Submitted:
14 January 2026
Posted:
14 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Molecular Sequence Evidence
2.1. Methodological Errors: Long-Branch Attraction (LBA)
2.1.1. Heterogeneity-Induced LBA
- Compositional heterogeneity arises from differences in base or amino acid usage across species, influenced by factors such as GC/AT bias or amino acid biosynthetic cost [14,15,16]. This can lead to convergent composition between distantly related taxa, potentially resulting in false groupings [14,15,17,18].
- Substitutional heterogeneity refers to variation in substitution rates and patterns (i.e., the probabilities of substitution between amino acids are not uniform) across sites [19,20,21]. Such heterogeneity can lead to saturation at rapidly evolving sites, causing the retained signal to reflect amino acid preferences rather than true phylogenetic relationships, and thereby introducing systematic errors due to convergent substitutions, also contributing to LBA [18,19,21].
2.1.2. Outgroup-Induced LBA
2.2. Biological Errors: Tree Incongruence and Gene Loss
2.2.1. “Observed” Factors: Observable Signals Within the Dataset
2.2.2. “Unobserved” Factors: Hidden Biases Beyond the Dataset
3. Rare Molecular Event Evidence
3.1. Fusion-with-Mixing in Nuclear Genomes: Anti-Convergent and Irreversible
3.2. Transfer of Mitochondrial Genes to the Nucleus: Irreversible but Lacking Anti-Convergence
4. Morphological and Embryological Evidence
4.1. Impact of Evolutionary Models on Criteria for Structural Complexity
4.1.1. Nervous Systems and Choanocytes: Independent Origins Disrupt Complexity-Based Inference
4.1.2. Germ Layers, Body Axes, and Symmetry: Secondary Loss Obscures Evolutionary History
4.2. Reassessment of Evidence and Its Implications for Morphological Complexity
4.2.1. Epithelial Tissue: Structural Complexity
4.2.2. Neuron-like and Muscle-like Functions: Physiological Complexity
5. Paleontological Evidence
5.1. Evidence Based on Constraining the Earliest Emergence
5.1.1. Traditional Fossil Evidence
- Characters criterion – whether the diagnostic traits used to assign a fossil to a particular group are well defined and phylogenetically informative;
- Diagnosis criterion – whether those characters are demonstrably present in the fossil;
- Age criterion – whether geochronological dating is available.
5.1.2. Molecular Fossil Evidence
5.2. Evidence Based on Reconstructing Evolutionary Trajectories
- The common ancestor of metazoans may already have been cylindrically symmetric [71], a body plan superficially similar to the hydroid form; moreover, skeletal biomineralization was widespread in the Ediacaran–Cambrian interval [126]. This raises the possibility that dinomischids represent either the metazoan ancestor itself or a case of convergence.
- The earliest dinomischid fossils postdate known ctenophores, whereas some controversial taxa such as Eoandromeda octobrachiata are older but exhibit more derived traits [102]. This paradox suggests both the potential existence of undiscovered basal lineages and the possibility that dinomischids were derived from ctenophores rather than ancestral to them.
- Because soft-bodied ctenophores have an exceptionally poor preservation potential, their role in early metazoan evolution remains obscure. This taphonomic gap further complicates phylogenetic interpretation. Overall, the evolutionary significance of dinomischid fossils remains ambiguous, and the signal they provide is limited.
6. Discussion
- explaining the causes of discrepancies among different disciplines;
- evaluating the resolving power of each methodological approach;
- exploring possible ways to integrate different lines of evidence; and
- assessing the overall tendencies of current evidence.
6.1. Why Are the Discrepancies Among Disciplines so Large?
- Accumulation of confounding evolutionary events. Over long evolutionary periods, extensive convergence and reversals inevitably occur, especially among lineages that have evolved independently since their divergence. This effect is especially striking in morphology, where even highly complex structures such as nervous systems have evolved independently and converged, while sponges appear to have undergone remarkably extensive secondary loss of characters. In molecular phylogenetics, such phenomena are even more common, giving rise to LBA and tree incongruence. In paleontology, the effect is smaller but still notable: traditional fossils may differ so greatly from crown groups that their relationships become obscure, while molecular fossils may be difficult to interpret due to uncertainties in biomarker specificity.
- Loss of evolutionary information. Although convergence can be viewed as a consequence of information loss, the mechanisms are distinct enough to warrant separate consideration. As seen in molecular sequences, saturation can erase phylogenetic signal, reducing sites to random noise. Large-scale gene or character loss across many taxa can similarly obliterate information, affecting both molecular and morphological analyses. In some cases, information loss arises directly from methodological constraints—for instance, molecular fossils preserve only chemical composition and concentration. Even traditional fossils, which appear to “fix” morphological characters in time, have effectively lost all molecular information. Unlike missing fossils, this kind of information loss affects entire clades or lineages, leaving no isolated remnants and thus representing a complete, irreversible loss of data.
- Destruction of physical evidence. This issue is specific to paleontology: over geological time, physical fossils may be lost or damaged through physicochemical processes. In contrast to information loss, the discovery of a single well-preserved fossil can sometimes recover evolutionary information through comparison. Nevertheless, fossil destruction likely occurs more frequently than true information loss. Although molecular sequence loss in fossils might mechanistically resemble physicochemical damage, its lineage-wide and irreversible nature justifies treating it as information loss rather than physical destruction.
- Asynchrony among disciplines. Evolutionary changes detected by one discipline may have little or no direct effect on others. For example, multiple minor-effect genetic changes may leave morphology largely unchanged, while a single regulatory mutation could drastically alter morphology but have little impact on sequence-based trees. Over long periods, this asynchronous accumulation amplifies discrepancies among data types. Notably, although the amplifying effect of time is not explicitly addressed, the issue of asynchrony among different lines of evidence had already been recognized [140].
6.2. How Far Back in Time Can Different Methods Reliably Resolve Evolutionary Relationships?
6.2.1. Factors extending the Resolution Limit
- Reducing the impact of confounding evolutionary events. This can be achieved when (I) such events are mechanistically improbable, (II) traits are insensitive to minor mutations, or (III) large data volumes dilute their effects.
- Preservation of evolutionary information. Facilitated when (I) evolutionary rates are low, avoiding saturation; (II) multiple coordinated changes are required for loss, making complete erasure unlikely; or (III) large datasets buffer against random loss.
- Ease of evidence acquisition. This includes (I) data readily obtainable from extant taxa (e.g., molecular sequences) and (II) ancient evidence that is both widespread and preservable (e.g., molecular fossils).
6.2.2. The Most Promising Approach: Highly Anti-Convergence and Highly Irreversible Marginal Instances (HACHIMI)
- Resistance to confounding events. By definition, such events are inherently robust to convergence and reversal. For example, in FWM, convergence would require similar chromosomal fusions involving entire linkage groups, followed by an intermixing of gene order between them, while reversal would necessitate implausibly precise re-separation of merged genes [40]; other potential alternative pathways are likewise highly improbable. Moreover, analyses of this type do not depend on exact sequences or gene order, making them insensitive to small-scale changes.
- Resistance to information loss. The rarity of these events prevents saturation and secondary loss, and some (e.g., FWM) require multiple concurrent changes to alter a single character, providing strong resistance to data loss.
- Accessibility of evidence. While not inherent to the definition, HACHIMI-type events are typically more readily detectable than traditional fossils, which are the main exception.
