Submitted:
23 January 2026
Posted:
26 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. Cell Annotation
2.3. Scissor Analysis
2.4. CIBERSORTx Analysis
2.5. WGCNA
2.6. Identification of Biomarkers Related to RF Progression in Key Cell Subtypes
2.7. Cell Communication and Pseudotime Trajectory Analyses
2.8. Functional Enrichment Analysis
2.9. Animal Model
2.10. Quantitative PCR (qPCR)
2.11. Histological Analysis
2.12. Drug Prediction and Molecular Docking
2.13. Statistical Analysis
3. Results
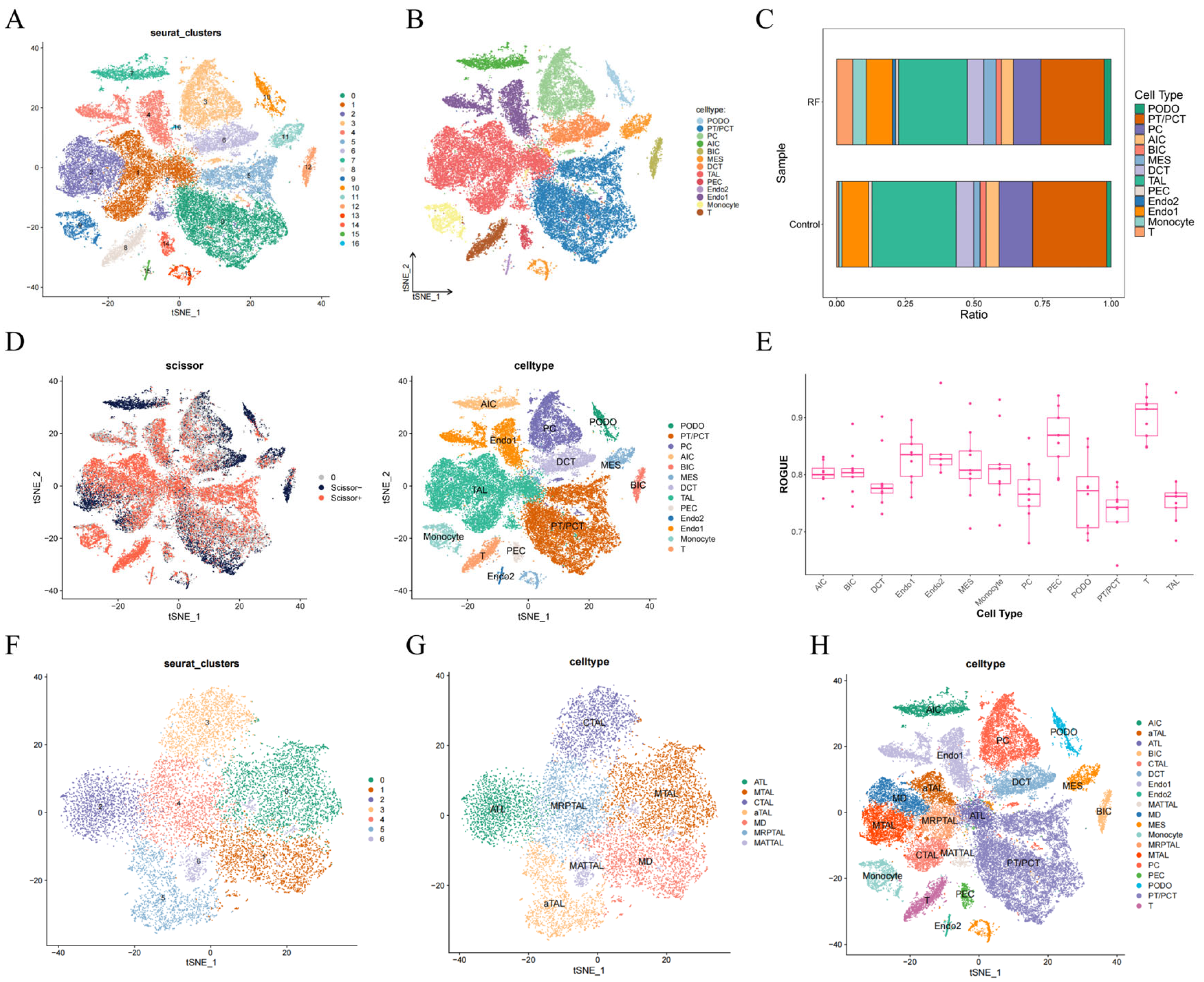
3.1. Single-Cell Atlas Reveal RF-Associated Renal Cell Heterogeneity and Functional Differentiation of Thick Ascending Limb (TAL) Cells
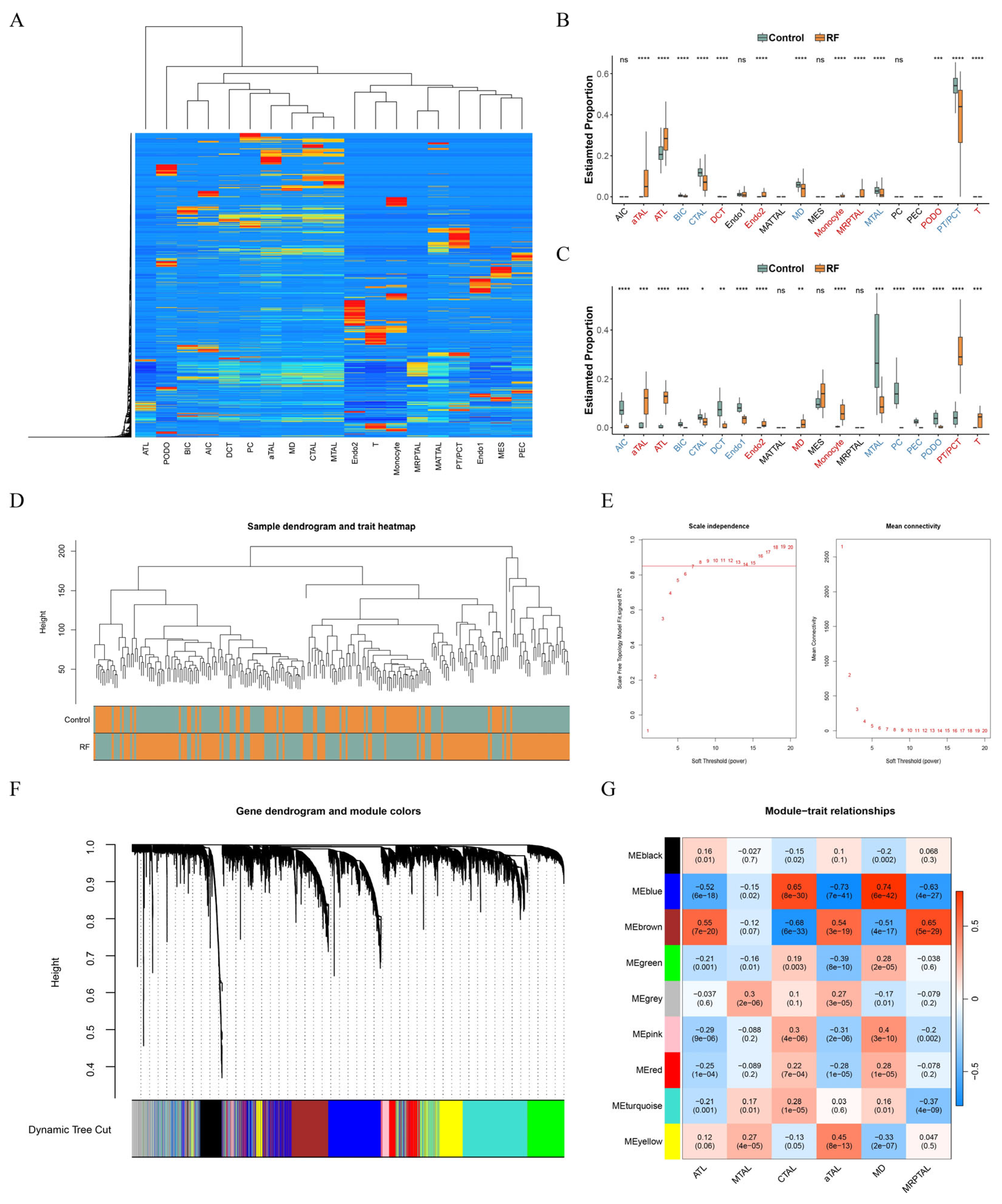
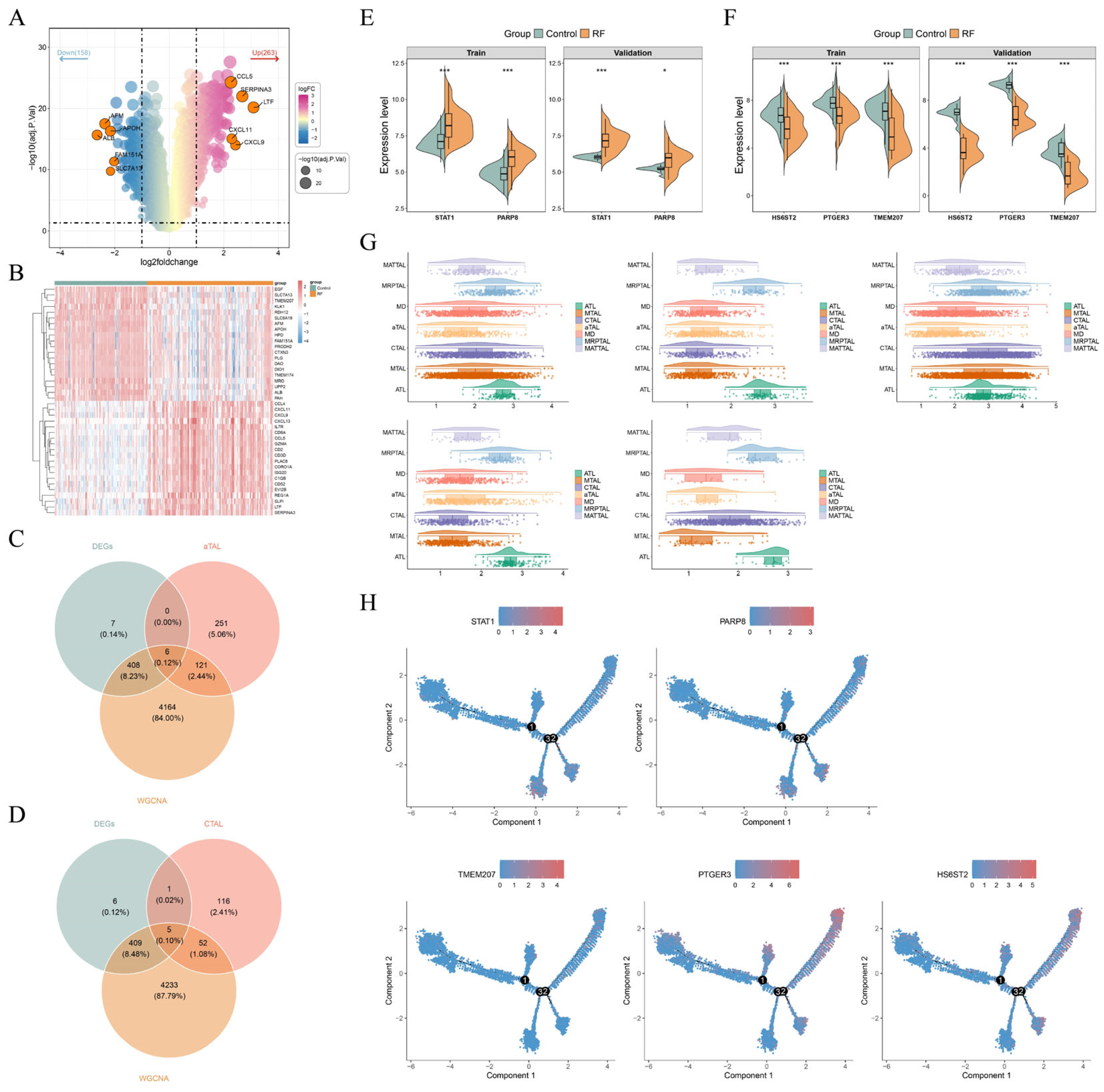
3.2. CTAL and aTAL Were Identified as the Key Cell Subtypes
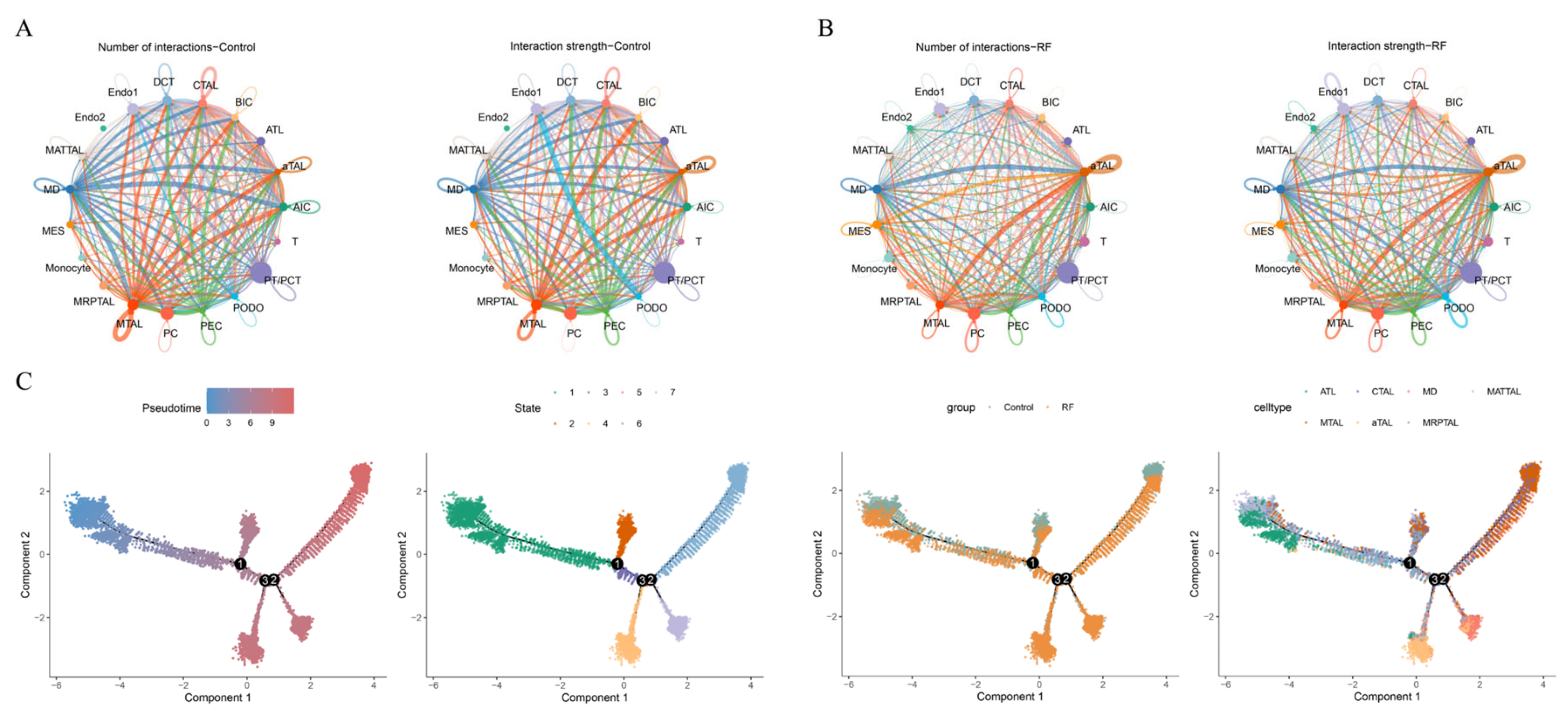
3.3. Cell Communication and Pseudotime Analysis of TAL Subtypes
3.4. STAT1, PARP8, HS6ST2, PTGER3, and TMEM207 Were Biomarkers for RF
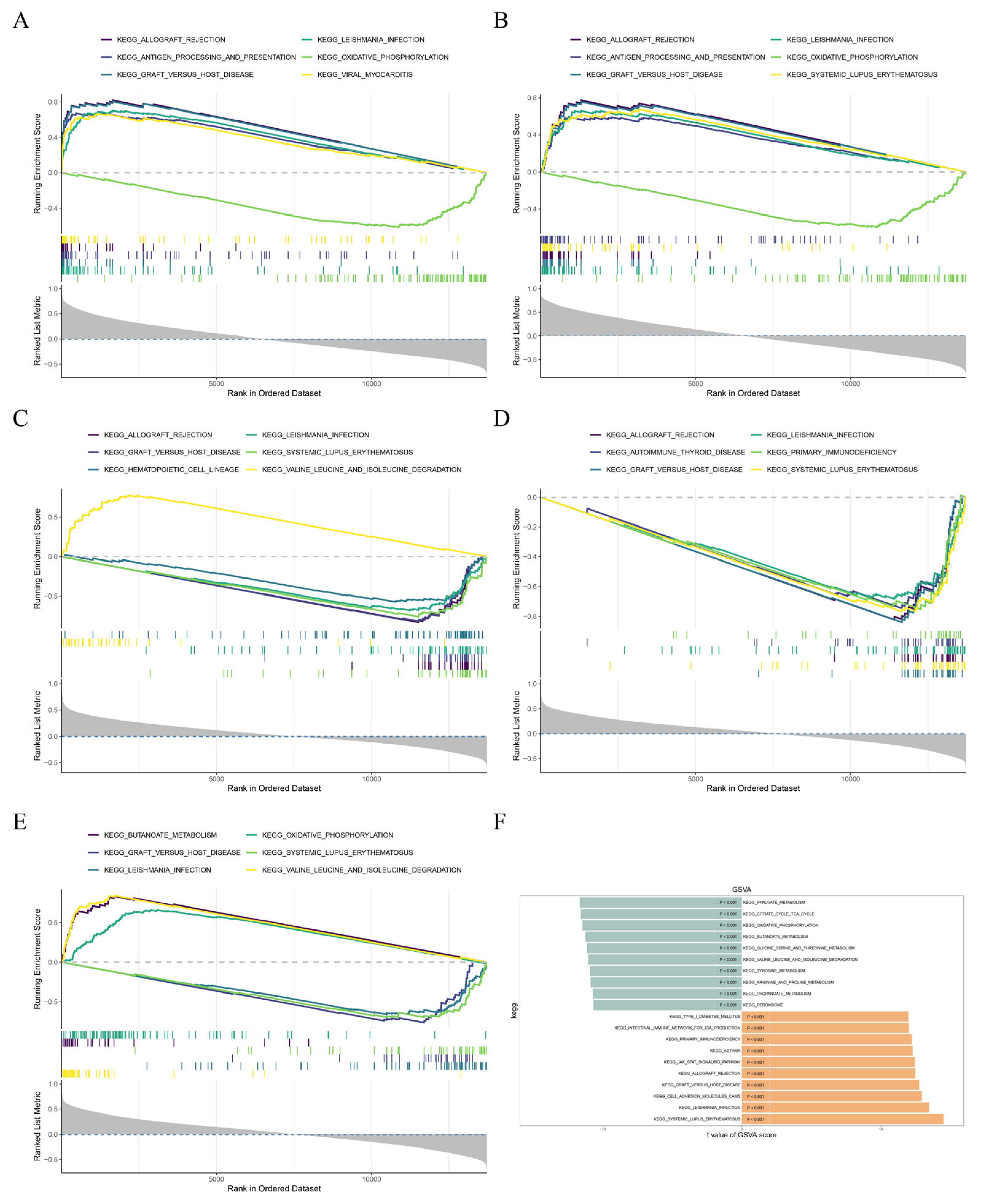
3.5. The Biomarkers Were Enriched in Metabolism and Immune Dysregulation Pathways
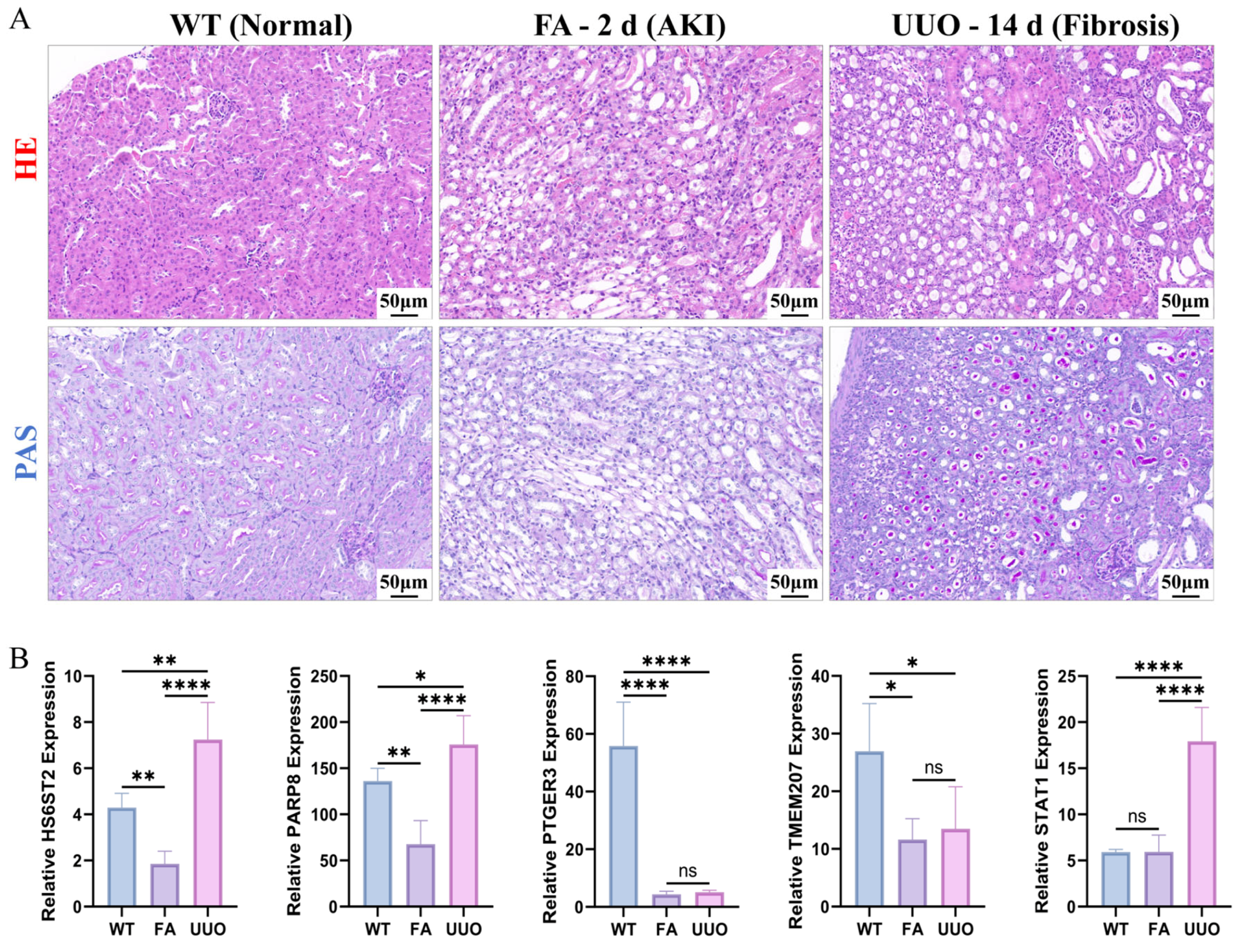
3.6. Experimental Validation of Elevated Expression of Hub Biomarkers in a Murine Model of Renal Fibrosis
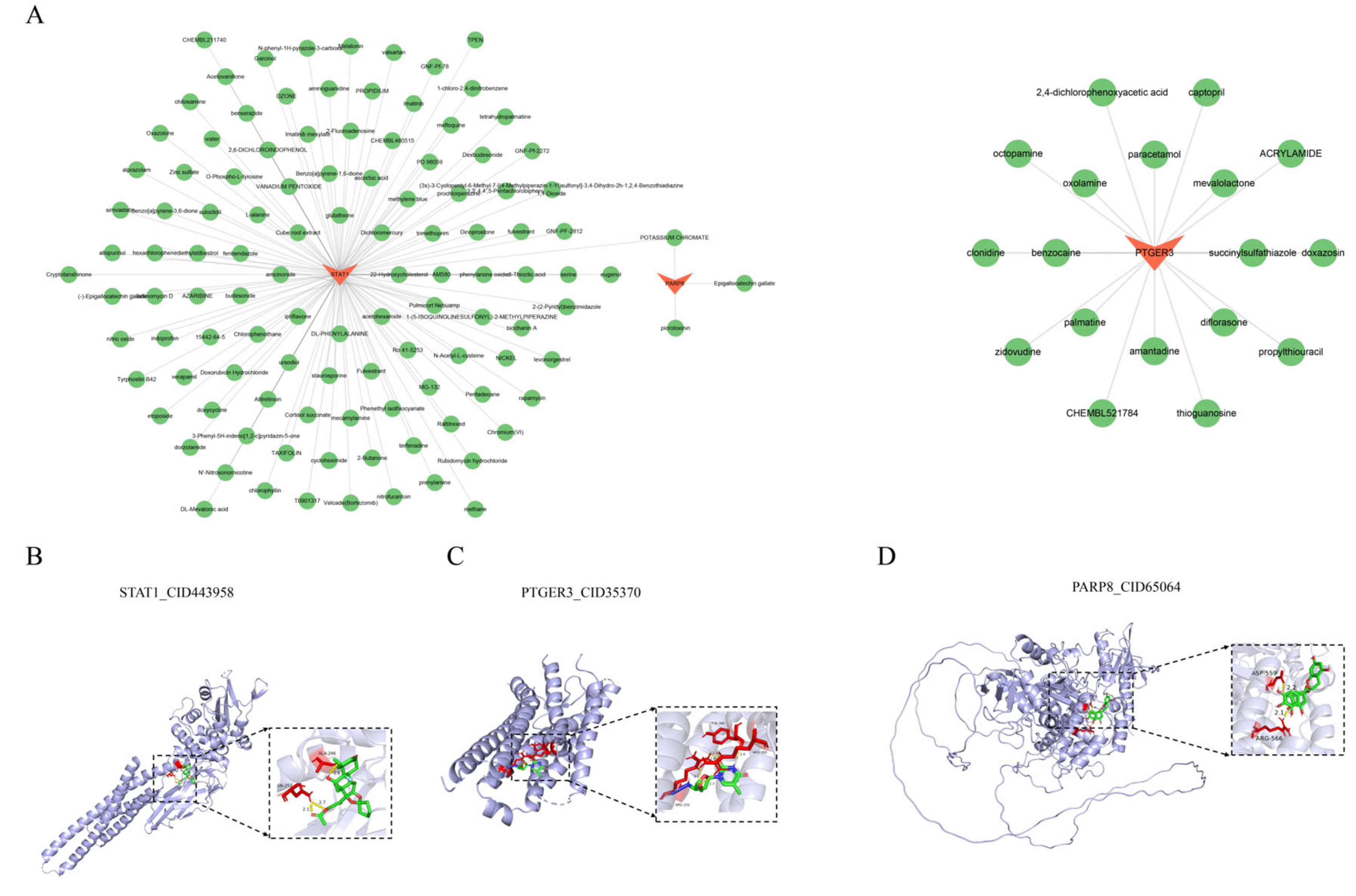
3.7. The Biomarkers Exhibited Strong Binding Affinity with Their Targeted Drugs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Huang, R.; Fu, P.; Ma, L. Kidney fibrosis: from mechanisms to therapeutic medicines. Signal Transduct. Target. Ther. 2023, 8, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Lv, W.; Booz, G.W.; Wang, Y.; Fan, F.; Roman, R.J. Inflammation and renal fibrosis: Recent developments on key signaling molecules as potential therapeutic targets. Eur. J. Pharmacol. 2018, 820, 65–76. [Google Scholar] [CrossRef] [PubMed]
