Submitted:
29 January 2026
Posted:
30 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Differential Metabolites Extraction and Analysis
2.2.1. Sample Preparation
2.2.2. UHPLC Conditions
2.2.3. Mass Spectrometry Detection Conditions
2.2.4. Metabolomics Data Analysis
2.3. Transcriptome Analysis
2.4. RT-qPCR Validation of DEGs Expression Levels
3. Results
3.1. Analysis of Metabolic Differences in Fowers Between PFs and WFs
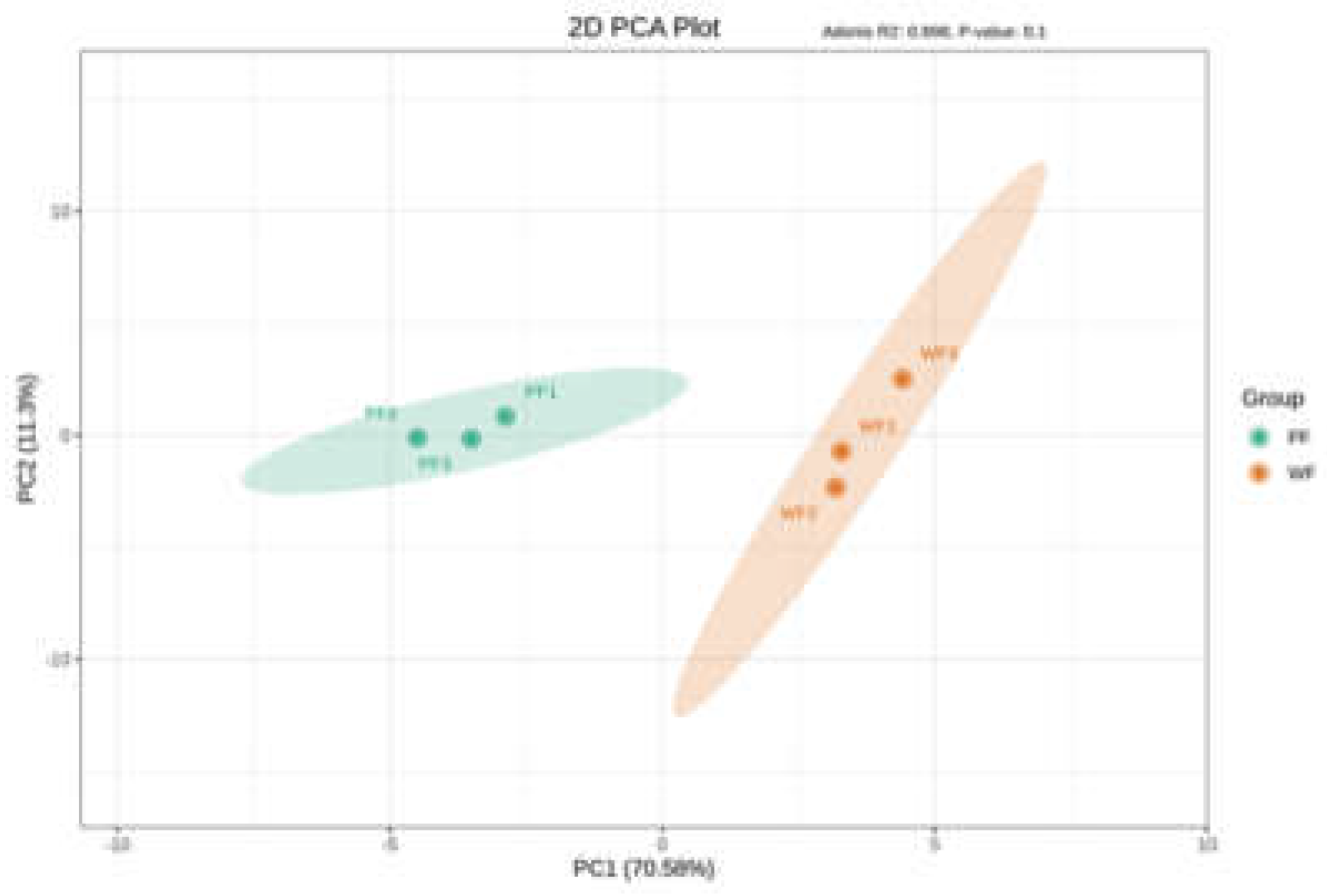
3.1.1. PCA Clusters Two Platycodon Flower Groups Distinctly
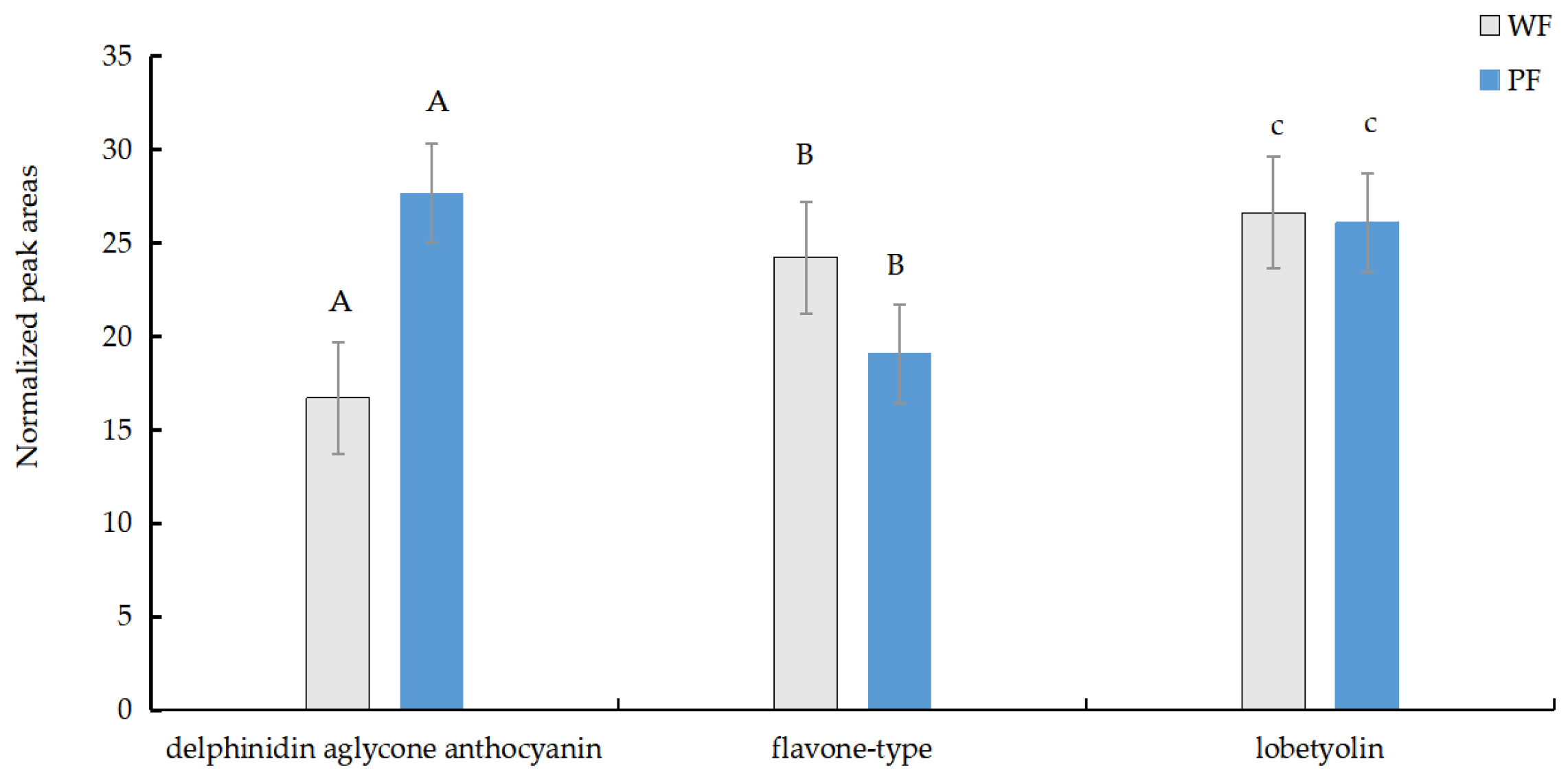
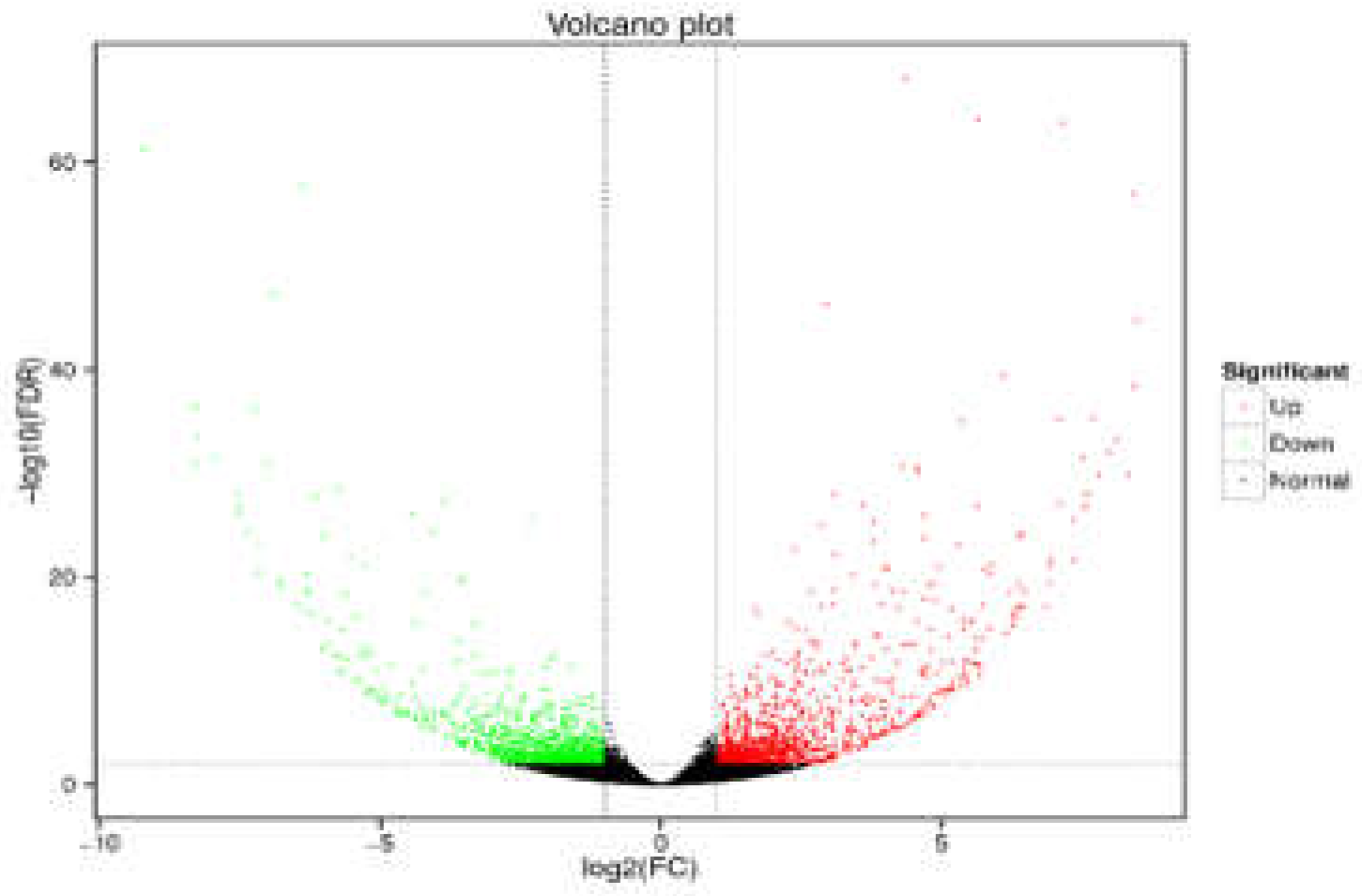
3.1.2. Identification of Differential Metabolites in PFs and WFs
| Index | Fold_Change | Log2FC | Regulation |
|---|---|---|---|
| A | 1504.16 | 10.55 | up |
| B | 2.04 | 1.03 | down |
| C | 1.70 | 0.77 | down |
3.2. Transcriptome Analysis of PFs and WFs
3.2.1. Sequencing and Quality Analysis
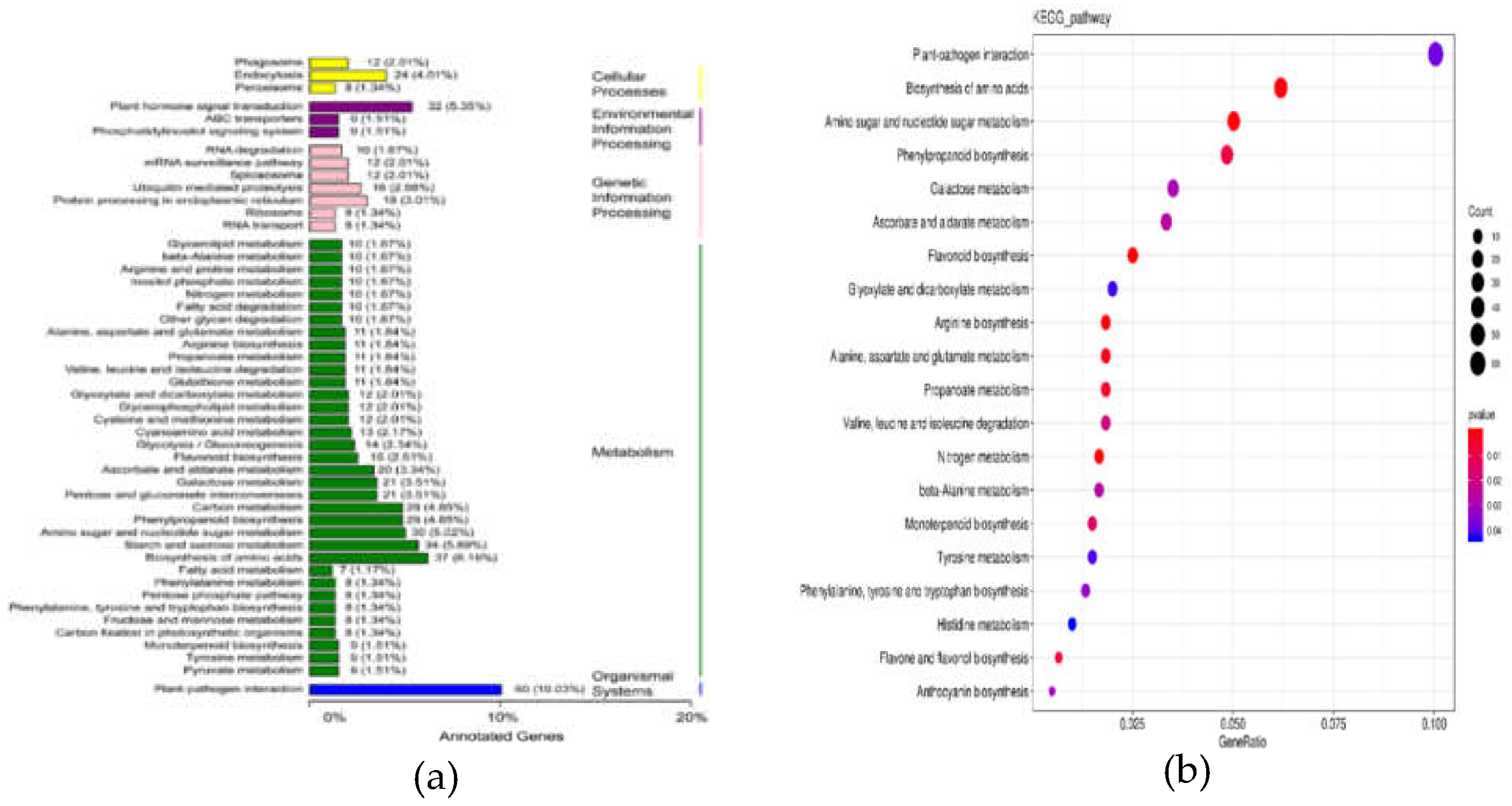
3.2.2. GO Analysis of DEGs
3.2.3. KEGG Analysis of DEGs
3.2.4. Key DEGs Regulating the Floral Color Formation
3.2.5. Key DEGs Regulating Transcription Factors Involved in Floral Color Formation
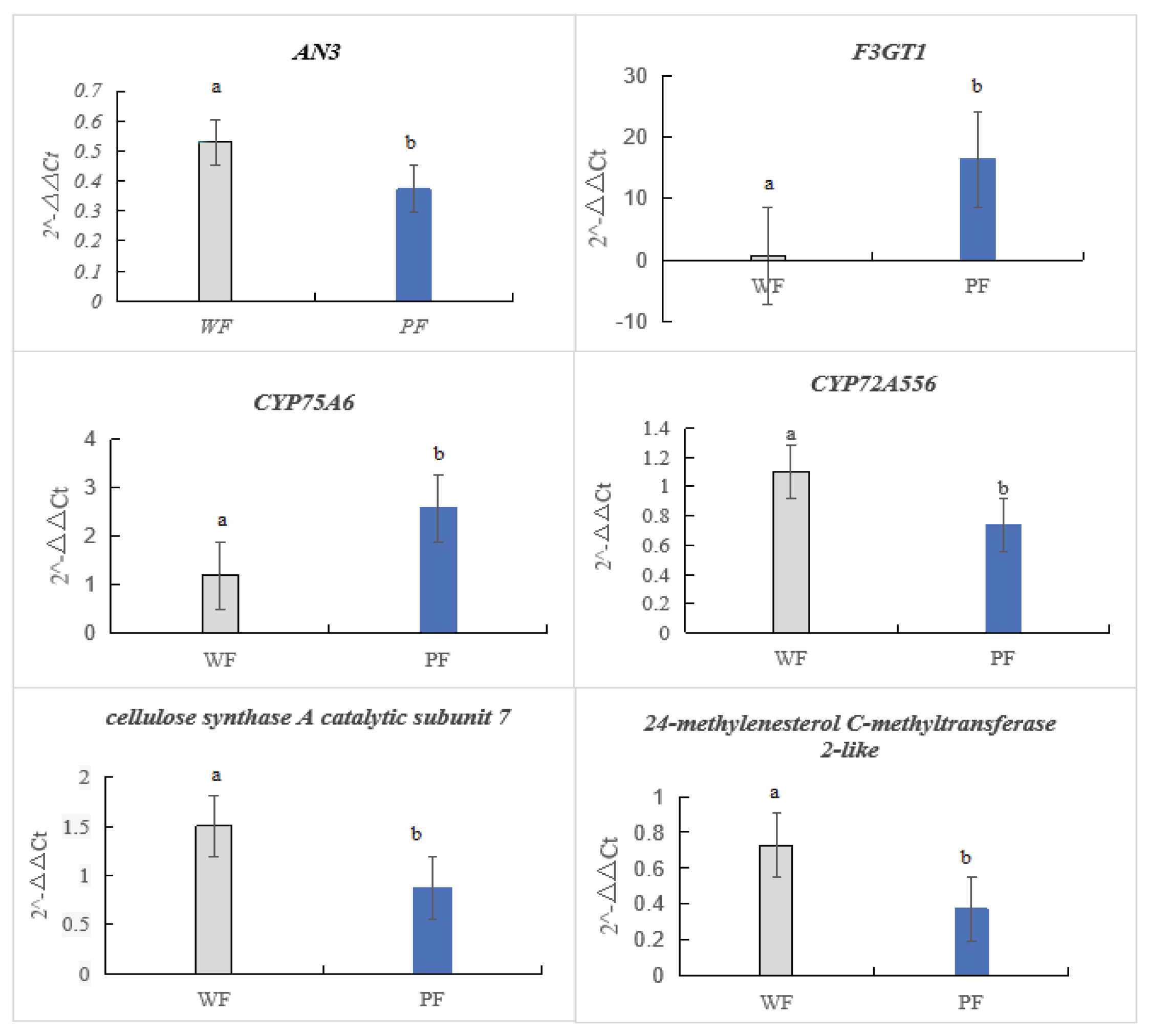
3.2.6. Validation of Transcriptome Sequencing via RT-qPCR
4. Discussion
4.1. Elucidation of the Molecular Mechanism Underlying PFs Color Formation in P.grandiflorus
4.2. Elucidation of the Molecular Mechanism Underlying WFs Color Formation in P.grandiflorus
4.3. Practical Applications of Floral Color Mechanisms in P.grandiflorus Breeding
5. Conclusions
Funding
References
