1. Introduction
Early guidance in sports has traditionally relied on a coach’s eye, observation during training, and basic fitness tests. These approaches, while valuable, often miss important biological differences between children, such as how quickly they recover, how their muscles respond to repeated loads, or how sensitive their joints and tendons are to stress. For health coaches and sports trainers working with young athletes, this can make it difficult to decide who needs extra rest, who can safely handle more training, and which type of training is likely to benefit each child the most.
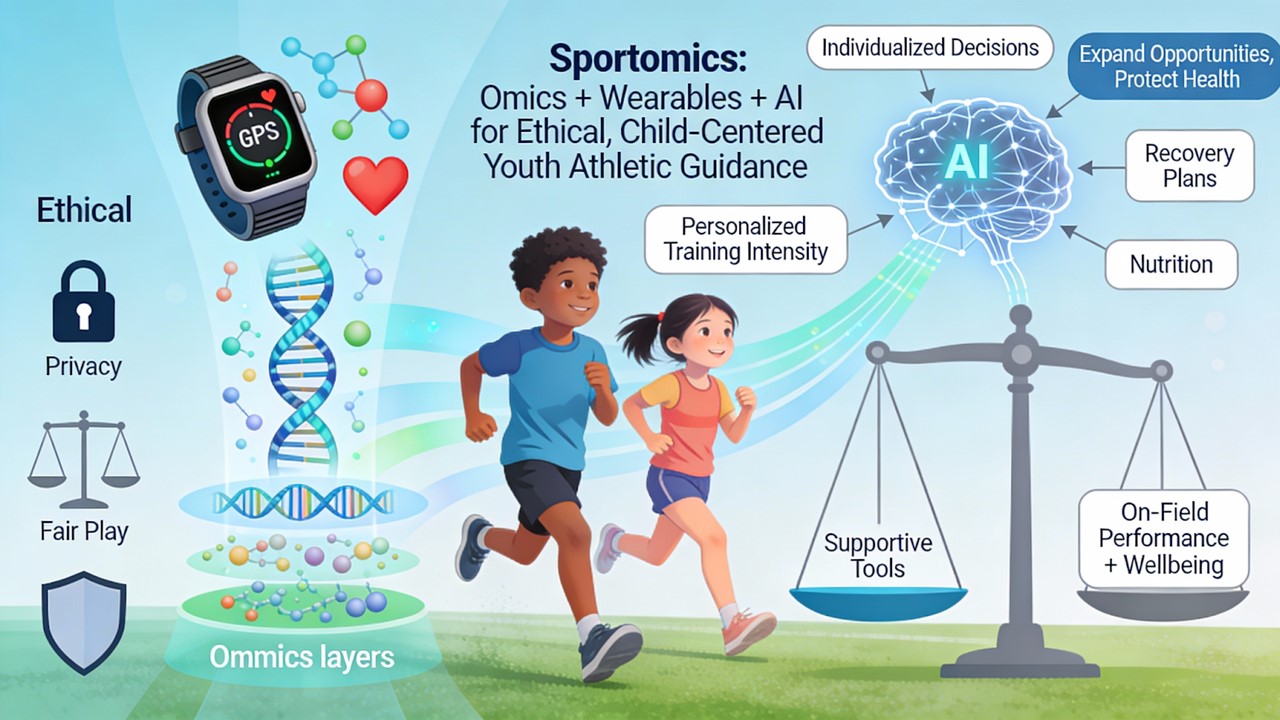
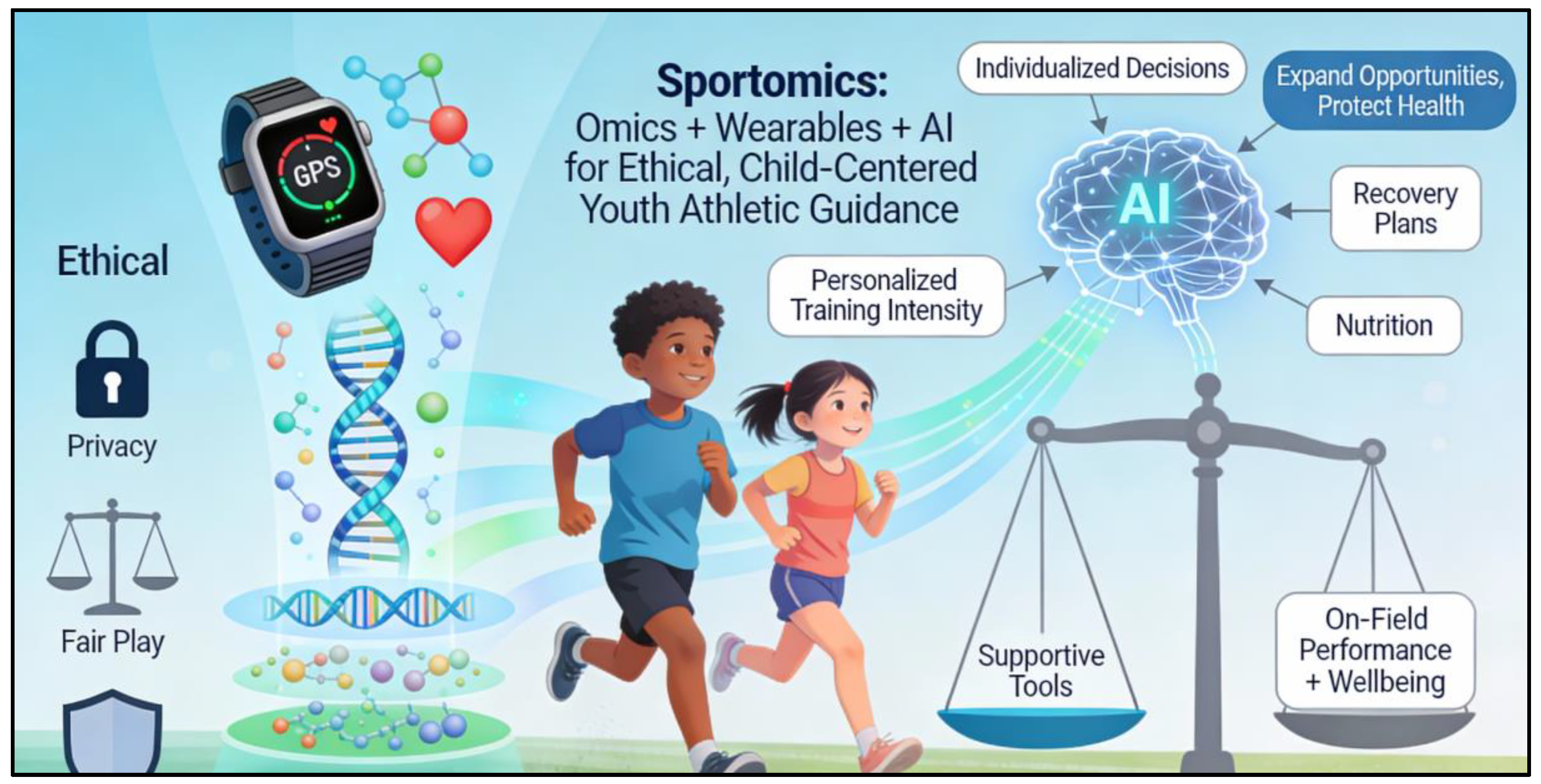
Sportomics offers a structured way to answer these questions by combining omics tools (genomics, proteomics, metabolomics) with data from wearables and Artificial Intelligence (AI). Instead of guessing, coaches and parents can use biological information to support decisions about training intensity, recovery, and nutrition in a way that is individualized and child-centred. Importantly, these tools are supportive, as they provide probabilities and tendencies that must always be interpreted alongside on-field performance, motivation, and overall wellbeing (Musat et al., 2024). (As shown in
Figure 1.)
At the same time, the use of genetic and AI-based tools in children raises ethical concerns around privacy, fairness, and the risk of labelling a child too early. Responsible sportomics requires that health coaches, parents, and academies use these technologies only to expand opportunities and protect health, never to limit or exclude a child based on a test result. This review summarizes key molecular techniques and AI-based tools relevant to youth sports, explains what they can and cannot tell us, and provides practical guidance for parents, coaches, and healthcare professionals on how to integrate sportomics ethically into training and talent development. (Collier et al., 2011).
