Submitted:
08 January 2026
Posted:
13 January 2026
You are already at the latest version
Abstract
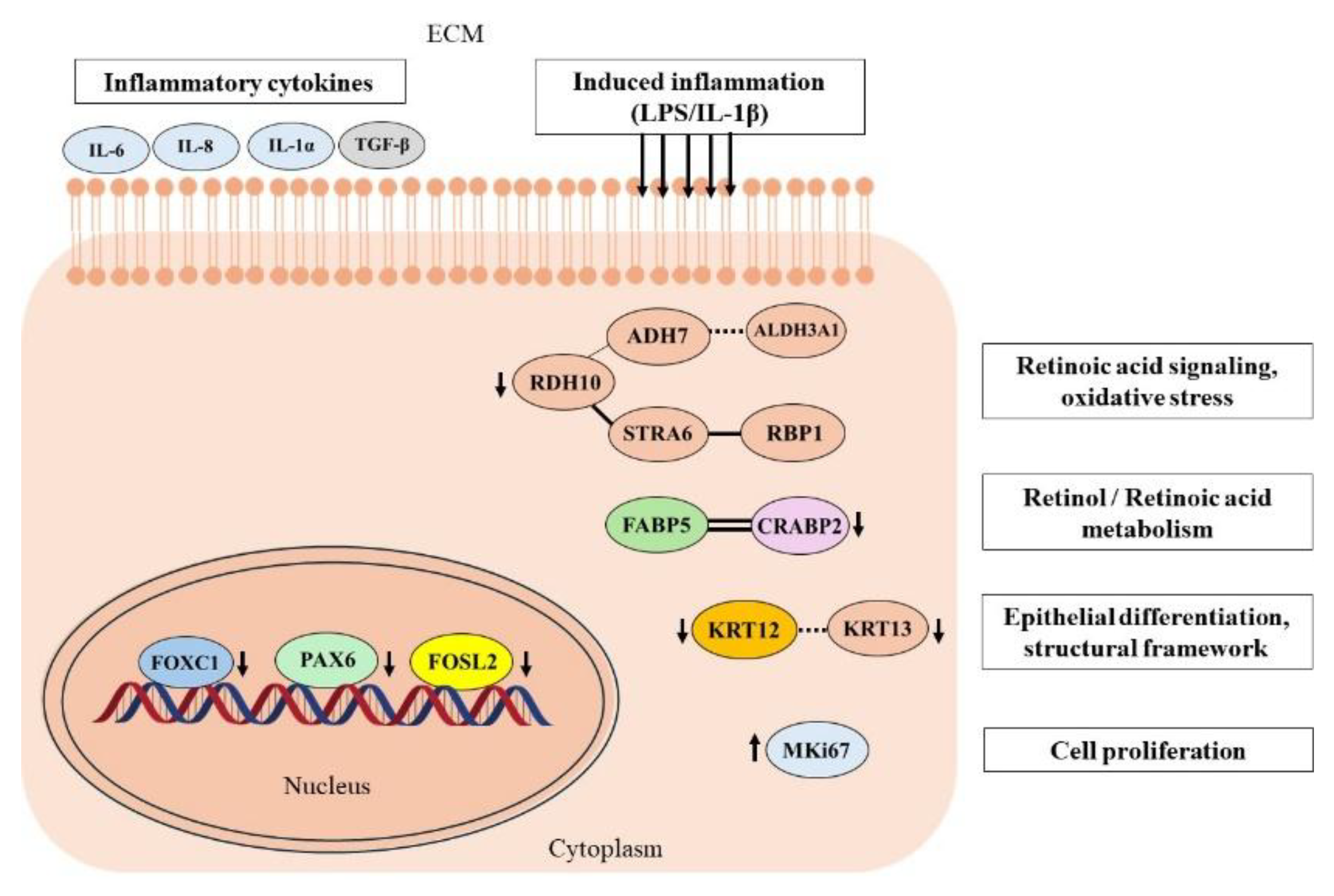
This study aimed to evaluate FOXC1-mediated regulatory mechanisms on gene and protein expression profiles in primary human limbal epithelial cells (pLECs), via siRNA knockdown; under basal and lipopolysaccharide (LPS) and interleukin-1β (IL-1β) induced inflammatory conditions. Gene expression was analysed for markers related to inflammation (CCL2, IL-6, IL-8, TNF-α, TGF-β), epithelial differentiation (KRT3, KRT12, KRT13, PAX6, FOXC1), cell proliferation and remodelling (FOSL2, MKi67, MMP2, VEGFA) and retinoic acid metabolism (ALDH3A1, CRABP2, CYP1B1, FABP5, RDH10, RBP1, STRA6). FOXC1 siRNA silencing in human pLECs significantly altered mRNA expression across multiple functional pathways, including inflammatory signaling (CCL2, IL-6, IL-8, IL-1α, VEGFA; p≤0.030), epithelial differentiation (KRT12, KRT13, PAX6; p≤0.045), cell proliferation and stress response (FOSL2, MKi67, VEGFA; p≤0.048) and retinoic acid metabolism (ALDH3A1, CRABP2, CYP1B1, FABP5, RDH10, STRA6; p≤0.037). Corresponding protein levels, evaluated by Western blotting and ELISA, were significantly modulated for the FABP5–CRABP2 axis, IL-6, IL-8, IL-1α, KRT12, KRT13, TGF-β, and RDH10 under different treatment conditions; (p≤0.045). FOXC1 maintains an anti-inflammatory, immune-quiescent state and coordinates TGF-β–mediated signaling, keratin expression, and retinoic acid metabolism to preserve corneal epithelial identity and homeostasis. Disruption of FOXC1 expression perturbs these pathways, potentially predisposing the ocular surface to fibrosis, lineage instability, and impaired regenerative capacity.