- Van der Kooi, C.J.; Vallejo-Marín, M.; Leonhardt, S. D. Mutualisms and (a) symmetry in plant-pollinator interactions. Current Biology 2021, 31, R91–R99. [Google Scholar] [CrossRef]
- Zhao, D; Tao, J; Han, C; et al. Flower color diversity was revealed by differential expression of flavonoid biosynthesis-related genes and flavonoid accumulation in herbaceous peony (Paeonia lactiflora Pall). Mol Biol Rep. 2012, 39, 11263–11275. [Google Scholar] [CrossRef]
- Zheng, XD; Pan, XX; Meng, XM; et al. Research progress in the determination of carotenoids in Lycium chinense by HPLC. China Fruit Vegetable 2013, 2013.10.013, 1027–1030. [Google Scholar]
- Tanaka, Y; Sasaki, N; Ohmiya, A. Biosynthesis of plant pigments: anthocyanins, betalains and carotenoids. Plant J. 2008, 54, 733–749. [Google Scholar] [CrossRef] [PubMed]
- Dong, N.Q.; Lin, H.X. Contribution of phenylpropanoid metabolism to plant development and plant–environment interactions. J. Integr. Plant Biol. 2021, 63, 180–209. [Google Scholar] [CrossRef] [PubMed]
- Shen, N.; Wang, T.; Gan, Q.; Liu, S.; Wang, L.; Jin, B. Plant flavonoids: classification, distribution, biosynthesis, and antioxidant activity. Food Chem. 2022, 383, 132531. [Google Scholar] [CrossRef]
- Erich, G. The evolution of flavonoids and their genes. In The Science of Flavonoids; Springer Science + Business Media, Inc: New York, 2006. [Google Scholar]
- da Silva, F.L.; Escribano-Bailón, M.T.; Pérez Alonso, J.J.; Rivas-Gonzalo, J.C.; Santos-Buelga, C. Anthocyanin pigment in strawberry. Lebensmittel-Wissenschaft und-Technologie 2007, 40, 374–382. [Google Scholar] [CrossRef]