1.2. Overview of Recent Advancements in Genomics
Recent progress in genomics has identified several genetic polymorphisms associated with traits towards higher performance. For instance, the ACE I/D variant influences cardiovascular efficiency and endurance capacity, while the ACTN3 R577X polymorphism affects muscle fiber composition and sprint performance (El Ouali et al., 2024; Ahmetov & Rogozkin, 2020). Similarly, COL1A1 and COL5A1 variants contribute to collagen synthesis and connective tissue resilience factors crucial for tissue integrity and injury prevention (Rodríguez-Larrad et al., 2020; Rodríguez-Romo & Peñalvo, 2022). Integrating these molecular insights with AI-based analytics enables the construction of predictive models capable of identifying genetic predispositions for strength, endurance, and injury risk (Claudino et al., 2021; Li et al., 2021). This integration enhances scientific understanding of performance variation and allows the design of individualized training and rehabilitation programs aligned with an athlete’s biological profile (Ahmetov et al., 2021; Nashef & Smith, 2023).
Across the world, sports organizations and academies are beginning to incorporate molecular insights and objective data into talent development pathways. The Olympic movement, major football clubs, and youth development programs increasingly rely on sportomics-based profiling to identify biological strengths and vulnerabilities (Goes et al., 2021; Huygens et al., 2022). In India, initiatives such as Khelo India and TOPS reflect a growing shift toward evidence-based approaches for nurturing emerging talent. By linking molecular information with real-world observations, sportomics helps parents and coaches appreciate each child’s unique physiological characteristics and support them more effectively (Lemmink et al., 2021; Hamilton & Hopkins, 2021).
1.3. Relation Between Sportomics and Digital Monitoring Devices
Wearable and Internet of Things (IoT) devices now form an essential component of this ecosystem. Smartwatches, GPS trackers, and motion sensors routinely collect data on heart rate, speed, sleep, and workload intensity. When these measurements are interpreted alongside molecular profiles and physiological markers, they provide meaningful insights into training load and recovery patterns (Piette et al., 2022; Babu et al., 2024).
For young athletes, such integration supports proactive adjustments before fatigue, overreaching, or overuse injuries (Muniz-Santos et al., 2021; Nascimento et al., 2022). Wearable-driven analytics supported by AI also allow parents to understand when rest is needed and how to structure recovery without undue strain (Khosravi et al., 2022; Claudino et al., 2022). Sportomics becomes especially valuable during childhood and adolescence, when the body is rapidly developing and more prone to stress (O’Connell et al., 2021; Ziemann et al., 2020).
1.4. Ethical Guidelines Responsible for Athletic Empowerment
Beyond performance improvement, sportomics supports an ethical and sustainable vision of youth sports. By making scientific tools more accessible, it promotes fairness and helps children from diverse backgrounds benefit from insights once limited to elite environments (Clarke et al., 2022; Williamson & Keller, 2023). However, responsible implementation is essential. Protecting genetic privacy, ensuring fairness in AI models, and preventing misuse of biological information remain important considerations (Khosravi et al., 2022; Martin et al., 2024). Parental involvement is central to maintaining transparency and trust throughout this process.
In summary, sportomics marks a major advancement in understanding youth athletic development. By connecting molecular individuality with practical decision-making, it empowers parents and coaches to guide children safely and effectively. As global sports systems continue shifting toward evidence-based practices, sportomics stands at the forefront of this transformation, supporting young athletes in exploring their potential while keeping health, wellbeing, and long-term development as the highest priorities (Zand et al., 2024; Roberts et al., 2023).
2. Key Molecular Techniques and Their Relevance
Modern sportomics relies on several molecular tools that help explain how a young athlete’s body responds to training, recovers from stress, and adapts over time, along with unique nutritional needs. These techniques provide objective biological information that complements traditional performance assessments. By understanding genomics, proteomics, and metabolomics together, parents and coaches can gain a clearer picture of an athlete’s strengths, vulnerabilities, and developmental needs. Each method captures a different layer of human biology, but when combined, they create a holistic view of performance potential (Sellami et al., 2020; Peake et al., 2020; de la Rosa et al., 2022).
For practitioners, it is useful to think of genomics, proteomics, and metabolomics as three complementary “layers” of information (
Table 1). Together, they help explain why two children with similar training loads might adapt differently or show different levels of fatigue and injury risk (Malsagova KA et al., 2024). These tools are not meant to rank or label children, but to give coaches and health professionals more precise information when adjusting volume, intensity, and recovery plans (Plaza-Diaz J et al.,2022).