- Ammirati, A.L. Chronic Kidney Disease. Rev Assoc Med Bras 2020, 66, s03–s09. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Fu, H.; Liu, Y. The fibrogenic niche in kidney fibrosis: components and mechanisms. Nat. Rev. Nephrol. 2022, 18, 545–557. [Google Scholar] [CrossRef]
- Guo, Y.; Cen, K.; Hong, K.; Mai, Y.; Jiang, M. Construction of a neural network diagnostic model for renal fibrosis and investigation of immune infiltration characteristics. Front. Immunol. 2023, 14, 1183088. [Google Scholar] [CrossRef]
- Jovic, D.; Liang, X.; Zeng, H.; Lin, L.; Xu, F.; Luo, Y. Single-cell RNA sequencing technologies and applications: A brief overview. Clin. Transl. Med. 2022, 12, e694. [Google Scholar] [CrossRef]
- Xie, S.; Cai, Y.; Chen, D.; Xiang, Y.; Cai, W.; Mao, J.; Ye, J. Single-cell transcriptome analysis reveals heterogeneity and convergence of the tumor microenvironment in colorectal cancer. Front. Immunol. 2023, 13, 1003419. [Google Scholar] [CrossRef]
- Valenzi, E.; Bulik, M.; Tabib, T.; Morse, C.; Sembrat, J.; Bittar, H.T.; Rojas, M.; Lafyatis, R. Single-cell analysis reveals fibroblast heterogeneity and myofibroblasts in systemic sclerosis-associated interstitial lung disease. Ann. Rheum. Dis. 2019, 78, 1379–1387. [Google Scholar] [CrossRef]
- Sun, D.; Guan, X.; Moran, A.E.; Wu, L.-Y.; Qian, D.Z.; Schedin, P.; Dai, M.-S.; Danilov, A.V.; Alumkal, J.J.; Adey, A.C.; et al. Identifying phenotype-associated subpopulations by integrating bulk and single-cell sequencing data. Nat. Biotechnol. 2021, 40, 527–538. [Google Scholar] [CrossRef]
- Newman, A.M.; Steen, C.B.; Liu, C.L.; Gentles, A.J.; Chaudhuri, A.A.; Scherer, F.; Khodadoust, M.S.; Esfahani, M.S.; Luca, B.A.; Steiner, D.; et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 2019, 37, 773–782. [Google Scholar] [CrossRef]