- Rausher, MD. Evolutionary transitions in floral color. International Journal of Plant Sciences 2008, 169, 7–21. [Google Scholar] [CrossRef]
- Li, Q; He, F; Zhu, B Q; Liu, B; Sun, R Z; Duan, C Q; Reeves, M J; Wang, J. Comparison of distinct transcriptional expression patterns of flavonoid biosynthesis in Cabernet Sauvignon grapes from east and west China. Plant Physiology and Biochemistry 2014, 84, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Mirco, M. Richard V. E, David S, Janine C Paul M. D, Anna S, Ross G.A, Roger P. H, Andrew C. A. Identification and characterization of F3GT1 and F3GGT1, two glycosyltransferases responsible for anthocyanin biosynthesis in red-fleshed kiwifruit (Actinidia chinensis). Plant J 2011, 65, 106–118. [Google Scholar]
- QI, H. Studies on the function and molecular mechanism of anthocyanin in the bluing capability of Hydrangea macrophylla sepals. Chinese Academy of Agricultural Sciences Thesis. Beijing, 2022. [Google Scholar]
- Koes, R; Verweij, W; Quattrocchio, F. Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. TRENDS in Plant Science 2005, 10, 236~242. [Google Scholar] [CrossRef]
- Zhang, L.; Wang, Y.; Yang, D.; Zhang, C.; Zhang, N.; Li, M.; Liu, Y. Platycodon grandiflorus-An ethnopharmacological, phytochemical and pharmacological review. J. Ethnopharmacol. 2015, 164, 147–161. [Google Scholar] [CrossRef]
- Flora of China. Editorial Committee of Flora of China; Science Press: Beijing, 1983.