6.2.3. Resolution Limits of Other Approaches
- Molecular sequence evidence. High rates of convergence and reversal generate severe conflicts (e.g., LBA), mitigated only partially by large datasets. Saturation and gene loss are common, and although sequences are easy to obtain from extant taxa, the method’s Resolution Limit remains shallow.
- Embryological and morphological evidence. While convergence and reversal occur, high-resolution imaging and molecular tools now enable detailed identification and differentiation of these traits (e.g., the nervous system of ctenophores). Because most morphological traits are polygenic, true loss is rare, and residual evidence can often be detected. However, data collection is labor-intensive and prone to incompleteness. Thus, despite a potentially deeper Resolution Limit, accumulation of high-quality data is slow and low-quality noise remains common.
- Traditional fossil evidence. Traditional fossils preserve limited, coarse-grained morphological information and rarely allow assessment of development or molecular traits. Crucially, specimens are rare. Consequently, their Resolution Limit is extremely shallow.
- Molecular fossil evidence. Biomarkers may be produced by multiple taxa, and specificity is difficult to confirm. Apart from chemical composition, nearly all information is lost. Despite their abundance and ease of sampling, molecular fossils have a very limited resolution capacity.
6.3. How Can Different Approaches be Integrated?
- Molecular sequences – encoding divergence times and gene evolutionary histories;
- HACHIMIs – rare events offering high-resolution support for specific nodes;
- Embryology and morphology – preserving phenotypic evolutionary trajectories;
- Paleontology – providing direct records of lineage existence and transitions.
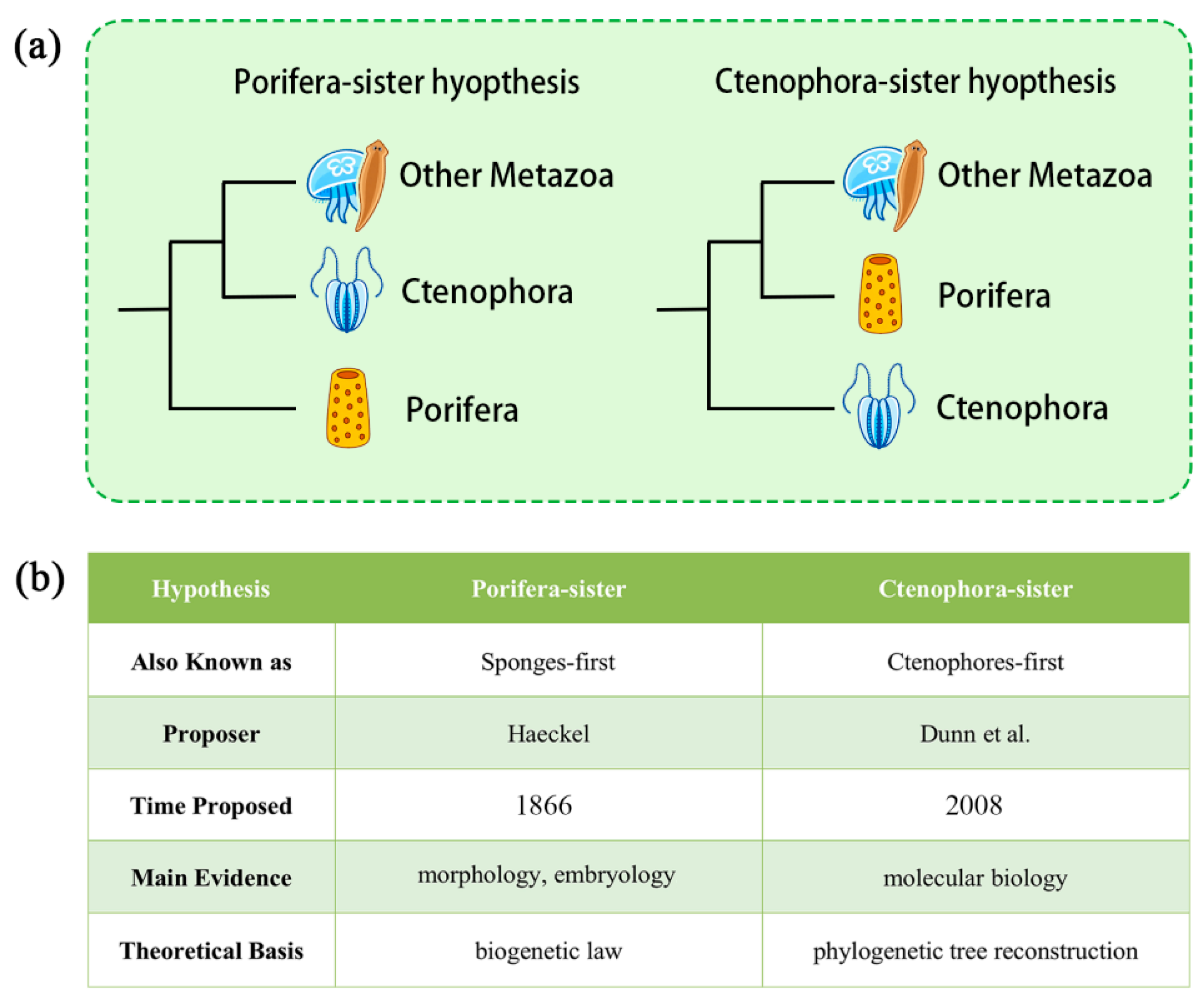
6.4. Which Hypothesis Is Currently Better Supported?
- HACHIMI evidence (FWM) supports the Ctenophora-sister hypothesis.