- Zhu, H.; Luo, H.; Skaug, B.; Tabib, T.; Li, Y.-N.; Tao, Y.; Matei, A.-E.; Lyons, M.A.; Schett, G.; Lafyatis, R.; et al. Fibroblast Subpopulations in Systemic Sclerosis: Functional Implications of Individual Subpopulations and Correlations with Clinical Features. J. Investig. Dermatol. 2023, 144, 1251–1261.e13. [Google Scholar] [CrossRef] [PubMed]
- McDaniels, J.M.; Shetty, A.C.; Rousselle, T.V.; Bardhi, E.; Maluf, D.G.; Mas, V.R. The cellular landscape of the normal kidney allograft: Main players balancing the alloimmune response. Front. Transplant. 2022, 1. [Google Scholar] [CrossRef]
- Modena, B.D.; Kurian, S.M.; Gaber, L.W.; Waalen, J.; Su, A.I.; Gelbart, T.; Mondala, T.S.; Head, S.R.; Papp, S.; Heilman, R.; et al. Gene Expression in Biopsies of Acute Rejection and Interstitial Fibrosis/Tubular Atrophy Reveals Highly Shared Mechanisms That Correlate With Worse Long-Term Outcomes. Am. J. Transplant. 2016, 16, 1982–1998. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Tolosa, M.F.; Zhang, T.; Goru, S.K.; Severino, L.U.; Misra, P.S.; McEvoy, C.M.; Caldwell, L.; Szeto, S.G.; Gao, F.; et al. Myofibroblast YAP/TAZ activation is a key step in organ fibrogenesis. J. Clin. Investig. 2022, 7. [Google Scholar] [CrossRef] [PubMed]
- Hao, Y.; Hao, S.; Andersen-Nissen, E.; Mauck, W.M., 3rd; Zheng, S.; Butler, A.; Lee, M.J.; Wilk, A.J.; Darby, C.; Zager, M.; et al. Integrated analysis of multimodal single-cell data. Cell 2021, 184, 3573–3587.e29. [Google Scholar] [CrossRef]
- Tsai, Y.-C.; Kuo, M.-C.; Huang, J.-C.; Chang, W.-A.; Wu, L.-Y.; Huang, Y.-C.; Chang, C.-Y.; Lee, S.-C.; Hsu, Y.-L. Single-cell transcriptomic profiles in the pathophysiology within the microenvironment of early diabetic kidney disease. Cell Death Dis. 2023, 14, 1–13. [Google Scholar] [CrossRef]
