Submitted:
23 January 2026
Posted:
26 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
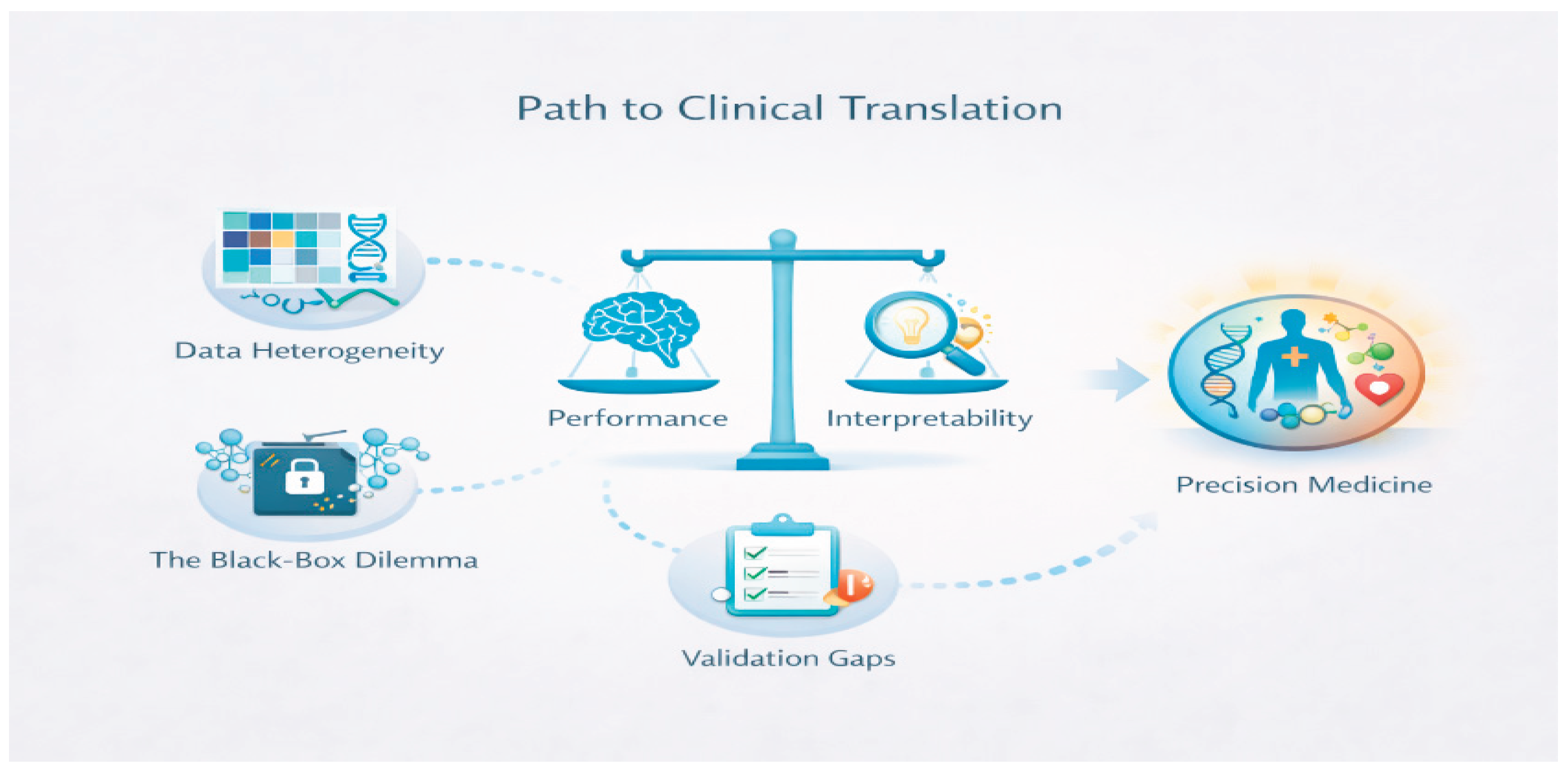
2. Characteristics of Multi-Omics Data and Core Challenges
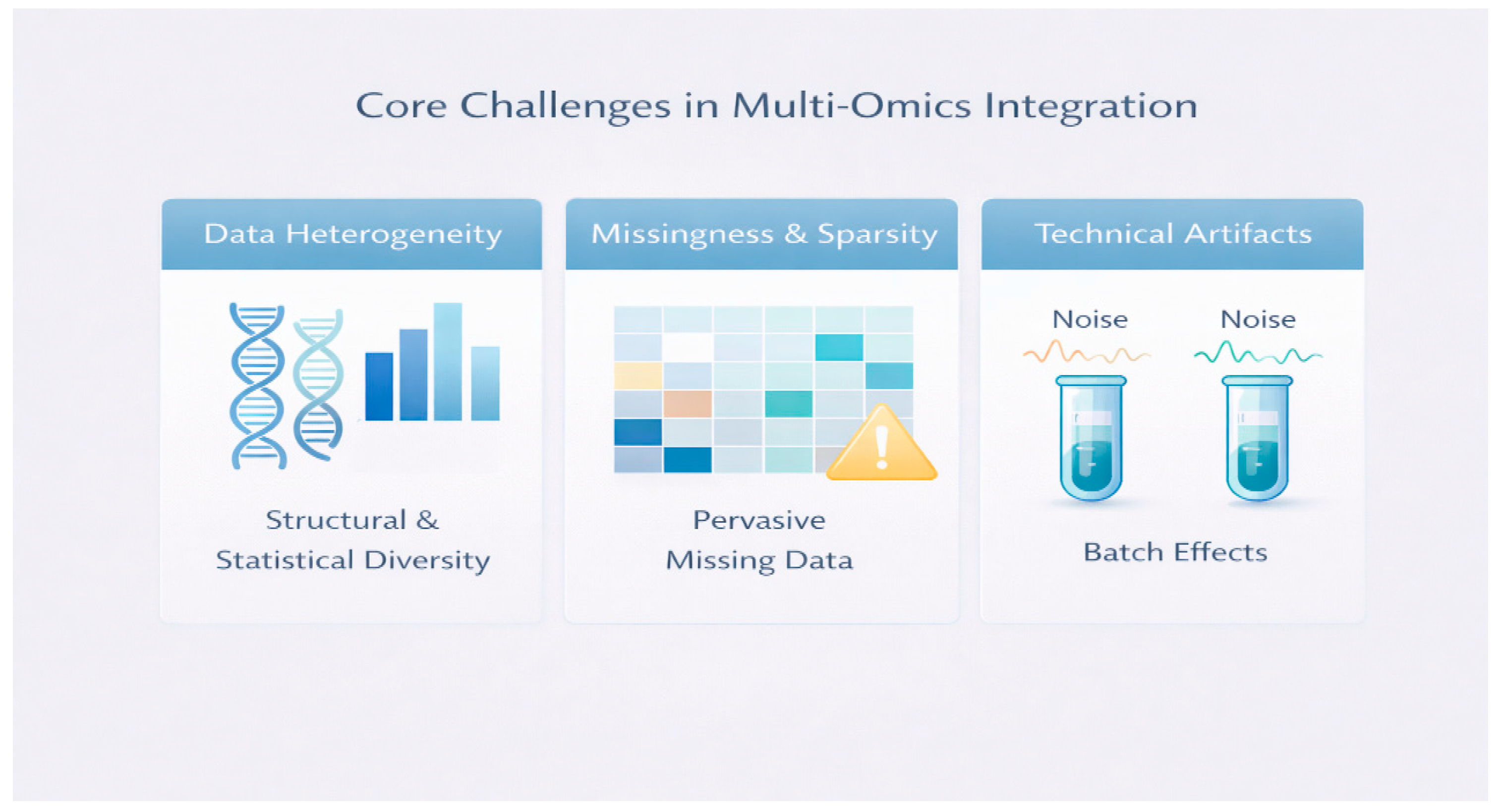
Data Heterogeneity and Statistical Distributions
Missing Data and Sparsity
Technical Artifacts and Batch Effects
3. Deep Learning Architectures for Multi-Omics Integration
Unsupervised Latent-Space Learning
Variational Autoencoders (VAEs)
Generative Models and Domain Adaptation
Network-Based Integration
Graph Neural Networks (GNNs)
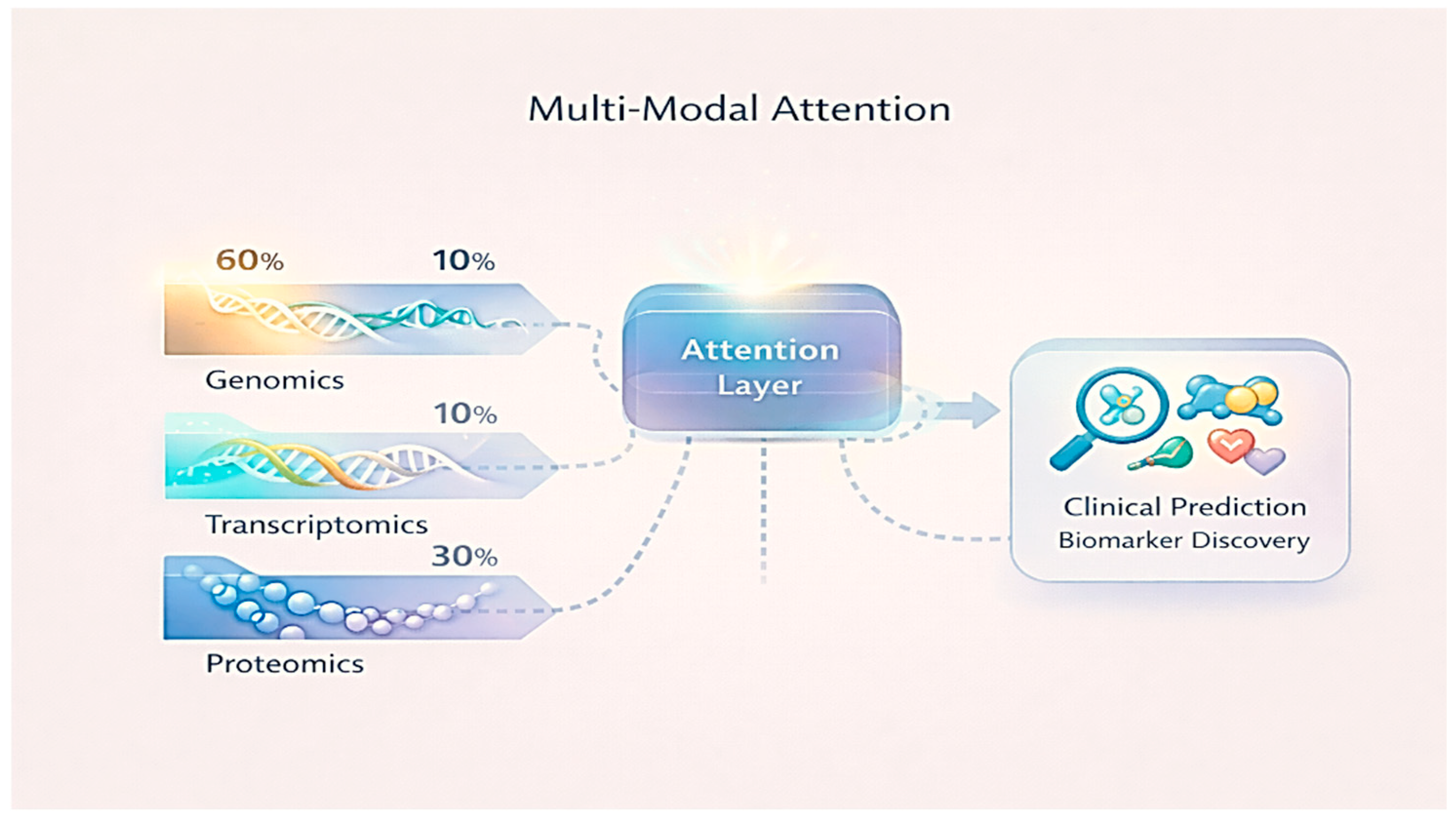
Advanced Supervised Strategies
| Category | Latent-Space Learning | Deconfounding Generative Models | Adversarial Generative Models | Network-Based Integration | Data-Driven Graph Models | Attention-Based Models | Ensemble Prognostic Models | Decentralized Learning Frameworks |
| Fusion Level | Representation-level (latent) | Representation-level (latent) | Feature / representation-level | Network-level | Network-level | Decision-level | Decision-level | Training-level |
| Core Architecture | VAEs, Denoising Autoencoders | Disentangled VAEs, Conditional VAEs | GANs | GNNs (GCN, GAT, GTN) | Learned relational graphs (e.g., MoRE-GNN) | Multi-head attention, Transformers | Autoencoders + classical ML (e.g., DeepProg) | Federated learning, Transfer learning |
| Handling of Missing Data | Explicit (latent-space imputation) | Explicit (signal–confounder separation) | Indirect (distribution alignment) | Limited | Partial | Partial | Indirect | Indirect |
| Use of Biological Priors | Limited / implicit | Partial (constraints) | None | Explicit (PPI, pathways) | Minimal / none | None | None | None |
| Primary Translational Applications | Disease subtyping, classification | Batch-corrected clustering, scMulti-omics | Batch correction, harmonization | Biomarker discovery, subtype analysis | Single-cell multi-omics integration | Prognosis, drug response prediction | Survival stratification, pan-cancer prognosis | Multi-center survival prediction |
| Key Strengths | Robust to noise and sparsity; scalable | Separates biological and technical variation | Improves cross-cohort robustness | Biologically interpretable modules | Adaptive, prior-free modeling | Dynamic modality weighting | Improved robustness via model diversity | Privacy-preserving, improved generalizability |