- Commonly Used Chinese Medicinal Materials; Science Press: Beijing, 1995.
- Ahn, K.S.; Noh, E.J.; Zhao, H.L.; Jung, S.H.; Kang, S.S.; Kim , Y.S. Inhibition of inducible nitric oxide synthase and cyclooxygenaseⅡ by Platycodon grandiflorum saponins via suppression of nuclear factor-κB activation in RAW 264. 7 cells. Life Sciences 2005, 76, 2315–2328. [Google Scholar]
- Lee, K J; You, H J; Park, S J; Kim, Y S; Chung, Y C; Jeong, T C; Jeong, H G. Hepatoprotective effects of Platycodon grandiflorum on acetaminophen-induced liver damage in mice. Cancer Letters 2001, 174, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Kyung J.L, Jae H.C, Hyung G. K, Eun H. Han, Yong P. H, Young C. L, Young C. C, Hye G. J. Protective effect of saponins derived from the roots of Platycodon grandiflorum against carbon tetrachloride induced hepatotoxicity in mice. Food and Chemical Toxicology 2008, 46, 1778–1785. [CrossRef] [PubMed]
- Zhang, H. Y; Hu, X. L; Li, H. J.; Dan, Z. Cultivation Techniques of Platycodon grandiflorus for Suitable Climate, High Quality, and High Yield. Shaanxi Agricultural Sciences 2011, (2), 255–257. [Google Scholar]
- Qi, S. Y; Yan, Y. Z; Jin, Z.; Wu, S. Q. Study on Differential Gene Expression during the Development of White and Purple Flowers of Platycodon grandifloras. Crops 2024, 06, 54–65. [Google Scholar]
- Lou, Q; Wang, L; Liu, H; et al. Anthocyanin profiles in flowers of grape hyacinth. Molecules 2017, 22, 688. [Google Scholar] [CrossRef]
- Winkel-Shirley, B. Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiology 2001, 126, 485–493. [Google Scholar] [CrossRef]