Keywords:
1. Introduction
2. Results
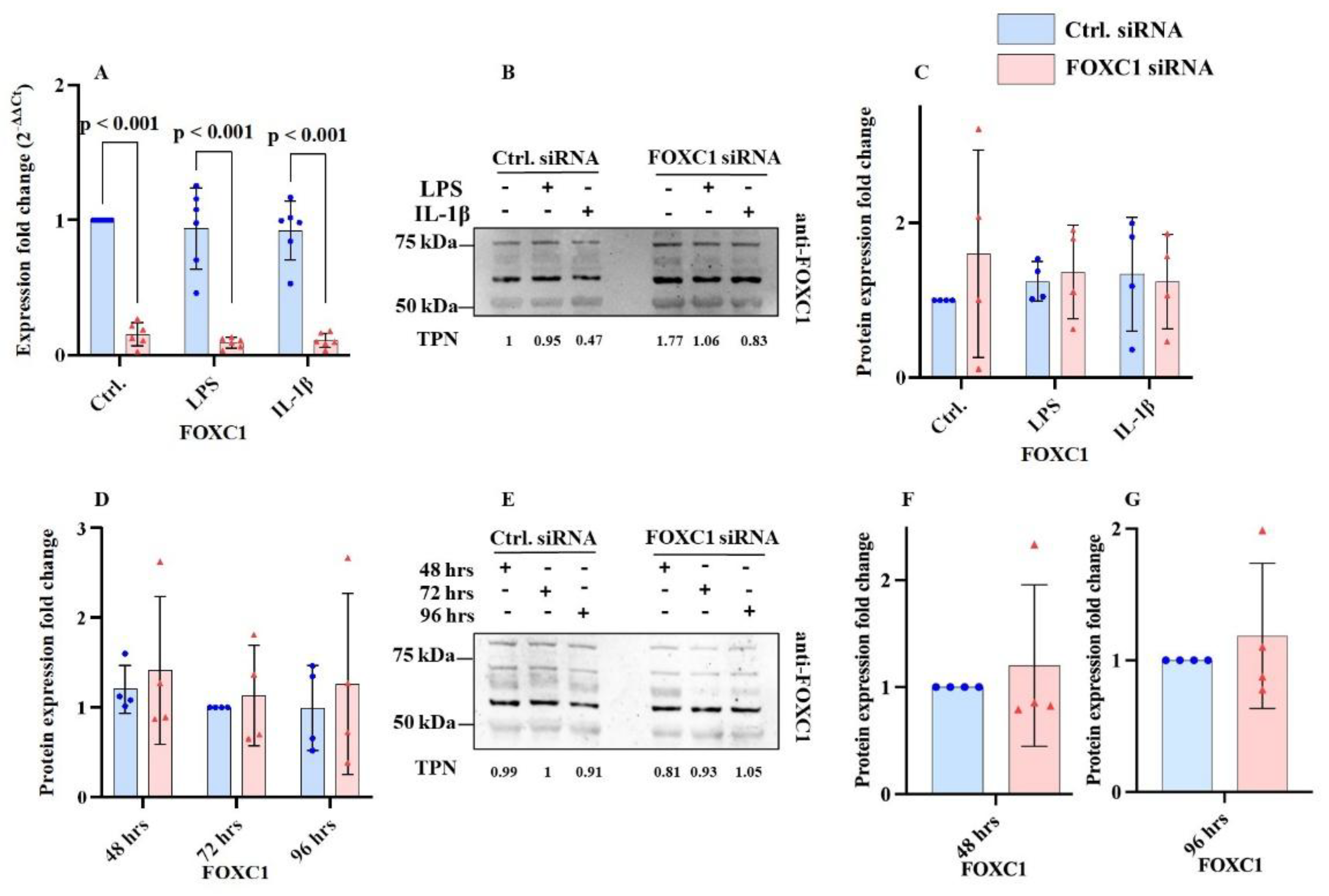
2.1. FOXC1 mRNA and Proteins Levels in pLECs
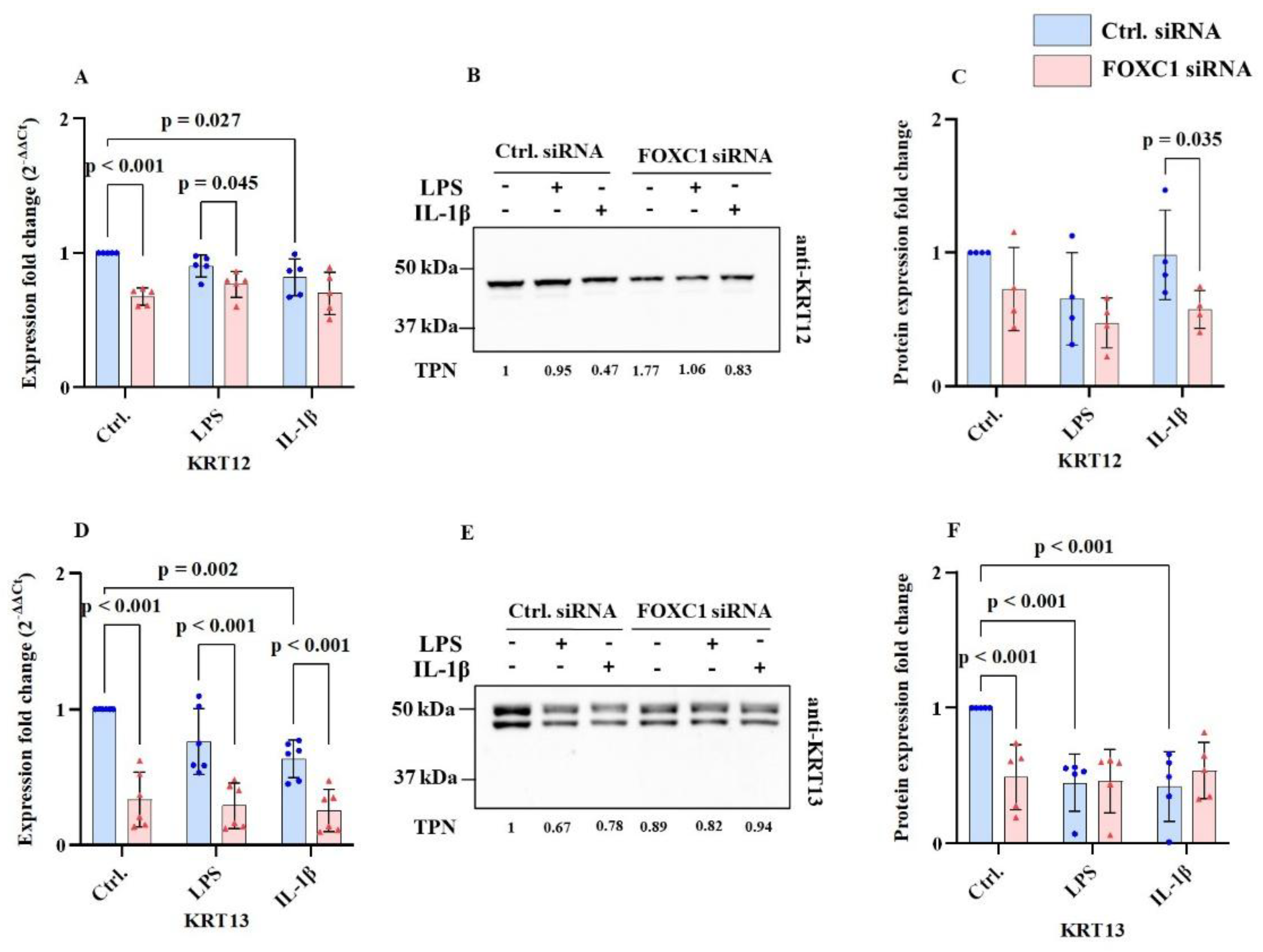
2.2. Cytokeratin mRNA and Protein Levels in pLECs
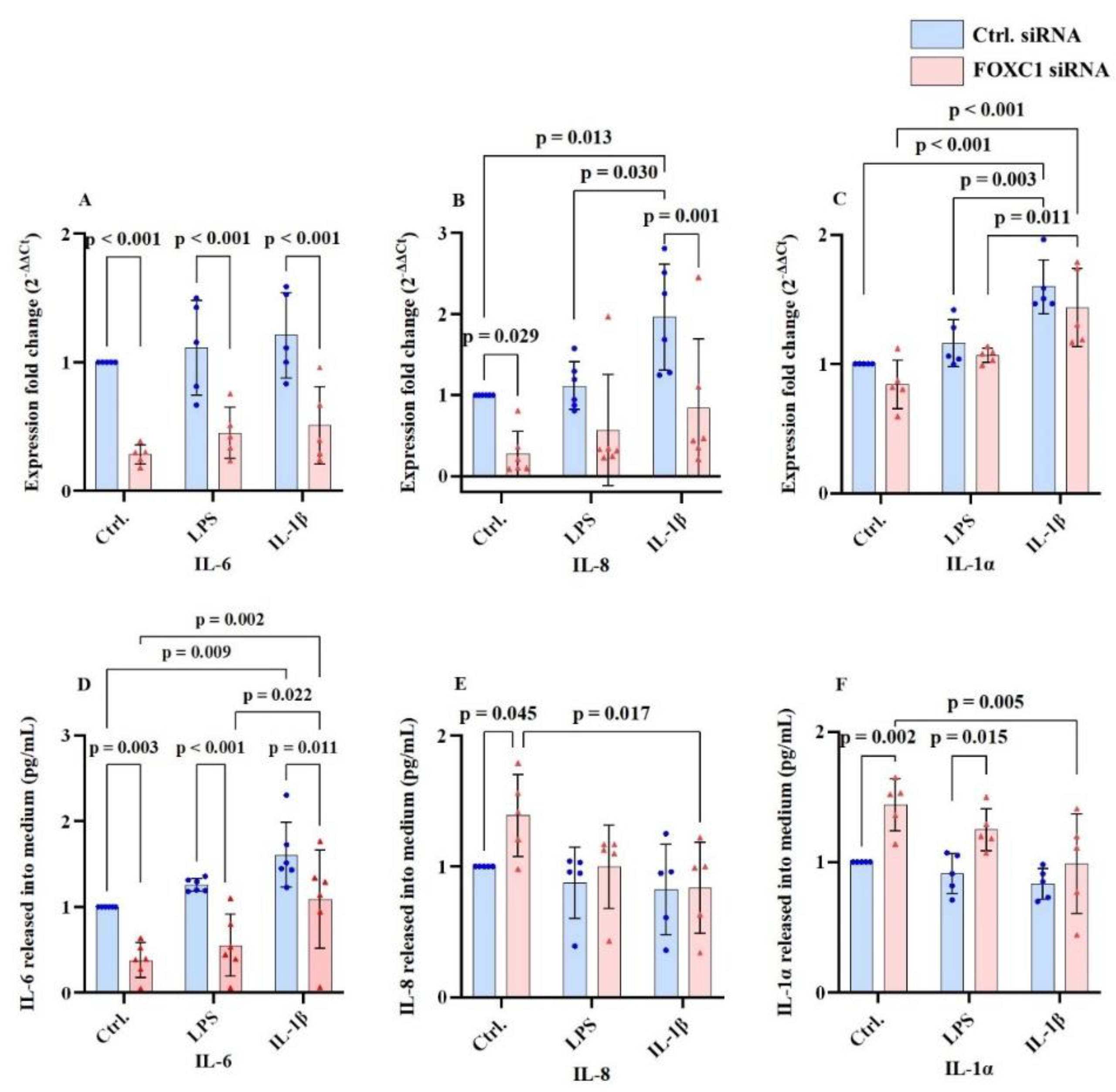
2.3. Interleukin mRNA and Protein Levels in pLECs
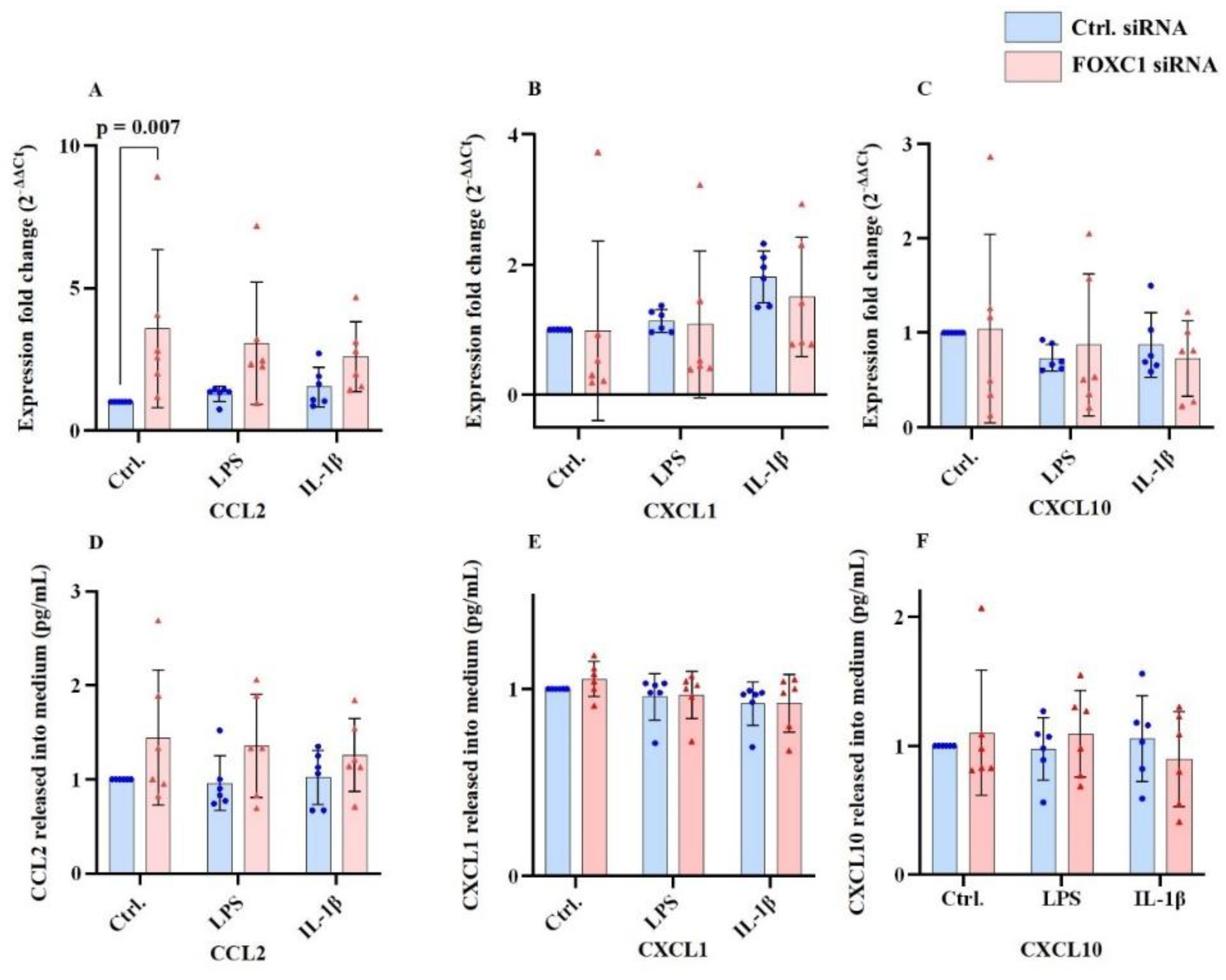
2.4. Chemokine Ligand mRNA and Protein Levels in pLECs
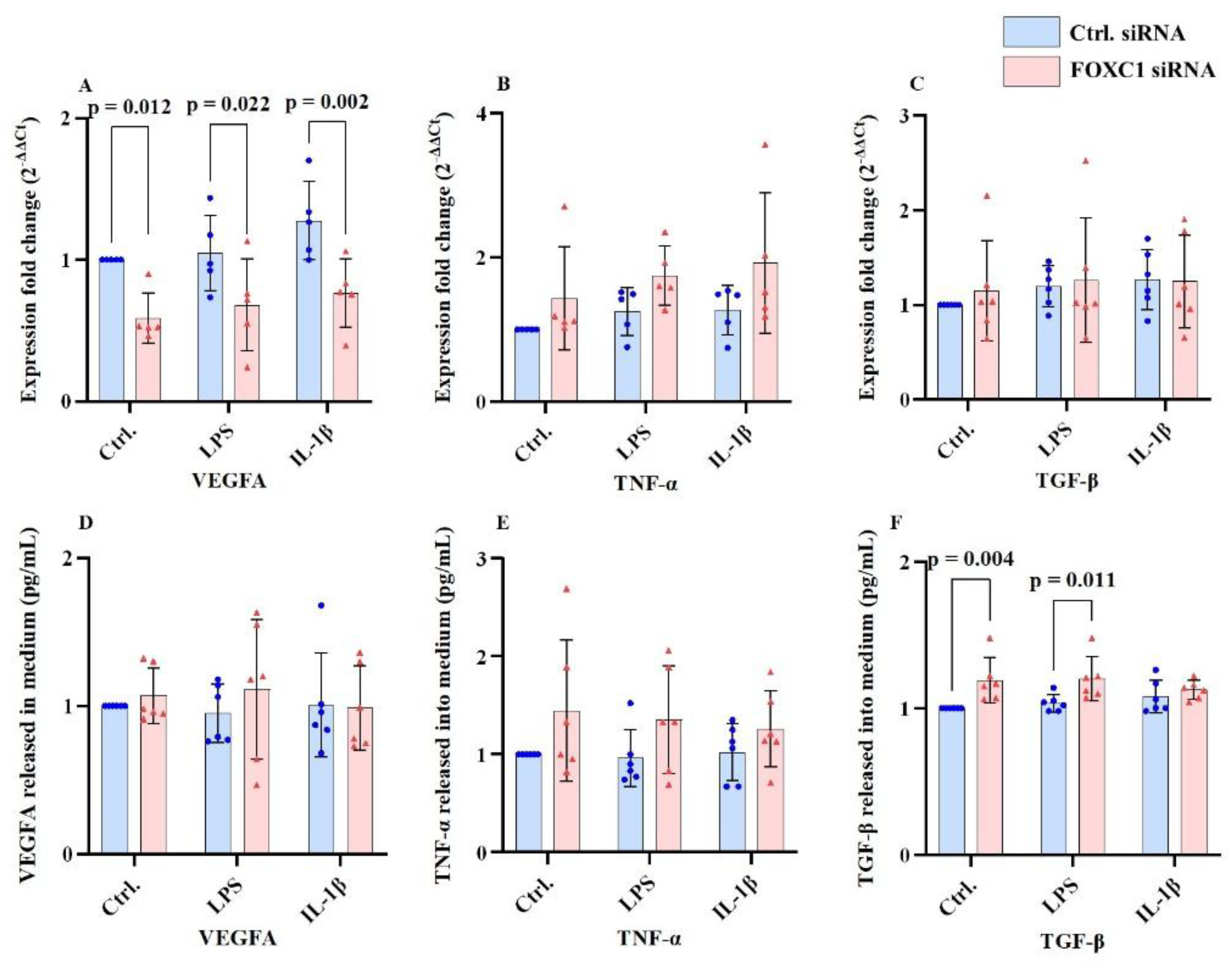
2.5. VEGFA, TNF-α and TGF-β mRNA and Proteins Levels in pLECs
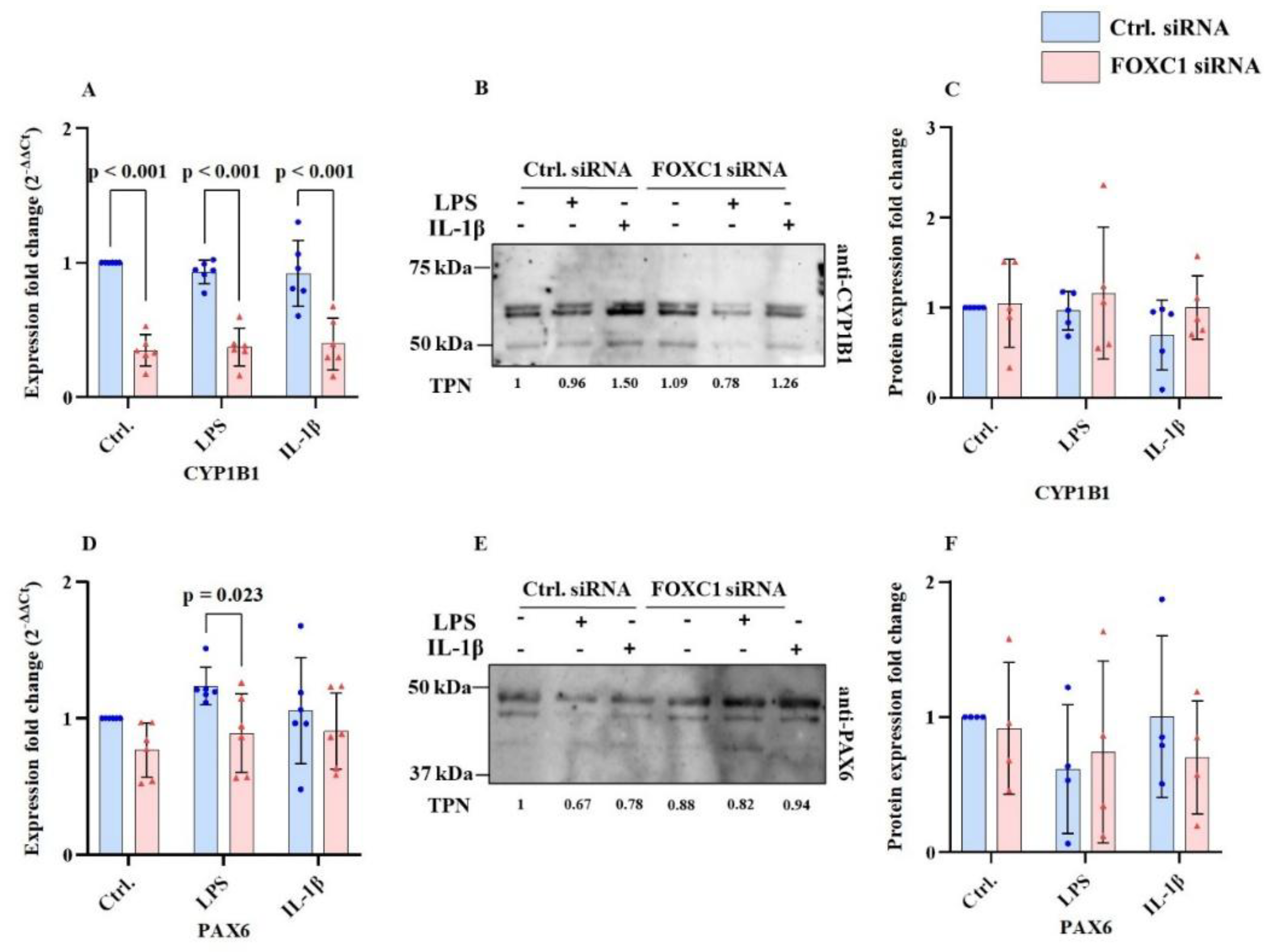
2.6. CYP1B1 and PAX6 mRNA and Proteins Levels in pLECs
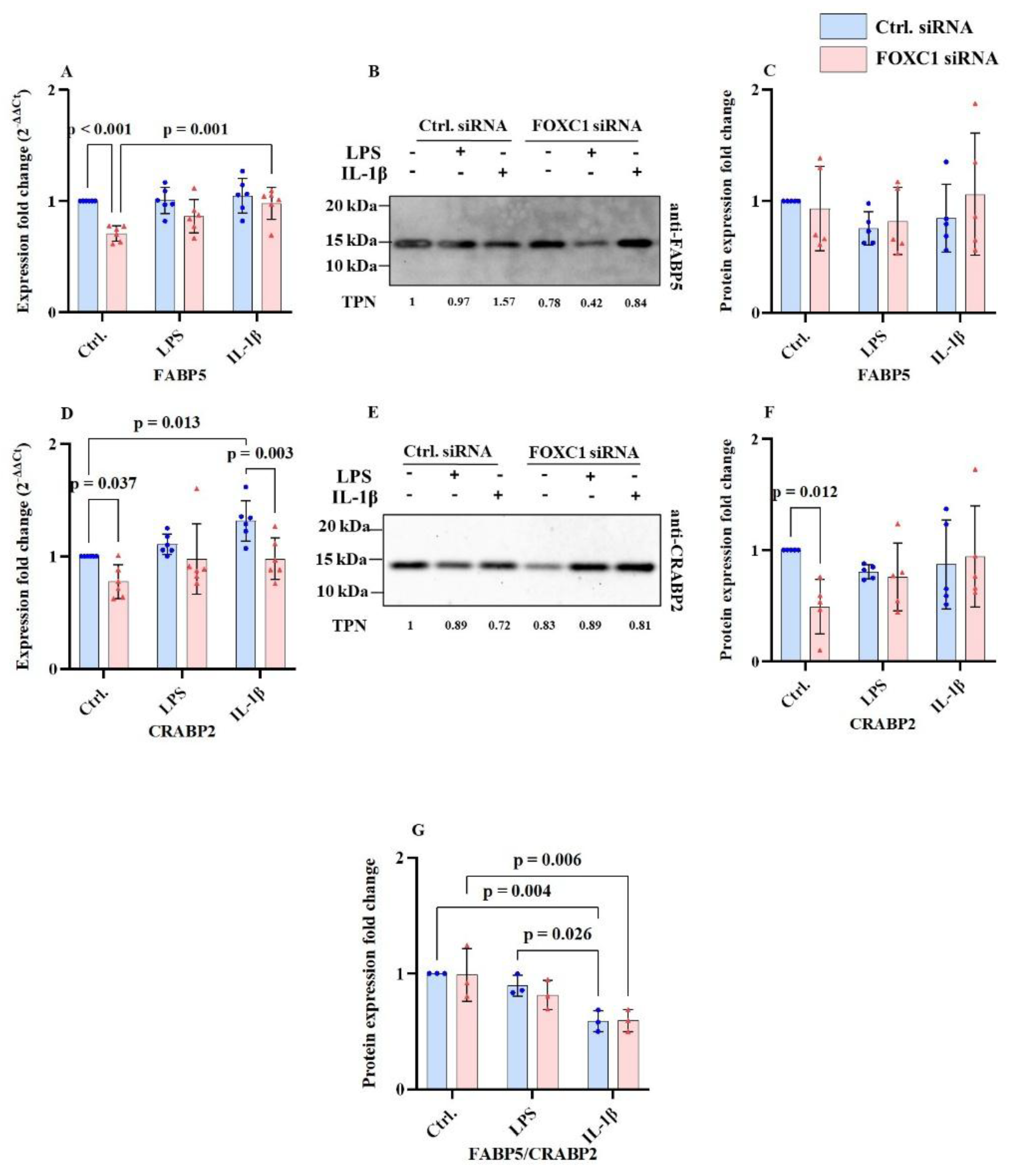
2.7. FABP5 and CRABP2 mRNA and Proteins Levels in pLECs
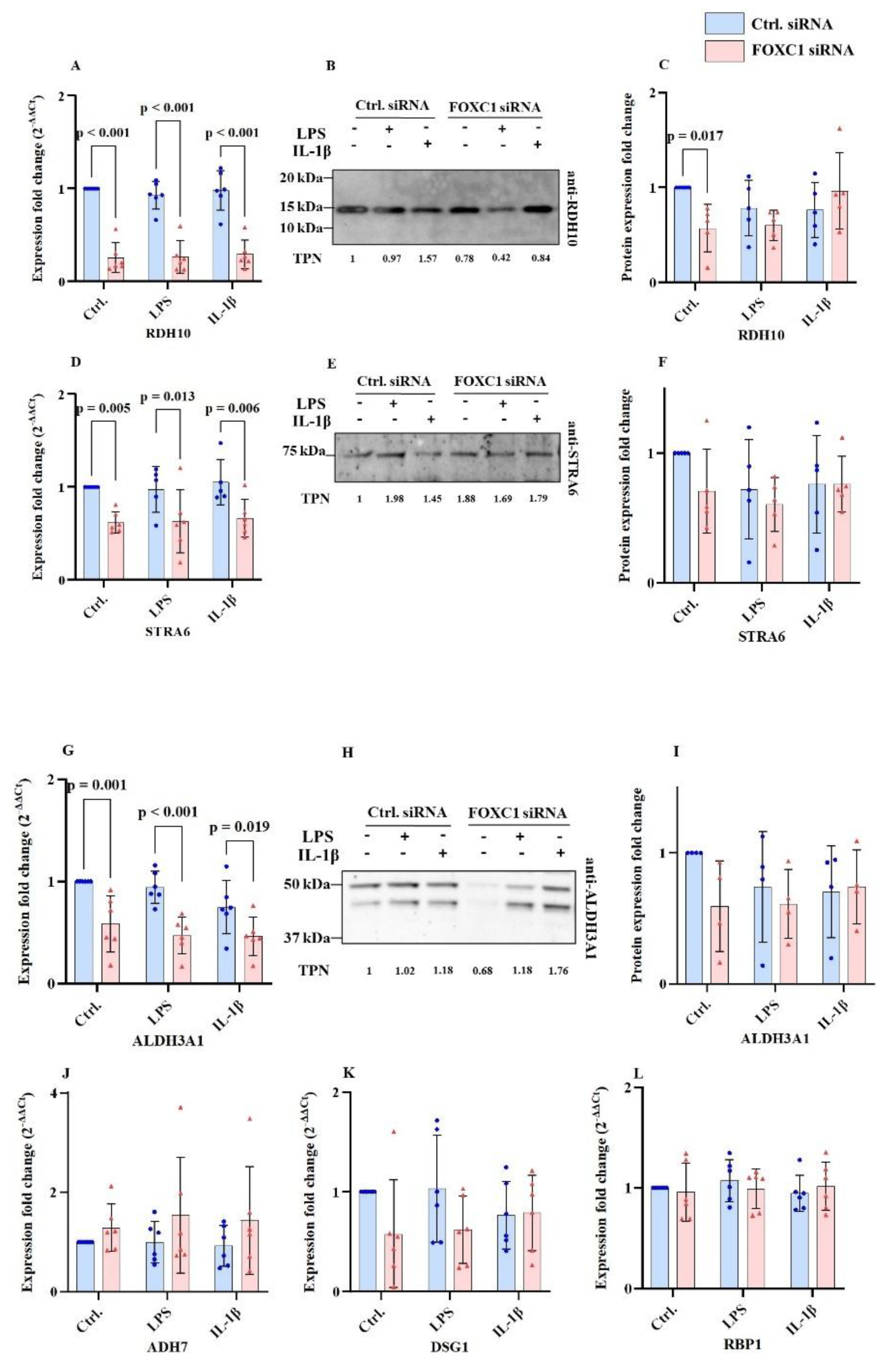
2.8. Further Retinoic Acid Signaling Pathway Related mRNA and Protein Levels in pLECs
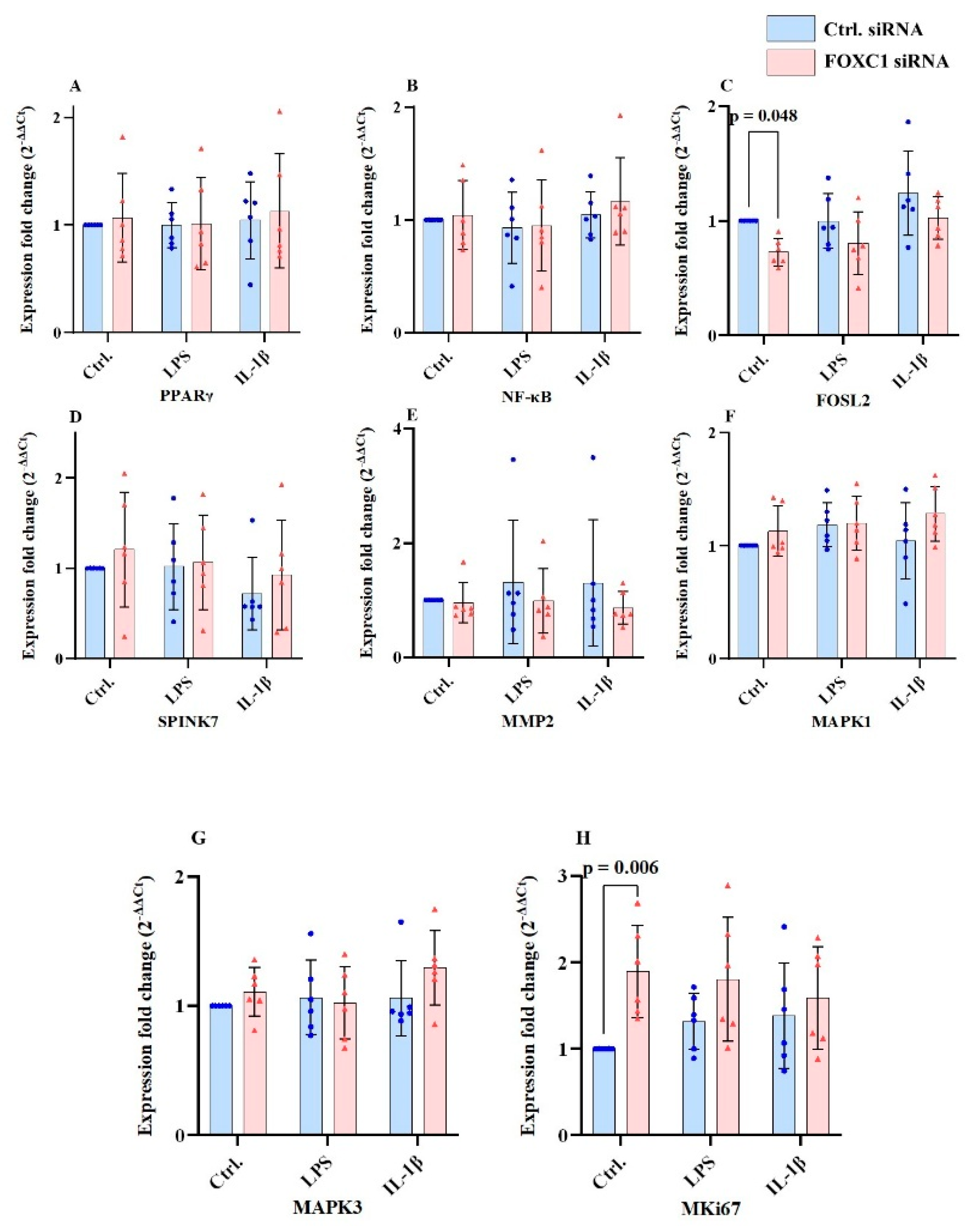
2.9. PPARγ, FOSL2, SPINK7, MMP2, MAPK1/ERK2, MAPK3 and Mki67 mRNA Levels in pLECs
3. Discussion
3.1. FOXC1 Transcriptional Efficiency, Protein Turnover Rate and Stability and Downstream Pathways
3.2. FOXC1 as a Regulator of Angiogenic Pathways and Chemokine and Cytokine Signaling
3.3. Implications of FOXC1 mRNA Silencing on pLEC Function, Differentiation and Corneal Homeostasis – via TGF-β Signaling and Epithelial Differentiation Markers
3.4. Understanding Retinoic Acid Metabolic Pathway and Its Link to Limbal Epithelium via FOXC1 siRNA Knockdown
4. Materials and Methods
4.1. Ethical Considerations
4.2. Cell Culture
4.2.1. siRNA Transfection of pLECs, and Inflammatory Treatment
4.3. RNA/Protein Extraction and cDNA Synthesis
4.4. Reverse Transcription Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
4.5. Western Blot Analysis
4.6. ELISA (Enzyme Linked Immunosorbent Assay) of Cell Culture Supernatant
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAK | Aniridia associated keratopathy |
| ADH7 | Alcohol dehydrogenase 7 |
| ALDH3A1 | Aldehyde dehydrogenase 3 Family member A1 |
| bp | base pairs |
| BSA | Bovine serum albumin |
| CCL2 | C-C motif chemokine ligand 2 |
| CXCL1 | C-X-C motif chemokine ligand 1 |
| CXCL10 | C-X-C motif chemokine ligand 10 |
| CRABP2 | Cellular retinoic acid binding protein 2 |
| CYP1B1 | Cytochrome P450 1B1 |
| Ct | Cycle threshold (in qPCR) |
| Ctrl | Control/ negative control |
| DSG1 | Desmoglein 1 |
| ECL | Enhanced chemiluminescence |
| ELISA | Enzyme-linked immuno sorbent assay |
| FABP5 | Fatty acid-binding protein 5 |
| FOSL2 | Fos like transcription protein 2 |