2.1. Genomics: Mapping the Blueprint of Potential
Genomics explores how variations in DNA influence traits relevant to sports such as endurance, strength, and injury resistance because specific genetic markers can affect muscle fiber composition, cardiovascular efficiency, metabolic pathways, and connective tissue integrity (Ahmetov & Fedotovskaya, 2020; Ginevičienė et al., 2022; Ahmetov & Rogozkin, 2020). Understanding these variations helps explain why athletes differ in performance potential and how they respond to training, recovery, and physical stress.
For example, below is the list of very important genes:
ACE (Angiotensin-Converting Enzyme) I/D polymorphism affects cardiovascular efficiency and endurance capacity (Ahmetov & Rogozkin, 2020; Rodríguez-Larrad et al., 2020).
ACTN3 R577X variant influences the balance between fast- and slow-twitch muscle fibers, shaping sprint versus endurance predispositions (El Ouali et al., 2024; Ahmetov et al., 2021).
COL1A1 and COL5A1 variants relate to collagen structure and ligament strength, indicating susceptibility to tendon or ligament injury (Rodríguez-Romo & Peñalvo, 2022; Kwasniewski et al., 2024).
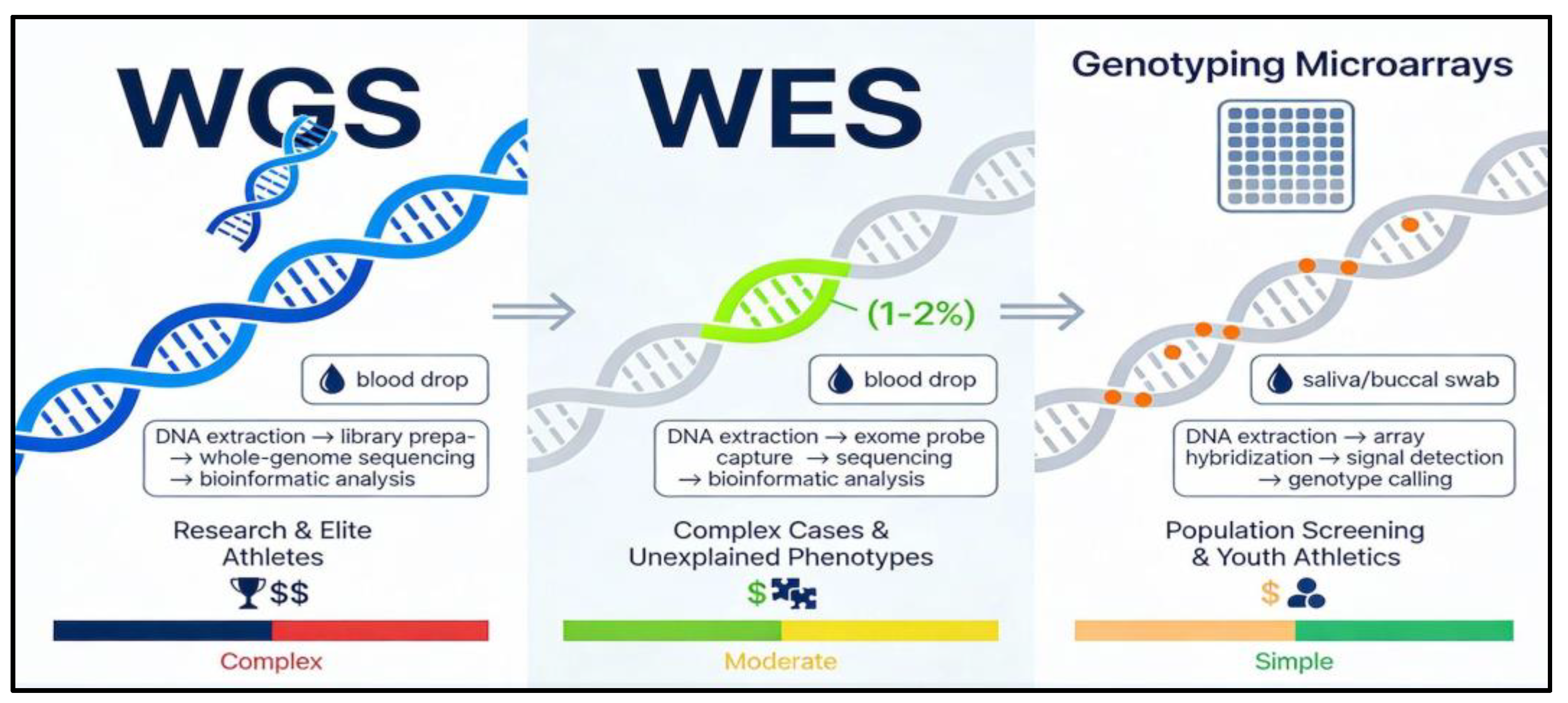
Genomic sequencing technologies have revolutionized sports genetics by enabling precise identification of performance-related variants in youth athletes, shifting from targeted single-gene tests to comprehensive profiling that informs personalized training and injury prevention. Whole Exome Sequencing (WES), Whole Genome Sequencing (WGS), and genotyping microarrays differ in scope, cost, and applicability, with each offering unique insights into polygenic traits like endurance, power, and recovery while requiring ethical considerations for pediatric use. These tools complement candidate gene studies (e.g., ACE, ACTN3) by detecting rare and regulatory variants, supporting evidence-based strategies in youth athletics without overemphasizing genetic determinism. (Boulygina EA et al.,2020).The detailed explanation is given below along with differences between each genetic technique discussed in
Table 2.