- Nakatsuka, T; Sasaki, N; Nishihara, M. Transcriptional regulators of flavonoid biosynthesis and their application to flower color modification in Japanese gentians. Plant Biotechnology 2014, 31, 389–399. [Google Scholar] [CrossRef]
- Xin, Y. K; Ke Yu, Z; Mei, Y. T; Xue, R. Y; Wen, Q. W. bHLH291 Regulates Anthocyanin Accumulation in Kiwifruit Under Bagging Treatment. Horti-culturae 2025, 11, 666. [Google Scholar]
- Yan, H. L; Pei, X. N; Zhang, H; Li, X; Zhang, X.X; Zha, M. H; Chiang, V.L.; Sederoff, R.R.; Zhao, X. Y. MYB-Mediated regulation of anthocyanin biosynthesis. International journal of molecular sciences 2021, 22, 3103. [Google Scholar] [CrossRef] [PubMed]
- Gao, J. H; Dou, Y. W; Wang, X. T; Zhang, D. L; Wei, M; Li, Y. Transcriptome analysis reveals the mechanism for blue-light–induced biosynthesis of delphinidin derivatives in harvested purple pepper fruit. Frontiers in Plant Science 2023, 1289120. [Google Scholar] [CrossRef]
- Luo, J; Chen, L. Q; Li, T. J; Ou, Z; Qu, Y. Analysis on effects of metal ions on the petal color of three species in Meconopsis based on transcriptome. Journal of Plant Resources and Environment 2023, 32, 16–27. [Google Scholar]
- Sun, Y; Hong, L; Huang, J R. Arabidopsis TT19 functions as a carrier to transport anthocyanin from the cytosol to tonoplasts. Molecular Plant. 2011, 5, 387–400. [Google Scholar] [CrossRef]
- Liu, G. L. Study on the color mechanism of Iris sanguinea Donn ex Horn. with different flower colors regulated by IsMYBL1 and IsMYBL2; Northeast Forestry University, 2024. [Google Scholar]
