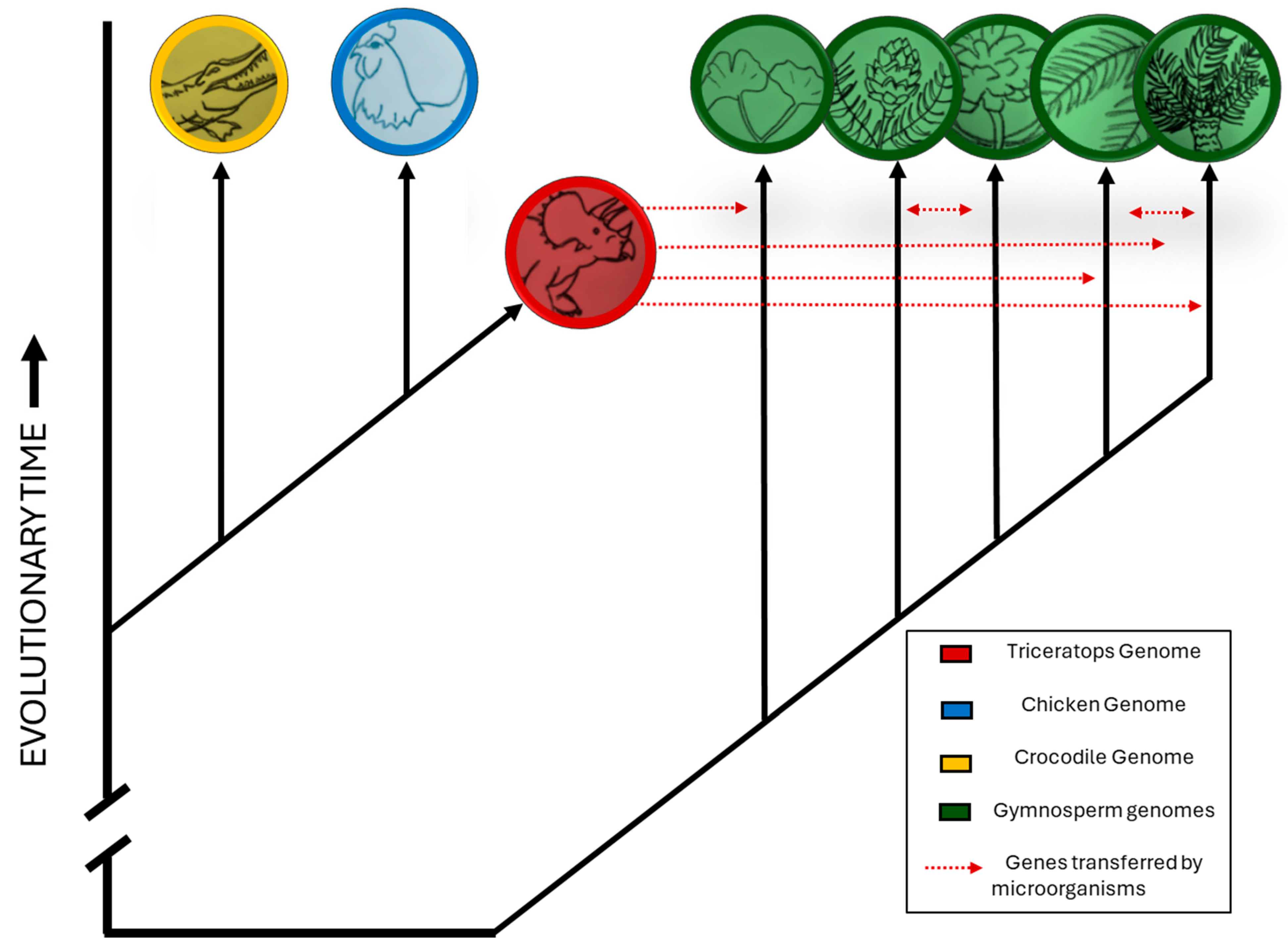
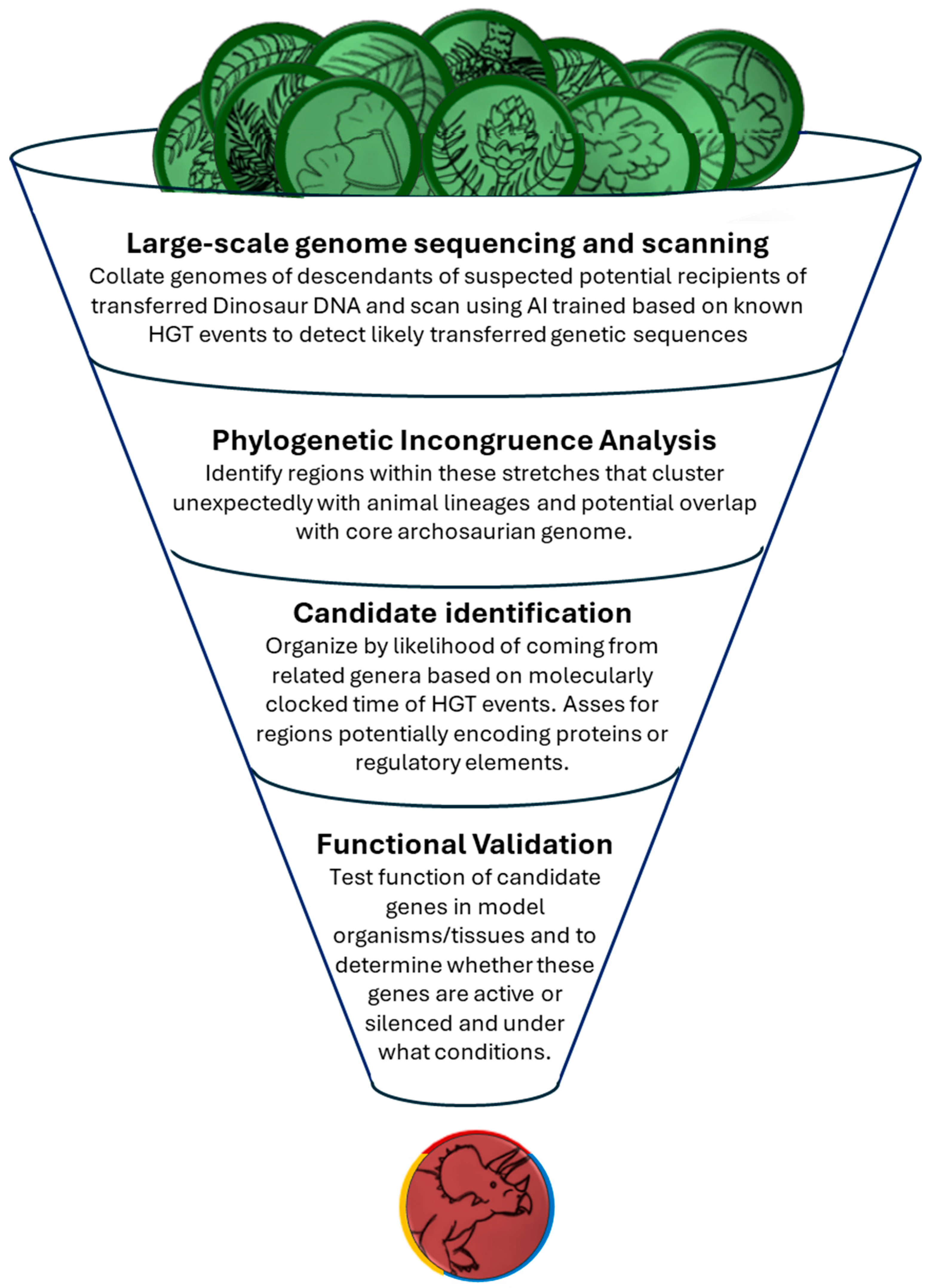
- Perform comprehensive genome sequencing of extant gymnosperm genera—particularly conifers—as well as relatively primitive (plesiomorphic) fungal and tick lineages.
- Screen the “dark genome” of these organisms—regions enriched with transposable elements and non-coding DNA—for sequences that appear to originate from non-plant sources.
- Cross-reference candidate sequences against known animal genomes and databases of horizontally transferred genes, focusing on segments plausibly derived from vertebrates.
- Apply molecular dating methods to estimate the timing of HGT events and evaluate whether these events plausibly occurred during the Mesozoic era.
- Use phylogenetic and comparative analyses to determine if these sequences correspond with known archosaur genomic features and to assess their likelihood of being dinosaurian in origin.