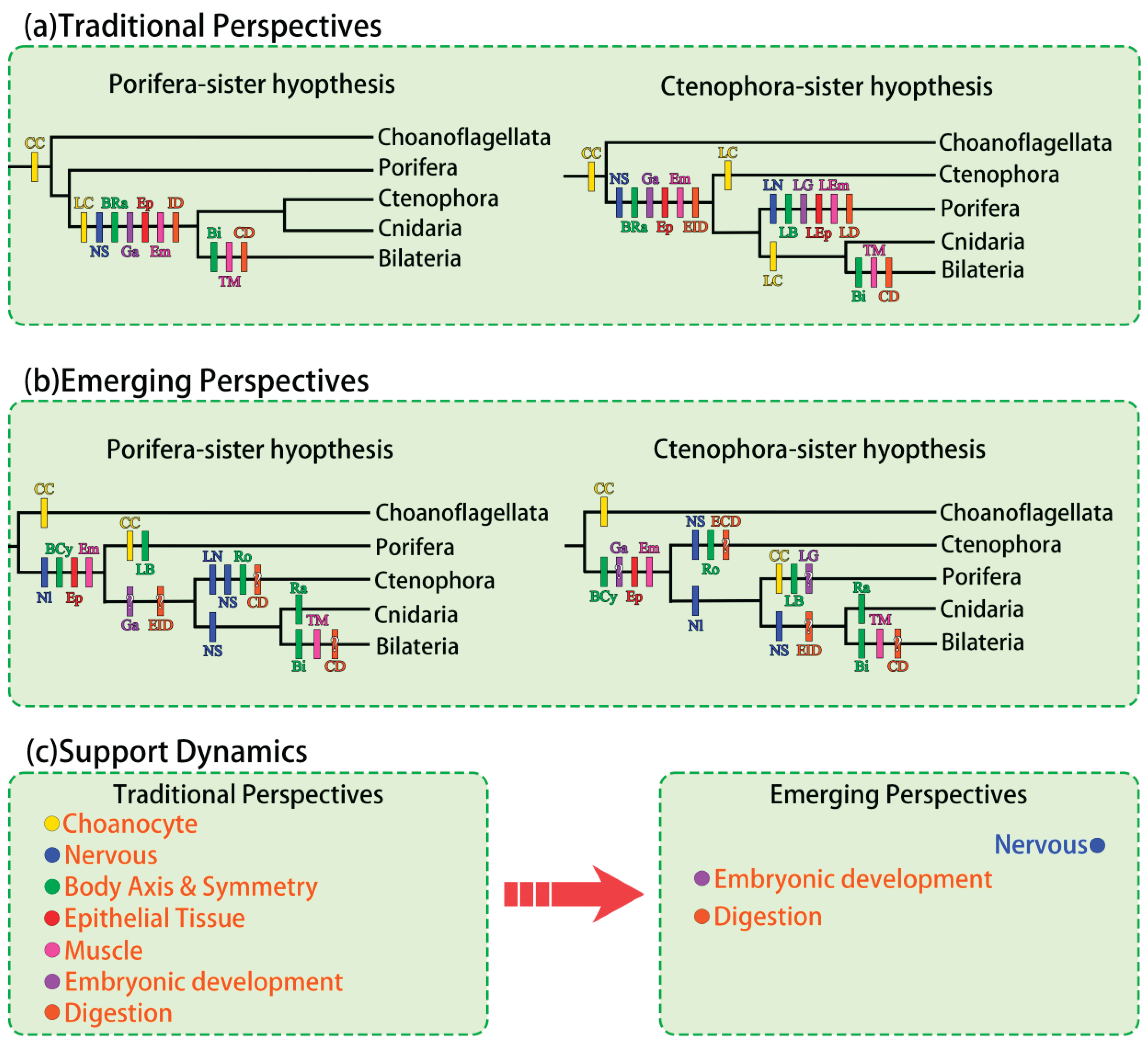
- Morphological and embryological evidence: recent high-quality studies tend to be neutral or mildly support the Ctenophora-sister hypothesis, whereas older, less-resolved evidence often supports the Porifera-sister hypothesis (Figure 2c). Overall, Porifera-sister evidence remains quantitatively more abundant, but Ctenophora-sister evidence exhibits greater methodological rigor and data reliability, indicating a trend increasingly favoring the latter.
- Molecular sequence evidence remains highly contentious but currently leans weakly toward the Porifera-sister hypothesis.
- Paleontological evidence (both traditional and molecular) is too fragmentary and uncertain to strongly support either side.
7. Conclusion
Funding
Data Availability Statement
Acknowledgements
Conflicts of Interest
Appendix A
| Year | Long-Branch Attraction (LBA) | Tree Incongruence | |||
|---|---|---|---|---|---|
| Model Complexity | Data Simplification | Outgroup Choice | “Observed” Factors | “Unobserved” Factors | |
| 2008 | First EST-based phylogeny supporting the Ctenophora-sister hypothesis [3] | ||||
| 2013 | First ctenophore whole-genome data further supporting Ctenophora-sister [12] | ||||
| 2015 | Site-heterogeneous models (e.g., CAT) show better fit than site-homogeneous models [6,7] |
Data excluding nuclear proteins mostly support Ctenophora-sister [6] | Homogeneous models show no effect of outgroup choice [6] | Correcting underestimation of gene loss restores Porifera-sister topology from Ryan et al.’s dataset [7] | |
| Excluding highly conserved proteins (e.g., ribosomal genes) exacerbates LBA [7] | Under CAT models, closely related outgroups recover Porifera-sister [7] | ||||
| 2017 | First use of amino acid recoding recovers Porifera-sister [23] | ||||
| 2020 | Widespread gene loss, especially in controversial lineages [37] | ||||
| 2021 | CAT models found to be overfitted; simpler site-heterogeneous models support Ctenophora-sister [8] | Amino acid recoding introduces biases by reducing information content [8] | Outgroup choice has no effect under simple site-heterogeneous models [8] | ||
| 2022 | Tree incongruence lower under Porifera-sister than Ctenophora-sister, mainly due to incomplete lineage sorting (ILS) [9] | ||||
| 2025 | Used only genes with consistent signals across concatenation and consensus analyses, recovering Porifera-sister [28] | ||||
References
- Haeckel, E. Generelle Morphologie der Organismen (2 Bde); G. Reimer, 1866. [Google Scholar] [CrossRef]
- Levit, G. S.; Hoßfeld, U.; Naumann, B.; Lukas, P.; Olsson, L. The biogenetic law and the Gastraea theory: From Ernst Haeckel's discoveries to contemporary views. Journal of Experimental Zoology Part B: Molecular and Developmental Evolution 2022, 338(1-2), 13–27. [Google Scholar] [CrossRef] [PubMed]
- Dunn, C. W., Hejnol, A., Matus, D. Q., Pang, K., Browne, W. E., Smith, S. A., ... & Giribet, G. Broad phylogenomic sampling improves resolution of the animal tree of life. Nature 2008, 452(7188), 745–749. [CrossRef]
- Dunn, C. W.; Giribet, G.; Edgecombe, G. D.; Hejnol, A. Animal phylogeny and its evolutionary implications. Annual Review of Ecology, Evolution, and Systematics 2014, 45(1), 371–395. [Google Scholar] [CrossRef]
- Dunn, C. W.; Leys, S. P.; Haddock, S. H. The hidden biology of sponges and ctenophores. Trends in Ecology & Evolution 2015, 30(5), 282–291. [Google Scholar] [CrossRef] [PubMed]
- Whelan, N. V.; Kocot, K. M.; Moroz, L. L.; Halanych, K. M. Error, signal, and the placement of Ctenophora sister to all other animals. Proceedings of the National Academy of Sciences 2015, 112(18), 5773–5778. [Google Scholar] [CrossRef] [PubMed]
- Pisani, D., Pett, W., Dohrmann, M., Feuda, R., Rota-Stabelli, O., Philippe, H., ... & Wörheide, G. Genomic data do not support comb jellies as the sister group to all other animals. Proceedings of the National Academy of Sciences 2015, 112(50), 15402–15407. [CrossRef]
- Li, Y.; Shen, X. X.; Evans, B.; Dunn, C. W.; Rokas, A. Rooting the animal tree of life. Molecular Biology and Evolution 2021, 38(10), 4322–4333. [Google Scholar] [CrossRef]
- Rosenzweig, B.; Kern, A.; Hahn, M. Accurate detection of incomplete lineage sorting via supervised machine learning. bioRxiv Preprint. 2022. [Google Scholar] [CrossRef]
- Krumbach, T. Ctenophora. In Handbuch der Zoologie; Krumbach, T., Starck, D., Eds.; W. de Gruyter: Berlin, 1925; pp. 905–995. [Google Scholar]
- Cavalier-Smith, T. M.; Allsopp, M. T. E. P.; Chao, E. E.; Boury-Esnault, N.; Vacelet, J. Sponge phylogeny, animal monophyly, and the origin of the nervous system: 18S rRNA evidence. Canadian Journal of Zoology 1996, 74(11), 2031–2045. [Google Scholar] [CrossRef]
- Ryan, J. F., Pang, K., Schnitzler, C. E., Nguyen, A. D., Moreland, R. T., Simmons, D. K., ... & Baxevanis, A. D. The genome of the ctenophore Mnemiopsis leidyi and its implications for cell type evolution. Science 2013, 342(6164), 1242592. [CrossRef]
- Felsenstein, J. Cases in which parsimony or compatibility methods will be positively misleading. Systematic Zoology 1978, 27(4), 401–410. [Google Scholar] [CrossRef]
- Lockhart, P. J.; Howe, C. J.; Bryant, D. A.; Beanland, T. J.; Larkum, A. W. D. Substitutional bias confounds inference of cyanelle origins from sequence data. Journal of Molecular Evolution 1992, 34(2), 153–162. [Google Scholar] [CrossRef] [PubMed]
- Foster, P. G.; Hickey, D. A. Compositional bias may affect both DNA-based and protein-based phylogenetic reconstructions. Journal of Molecular Evolution 1999, 48(3), 284–290. [Google Scholar] [CrossRef]
- Akashi, H.; Gojobori, T. Metabolic efficiency and amino acid composition in the proteomes of Escherichia coli and Bacillus subtilis. Proceedings of the National Academy of Sciences 2002, 99(6), 3695–3700. [Google Scholar] [CrossRef]