- Stushnoff, C.; Ducreux, L. J.; Hancock, R. D.; Hedley, P. E.; Holm, D. G.; McDougall, G. J. Flavonoid profiling and transcriptome analysis reveals new gene-metabolite correlations in tubers of Solanum tuberosum L. J. Exp. Bot. 2010, 61, 1225–1238. [Google Scholar] [CrossRef]
- Lloyd, A; Brockman, A; Aguirre, L; Campbell, A; Bean, A; Cantero, A; Gonzalez, A. Advances in the MYB-bHLH-WD repeat (MBW) pigment regulatory model: addition of a WRKY factor and Co-option of an anthocyanin MYB for Betalain Regulation. Plant Cell Physiology 2017, 58, 1431–1441. [Google Scholar] [CrossRef] [PubMed]
- Wang, L. H; Tang, W; Hu, Y. W; Zhang, Y. B; Sun, J. Q; Guo, X. H; et al. A MYB/bHLH complex regulates tissue-specific anthocyanin biosynthesis in the inner pericarp of red-centered kiwifruit Actinidia chinensis cv. Hongyang. The Plant journal: for cell and molecular biology 2019, 99, 359–378. [Google Scholar] [CrossRef] [PubMed]
- He, Y.J; Wang, Z. W; Ge, H. Y; Liu, Y; Chen, H. Weighted gene co-expression network analysis identifies genes related to anthocyanin biosynthesis and functional verification of hub gene SmWRKY44. Plant Science 2021, 309, 110935. [Google Scholar] [CrossRef]
- Jie, Yang. Transcriptional regulation mechanism of anthocyanin biosynthesis in oriental hybrid lily; Beijing Forestry University: Beijing, 2022. [Google Scholar]
- Lu, M. Y. Discovery and functional validation of the key gene PgF3', 5'H in the biosynthetic pathway of Platycodon grandiflorus blue flower. Master’s Thesis, Anhui University of Chinese Medicine, 2023. [Google Scholar]