| Key Limitations | Limited interpretability of latent features | Increased complexity; confounder specification | Training instability; low interpretability | Sensitive to prior knowledge quality | Reduced biological interpretability | Attention ≠ causality | Complex deployment; limited transparency | Communication overhead; data heterogeneity |
4. Translational Applications of Multi-Omics Deep Learning
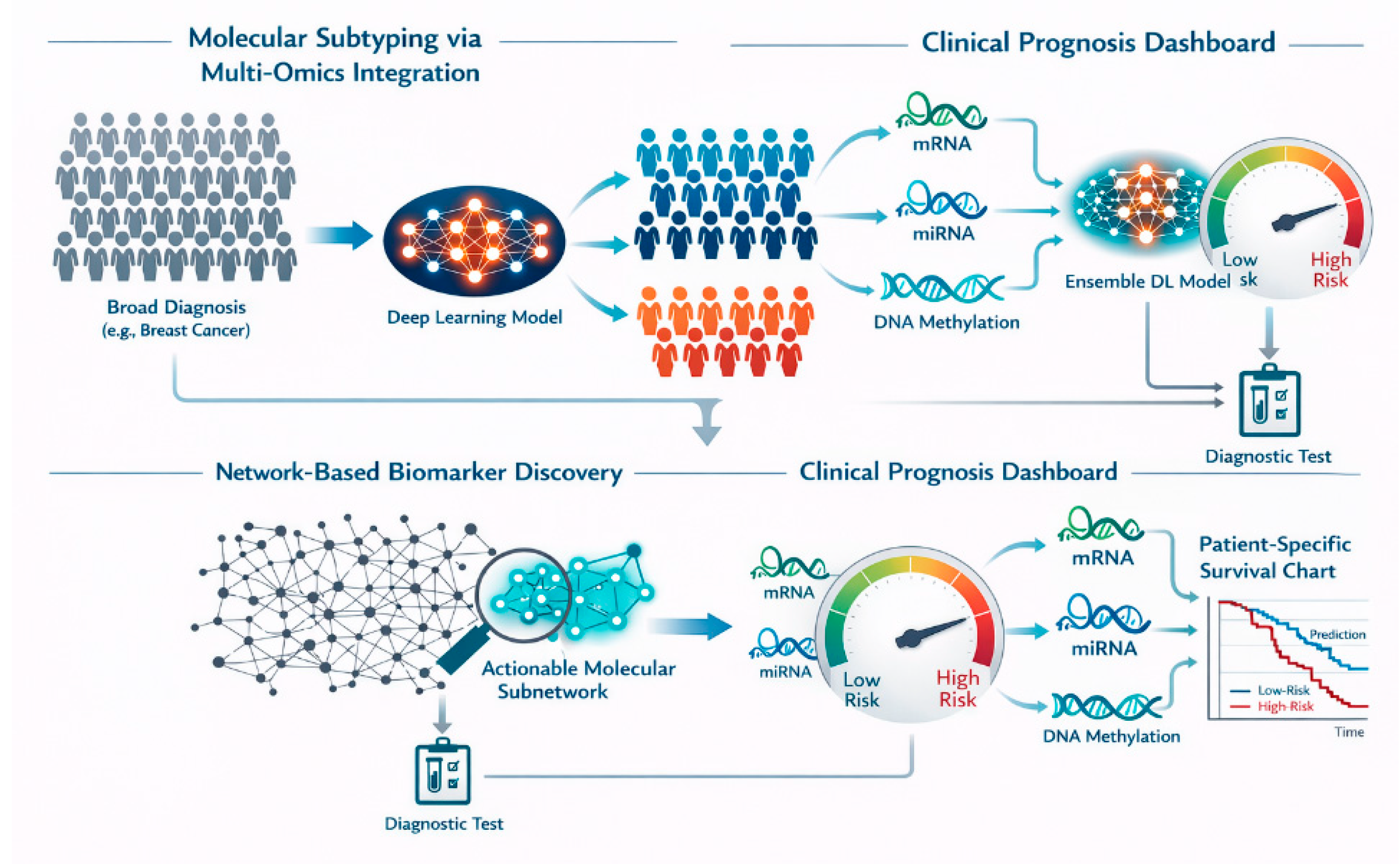
Molecular Subtyping and Disease Classification
Robust Biomarker Discovery
Prognosis and Survival Prediction
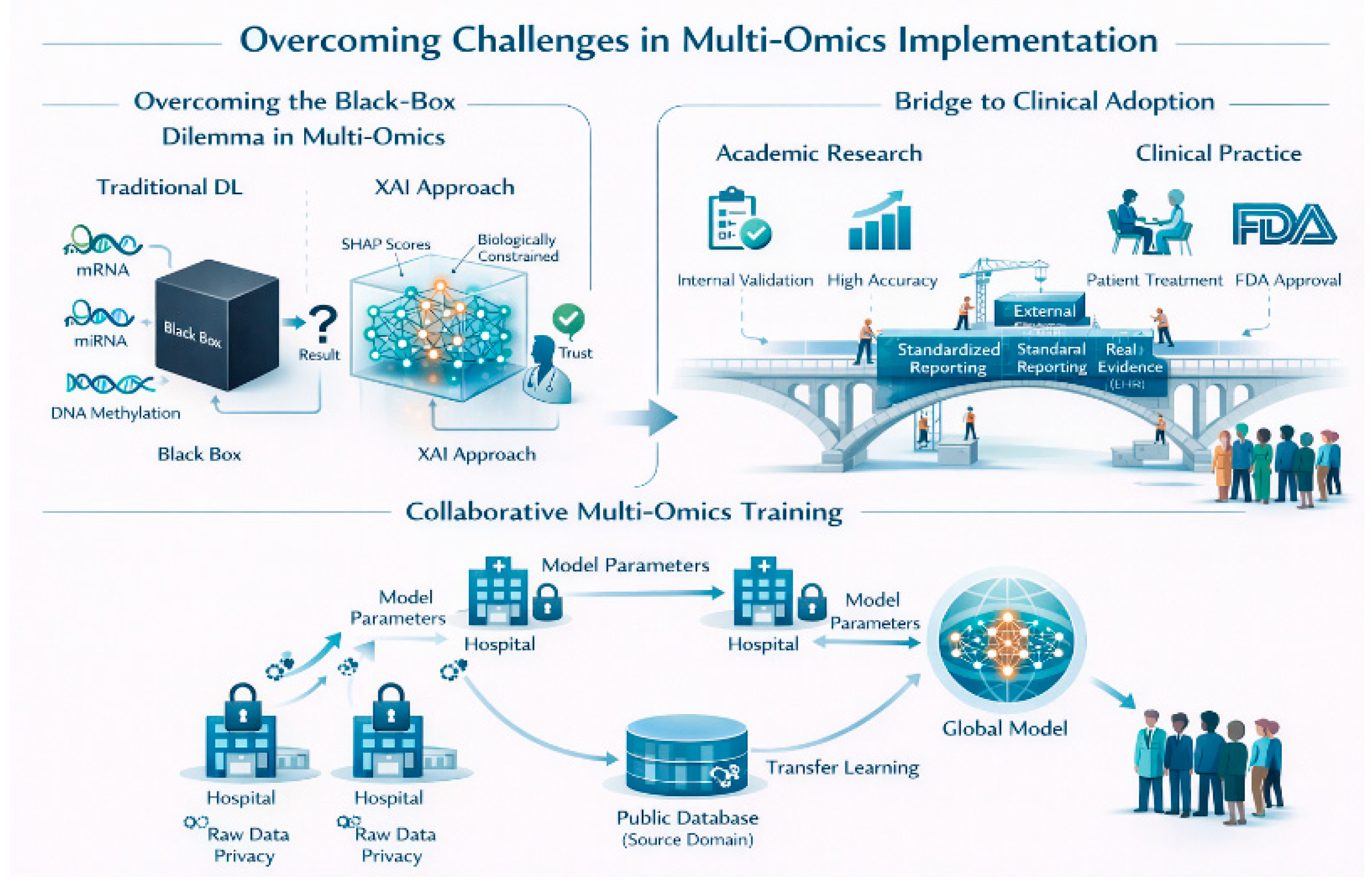
5. Open Challenges and Future Directions
Interpretability–Performance Trade-off