- Wilson, P.C.; Wu, H.; Kirita, Y.; Uchimura, K.; Ledru, N.; Rennke, H.G.; Welling, P.A.; Waikar, S.S.; Humphreys, B.D. The single-cell transcriptomic landscape of early human diabetic nephropathy. Proc. Natl. Acad. Sci. USA 2019, 116, 19619–19625. [Google Scholar] [CrossRef]
- Lu, Y.-A.; Liao, C.-T.; Raybould, R.; Talabani, B.; Grigorieva, I.; Szomolay, B.; Bowen, T.; Andrews, R.; Taylor, P.R.; Fraser, D. Single-Nucleus RNA Sequencing Identifies New Classes of Proximal Tubular Epithelial Cells in Kidney Fibrosis. J. Am. Soc. Nephrol. 2021, 32, 2501–2516. [Google Scholar] [CrossRef]
- Chen, Z.; Ye, L.; Zhu, M.; Xia, C.; Fan, J.; Chen, H.; Li, Z.; Mou, S. Single cell multi-omics of fibrotic kidney reveal epigenetic regulation of antioxidation and apoptosis within proximal tubule. Cell. Mol. Life Sci. 2024, 81, 1–16. [Google Scholar] [CrossRef]
- Doke, T.; Abedini, A.; Aldridge, D.L.; Yang, Y.-W.; Park, J.; Hernandez, C.M.; Balzer, M.S.; Shrestra, R.; Coppock, G.; Rico, J.M.I.; et al. Single-cell analysis identifies the interaction of altered renal tubules with basophils orchestrating kidney fibrosis. Nat. Immunol. 2022, 23, 947–959. [Google Scholar] [CrossRef]
- McGinnis, C.S.; Murrow, L.M.; Gartner, Z.J. DoubletFinder: Doublet Detection in Single-Cell RNA Sequencing Data Using Artificial Nearest Neighbors. Cell Syst. 2019, 8, 329–337.e4. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Li, C.; Li, Z.; Wang, D.; Ren, X.; Zhang, Z. An entropy-based metric for assessing the purity of single cell populations. Nat. Commun. 2020, 11, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Lake, B.B.; Menon, R.; Winfree, S.; Hu, Q.; Ferreira, R.M.; Kalhor, K.; Barwinska, D.; Otto, E.A.; Ferkowicz, M.; Diep, D.; et al. An atlas of healthy and injured cell states and niches in the human kidney. Nature 2023, 619, 585–594. [Google Scholar] [CrossRef] [PubMed]