Finding a Dinosaur in a Haystack
Bully for Brontosaurus
Summary and Conclusions
Data Availability Statement
References
- Abby, S.S.; Tannier, E.; Gouy, M.; Daubin, V. Detecting lateral gene transfers by statistical reconciliation of phylogenetic forests. BMC Bioinform. 2014, 11, 324–324. [CrossRef]
- Alexander, W.G.; Wisecaver, J.H.; Rokas, A.; Hittinger, C.T. Horizontally acquired genes in early-diverging pathogenic fungi enable the use of host nucleosides and nucleotides. Proc. Natl. Acad. Sci. 2016, 113, 4116–4121. [CrossRef]
- Allentoft, M.E.; Collins, M.; Harker, D.; Haile, J.; Oskam, C.L.; Hale, M.L.; Campos, P.F.; Samaniego, J.A.; Gilbert, M.T.P.; Willerslev, E.; et al. The half-life of DNA in bone: measuring decay kinetics in 158 dated fossils. Proc. R. Soc. B: Biol. Sci. 2012, 279, 4724–4733. [CrossRef]
- Asara, J.M.; Schweitzer, M.H.; Freimark, L.M.; Phillips, M.; Cantley, L.C. Protein Sequences from Mastodon and Tyrannosaurus Rex Revealed by Mass Spectrometry. Science 2007, 316, 280–285. [CrossRef]
- Bergthorsson, U.; Richardson, A.O.; Young, G.J.; Goertzen, L.R.; Palmer, J.D. Massive horizontal transfer of mitochondrial genes from diverse land plant donors to the basal angiosperm Amborella. Proc. Natl. Acad. Sci. 2004, 101, 17747–17752. [CrossRef]
- Bhullar, B.-A.S.; Morris, Z.S.; Sefton, E.M.; Tok, A.; Tokita, M.; Namkoong, B.; Camacho, J.; Burnham, D.A.; Abzhanov, A. A molecular mechanism for the origin of a key evolutionary innovation, the bird beak and palate, revealed by an integrative approach to major transitions in vertebrate history. Evolution 2015, 69, 1665–1677. [CrossRef]
- Biswas, C., & Johri, B. M. (2013). The gymnosperms. Springer Science & Business Media.
- Bock, R. The give-and-take of DNA: horizontal gene transfer in plants. Trends Plant Sci. 2010, 15, 11–22. [CrossRef]
- Boothby, T.C.; Tenlen, J.R.; Smith, F.W.; Wang, J.R.; Patanella, K.A.; Nishimura, E.O.; Tintori, S.C.; Li, Q.; Jones, C.D.; Yandell, M.; et al. Evidence for extensive horizontal gene transfer from the draft genome of a tardigrade. Proc. Natl. Acad. Sci. 2015, 112, 15976–15981. [CrossRef]
- Boto, L. Horizontal gene transfer in evolution: facts and challenges. Proc. R. Soc. B: Biol. Sci. 2014, 277, 819–827. [CrossRef]
- Bromham, L.; Duchêne, S.; Hua, X.; Ritchie, A.M.; Duchêne, D.A.; Ho, S.Y.W. Bayesian molecular dating: opening up the black box. Biol. Rev. 2020, 93, 1165–1191. [CrossRef]
- Brusatte, S.L.; Carr, T.D. The phylogeny and evolutionary history of tyrannosauroid dinosaurs. Sci. Rep. 2016, 6, 20252. [CrossRef]
- Carroll, S. B. (2005). Endless forms most beautiful. W. W. Norton & Company.
- Cediel-Becerra, J.D.D.; Cumsille, A.; Guerra, S.; Ding, Y.; de Crécy-Lagard, V.; Chevrette, M.G. Targeted genome mining with GATOR-GC maps the evolutionary landscape of biosynthetic diversity. Nucleic Acids Res. 2025, 53. [CrossRef]
- Crisp, A.; Boschetti, C.; Perry, M.; Tunnacliffe, A.; Micklem, G. Expression of multiple horizontally acquired genes is a hallmark of both vertebrate and invertebrate genomes. Genome Biol. 2015, 16, 50–50. [CrossRef]
- Dawkins, R. (2024). The Genetic Book of the Dead: A Darwinian Reverie. Yale University Press.
- Dennett, D. C. (2017). From bacteria to Bach and back: The evolution of minds. W. W. Norton & Company.