- Jermiin, L. S.; Ho, S. Y.; Ababneh, F.; Robinson, J.; Larkum, A. W. The biasing effect of compositional heterogeneity on phylogenetic estimates may be underestimated. Systematic Biology 2004, 53(4), 638–643. [Google Scholar] [CrossRef]
- Ho, S. Y.; Jermiin, L. S. Tracing the decay of the historical signal in biological sequence data. Systematic Biology 2004, 53(4), 623–637. [Google Scholar] [CrossRef]
- Yang, Z. Maximum-likelihood estimation of phylogeny from DNA sequences when substitution rates differ over sites. Molecular Biology and Evolution 1993, 10(6), 1396–1401. [Google Scholar] [CrossRef]
- Halpern, A. L.; Bruno, W. J. Evolutionary distances for protein-coding sequences: modeling site-specific residue frequencies. Molecular Biology and Evolution 1998, 15(7), 910–917. [Google Scholar] [CrossRef] [PubMed]
- Lartillot, N.; Philippe, H. A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Molecular Biology and Evolution 2004, 21(6), 1095–1109. [Google Scholar] [CrossRef] [PubMed]
- Redmond, A. K.; McLysaght, A. Evidence for sponges as sister to all other animals from partitioned phylogenomics with mixture models and recoding. Nature Communications 2021, 12(1), 1783. [Google Scholar] [CrossRef]
- Feuda, R.; Dohrmann, M.; Pett, W.; Philippe, H.; Rota-Stabelli, O.; Lartillot, N.; ... & Pisani, D. Improved modeling of compositional heterogeneity supports sponges as sister to all other animals. Current Biology 2017, 27(24), 3864–3870. [CrossRef]
- Susko, E.; Roger, A. J. On reduced amino acid alphabets for phylogenetic inference. Molecular Biology and Evolution 2007, 24(9), 2139–2150. [Google Scholar] [CrossRef]
- Hernandez, A. M.; Ryan, J. F. Six-state amino acid recoding is not an effective strategy to offset compositional heterogeneity and saturation in phylogenetic analyses. Systematic Biology 2021, 70(6), 1200–1212. [Google Scholar] [CrossRef] [PubMed]
- Whelan, N. V.; Halanych, K. M. Available data do not rule out Ctenophora as the sister group to all other Metazoa. Nature Communications 2023, 14(1), 711. [Google Scholar] [CrossRef]
- Shen, X. X.; Hittinger, C. T.; Rokas, A. Contentious relationships in phylogenomic studies can be driven by a handful of genes. Nature Ecology & Evolution 2017, 1(5), 0126. [Google Scholar] [CrossRef] [PubMed]
- Steenwyk, J. L.; King, N. Integrative phylogenomics positions sponges at the root of the animal tree. Science 2025, 390(6774), 751–756. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, W. C. Nucleic acid sequence phylogeny and random outgroups. Cladistics 1990, 6(4), 363–367. [Google Scholar] [CrossRef]
- Bergsten, J. A review of long-branch attraction. Cladistics 2005, 21(2), 163–193. [Google Scholar] [CrossRef]
- Pamilo, P.; Nei, M. Relationships between gene trees and species trees. Molecular Biology and Evolution 1988, 5(5), 568–583. [Google Scholar] [CrossRef]
- Maddison, W. P. Gene trees in species trees. Systematic Biology 1997, 46(3), 523–536. [Google Scholar] [CrossRef]
- Jeffroy, O.; Brinkmann, H.; Delsuc, F.; Philippe, H. Phylogenomics: the beginning of incongruence? Trends in Genetics 2006, 22(4), 225–231. [Google Scholar] [CrossRef] [PubMed]
- Hotopp, J. C. D. Horizontal gene transfer between bacteria and animals. Trends in Genetics 2011, 27(4), 157–163. [Google Scholar] [CrossRef]
- Bjornson, S.; Upham, N.; Verbruggen, H.; Steenwyk, J. Phylogenomic inference, divergence-time calibration, and methods for characterizing reticulate evolution Preprint. Preprints 2023. [Google Scholar] [CrossRef]
- Roure, B.; Baurain, D.; Philippe, H. Impact of missing data on phylogenies inferred from empirical phylogenomic data sets. Molecular Biology and Evolution 2013, 30(1), 197–214. [Google Scholar] [CrossRef]
- Fernández, R.; Gabaldón, T. Gene gain and loss across the metazoan tree of life. Nature Ecology & Evolution 2020, 4(4), 524–533. [Google Scholar] [CrossRef] [PubMed]
- Pagel, M.; Meade, A.; Barker, D. Bayesian estimation of ancestral character states on phylogenies. Systematic Biology 2004, 53(5), 673–684. [Google Scholar] [CrossRef]
- Renwick, J. H. The mapping of human chromosomes. Annual Review of Genetics 1971, 5(1), 81–120. [Google Scholar] [CrossRef]
- Simakov, O., Bredeson, J., Berkoff, K., Marletaz, F., Mitros, T., Schultz, D. T., ... & Rokhsar, D. S. Deeply conserved synteny and the evolution of metazoan chromosomes. Science Advances 2022, 8(5), eabi5884. [CrossRef]
- Schultz, D. T.; Haddock, S. H.; Bredeson, J. V.; Green, R. E.; Simakov, O.; Rokhsar, D. S. Ancient gene linkages support ctenophores as sister to other animals. Nature 2023, 618(7963), 110–117. [Google Scholar] [CrossRef]
- Green, R. E., Krause, J., Briggs, A. W., Maricic, T., Stenzel, U., Kircher, M., ... & Pääbo, S. A draft sequence of the Neandertal genome. Science 2010, 328(5979), 710–722. [CrossRef]
- Crisp, A.; Boschetti, C.; Perry, M.; Tunnacliffe, A.; Micklem, G. Expression of multiple horizontally acquired genes is a hallmark of both vertebrate and invertebrate genomes. Genome Biology 2015, 16(1), 50. [Google Scholar] [CrossRef]
- Steenwyk, J. L.; King, N. The promise and pitfalls of synteny in phylogenomics. PLoS Biology 2024, 22(5), e3002632. [Google Scholar] [CrossRef]
- Jacobs, H. T.; Posakony, J. W.; Grula, J. W.; Roberts, J. W.; Xin, J. H.; Britten, R. J.; Davidson, E. H. Mitochondrial DNA sequences in the nuclear genome of Strongylocentrotus pupuratus. Journal of Molecular Biology 1983, 165(4), 609–632. [Google Scholar] [CrossRef]
- Farrelly, F.; Butow, R. A. Rearranged mitochondrial genes in the yeast nuclear genome. Nature 1983, 301(5898), 296–301. [Google Scholar] [CrossRef] [PubMed]
- Lopez, J. V.; Yuhki, N.; Masuda, R.; Modi, W.; O'Brien, S. J. Numt, a recent transfer and tandem amplification of mitochondrial DNA to the nuclear genome of the domestic cat. Journal of Molecular Evolution 1994, 39(2), 174–190. [Google Scholar] [CrossRef]