| FOXC1 | Forkhead Box C1 |
| HRP | Horseradish peroxidase |
| IL-1α | Interleukin 1α |
| IL-1β | Interleukin 1β |
| IL-6 | Interleukin 6 |
| IL-8 | Interleukin 8 |
| KEGG | Kyoto Encyclopaedia of Genes and Genomes |
| KGM | Keratinocyte growth medium |
| KRT3 | Keratin, type II cytoskeletal 3 |
| KRT12 | Keratin, type II cytoskeletal 12 |
| KRT13 | Keratin, type I cytoskeletal 13 |
| KRT19 | Keratin, type I cytoskeletal 19 |
| LECs | Limbal epithelial cells |
| LSCs | Limbal stem cells |
| LSCD | Limbal stem cell deficiency |
| LPS | lipopolysaccharide |
| μL | microliter |
| mL | milliliter |
| MAPK1 | Mitogen-activated protein kinase1 |
| MAPK3 | Mitogen-activated protein kinase3 |
| MKi67 | Marker of proliferation Ki-67 |
| MMP2 | Matrix metalloproteinase-2 Protein (type IV collagenase) |
| NFkβ | Nuclear factor kappa-light-chain-enhancer of activated B cells |
| OD | Optical density |
| PAX6 | Paired box protein Pax-6 |
| pmol | picomole |
| PPARγ | Peroxisome proliferator-activated receptor gamma |
| pg | picogram |
| qRT-PCR | quantitative Reverse Transcription Polymerase Chain Reaction |
| RBP1 | Retinol binding protein 1 |
| RDH10 | Retinol dehydrogenase 10 |
| SD | Standard deviation |
| SDS-PAGE | Sodium Dodecyl Sulfate–Polyacrylamide Gel Electrophoresis |
| SPINK7 | Serine Peptidase Inhibitor, Kazal Type |
| STRA6 | Signaling Receptor And Transporter Of Retinol 6 |
| STRING | Search Tool for Retrieval of Interacting Genes/Proteins |
| siRNA | Small interfering RNA |
| TBP | TATA box binding protein |
| TMB | 3,3′,5,5′-tetramethylbenzidine |
| TNF-α | Tumor Necrosis Factor-alpha |
| TGF-β | Transforming Growth Factor beta |
| TPN | Total protein normalisation |
| V | Volts |
| VEGFA | Vascular endothelial growth factorA |
References
- Schlötzer-Schrehardt, U; Latta, L; Gießl, A; et al. Dysfunction of the limbal epithelial stem cell niche in aniridia-associated keratopathy: Limbal niche in aniridia-associated keratopathy. Ocular Surface 2021, 21, 160–173. [Google Scholar] [CrossRef]
- Smits, JGA; Cunha, DL; Amini, M; et al. Identification of the regulatory circuit governing corneal epithelial fate determination and disease. PLoS Biol 2023, 21. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, H; Xue, Y; Lin, Y; et al. WNT7A and PAX6 define corneal epithelium homeostasis and pathogenesis. Nature 2014, 511, 358–361. [Google Scholar] [CrossRef]