Data Scarcity and Generalizability through Collaborative Learning
Standardized Validation and Clinical Adoption
6. Conclusions
References
- Wang, R.C.; Wang, Z. Precision medicine: Disease subtyping and tailored treatment. Cancers 2023, 15(15), 3837. [Google Scholar] [CrossRef]
- Athieniti, E.; Spyrou, G.M. A guide to multi-omics data collection and integration for translational medicine. Comput. Struct. Biotechnol. J. 2023, 21, 134–149. [Google Scholar] [CrossRef]
- Ballard, J.L.; Wang, Z.; Li, S.; Han, K.; Long, Q. Deep learning-based approaches for multi-omics data integration and analysis. BioData Min. 2024, 17(1), 22. [Google Scholar] [CrossRef] [PubMed]
- A. Yetgin, Revolutionizing multi-omics analysis with artificial intelligence and data processing. Quant. Biol. 2025, 13(3), 232–245.
- Ishida, J.P.; Huang, S.C.; Trostle, J.; Wang, K. A review of multi-omics data integration through deep learning approaches for disease diagnosis, prognosis, and treatment. Front. Genet. 2023, 14, 1199087. [Google Scholar] [CrossRef]
- Chen, C.; Wang, J.; Pan, D.; Lv, Y.; Ding, Y.; Sun, S.; Fan, D.; Liu, W.; Wei, R. Applications of multi-omics analysis in human diseases. MedComm 2023, 4(4), e315. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Sun, T.; Jin, D. Review on the computational burden of single-cell multi-omics data: Sparsity, noise, and scalability. Quant. Biol. 2023, 11(4), 301–315. [Google Scholar]
- Nayak, S.; Khilar, P.M.; Roul, R.K. Deep generative models for multi-omics data imputation: A review, J. Supercomput 2025.