- Mundy, N. I.; Pissinatti, A.; Woodruff, D. S. Multiple nuclear insertions of mitochondrial cytochrome b sequences in callitrichine primates. Molecular Biology and Evolution 2000, 17(7), 1075–1080. [Google Scholar] [CrossRef] [PubMed]
- Williams, S. T.; Knowlton, N. Mitochondrial pseudogenes are pervasive and often insidious in the snapping shrimp genus Alpheus. Molecular Biology and Evolution 2001, 18(8), 1484–1493. [Google Scholar] [CrossRef] [PubMed]
- Lü, X. M.; Fu, Y. X.; Zhang, Y. P. Evolution of mitochondrial cytochrome b pseudogene in genus Nycticebus. Molecular Biology and Evolution 2002, 19(12), 2337–2341. [Google Scholar] [CrossRef]
- Pett, W.; Ryan, J. F.; Pang, K.; Mullikin, J. C.; Martindale, M. Q.; Baxevanis, A. D.; Lavrov, D. V. Extreme mitochondrial evolution in the ctenophore Mnemiopsis leidyi: insight from mtDNA and the nuclear genome. Mitochondrial DNA 2011, 22(4), 130–142. [Google Scholar] [CrossRef]
- Thorsness, P. E.; Fox, T. D. Escape of DNA from mitochondria to the nucleus in Saccharomyces cerevisiae. Nature 1990, 346(6282), 376–379. [Google Scholar] [CrossRef]
- Knoop, V.; Unseld, M.; Marienfeld, J.; Brandt, P.; Sünkel, S.; Ullrich, H.; Brennicke, A. copia-, gypsy-and LINE-like retrotransposon fragments in the mitochondrial genome of Arabidopsis thaliana. Genetics 1996, 142(2), 579–585. [Google Scholar] [CrossRef]
- Kubo, T.; Nishizawa, S.; Sugawara, A.; Itchoda, N.; Estiati, A.; Mikami, T. The complete nucleotide sequence of the mitochondrial genome of sugar beet (Beta vulgaris L.) reveals a novel gene for tRNACys (GCA). Nucleic Acids Research 2000, 28(13), 2571–2576. [Google Scholar] [CrossRef]
- Notsu, Y., Masood, S., Nishikawa, T., Kubo, N., Akiduki, G., Nakazono, M., ... & Kadowaki, K. The complete sequence of the rice (Oryza sativa L.) mitochondrial genome: frequent DNA sequence acquisition and loss during the evolution of flowering plants. Molecular Genetics and Genomics 2002, 268(4), 434–445. [CrossRef]
- Rodríguez-Moreno, L.; González, V. M.; Benjak, A.; Martí, M. C.; Puigdomènech, P.; Aranda, M. A.; Garcia-Mas, J. Determination of the melon chloroplast and mitochondrial genome sequences reveals that the largest reported mitochondrial genome in plants contains a significant amount of DNA having a nuclear origin. BMC Genomics 2011, 12(1), 424. [Google Scholar] [CrossRef]
- Martin, W.; Herrmann, R. G. Gene transfer from organelles to the nucleus: how much, what happens, and why? Plant Physiology 1998, 118(1), 9–17. [Google Scholar] [CrossRef]
- Burkhardt, P., Colgren, J., Medhus, A., Digel, L., Naumann, B., Soto-Angel, J. J., ... & Kittelmann, M. Syncytial nerve net in a ctenophore adds insights on the evolution of nervous systems. Science 2023, 380(6642), 293–297. [CrossRef]
- Moroz, L. L., Kocot, K. M., Citarella, M. R., Dosung, S., Norekian, T. P., Povolotskaya, I. S., ... & Kohn, A. B. The ctenophore genome and the evolutionary origins of neural systems. Nature 2014, 510(7503), 109–114. [CrossRef]
- Moroz, L. L. Convergent evolution of neural systems in ctenophores. Journal of Experimental Biology 2015, 218(4), 598–611. [Google Scholar] [CrossRef]
- Nielsen, C. Early animal evolution: a morphologist's view. Royal Society Open Science 2019, 6(7), 190638. [Google Scholar] [CrossRef] [PubMed]
- Clark, H. J. Conclusive proofs of the animality of the ciliate sponges, and of their affinities with the Infusoria flagellata. American Journal of Science 1866, 2(126), 320–324. [Google Scholar] [CrossRef]
- Carr, M.; Leadbeater, B. S.; Hassan, R.; Nelson, M.; Baldauf, S. L. Molecular phylogeny of choanoflagellates, the sister group to Metazoa. Proceedings of the National Academy of Sciences 2008, 105(43), 16641–16646. [Google Scholar] [CrossRef] [PubMed]
- Lyons, K. M. Evolutionary implications of collar cell ectoderm in a coral planula. Nature 1973, 245(5419), 50–51. [Google Scholar] [CrossRef]
- Nerrevang, A.; Wingstrand, K. G. On the occurrence and structure of choanocyte-like cells in some echinoderms. Acta Zoologica 1970, 51(3), 249–270. [Google Scholar] [CrossRef]
- Mah, J. L.; Christensen-Dalsgaard, K. K.; Leys, S. P. Choanoflagellate and choanocyte collar-flagellar systems and the assumption of homology. Evolution & Development 2014, 16(1), 25–37. [Google Scholar] [CrossRef]
- Burton, P. M. Insights from diploblasts; the evolution of mesoderm and muscle. Journal of Experimental Zoology Part B: Molecular and Developmental Evolution 2008, 310(1), 5–14. [Google Scholar] [CrossRef]
- Steinmetz, P. R., Kraus, J. E., Larroux, C., Hammel, J. U., Amon-Hassenzahl, A., Houliston, E., ... & Technau, U. Independent evolution of striated muscles in cnidarians and bilaterians. Nature 2012, 487(7406), 231–234. [CrossRef]
- Degnan, B. M.; Leys, S. P.; Larroux, C. Sponge development and antiquity of animal pattern formation. Integrative and Comparative Biology 2005, 45(2), 335–341. [Google Scholar] [CrossRef] [PubMed]
- Ereskovsky, A. V.; Dondua, A. K. The problem of germ layers in sponges (Porifera) and some issues concerning early metazoan evolution. Zoologischer Anzeiger-A Journal of Comparative Zoology 2006, 245(2), 65–76. [Google Scholar] [CrossRef]
- Manuel, M. Early evolution of symmetry and polarity in metazoan body plans. Comptes Rendus Biologies 2009, 332(2-3), 184–209. [Google Scholar] [CrossRef]
- Leys, S. P.; Riesgo, A. Epithelia, an evolutionary novelty of metazoans. Journal of Experimental Zoology Part B: Molecular and Developmental Evolution 2011, 318(6), 438–447. [Google Scholar] [CrossRef]
- Revel, J. P. Fine structure of intercellular contacts in the sponge, Microciona prolifera. Biol Bull 1966, 131, 402. [Google Scholar]
- Tsukita, S.; Furuse, M. Occludin and claudins in tight-junction strands: leading or supporting players? Trends in Cell Biology 1999, 9(7), 268–273. [Google Scholar] [CrossRef] [PubMed]