- Zhu, L; Chen, C; Wu, S; et al. PAX6-WNK2 Axis Governs Corneal Epithelial Homeostasis. Invest Ophthalmol Vis Sci 2024, 65, 40. [Google Scholar] [CrossRef]
- Ouyang, J; Shen, YC; Yeh, LK; et al. Pax6 overexpression suppresses cell proliferation and retards the cell cycle in corneal epithelial cells. Invest Ophthalmol Vis Sci 2006, 47, 2397–2407. [Google Scholar] [CrossRef]
- Li, M; Zhu, L; Liu, J; et al. Loss of FOXC1 contributes to the corneal epithelial fate switch and pathogenesis. Signal Transduct Target Ther 2021, 6. [Google Scholar] [CrossRef]
- Latta, L; Figueiredo, FC; Ashery-Padan, R; et al. Pathophysiology of aniridia-associated keratopathy: Developmental aspects and unanswered questions. Ocular Surface 2021, 22, 245–266. [Google Scholar] [CrossRef]
- Lewis, CJ; Hedberg-Buenz, A; DeLuca, AP; et al. Primary congenital and developmental glaucomas. Hum Mol Genet 2017, 26, R28–R36. [Google Scholar] [CrossRef] [PubMed]
- Seo, S; Singh, HP; Lacal, PM; et al. Forkhead box transcription factor FoxC1 preserves corneal transparency by regulating vascular growth. Proc Natl Acad Sci U S A 2012, 109, 2015–2020. [Google Scholar] [CrossRef] [PubMed]
- Wang, X; Shan, X; Gregory-Evans, CY. A mouse model of aniridia reveals the in vivo downstream targets of Pax6 driving iris and ciliary body development in the eye. Biochim Biophys Acta Mol Basis Dis 2017, 1863, 60–67. [Google Scholar] [CrossRef]
- Omatsu, Y; Seike, M; Sugiyama, T; et al. Foxc1 is a critical regulator of haematopoietic stem/progenitor cell niche formation. Nature 2014, 508, 536–540. [Google Scholar] [CrossRef]
- Yang, Y; Li, W; Yang, H; et al. Research progress on the regulatory mechanisms of FOXC1 expression in cancers and its role in drug resistance. Gene 2024, 897. [Google Scholar] [CrossRef] [PubMed]
- Arcot Sadagopan, K; Liu, GT; Capasso, JE; et al. Anirdia-like phenotype caused by 6p25 dosage aberrations. Am J Med Genet A 2015, 167, 524–528. [Google Scholar] [CrossRef]
- Micheal, S; Siddiqui, SN; Zafar, SN; et al. A novel homozygous mutation in FOXC1 causes axenfeld Rieger syndrome with congenital glaucoma. PLoS One 2016, 11. [Google Scholar] [CrossRef]
- Ito, YA; Footz, TK; Berry, FB; et al. Severe molecular defects of a novel FOXC1 W152G mutation result in aniridia. Invest Ophthalmol Vis Sci 2009, 50, 3573–3579. [Google Scholar] [CrossRef]
- François, M; Ramchandran, R. Studies on Axenfeld-Rieger syndrome patients and mice reveal Foxc1’s role in corneal neovascularization. Proc Natl Acad Sci U S A 2012, 109, 1818–1819. [Google Scholar] [CrossRef]