- Muto, Y.; Wilson, P.C.; Ledru, N.; Wu, H.; Dimke, H.; Waikar, S.S.; Humphreys, B.D. Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney. Nat. Commun. 2021, 12, 1–17. [Google Scholar] [CrossRef]
- Langfelder, P.; Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Trapnell, C.; Cacchiarelli, D.; Grimsby, J.; Pokharel, P.; Li, S.; Morse, M.; Lennon, N.J.; Livak, K.J.; Mikkelsen, T.S.; Rinn, J.L. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 2014, 32, 381–386. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. clusterProfiler: An R Package for Comparing Biological Themes Among Gene Clusters. OMICS J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Saikia, S.; Bordoloi, M. Molecular Docking: Challenges, Advances and its Use in Drug Discovery Perspective. Curr. Drug Targets 2019, 20, 501–521. [Google Scholar] [CrossRef]
- Ouyang, Q.; Wang, C.; Sang, T.; Tong, Y.; Zhang, J.; Chen, Y.; Wang, X.; Wu, L.; Wang, X.; Liu, R.; et al. Depleting profibrotic macrophages using bioactivated in vivo assembly peptides ameliorates kidney fibrosis. Cell. Mol. Immunol. 2024, 21, 826–841. [Google Scholar] [CrossRef]
- Xia, W.; He, Y.; Gan, Y.; Zhang, B.; Dai, G.; Ru, F.; Jiang, Z.; Chen, Z.; Chen, X. Long Non-coding RNA: An Emerging Contributor and Potential Therapeutic Target in Renal Fibrosis. Front. Genet. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Mount, D.B. Thick Ascending Limb of the Loop of Henle. Clin. J. Am. Soc. Nephrol. 2014, 9, 1974–1986. [Google Scholar] [CrossRef] [PubMed]
- Layton, A.T.; Edwards, A. Tubuloglomerular Feedback Signal Transduction in a Short Loop of Henle. Bull. Math. Biol. 2009, 72, 34–62. [Google Scholar] [CrossRef] [PubMed]
- Simon, N.; Hertig, A. Alteration of Fatty Acid Oxidation in Tubular Epithelial Cells: From Acute Kidney Injury to Renal Fibrogenesis. Front. Med. 2015, 2, 52. [Google Scholar] [CrossRef]