- Leys, S. P.; Nichols, S. A.; Adams, E. D. Epithelia and integration in sponges. Integrative and Comparative Biology 2009, 49(2), 167–177. [Google Scholar] [CrossRef]
- Fahey, B.; Degnan, B. M. Origin of animal epithelia: insights from the sponge genome. Evolution & Development 2010, 12(6), 601–617. [Google Scholar] [CrossRef]
- Dickinson, D. J.; Nelson, W. J.; Weis, W. I. A polarized epithelium organized by β-and α-catenin predates cadherin and metazoan origins. Science 2011, 331(6022), 1336–1339. [Google Scholar] [CrossRef] [PubMed]
- Abedin, M.; King, N. The premetazoan ancestry of cadherins. Science 2008, 319(5865), 946–948. [Google Scholar] [CrossRef] [PubMed]
- Mackie, G. O.; Lawn, I. D.; De Ceccatty, M. P. Studies on hexactinellid sponges. II. Excitability, conduction and coordination of responses in Rhabdocalyptus dawsoni (Lambe, 1873). Philosophical Transactions of the Royal Society of London. B, Biological Sciences 1983, 301(1107), 401–418. [Google Scholar] [CrossRef]
- Leys, S. P. Elements of a ‘nervous system’in sponges. Journal of Experimental Biology 2015, 218(4), 581–591. [Google Scholar] [CrossRef]
- Elliott, G. R.; Leys, S. P. Evidence for glutamate, GABA and NO in coordinating behaviour in the sponge, Ephydatia muelleri (Demospongiae, Spongillidae). Journal of Experimental Biology 2010, 213(13), 2310–2321. [Google Scholar] [CrossRef]
- Ellwanger, K.; Nickel, M. Neuroactive substances specifically modulate rhythmic body contractions in the nerveless metazoon Tethya wilhelma (Demospongiae, Porifera). Frontiers in Zoology 2006, 3(1), 7. [Google Scholar] [CrossRef]
- Ludeman, D. A.; Farrar, N.; Riesgo, A.; Paps, J.; Leys, S. P. Evolutionary origins of sensation in metazoans: functional evidence for a new sensory organ in sponges. BMC Evolutionary Biology 2014, 14(1), 3. [Google Scholar] [CrossRef] [PubMed]
- Leys, S. P.; Degnan, B. M. Cytological basis of photoresponsive behavior in a sponge larva. The Biological Bulletin 2001, 201(3), 323–338. [Google Scholar] [CrossRef]
- Leys, S. P.; Cronin, T. W.; Degnan, B. M.; Marshall, J. N. Spectral sensitivity in a sponge larva. Journal of Comparative Physiology A 2002, 188(3), 199–202. [Google Scholar] [CrossRef] [PubMed]
- Wong, E.; Anggono, V.; Williams, S. R.; Degnan, S. M.; Degnan, B. M. Phototransduction in a marine sponge provides insights into the origin of animal vision. iScience 2022, 25(6). [Google Scholar] [CrossRef]
- Elliott, G. R.; Leys, S. P. Coordinated contractions effectively expel water from the aquiferous system of a freshwater sponge. Journal of Experimental Biology 2007, 210(21), 3736–3748. [Google Scholar] [CrossRef]
- Nickel, M. Kinetics and rhythm of body contractions in the sponge Tethya wilhelma (Porifera: Demospongiae). Journal of Experimental Biology 2004, 207(26), 4515–4524. [Google Scholar] [CrossRef]
- Philippe, H., Derelle, R., Lopez, P., Pick, K., Borchiellini, C., Boury-Esnault, N., ... & Manuel, M. Phylogenomics revives traditional views on deep animal relationships. Current Biology 2009, 19(8), 706–712. [CrossRef]
- Eitel, M. S. Phylogenetic position, biodiversity, phylogeography and biology of the Placozoa (Doctoral dissertation), Gottfried Wilhelm Leibniz Universität Hannover. 2010. [Google Scholar]
- Smith, C. L., Varoqueaux, F., Kittelmann, M., Azzam, R. N., Cooper, B., Winters, C. A., ... & Reese, T. S. Novel cell types, neurosecretory cells, and body plan of the early-diverging metazoan Trichoplax adhaerens. Current Biology 2014, 24(14), 1565–1572. [CrossRef]
- Presnell, J. S.; Vandepas, L. E.; Warren, K. J.; Swalla, B. J.; Amemiya, C. T.; Browne, W. E. The presence of a functionally tripartite through-gut in Ctenophora has implications for metazoan character trait evolution. Current Biology 2016, 26(20), 2814–2820. [Google Scholar] [CrossRef]
- Silva, F B D; Muschner, V C; Bonatto, S L. Phylogenetic position of Placozoa based on large subunit (LSU) and small subunit (SSU) rRNA genes. Genetics and Molecular Biology 2007, 30, 127–132. [Google Scholar] [CrossRef]
- Telford, M. J.; Budd, G. E.; Philippe, H. Phylogenomic insights into animal evolution. Current Biology 2015, 25(19), R876–R887. [Google Scholar] [CrossRef]
- Antcliffe, J. B.; Callow, R. H.; Brasier, M. D. Giving the early fossil record of sponges a squeeze. Biological Reviews 2014, 89(4), 972–1004. [Google Scholar] [CrossRef]
- Benton, M. J.; Donoghue, P. C. J.; Asher, R. J. Calibrating and constraining molecular clocks. In The timetree of life; Hedges, S. B., Kumar, S., Eds.; Oxford University Press, 2009; pp. 35–86. [Google Scholar] [CrossRef]
- Wang, X., Liu, A. G., Chen, Z., Wu, C., Liu, Y., Wan, B., ... & Xiao, S. A late-Ediacaran crown-group sponge animal. Nature 2024, 630(8018), 905–911. [CrossRef]
- Condon, D.; Zhu, M.; Bowring, S.; Wang, W.; Yang, A.; Jin, Y. U-Pb ages from the neoproterozoic Doushantuo Formation, China. Science 2005, 308(5718), 95–98. [Google Scholar] [CrossRef]
- Huang, T.; Chen, D.; Ding, Y.; Zhou, X.; Zhang, G. SIMS U-Pb zircon geochronological and carbon isotope chemostratigraphic constraints on the Ediacaran-Cambrian boundary succession in the Three Gorges area, South China. Journal of Earth Science 2020, 31(1), 69–78. [Google Scholar] [CrossRef]
- Steiner, M.; Li, G.; Qian, Y.; Zhu, M. Lower Cambrian small shelly fossils of northern Sichuan and southern Shaanxi (China), and their biostratigraphic importance. Geobios 2004, 37(2), 259–275. [Google Scholar] [CrossRef]
- Chen, J. Y., Schopf, J. W., Bottjer, D. J., Zhang, C. Y., Kudryavtsev, A. B., Tripathi, A. B., ... & Yang, Y. Raman spectra of a Lower Cambrian ctenophore embryo from southwestern Shaanxi, China. Proceedings of the National Academy of Sciences 2007, 104(15), 6289–6292. [CrossRef]