- Le Maitre, N.C.; Pirie, M.D.; Bellstedt, D.U. Floral Color, Anthocyanin Synthesis Gene Expression and Control in Cape Erica Species. Frontiers in Plant Science 2019, 11, 1565. [Google Scholar] [CrossRef]
- Liu, H. F; Chen, F. B. Candidate genes in red pigment biosynthesis of a red-fleshed radish cultivar (Raphanus sativus L.) as revealed by transcriptome analysis. Biochemical Systematics and Ecology 2019, 86, 103933. [Google Scholar] [CrossRef]
- Ma, Y. T; Ma, L; Xu, L; Wei, R. N; Chen, G. P; Dang, J. H; Chen, Z; Ma, S. Y; Li, S. Research on the Mechanism of Growth of Codonopsis pilosula (Franch.) Nannf. Root Responding to Phenolic Stress Induced by Benzoic Acid. Inter-national Journal of Molecular Sciences 2024, 25, 11007. [Google Scholar] [CrossRef] [PubMed]
| Sample | Total | Clean reads number | Error rate (%) |
Mapped ratio (%) |
Q20 base percentage(%) | Q30 base percentage(%) |
|---|---|---|---|---|---|---|
| WFs-1 | 48,343,934 | 24,171,967 | <0.05 | 93.28% | 97.95 | 94.21 |
| WFs-2 | 42,370,336 | 21,185,168 | <0.05 | 92.55% | 97.94 | 94.21 |
| WFs-3 | 39,492,676 | 19,746,338 | <0.05 | 93.87% | 98.09 | 94.57 |
| PFs-1 | 55,798,134 | 27,899,067 | <0.05 | 92.59% | 97.83 | 93.98 |
| PFs-2 | 43,820,534 | 27,899,067 | <0.05 | 93.01% | 97.9 | 94.16 |
| PFs-3 | 43,820,534 | 21,655,204 | <0.05 | 93.30% | 97.99 | 94.35 |
| DEG Set | Total | COG | GO | KEGG | KOG | NR | Pfam | Swiss-Prot | eggNOG |
|---|---|---|---|---|---|---|---|---|---|
| WFs vs. PFs | 1655 | 604 | 1388 | 1197 | 926 | 1632 | 1334 | 1224 | 1433 |