WES reads the protein-coding regions of the genome (the “exome”), which contains most known disease-causing and many performance-related variants. It can detect rare variants in genes affecting muscle development, connective tissue integrity, cardiac function, or metabolism that are missed by single-gene tests. For a young athlete with repeated unexplained injuries or unusual fatigue, WES may help uncover underlying biological contributors. A blood or saliva sample is used. DNA is extracted, the exonic regions are captured with probes, and then sequenced at high depth. Bioinformatic pipelines identify variants, which must be interpreted by specialists (clinical geneticists or sports medicine experts) (Bojarczuk A.,2024)
- 2.
Whole Genome Sequencing (WGS)
WGS reads almost the entire genome, including non-coding regulatory regions. It offers the most comprehensive view and can identify structural variants or regulatory changes that may subtly influence muscle function, heart performance, or recovery. This level is mainly relevant for research projects or for very high-performance environments, because interpretation is complex and cost is higher. Similar to WES but with broader coverage; advanced bioinformatics are required, and careful ethical review is needed before using it in children (Konopka MJ et al.,2023).
- 3.
Genotyping Microarrays
Microarrays test hundreds to thousands of pre-selected, well-studied variants in one run (for example, ACE, ACTN3, COL1A1 and other sport-relevant SNPs).They are currently the most practical and economical option for youth athletic settings because they focus on variants with published evidence and generate easy-to-interpret reports. They are suitable for identifying tendencies for endurance vs power, recovery profile, and soft-tissue injury susceptibility. A saliva or buccal swab sample is applied to a chip containing probes; hybridization patterns are read and converted into genotype calls.
For children and adolescents, genotyping panels are generally preferred because they focus on well-validated markers and can be paired with clear counselling about limitations, while more extensive sequencing should be reserved for carefully designed clinical or research contexts with robust ethical safeguards (Camporesi S et al.,2016). Comparative overview of Whole Exome Sequencing (WES), Whole Genome Sequencing (WGS), and genotyping microarray panels for youth sports genomics, highlighting coverage, typical sport-specific applications, cost/complexity, and suitability in children as shown in
Figure 2.
These genetic variants form an interconnected network influencing athletic performance across endurance (ACE I, BDKRB29, PPARGC1A Gly), power/sprint (ACTN3 R, ACE D), injury resilience (COL5A1 T, COL1A1 T), recovery dynamics (IL6 C, TNF-α G), and metabolic adaptation (HIF1A Ser, MCT1 T), enabling polygenic risk scores for personalized youth training protocols that balance genetic predispositions with associated traits as shown in
Table 3.
From a health coach or trainer’s perspective, genomic tests can be understood in simple, practical terms:
Endurance vs power tendency: Variants in genes such as ACE and ACTN3 do not decide which sport a child “must” play, but they can indicate whether the body is naturally more efficient at long-duration, oxygen-based work or at short, explosive efforts. This helps in deciding how much of the weekly training load should be endurance-focused versus power-focused, not in excluding any sport (Konopka MJ et al.2023).
Soft-tissue and ligament vulnerability: Variants in collagen-related genes (for example COL1A1, COL5A1) may signal that an athlete’s tendons and ligaments need more gradual loading, longer warm-ups, and a stronger focus on technique and neuromuscular control to prevent injuries ( Bojarczuk A; 2024).