- Sipos, A.; Vargas, S.; Peti-Peterdi, J. Direct demonstration of tubular fluid flow sensing by macula densa cells. Am. J. Physiol. Physiol. 2010, 299, F1087–F1093. [Google Scholar] [CrossRef]
- Pihl, L.; Persson, P.; Fasching, A.; Hansell, P.; DiBona, G.F.; Palm, F. Insulin induces the correlation between renal blood flow and glomerular filtration rate in diabetes: implications for mechanisms causing hyperfiltration. Am. J. Physiol. Integr. Comp. Physiol. 2012, 303, R39–R47. [Google Scholar] [CrossRef]
- Sällström, J.; Carlström, M.; Olerud, J.; Fredholm, B.B.; Kouzmine, M.; Sandler, S.; Persson, A.E.G. High-protein-induced glomerular hyperfiltration is independent of the tubuloglomerular feedback mechanism and nitric oxide synthases. Am. J. Physiol. Integr. Comp. Physiol. 2010, 299, R1263–R1268. [Google Scholar] [CrossRef]
- Wisman, M.; Nizamoglu, M.; Noordhoek, J.A.; Timens, W.; Burgess, J.K.; Heijink, I.H. Dysregulated cross-talk between alveolar epithelial cells and stromal cells in idiopathic pulmonary fibrosis reduces epithelial regenerative capacity. Front. Med. 2023, 10, 1182368. [Google Scholar] [CrossRef]
- Gonzalez-Vicente, A.; Saez, F.; Monzon, C.M.; Asirwatham, J.; Garvin, J.L. Thick Ascending Limb Sodium Transport in the Pathogenesis of Hypertension. Physiol. Rev. 2019, 99, 235–309. [Google Scholar] [CrossRef]
- Li, M.; Xu, Y.; Liang, J.; Lin, H.; Qi, X.; Li, F.; Han, P.; Gao, Y.; Yang, X. USP22 deficiency in melanoma mediates resistance to T cells through IFNγ-JAK1-STAT1 signal axis. Mol. Ther. 2021, 29, 2108–2120. [Google Scholar] [CrossRef]
- Fu, Y.; Xiang, Y.; Wang, Y.; Liu, Z.; Yang, D.; Zha, J.; Tang, C.; Cai, J.; Chen, G.; Dong, Z. The STAT1/HMGB1/NF-κB pathway in chronic inflammation and kidney injury after cisplatin exposure. Theranostics 2023, 13, 2757–2773. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Dai, X.; Yang, S.; Peng, Y.; Hou, F.; Zhou, Q. High salt aggravates renal inflammation via promoting pro-inflammatory macrophage in 5/6-nephrectomized rat. Life Sci. 2021, 274, 119109. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Lim, J.Y.; Kim, J. Mechanical stretch induces angiotensinogen expression through PARP1 activation in kidney proximal tubular cells. Vitr. Cell. Dev. Biol. - Anim. 2014, 51, 72–78. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Padanilam, B.J. Loss of poly(ADP-ribose) polymerase 1 attenuates renal fibrosis and inflammation during unilateral ureteral obstruction. Am. J. Physiol. Physiol. 2011, 301, F450–F459. [Google Scholar] [CrossRef]
- Yoon, S.P.; Kim, J. Poly(ADP-ribose) polymerase 1 activation links ischemic acute kidney injury to interstitial fibrosis. J. Physiol. Sci. 2015, 65, 105–111. [Google Scholar] [CrossRef]
- Cheng, L.; Tu, C.; Min, Y.; He, D.; Wan, S.; Xiong, F. MiR-194 targets Runx1/Akt pathway to reduce renal fibrosis in mice with unilateral ureteral obstruction. Int. Urol. Nephrol. 2020, 52, 1801–1808. [Google Scholar] [CrossRef]
- John, A.S.P.; Kundu, S.; Pushpakumar, S.; Fordham, M.; Weber, G.; Mukhopadhyay, M.; Sen, U. GYY4137, a Hydrogen Sulfide Donor Modulates miR194-Dependent Collagen Realignment in Diabetic Kidney. Sci. Rep. 2017, 7, 1–20. [Google Scholar] [CrossRef]
- Lucarini, L.; Durante, M.; Lanzi, C.; Pini, A.; Boccalini, G.; Calosi, L.; Moroni, F.; Masini, E.; Mannaioni, G. HYDAMTIQ, a selective PARP-1 inhibitor, improves bleomycin-induced lung fibrosis by dampening the TGF-β/SMAD signalling pathway. J. Cell. Mol. Med. 2016, 21, 324–335. [Google Scholar] [CrossRef]
- Huang, D.; Wang, Y.; Wang, L.; Zhang, F.; Deng, S.; Wang, R.; Zhang, Y.; Huang, K. Poly(ADP-ribose) Polymerase 1 Is Indispensable for Transforming Growth Factor-β Induced Smad3 Activation in Vascular Smooth Muscle Cell. PLOS ONE 2011, 6, e27123. [Google Scholar] [CrossRef]