- Han, B; Bhowmick, N; Qu, Y; et al. FOXC1: An emerging marker and therapeutic target for cancer. Oncogene 2017, 36, 3957–3963. [Google Scholar] [CrossRef] [PubMed]
- Yin, L; Li, Q; Mrdenovic, S; et al. KRT13 promotes stemness and drives metastasis in breast cancer through a plakoglobin/c-Myc signaling pathway. Breast Cancer Research 2022, 24. [Google Scholar] [CrossRef]
- Tibrewal, S; Ratna, R; Gour, A; et al. Clinical and molecular aspects of congenital aniridia - A review of current concepts. Indian J Ophthalmol 2022, 70, 2280–2292. [Google Scholar] [CrossRef] [PubMed]
- Samant, M; Chauhan, BK; Lathrop, KL; et al. Congenital aniridia: Etiology, manifestations and management. Expert Rev Ophthalmol 2016, 11, 135–144. [Google Scholar] [CrossRef]
- Landsend, ECS; Utheim, ØA; Pedersen, HR; et al. The level of inflammatory tear cytokines is elevated in congenital Aniridia and associated with meibomian gland dysfunction. Invest Ophthalmol Vis Sci 2018, 59, 2197–2204. [Google Scholar] [CrossRef]
- Ihnatko, R; Edén, U; Lagali, N; et al. Analysis of protein composition and protein expression in the tear fluid of patients with congenital aniridia. J Proteomics 2013, 94, 78–88. [Google Scholar] [CrossRef]
- Latta, L; Nordström, K; Stachon, T; et al. Expression of retinoic acid signaling components ADH7 and ALDH1A1 is reduced in aniridia limbal epithelial cells and a siRNA primary cell based aniridia model. Exp Eye Res 2019, 179, 8–17. [Google Scholar] [CrossRef]
- Latta, L; Knebel, I; Bleil, C; et al. Similarities in DSG1 and KRT3 down regulation through retinoic acid treatment and PAX6 knockdown related expression profiles: Does PAX6 affect RA signaling in limbal epithelial cells? Biomolecules 2021, 11. [Google Scholar] [CrossRef] [PubMed]
- Katiyar, P; Stachon, T; Fries, FN; et al. Decreased FABP5 and DSG1 protein expression following PAX6 knockdown of differentiated human limbal epithelial cells. Exp Eye Res 2022, 215. [Google Scholar] [CrossRef] [PubMed]
- Kundu, S; Amini, M; Stachon, T; et al. Effect of isolated keratin 3 knockdown on gene expression of primary limbal epithelial cells without and with inflammatory stimuli. Annals of Anatomy 2025, 260. [Google Scholar] [CrossRef] [PubMed]
- Schwanhüusser, B; Busse, D; Li, N; et al. Global quantification of mammalian gene expression control. Nature 2011, 473, 337–342. [Google Scholar] [CrossRef]
- Perl, K; Ushakov, K; Pozniak, Y; et al. Reduced changes in protein compared to mRNA levels across non-proliferating tissues. BMC Genomics 2017, 18. [Google Scholar] [CrossRef]