Recovery profile: Markers linked to inflammation and oxidative stress regulation (such as IL6 or TNF-α polymorphisms) can help explain why some children “bounce back” quickly after hard sessions, while others feel sore and fatigued longer. Coaches can respond by adjusting recovery days, sleep education, and nutrition strategies for each child (Malsagova et al.,2024).
However, it is essential to stress that these tests show probabilities, not promises. They should never be used to deny a child access to a sport, to set rigid expectations, or to make long-term career decisions. Instead, they are best used as one more piece of information to fine-tune training and to protect health. (Vlahovich et al., 2017). Ethically, the use of genetic testing in minors should follow three principles: (Indian Council of Medical Research, 2018).
The child’s best interest must come first (health and enjoyment before performance).
Results should always be explained in language that avoids labels such as “talented” or “untalented”, which can harm self-confidence.
Data must be stored securely, with parental consent and the possibility for the young person to withdraw their data when they are older.
The genetic testing shows an important but partial picture, i.e. only the inherited potential of the target person. To understand how the body reacts during training, we must examine the real-time biological responses captured by proteomics.
2.2. Proteomics: Understanding Muscle Recovery and Fatigue
Proteomic profiling studies the complete set of proteins expressed in the body under different physiological states.
For health coaches, this can be translated into a few key questions:
How much muscle damage did yesterday’s session cause?
Is the athlete’s inflammation at a normal adaptive level or trending towards overuse?
Are stress hormones balanced, or is the child showing signs of chronic overload?
In practice, these questions can be addressed by multi-marker laboratory panels that measure enzymes like creatine kinase (CK) and lactate dehydrogenase (LDH), muscle-related proteins such as myoglobin, inflammatory cytokines (for example IL-6, TNF-α), and sometimes hormone ratios such as testosterone to cortisol. (Harshman et al., 2018). Monitoring proteins related to muscle repair, inflammation, and oxidative stress helps assess recovery and readiness (Cameron-Smith et al., 2021; El-Sayed & Bouhlel, 2020; Powers & Jackson, 2020)
Table 4).
For youth athletes, it is usually sufficient to perform such panels at pre-season baseline and then at selected time points (for example, after several weeks of intensified training or when the child reports unusual fatigue). The results should never be interpreted in isolation, but always together with symptoms (sleep, mood, soreness), training diary, and wearable data. (Musat et al., 2024). Because these tests involve blood sampling and sensitive biological data, they should only be carried out through accredited laboratories, under the supervision of qualified healthcare professionals. Direct-to-consumer services that promise performance prediction without proper counselling should be avoided in children, in line with international ethical recommendations. (Camporesi & McNamee, 2016).
Proteomics insights guide individualized training loads, rest periods, and nutrition, reducing the risk of overuse injuries common in young athletes (Muniz-Santos et al., 2021). Yet proteins represent only one layer of the body’s response. To understand how nutrients are used, how energy is produced, and how fatigue develops, metabolomics provides the next level of insight.
2.3. Metabolomics: Tracking Energy and Nutrition Dynamics
Metabolomics examines the small molecules produced during metabolism. These markers reveal how efficiently an athlete’s body converts nutrients into energy and manages oxidative stress (Sellami et al., 2020; Bassini et al., 2022). For parents, metabolomic assessments can help refine diet plans, ensuring optimal carbohydrate use, hydration, and micronutrient balance for training and recovery (Nashef & Smith, 2023; Rodríguez-Romo & Peñalvo, 2022). When combined with genomics and proteomics, metabolomic patterns complete the biological picture but the data are complex. This is where integrative analytics helps connect all layers into meaningful, actionable insights (de la Rosa et al., 2022).
Sportomics testing services available through ICMR-recommended laboratories encompass genomic profiling (ACTN3, ACE, COL5A1 variants via saliva/blood with 20-40+ markers), targeted single-gene genotyping (e.g., ACTN3 R577X for fast-twitch assessment), integrated wellness-genomics platforms combining athletic traits with nutrigenomics (40+ traits, saliva/blood), proteomics panels quantifying muscle damage and inflammatory markers (CK, LDH, IL-6 from serum/plasma), metabolomics analysis of training biomarkers (lactate, BCAAs from blood/urine), and integrated performance physiology combining VO₂max testing with biochemical profiling and wearable technology integration. As shown in
Table 5.