- Drummond, A.J.; Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 2007, 7, 1–8. [CrossRef]
- Hotopp, J.C.D. Horizontal gene transfer between bacteria and animals. Trends Genet. 2011, 27, 157–163. [CrossRef]
- Edsinger, E.; Moroz, L.L. Genomic hotspots: localized chromosome gene expansions identify lineage-specific innovations as targets for functional biodiversity and predictions of stress resilience. Front. Mar. Sci. 2024, 11, 1434130. [CrossRef]
- Feschotte, C.; Pritham, E.J. DNA Transposons and the Evolution of Eukaryotic Genomes. Annu. Rev. Genet. 2007, 41, 331–368. [CrossRef]
- Gilbert, C., Schaack, S., Pace II, J. K., Brindley, P. J., & Feschotte, C. (2010). A role for host–parasite interactions in the horizontal transfer of transposons across phyla. Nature, 464(7293), 1347-1350.
- Go, D.; Yeon, G.-H.; Park, S.J.; Lee, Y.; Koh, H.G.; Koo, H.; Kim, K.H.; Jin, Y.-S.; Sung, B.H.; Kim, J. Integration of metabolomics and other omics: from microbes to microbiome. Appl. Microbiol. Biotechnol. 2024, 108, 1–7. [CrossRef]
- González Villa, T., & Vinas, M. (2019). Horizontal gene transfer: breaking borders between living kingdoms. Springer.
- Gould, S. J. (1983). Hen’s teeth and horse’s toes: Further reflections in natural history. New York, NY: W. W. Norton & Company.
- Gould, S. J. (1995). Dinosaur in a Haystack: Reflections in Natural History. Harvard University Press.
- Green, R.E.; Braun, E.L.; Armstrong, J.; Earl, D.; Nguyen, N.; Hickey, G.; Vandewege, M.W.; John, J.A.S.; Capella-Gutiérrez, S.; Castoe, T.A.; et al. Three crocodilian genomes reveal ancestral patterns of evolution among archosaurs. Science 2014, 346, 1254449. [CrossRef]
- Gulia-Nuss, M.; Nuss, A.B.; Meyer, J.M.; Sonenshine, D.E.; Roe, R.M.; Waterhouse, R.M.; Sattelle, D.B.; de la Fuente, J.; Ribeiro, J.M.; Megy, K.; et al. Genomic insights into the Ixodes scapularis tick vector of Lyme disease. Nat. Commun. 2016, 7, 1–13. [CrossRef]
- Griffin, D. K., Hassan, S., & Trifonov, V. (2019). Time lapse: A glimpse into prehistoric genomics. Genes, 10(6), 464.
- Hedlund, J. Risky business: safety regulations, risk compensation, and individual behavior. Inj. Prev. 2000, 6, 82–89. [CrossRef]
- Ho, S.Y.W.; Duchêne, S. Molecular-clock methods for estimating evolutionary rates and timescales. Mol. Ecol. 2014, 23, 5947–5965. [CrossRef]
- Horner, J., & Gorman, J. (2009). How to build a dinosaur: The new science of reverse evolution. Penguin.
- Kambayashi, C.; Kakehashi, R.; Sato, Y.; Mizuno, H.; Tanabe, H.; Rakotoarison, A.; Künzel, S.; Furuno, N.; Ohshima, K.; Kumazawa, Y.; et al. Geography-Dependent Horizontal Gene Transfer from Vertebrate Predators to Their Prey. Mol. Biol. Evol. 2022, 39. [CrossRef]
- Klompen, J. S. H., Black, W., Keirans, J. E., & Oliver Jr, J. H. (1996). Evolution of ticks. Annual review of entomology, 41(1), 141-161.
- Leitch, I.J.; Leitch, A.R. (2012). Genome size diversity and evolution in land plants. In Plant genome diversity Volume 2: Physical structure, behaviour and evolution of plant genomes (pp. 307-322). Vienna: Springer Vienna.
- Mans, B.J.; de Klerk, D.; Pienaar, R.; Latif, A.A. Nuttalliella namaqua: A Living Fossil and Closest Relative to the Ancestral Tick Lineage: Implications for the Evolution of Blood-Feeding in Ticks. PLOS ONE 2011, 6, e23675. [CrossRef]
- Marshall, C. R., Ward, P. D., & Smith, F. A. (2021). Absolute abundance and preservation rate of Tyrannosaurus rex. Science, 372(6539), 284–287.
- Naranjo-Ortiz, M.A.; Gabaldón, T. Fungal evolution: major ecological adaptations and evolutionary transitions. Biol. Rev. 2019, 94, 1443–1476. [CrossRef]
- National Human Genome Research Institute. (2010.). Why mouse matters. Retrieved August 21, 2025, from https://www.genome.gov/10001345/importance-of-mouse-genome.