- Rana, M.; Seneviratne, S.; O’Connell, M.J. A technical review of multi-omics data integration methods: From classical statistical to deep generative approaches, Brief. Bioinform. 2025, 26(2), bbaf355. [Google Scholar]
- Kalafut, N.C.; Huang, X.; Wang, D. Joint variational autoencoders for multimodal imputation and embedding. Nat. Mach. Intell. 2023, 5(6), 631–642. [Google Scholar] [CrossRef]
- Liu, J.; Zhang, M.; Li, G. Disentangled representation learning in VAEs for separating biological and technical variance in scRNA-seq. Nat. Methods 2025, 22(1), 11–19. [Google Scholar]
- Huang, Y.; Li, Q.; Wang, J. Deep generative models for synthetic multi-omics data augmentation. Cell Syst. 2024, 15(3), 221–235. [Google Scholar]
- Li, J.; Liu, B.; Wang, Y.; Zhu, Y.; Liu, Q. A multi-omics integration model based on graph convolutional network to analyze and classify cancer subtypes. Front. Genet. 2022, 13, 994087. [Google Scholar]
- Cheng, Y.; Zhao, Y.; Li, M. A hierarchical graph neural network integrating pathways for multi-omics analysis. Bioinform. Adv. 2024, 4(1), vbad150. [Google Scholar]
- Kim, K.; Lee, M.; Park, H. Graph attention networks for robust multi-omics drug sensitivity prediction. Nat. Comput. Sci. 2024, 4(5), 450–461. [Google Scholar]
- Ren, Y.; Gao, Y.; Du, W.; Qiao, W.; Li, W.; Yang, Q.; Liang, Y.; Li, G. Classifying breast cancer using multi-view graph neural network based on multi-omics data. Front. Genet. 2024, 15, 1363896. [Google Scholar] [CrossRef]
- Tan, S.; Yang, C.; Goudie, M.; Zhu, H.; Shi, Q. Enhancing non-small cell lung cancer survival prediction through multi-omics integration using graph attention network. Medicina 2024, 60(1), 2178. [Google Scholar]
- Wang, Y.; Li, Q.; Zhang, H. Benchmarking deep learning models for multi-omics integration on pan-cancer datasets. Genome Med. 2024, 16(1), 78. [Google Scholar]
- Park, T.; Kim, S.; Lee, J. Multi-omics model for early prediction of Alzheimer’s disease using deep learning. Alzheimers Dement. 2024, 20(4), 1–12. [Google Scholar]
- Wen, Y.; Zheng, L.; Leng, D.; Dai, C.; Lu, J.; Zhang, Z.; He, S.; Bo, X. Deep learning-based multiomics data integration methods for biomedical application. Adv. Intell. Syst. 2023, 5(5), 2200247. [Google Scholar] [CrossRef]
- Hussein, A.; Prasad, M.; Braytee, A. Explainable AI methods for multi-omics analysis: A survey. ACM Trans. Comput. Surv. 2024, 57(1), 1–39. [Google Scholar]
- Zhu, X.; Zhang, Y.; Wang, J. Explainable AI methods for multi-omics analysis: A survey. J. Biomed. Inform. 2024, 15(1), 384973895. [Google Scholar]
- Xie, X.; Liu, B.; Zhang, S. SHAP-based feature importance for graph neural networks in multi-omics biomarker discovery. Bioinformatics 2025, 41(2), btad899. [Google Scholar]
- Han, K.; Chen, Y.; Lee, S. Visible neural networks for multi-omics integration: A critical review. Front. Artif. Intell. 2025, 8, 1595291. [Google Scholar] [CrossRef]
- Andreux, M.; Wang, Y.; Yu, X.; N’guessan, A.; El-Haddad, R.; Sénécal, F.; Alimonti, J. Federated transfer learning with differential privacy for multi-omics survival analysis, Brief. Bioinform 2025, 26(2), bbaf555. [Google Scholar]
- Chen, S.; Li, M.; Zhao, H. AFEI: Adaptive optimized vertical federated learning for heterogeneous multi-omics data integration, Brief. Bioinform 2023, 24(10), bbac341. [Google Scholar]
- Zhang, Y.; Liu, Y.; Wang, B. Federated transfer learning with differential privacy for multi-omics survival analysis, Brief. Bioinform. 2025, 26(2), 390793804. [Google Scholar]
- Li, Q.; Zhang, Y.; Liu, B. Ethical and privacy challenges in federated learning for genomic data. Sci. Adv. 2023, 9(38), eadh4775. [Google Scholar]
- Cui, R.; Han, K.; Long, Q. A systematic review of transfer learning methods for multi-omics data. Bioinformatics 2023, 39(12), btad732. [Google Scholar]
- Qiao, W.; Wang, Y.; Zhang, X.; Zhou, Y. Foundation models in biology: A review on multi-omics data. Cell Rep. Phys. Sci. 2023, 4(10), 101569. [Google Scholar]
- Song, Y.; Zhang, W.; Sun, Y. Integrating electronic health records as a clinical ‘omic’ layer using deep learning. Nat. Biomed. Eng. 2024, 8(2), 150–162. [Google Scholar]
- Zhao, Y.; Shao, J.; Asmann, Y.W. Assessment and optimization of explainable machine learning models applied to transcriptomic data. Genomics Proteomics Bioinformatics 2022, 20(4), 899–911. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