3. Integrative Analytics: The Role of AI and Machine Learning
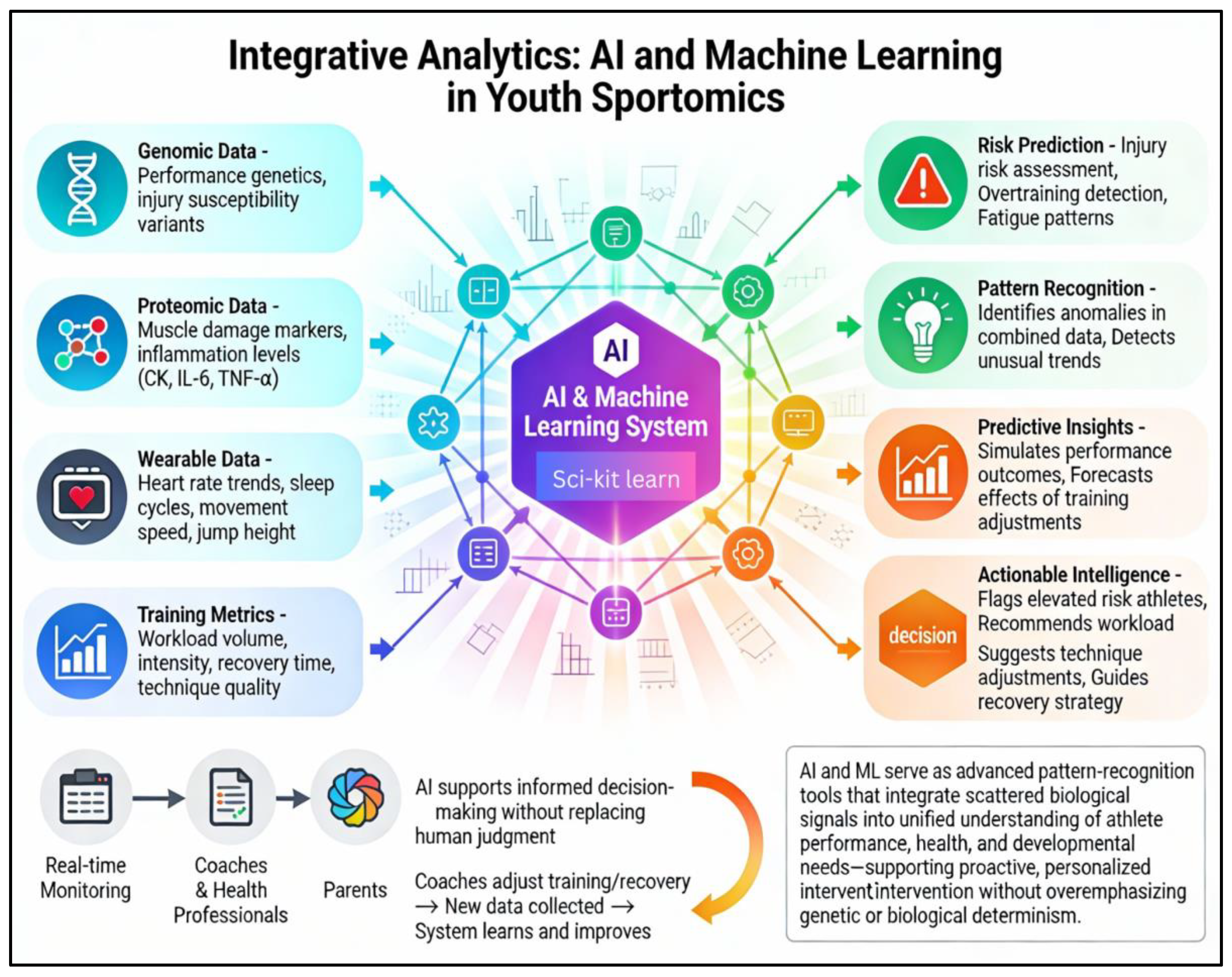
AI and ML enable integration of genomics, proteomics, and physiological data into cohesive, predictive systems. For coaches and health professionals, these systems can be thought of as advanced pattern-recognition tools. They do not replace coaching judgement; instead, they analyze large volumes of data, such as heart-rate trends, sleep duration, changes in speed or jump height, and simple blood markers to alert the staff when something looks unusual. For example, if a child’s training load, sleep quality and inflammatory markers together match patterns that previously preceded injuries in similar athletes, the AI system can flag the athlete as being at elevated risk so that coaches can proactively adjust workload, technique, or recovery strategies. (Wang et al., 2025).