| Gene Name | log₂FC | FDR | Regulation | Function Annotation |
|---|---|---|---|---|
| CYP98A2 | -2.41 | 4.33E-06 | down | 5-O-(4-coumaroyl)-D-quinate 3'-monooxygenase |
| BGLU18 | -2.03 | 4.49E-09 | down | Beta-glucosidase 18 |
| CYP73A12 | -2.53 | 0.000734427 | down | Trans-cinnamate 4-monooxygenase |
| AN3 | 6.15 | 1.99E-28 | up | naringenin,2-oxoglutarate 3-dioxygenase |
| DFR | 5.75 | 3.78E-29 | up | dihydroflavonol 4-reductase |
| CYP75A6 | 6.03 | 1.17E-13 | up | flavonoid-3',5'-hydroxylase |
| UGT29 | 2.44 | 6.75E-09 | up | UDP-glucosyltransferase 29-like |
| 3MAT | 2.02 | 0.001172479 | up | anthocyanin 3-O-glucoside-6''-O-malonyltransferase |
| F3GT1 | 6.32 | 1.51E-19 | up | anthocyanidin 3-O-glucosyltransferase |
| RT | 4.43 | 1.68E-10 | up | delphinidin 3-O-glucoside |
| HMA5 | 8.22 | 2.91E-34 | up | copper-transporting ATPase HMA5 |
| VPS35B | 4.89 | 1.15426E-08 | up | Vacuolar protein sorting-associated protein 35B |
| Group | Gene Name | log₂FC | FDR | Regulation | Function Annotation | |
|---|---|---|---|---|---|---|
| PFs | MYB4 | 4.71 | 1.23E-07 | up | Myb transcription factor C2 repressor motif protein | |
| bHLH2 | 2.02 | 0.0067 | up | bHLH-MYC and R2R3-MYB transcription factors N-terminal | ||
| WRKY44 | 4.23 | 9.04E-12 | up | WRKY transcription factor 44 | ||
| ARPC1A | 3.13 | 2.78E-07 | up | Anaphase-promoting complex subunit 4 WD40 domain | ||
| WFs | HARBI1 | 4.12 | 1.03E-09 | up | Myb/SANT-like DNA-binding domain | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