- Vogel, C.; Marcotte, E. M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nature Reviews Genetics 2012, 13(4), 227–232. [Google Scholar] [CrossRef]
- Koo, H-Y; Kume, T. FoxC1-dependent regulation of vascular endothelial growth factor signaling in corneal avascularity. Trends Cardiovasc Med 2013, 23(1), 1–4. [Google Scholar] [CrossRef]
- Bhakuni, T; Norden, PR; Ujiie, N; et al. FOXC1 regulates endothelial CD98 (LAT1/4F2hc) expression in retinal angiogenesis and blood-retina barrier formation. Nat Commun 2024, 15. [Google Scholar] [CrossRef]
- Ray, T; Ryusaki, T; Ray, PS. Therapeutically Targeting Cancers That Overexpress FOXC1: A Transcriptional Driver of Cell Plasticity, Partial EMT, and Cancer Metastasis. Front Oncol 2021, 11, 721959. [Google Scholar] [CrossRef]
- Liu, Y; Chen, S; Tian, M; et al. FOXC1 restrains NF-κB-mediated interleukin-1β transcription in breast cancer. MedComm (Beijing) 2023, 4(6), e440. [Google Scholar] [CrossRef]
- Suwara, MI; Green, NJ; Borthwick, LA; et al. IL-1a released from damaged epithelial cells is sufficient and essential to trigger inflammatory responses in human lung fibroblasts. Mucosal Immunol 2014, 7, 684–693. [Google Scholar] [CrossRef]
- Zhou, Y; Kato, H; Asanoma, K; et al. Identification of FOXC1 as a TGF-beta1 responsive gene and its involvement in negative regulation of cell growth. Genomics 2002, 80(5), 465–72. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H; Nie, J; Bao, Z; et al. FOXC1 promotes EMT and colorectal cancer progression by attracting M2 macrophages via the TGF-β/Smad2/3/snail pathway. Cell Signal 2025, 130. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y; Yeh, LK; Liu, H; et al. Targeted overexpression of TGF-α in the corneal epithelium of adult transgenic mice induces changes in anterior segment morphology and activates noncanonical Wnt Signaling. Invest Ophthalmol Vis Sci 2013, 54, 1829–1837. [Google Scholar] [CrossRef]
- Yang, Z; Jiang, S; Cheng, Y; et al. FOXC1 in cancer development and therapy: deciphering its emerging and divergent roles. Ther Adv Med Oncol 2017, 9, 797–816. [Google Scholar] [CrossRef]
- Latta, L; Viestenz, A; Stachon, T; et al. Human aniridia limbal epithelial cells lack expression of keratins K3 and K12. Exp Eye Res 2018, 167, 100–109. [Google Scholar] [CrossRef]
- Siegenthaler, JA; Ashique, AM; Zarbalis, K; et al. Retinoic Acid from the Meninges Regulates Cortical Neuron Generation. Cell 2009, 139, 597–609. [Google Scholar] [CrossRef] [PubMed]