- Zhao, Y., Vinther, J., Parry, L. A., Wei, F., Green, E., Pisani, D., ... & Cong, P. Cambrian sessile, suspension feeding stem-group ctenophores and evolution of the comb jelly body plan. Current Biology 2019, 29(7), 1112–1125. [CrossRef]
- Wang, W.; Zhou, M.; Chu, Z.; Xu, J.; Li, C.; Luo, T.; Guo, J. Constraints on the Ediacaran-Cambrian boundary in deep-water realm in South China: Evidence from zircon CA-ID-TIMS U-Pb ages from the topmost Liuchapo Formation. Science China Earth Sciences 2020, 63(8), 1176–1187. [Google Scholar] [CrossRef]
- Compston, W.; Zhang, Z.; Cooper, J. A.; Guogan, M.; Jenkins, R. J. F. Further SHRIMP geochronology on the early Cambrian of South China. American Journal of Science 2008, 308, 399–420. [Google Scholar] [CrossRef]
- Ou, Q.; Xiao, S.; Han, J.; Sun, G.; Zhang, F.; Zhang, Z.; Shu, D. A vanished history of skeletonization in Cambrian comb jellies. Science Advances 2015, 1(6), e1500092. [Google Scholar] [CrossRef]
- Tang, T. F.; Zhang, J. M.; Jiang, X. Discovery and significance of the Late Sinian fauna from western Hunan and Hubei. Acta Stratigraphica Sinica 1978, 2, 32–45. [Google Scholar]
- Hahn, G.; Pflug, H. D. Polypenartige Organismen aus dem Jung-Präkambrium (Nama-Gruppe) von Namibia. Geologica et Palaeontologica 1985, (19), 1–13. [Google Scholar]
- Steiner, M.; Mehl, D.; Reitner, J.; Erdtmann, B. D. Oldest entirely preserved sponges and other fossils from the lowermost Cambrian and a new facies reconstruction of the Yangtze Platform (China). Berliner Geowissenschaftliche Abhandlungen 1993, 9, 293–329. [Google Scholar] [CrossRef]
- Gehling, J. G.; Rigby, J. K. Long expected sponges from the Neoproterozoic Ediacara fauna of South Australia. Journal of Paleontology 1996, 70(2), 185–195. [Google Scholar] [CrossRef]
- Steiner, M.; Reitner, J. Evidence of organic structures in Ediacara-type fossils and associated microbial mats. Geology 2001, 29(12), 1119–1122. [Google Scholar] [CrossRef]
- Clapham, M. E.; Narbonne, G. M.; Gehling, J. G.; Greentree, C.; Anderson, M. M. Thectardis avalonensis: a new Ediacaran fossil from the Mistaken Point biota, Newfoundland. Journal of Paleontology 2004, 78(6), 1031–1036. [Google Scholar] [CrossRef]
- German, T. N.; Podkovyrov, V. N. Records of a new spongelike group in the Riphean biota. Paleontological Journal 2012, 46(3), 219–227. [Google Scholar] [CrossRef]
- Tang, F.; Bengtson, S.; Wang, Y.; Wang, X. L.; Yin, C. Y. Eoandromeda and the origin of Ctenophora. Evolution & Development 2011, 13(5), 408–414. [Google Scholar] [CrossRef]
- Harvey, T. H., Williams, M., Condon, D. J., Wilby, P. R., Siveter, D. J., Rushton, A. W., ... & Gabbott, S. E. A refined chronology for the Cambrian succession of southern Britain. Journal of the Geological Society 2011, 168(3), 705–716. [CrossRef]
- Peters, K. E.; Walters, C. C.; Moldowan, J. M. The biomarker guide; Cambridge University Press, 2005; Vol. 1. [Google Scholar] [CrossRef]
- McCaffrey, M. A.; Moldowan, J. M.; Lipton, P. A.; Summons, R. E.; Peters, K. E.; Jeganathan, A.; Watt, D. S. Paleoenvironmental implications of novel C30 steranes in Precambrian to Cenozoic age petroleum and bitumen. Geochimica et Cosmochimica Acta 1994, 58(1), 529–532. [Google Scholar] [CrossRef]
- Love, G. D., Grosjean, E., Stalvies, C., Fike, D. A., Grotzinger, J. P., Bradley, A. S., ... & Summons, R. E. Fossil steroids record the appearance of Demospongiae during the Cryogenian period. Nature 2009, 457(7230), 718–721. [CrossRef]
- Zumberge, J. A., Love, G. D., Cárdenas, P., Sperling, E. A., Gunasekera, S., Rohrssen, M., ... & Summons, R. E. Demosponge steroid biomarker 26-methylstigmastane provides evidence for Neoproterozoic animals. Nature Ecology & Evolution 2018, 2(11), 1709–1714. [CrossRef]
- Nettersheim, B. J., Brocks, J. J., Schwelm, A., Hope, J. M., Not, F., Lomas, M., ... & Hallmann, C. Putative sponge biomarkers in unicellular Rhizaria question an early rise of animals. Nature Ecology & Evolution 2019, 3(4), 577–581. [CrossRef]
- Shawar, L.; Love, G. D.; Uveges, B. T.; Zumberge, J. A.; Cárdenas, P.; Giner, J. L.; Summons, R. E. Chemical characterization of C31 sterols from sponges and Neoproterozoic fossil sterane counterparts. Proceedings of the National Academy of Sciences 2025, 122(41), e2503009122. [Google Scholar] [CrossRef] [PubMed]
- Gold, D. A.; Grabenstatter, J.; De Mendoza, A.; Riesgo, A.; Ruiz-Trillo, I.; Summons, R. E. Sterol and genomic analyses validate the sponge biomarker hypothesis. Proceedings of the National Academy of Sciences 2016, 113(10), 2684–2689. [Google Scholar] [CrossRef]
- Gong, Z.; Li, M. Astrochronology of the Ediacaran Shuram carbon isotope excursion, Oman. Earth and Planetary Science Letters 2020, 547, 116462. [Google Scholar] [CrossRef]
- Kayal, E., Bentlage, B., Sabrina Pankey, M., Ohdera, A. H., Medina, M., Plachetzki, D. C., ... & Ryan, J. F. Phylogenomics provides a robust topology of the major cnidarian lineages and insights on the origins of key organismal traits. BMC Evolutionary Biology 2018, 18(1), 68. [CrossRef]
- Tamm, S. L. Cilia and the life of ctenophores. Invertebrate Biology 2014, 133(1), 1–46. [Google Scholar] [CrossRef]
- Parry, L. A., Lerosey-Aubril, R., Coleman, R., Mouro, L. D., Gaines, R. R., Skabelund, J., ... & Ortega-Hernández, J. The oldest diverse jellyfish fauna reinterpreted as sessile polypoid dinomischids (stem-group Ctenophora). Journal of Systematic Palaeontology 2025, 23(1), 2545834. [CrossRef]
- Wood, R. Exploring the drivers of early biomineralization. Emerging Topics in Life Sciences 2018, 2(2), 201–212. [Google Scholar] [CrossRef]
- Komai, T. Studies on two aberrant ctenophores: Coeloplana and Gastrodes; Published by the author, 1922. [Google Scholar] [CrossRef]
- Franc, J. M. Organization and function of ctenophore colloblasts: an ultrastructural study. The Biological Bulletin 1978, 155(3), 527–541. [Google Scholar] [CrossRef]