- Stöcker, G.; Stickeler, E.; Switalla, S.; Fischer, D.-C.; Greiling, H.; Haubeck, H.-D. Development of an Enzyme Immunoassay Specific for a Core Protein Epitope of a Novel Small Basement Membrane Associated Heparan Sulphate Proteoglycan from Human Kidney. Clin. Chem. Lab. Med. 1997, 35, 95–100. [Google Scholar] [CrossRef]
- Yu, Y.; Jia, Y.-Y.; Wang, M.; Mu, L.; Li, H.-J. PTGER3 and MMP-2 play potential roles in diabetic nephropathy via competing endogenous RNA mechanisms. BMC Nephrol. 2021, 22, 1–11. [Google Scholar] [CrossRef]
- Nakamoto, S.; Ito, Y.; Nishizawa, N.; Goto, T.; Kojo, K.; Kumamoto, Y.; Watanabe, M.; Narumiya, S.; Majima, M. EP3 signaling in dendritic cells promotes liver repair by inducing IL-13-mediated macrophage differentiation in mice. FASEB J. 2020, 34, 5610–5627. [Google Scholar] [CrossRef]
- Chen, S.; Song, X.; Xiao, Q.; Wang, L.; Zhu, X.; Zou, Y.; Li, G. Knockdown of TMEM30A in renal tubular epithelial cells leads to reduced glucose absorption. BMC Nephrol. 2023, 24, 1–8. [Google Scholar] [CrossRef]
| Gene | Forward primer(5′-3′) | Reverse primer(5′-3′) |
|---|---|---|
| STAT1 | TCACAGTGGTTCGAGCTTCAG | GCAAACGAGACATCATAGGCA |
| PARP8 | TAAATCGCACAAACTTTTGGGC | TCTCCAGAACAAGATCGAGTCAA |
| HS6ST2 | ACCGGGGAAGTCAGAAGCA | CTCTACGCTCCCTATGTAGTCAT |
| PTGER3 | CCGGAGCACTCTGCTGAAG | CCCCACTAAGTCGGTGAGC |
| TMEM207 | TGCTCTCGGATCTATCCTGTG | ATTCCGCACCTTTTCAGCCA |
| mRps16 | CGTGCTTGTGCTCGGAGCTA | GCTCCTTGCCCAGAAGCAAA |
| Symbol | UniProt Accession | Molecule Name | CID | Affinity (kcal/mol) |
hydrogen bonds |
|---|---|---|---|---|---|
| STAT1 | P42224 | Amcinonide | CID443958 | -7.13 | 3 |
| PTGER3 | P43115 | zidovudine | CID35370 | -6.64 | 5 |
| PARP8 | Q8N3A8 | Epigallocatechin Gallate | CID65064 | -7.57 | 2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