- Como, CN; O’Rourke, R; Winkler, C; et al. Meningeal-derived retinoic acid regulates neurogenesis via suppression of Notch and Sox2. Cell Rep 2025, 44. [Google Scholar] [CrossRef]
- Celik, SD; Ates, O. Analysis of CRABP2 and FABP5 genes in primary and recurrent pterygium tissues. Mol Biol Rep 2020, 47, 6105–6110. [Google Scholar] [CrossRef] [PubMed]
- Ramachandran, R; Ibragimova, S; Woods, LM; et al. Conserved role of FOXC1 in TNBC is parallel to FOXA1 in ER+ breast cancer. iScience 2024, 27. [Google Scholar] [CrossRef] [PubMed]
- Tang, X; Liang, Y; Sun, G; et al. Upregulation of CRABP2 by TET1-mediated DNA hydroxymethylation attenuates mitochondrial apoptosis and promotes oxaliplatin resistance in gastric cancer. Cell Death Dis. 2022, 13(10), 848. [Google Scholar] [CrossRef] [PubMed]
| Donors | Gender (42.85% Male) | Age (Years)76.16 ±6.87 (67-92) |
| Donor 1 | Male | 67 |
| Donor 2 | Male | 68 |
| Donor 3 | Female | 72 |
| Donor 4 | Male | 77 |
| Donor 5 | Female | 81 |
| Donor 6 | Female | 92 |
| Donor 7 | Female | unknown |
| Target mRNA transcript with transcript ID (source: NCBI) | Catalogue No. | Amplicon size (bp) |
| ADH7: NM_000673 | QT00000217 | 87 bp |
| ALDH3A1: NM_000691 | QT0240193 | 121 bp |
| CCL2: NM_002982 | QT00212730 | 60 bp |
| CRABP2: NM_001199723 | QT00063434 | 140 bp |
| CXCL1: NM_001511 | QT00199752 | 120 bp |
| CXCL10: NM_001565 | QT01003065 | 129 bp |
| CYP1B1: NM_000104 | QT00209496 | 114 bp |
| DSG1: NM_001942 | QT00001617 | 96 bp |
| FABP5: NM_001444 | QT00225561 | 97 bp |
| FOSL2: NM_005253, XM_005264231, XM_006711976, XM_006711977 | QT01000881 | 126 bp, 177 bp, 229 bp |
| FOXC1: NM_001453 | QT00217161 | 109 bp |
| IL-1α: NM_000575 | QT00001127 | 74 bp |
| IL-6: NM_000600, XM_005249745 | QT00083720 | 107 bp |
| IL-8: NM_000584 | QT00000322 | 102 bp |
| KRT3: NM_057088, XM_006719393, XM_005268859, XM_006719392 | QT00050365 | 118 bp |
| KRT12: NM_000223 | QT00011949 | 104 bp |
| KRT13: NM_002274, NM_153490 | QT00068747 | 60 bp |
| KRT19: NM_002276 | QT00081137 | 117 bp |
| MAPK1: NM_002745, NM_138957 | QT00065933 | 118 bp |
| MAPK3: NM_001109891, NM_002746 | QT02589314 | 180 bp |
| MKi67: NM_001145966, NM_002417, XM_006717864 | QT00014203 | 86 bp |
| MMP2: NM_004530 | QT02395778 | 95 bp |
| NFkβ: (RELA 2): NM_001243984, NM_001243985, NM_001145138, NM_021975 | QT02324308 | 136 bp |