- Nystedt, B.; Street, N.R.; Wetterbom, A.; Zuccolo, A.; Lin, Y.-C.; Scofield, D.G.; Vezzi, F.; Delhomme, N.; Giacomello, S.; Alexeyenko, A.; et al. The Norway spruce genome sequence and conifer genome evolution. Nature 2013, 497, 579–584. [CrossRef]
- O’cOnnor, R.E.; Romanov, M.N.; Kiazim, L.G.; Barrett, P.M.; Farré, M.; Damas, J.; Ferguson-Smith, M.; Valenzuela, N.; Larkin, D.M.; Griffin, D.K. Reconstruction of the diapsid ancestral genome permits chromosome evolution tracing in avian and non-avian dinosaurs. Nat. Commun. 2018, 9, 1–9. [CrossRef]
- Peñalver, E.; Arillo, A.; Delclòs, X.; Peris, D.; Grimaldi, D.A.; Anderson, S.R.; Nascimbene, P.C.; la Fuente, R.P.-D. Ticks parasitised feathered dinosaurs as revealed by Cretaceous amber assemblages. Nat. Commun. 2017, 8, 1924–1924. [CrossRef]
- Penney, D.; Wadsworth, C.; Fox, G.; Kennedy, S.L.; Preziosi, R.F.; A Brown, T. Absence of Ancient DNA in Sub-Fossil Insect Inclusions Preserved in ‘Anthropocene’ Colombian Copal. PLOS ONE 2013, 8, e73150. [CrossRef]
- Peris, D.; Janssen, K.; Barthel, H.J.; Bierbaum, G.; Delclòs, X.; Peñalver, E.; Solórzano-Kraemer, M.M.; Jordal, B.H.; Rust, J. DNA from resin-embedded organisms: Past, present and future. PLOS ONE 2020, 15, e0239521. [CrossRef]
- Perkel, J.M. Beyond AlphaFold: how AI is decoding the grammar of the genome. Nature 2025, 644, 829–832. [CrossRef]
- Prabahar, A. (2022). Integration of Transcriptomics Data and Metabolomic Data Using Biomedical Literature Mining and Pathway Analysis. In Biomedical Text Mining (pp. 301-316). New York, NY: Springer US.
- Peltzman, S. The Effects of Automobile Safety Regulation. J. Politi- Econ. 1975, 83, 677–725. [CrossRef]
- Rasmussen, M.D.; Kellis, M. Unified modeling of gene duplication, loss, and coalescence using a locus tree. Genome Res. 2012, 22, 755–765. [CrossRef]
- Redecker, D.; Kodner, R.; Graham, L.E. Glomalean Fungi from the Ordovician. Science 2000, 289, 1920–1921. [CrossRef]
- Richards, T.A.; Leonard, G.; Soanes, D.M.; Talbot, N.J. Gene transfer into the fungi. Fungal Biol. Rev. 2011, 25, 98–110. [CrossRef]
- Richardson, S.R.; Doucet, A.J.; Kopera, H.C.; Moldovan, J.B.; Garcia-Perez, J.L.; Moran, J.V.; Chandler, M.; Gellert, M.; Lambowitz, A.M.; Rice, P.A.; et al.
- Royal Veterinary College (RVC). (2024). Scientists use modern species to discover genome structure of dinosaurs. [Press release]. https://www.rvc.ac.uk/news-and-events/rvc-news/scientists-use-modern-species-to-discover-genome-structure-of-dinosaurs.
- Szöllősi, G.J.; Tannier, E.; Daubin, V.; Boussau, B. The Inference of Gene Trees with Species Trees. Syst. Biol. 2015, 64, e42–e62. [CrossRef]
- Wani, A.K.; Akhtar, N.; Sher, F.; Navarrete, A.A.; Américo-Pinheiro, J.H.P. Microbial adaptation to different environmental conditions: molecular perspective of evolved genetic and cellular systems. Arch. Microbiol. 2022, 204, 1–16. [CrossRef]
| 1 | Tyrannosaurus and birds share a common ancestor approximately 150-160 mya. This means the lineage giving rise to modern birds has had, say 150 million years to evolve differences from the common ancestor, while the Tyrannosaurus lineage had 85 million years (because it went extinct 65 mya). Thus, some 210 million years of evolutionary change separates Tyrannosaurus Rex from chickens. Humans and mice share a common ancestor around 90 mya and so have had about 180 million years to evolve differences. If we take it that approximately 85 per cent of expressed DNA in human and mice are identical (National Human Genome Research Institute, 2010), then 80 per cent does not seem an outlandish estimate. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