- Harbison, G. R. On the classification and evolution of the Ctenophora. In The origin and relationships of lower invertebrates; Morris, S. C., et al., Eds.; Oxford University Press, 1985; pp. 78–100. [Google Scholar]
- David, C. N., Özbek, S., Adamczyk, P., Meier, S., Pauly, B., Chapman, J., ... & Holstein, T. W. Evolution of complex structures: minicollagens shape the cnidarian nematocyst. Trends in Genetics 2008, 24(9), 431–438. [CrossRef]
- Colin, S. P.; Costello, J. H.; Hansson, L. J.; Titelman, J.; Dabiri, J. O. Stealth predation and the predatory success of the invasive ctenophore Mnemiopsis leidyi. Proceedings of the National Academy of Sciences 2010, 107(40), 17223–17227. [Google Scholar] [CrossRef]
- Jager, M.; Chiori, R.; Alié, A.; Dayraud, C.; Quéinnec, E.; Manuel, M. New insights on ctenophore neural anatomy: immunofluorescence study in Pleurobrachia pileus (Müller, 1776). Journal of Experimental Zoology Part B: Molecular and Developmental Evolution 2011, 316(3), 171–187. [Google Scholar] [CrossRef] [PubMed]
- Dayraud, C.; Alié, A.; Jager, M.; Chang, P.; Le Guyader, H.; Manuel, M.; Quéinnec, E. Independent specialisation of myosin II paralogues in muscle vs. non-muscle functions during early animal evolution: a ctenophore perspective. BMC Evolutionary Biology 2012, 12(1), 107. [Google Scholar] [CrossRef]
- Tamm, S. L. Defecation by the ctenophore Mnemiopsis leidyi occurs with an ultradian rhythm through a single transient anal pore. Invertebrate Biology 2019, 138(1), 3–16. [Google Scholar] [CrossRef]
- Norekian, T. P.; Moroz, L. L. Neuromuscular organization of the Ctenophore Pleurobrachia bachei. Journal of Comparative Neurology 2019, 527(2), 406–436. [Google Scholar] [CrossRef]
- Wickett, N. J., Mirarab, S., Nguyen, N., Warnow, T., Carpenter, E., Matasci, N., ... & Leebens-Mack, J. Phylotranscriptomic analysis of the origin and early diversification of land plants. Proceedings of the National Academy of Sciences 2014, 111(45), E4859–E4868. [CrossRef]
- Puttick, M. N., Morris, J. L., Williams, T. A., Cox, C. J., Edwards, D., Kenrick, P., ... & Donoghue, P. C. The interrelationships of land plants and the nature of the ancestral embryophyte. Current Biology 2018, 28(5), 733–745. [CrossRef]
- Sousa, F.; Civáň, P.; Foster, P. G.; Cox, C. J. The chloroplast land plant phylogeny: analyses employing better-fitting tree-and site-heterogeneous composition models. Frontiers in Plant Science 2020, 11, 1062. [Google Scholar] [CrossRef] [PubMed]
- Donoghue, P. C.; Harrison, C. J.; Paps, J.; Schneider, H. The evolutionary emergence of land plants. Current Biology 2021, 31(19), R1281–R1298. [Google Scholar] [CrossRef]
- Huelsenbeck, J. P.; Bull, J. J.; Cunningham, C. W. Combining data in phylogenetic analysis. Trends in Ecology & Evolution 1996, 11(4), 152–158. [Google Scholar] [CrossRef] [PubMed]
- Hillis, D. M.; Moritz, C.; Mable, B. K. Molecular systematics; Sinauer, 1996; Vol. 23. [Google Scholar]
- King, N.; Rokas, A. Embracing uncertainty in reconstructing early animal evolution. Current Biology 2017, 27(19), R1081–R1088. [Google Scholar] [CrossRef]
- Yang, Z.; Rannala, B. Bayesian estimation of species divergence times under a molecular clock using multiple fossil calibrations with soft bounds. Molecular Biology and Evolution 2006, 23(1), 212–226. [Google Scholar] [CrossRef]
- Ho, S. Y.; Phillips, M. J. Accounting for calibration uncertainty in phylogenetic estimation of evolutionary divergence times. Systematic Biology 2009, 58(3), 367–380. [Google Scholar] [CrossRef] [PubMed]
- Warnock, R.; Yang, Z.; Donoghue, P. C. Exploring uncertainty in the calibration of the molecular clock. Biology Letters 2012, 8(1), 156–159. [Google Scholar] [CrossRef]
- Koch, N. M.; Garwood, R. J.; Parry, L. A. Fossils improve phylogenetic analyses of morphological characters. Proceedings of the Royal Society B 2021, 288(1950), 20210044. [Google Scholar] [CrossRef]
- Asher, R. J.; Smith, M. R. Phylogenetic signal and bias in paleontology. Systematic Biology 2022, 71(4), 986–1008. [Google Scholar] [CrossRef]
- Keating, J. N.; Garwood, R. J.; Sansom, R. S. Phylogenetic congruence, conflict and consilience between molecular and morphological data. BMC Ecology and Evolution 2023, 23(1), 30. [Google Scholar] [CrossRef]
- Turner, A. H.; Pritchard, A. C.; Matzke, N. J. Empirical and Bayesian approaches to fossil-only divergence times: a study across three reptile clades. PloS one 2017, 12(2), e0169885. [Google Scholar] [CrossRef] [PubMed]
| Type | Detail | ||
|---|---|---|---|
| Mechanism | Mitigating Factors | Mainly Affected Aspects1 | |
| Confounding Events | convergence, reversal |
low conv. & rev. potential, robust to micro-variation, extensive dataset |
molecular sequence***, morphology & embryology*, traditional fossil*, molecular fossil**** |
| Information Loss | saturation effect, gene loss, information reduction |
low evolutionary rate, multi-step loss required, extensive dataset |
molecular sequence***, morphology & embryology*, traditional fossil**, molecular fossil**** |
| Evidence Destruction | physicochemical destruction | extant, broad & stable record |
traditional fossil****, molecular fossil* |
| Rank & Type | Evaluation Dimension | Remark | |||
|---|---|---|---|---|---|
| Confounding Events | Information Loss | Accessibility | |||
| No.1 | HACHIMI | very rare | very rare | (very) easy1 | most promising |
| No.2 | Morph. & Embryo. | distinguishable | distinguishable | easy | currently underestimated |
| No.32 | Molecular Sequence | frequent | frequent | very easy | requires error correction |
| No.4 | Traditional Fossil | hard to discern | significant | very hard | — |
| No.5 | Molecular Fossil | indistinguishable | almost all | easy | — |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