| PAX6: NM_000280, NM_001127612, NM_001604, NM_001258462, NM_001258463, NM_001258464, NM_001258465 | QT00071169 | 113 bp |
| PPARγ: NM_005037, NM_015869, NM_138711, NM_138712, XM_006713208 | QT00029841 | 113 bp |
| RBP1: NM_002899, NM_001130992, NM_001130993 | QT01850296 | 126 bp |
| RDH10: NM_172037 | QT00029176 | 107 bp |
| SPINK7: NM_032566 | QT00039585 | 126 bp |
| STRA6: NM_001142618, NM_022369, NM_001142617, NM_001199042, NM_001142619, NM_001199041, NM_001199040 | QT00006748 | 74 bp |
| TBP: NM_001172085, NM_003194 | QT00000721 | 132 bp |
| TNF-α: NM_000594 | QT00029162 | 98 bp |
| TGF-β: NM_000594 | QT00029162 | 98 bp |
| VEGA: NM_001025366, NM_001025367, NM_001025368, NM_001033756, NM_001171623, NM_001171624, NM_001171625, NM_001171626, NM_001171629, NM_003376, NM_001287044 | QT01010184 | 150 bp 204 bp, 222 bp, 273 bp |
|
Antibody |
Source |
Molecular weight (kDa) |
Catalog No./ Manufacturer |
Dilution |
| ALDH3A1 | Rabbit, polyclonal | 48 kDa | 15578-1-AP, Proteintech, USA | 1:3000 |
| CRABP2 | Mouse, monoclonal | 14 kDa | 10225-1-AP, Proteintech, USA | 1:2500 |
| CYP1B1 | Rabbit, polyclonal | 61 kDa | 18505-1-AP, Proteintech, USA | 1:250 |
| FABP5 | Rabbit, polyclonal | 15 kDa | 12348-1-AP, Proteintech, USA | 1:1000 |
| FOSL2 | Mouse, monoclonal | 50 kDa | TA809660S, OriGene Technologies, USA | 1:2000 |
| (i) FOXC1 (ii) FOXC1 |
Rabbit, polyclonal Mouse, monoclonal |
72 kDa 55 kDa |
30082-1-AP, Proteintech, USA MAB6329-SP, R&D Biotechne, USA |
1:500 1:500 |
| KRT12 | Mouse, monoclonal | 54 kDa | sc-515882, Santa Cruz Biotechnology, USA | 1:100 |
| KRT13 | Mouse, monoclonal | 52 kDa | sc-101460, Santa Cruz Biotechnology, USA | 1:200 |
| PAX6 | Rabbit, polyclonal | 46-48 kDa | AB-2237, Merck, Germany | 1:1000 |
| RDH10 | Rabbit, polyclonal | 39 kDa | 14644-1-AP, Proteintech, USA | 1:1000 |
| STRA6 | Rabbit, polyclonal | 73-78 kDa | 22001-1-AP, Proteintech, USA | 1:2500 |
| Detected protein | Catalog No./Lot No. | Range of detection |
| CCL2 | DY279-05 / P456545 | 1000 pg/mL - 7.8 pg/mL |
| CXCL1 | DY275-05 / P436085 | 2000 pg/mL - 15.65 pg/mL |
| CXCL10 | DY266-05 / P432346 | 2000 pg/mL - 15.65 pg/mL |
| IL-1α | DY200-05 / P410705 | 500 pg/mL - 3.90 pg/mL |
| IL-6 | DY206-05 / P437281 | 600 pg/mL - 4.7 pg/mL |
| IL-8 | DY208-05 / P445124 | 2000 pg/mL - 15.65 pg/mL |
| TNF-α | DY210-05 / P452568 | 1000 pg/mL - 7.8 pg/mL |
| TGF-β | DY240-05 / P424503 | 2000 pg/mL - 15.65 pg/mL |
| VEGFA | DY293B-05 /P394527 | 2000 pg/mL - 15.65 pg/mL |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