Using tools such as TensorFlow and scikit-learn, these methods can identify patterns linked to fatigue, predict injury risks, and simulate how performance may change with different training adjustments (Ghosh et al., 2023; Pietraszewski et al., 2025; Li et al., 2021). (as shown in
Figure 3); When paired with wearable-generated data (heart rate, speed, sleep cycles, workload), these systems offer practical, real-time insights accessible to parents and coaches, supporting informed and safe decision-making for young athletes (Piette et al., 2022; Khosravi et al., 2022; Roberts et al., 2023).
3.1. AI-Enabled Innovations in Training Optimization and Injury Prevention
Recent technological advances have strengthened the field of sportomics by providing precise, data-driven insights that complement molecular information and support a comprehensive understanding of athletic performance (Baca & Kornfeind, 2021; Pereira et al., 2020). Modern biomechanical assessment tools now allow detailed tracking of joint angles, running mechanics, and impact forces (Muro-De-La-Herran et al., 2022). Such systems are increasingly used in sports academies for football, cricket, tennis, and athletics. By identifying inefficient or potentially harmful movement patterns, these tools help guide timely adjustments in technique, training intensity, and posture to reduce the risk of overuse injuries (Philip et al., 2023).
Advances in wearable technology also enable continuous monitoring of training load and fatigue (Fuller et al., 2022; Mehta et al., 2023). By combining information such as heart-rate trends, sleep quality, exertion patterns, and recovery markers, these systems help determine when an athlete is approaching a fatigue threshold. This is particularly important for children and adolescents, whose developing musculoskeletal systems require carefully balanced and age-appropriate workloads (Bergeron, 2018). In the clinical setting, enhanced imaging and digital assessment platforms support early detection of soft-tissue stress and contribute to personalized rehabilitation planning (Kobayashi et al., 2021). They guide structured and gradual return-to-play programs, ensuring safe recovery and long-term musculoskeletal health.
These technological innovations also contribute to more responsible talent development. By evaluating growth patterns, coordination, reaction time, and skill progression (Lloyd & Oliver, 2019), they allow each child’s strengths and developmental needs to be recognized without imposing premature specialization or high-pressure expectations. Together, these approaches highlight how modern technological tools reinforce the foundation of sportomics. When integrated with insights from genomics, proteomics, and metabolomics, they transform complex biological data into practical, actionable information (Freitas et al., 2021). This supports performance enhancement while prioritizing health and well-being, ensuring that training decisions align with the long-term development of young athletes.
4. Building Trust: Making AI a Reliable Partner in Sports
To maintain trust, AI tools used in youth sports should be transparent and explainable. Instead of simply labelling an athlete as “high risk”, well-designed systems show which factors are contributing most for example, “rapid increase in weekly training load”, “reduced sleep over the last 7 days”, and “elevated inflammatory markers”. This helps coaches and parents understand the reasoning, discuss it with the child, and decide on specific, constructive changes. It is ethically important that AI is used only as a decision-support tool, with final decisions always made by humans who consider the child’s feelings, goals, and overall context. Data inputs must be high-quality, privacy-protected, and used only for the athlete’s benefit. (Kim et al., in press).
As sportomics evolves, combining molecular insights with advanced analytical tools, parents and coaches often wonder how much they should rely on these technologies (Thorpe et al., 2022). Molecular data can explain how a child grows, recovers, or responds to training, while performance-monitoring systems help translate everyday training signals into simple, useful information (Buchheit, 2020). Yet no system is flawless; measurements may differ between devices, and numbers alone cannot capture a young athlete’s emotions, confidence, or daily well-being. For this reason, technology should act as a supportive partner rather than the final authority. When scientific information, whether related to genetics, metabolism, movement patterns, or training load, is interpreted alongside real-life experience, it becomes far more meaningful (Stokes et al., 2020).
Coaches, physiologists, and parents bring essential context by understanding the child’s behaviour, motivation, and readiness, which no device can measure (Halperin, 2018).
Table 6 discusses all these points and by blending reliable scientific processes with human observation, families can make training decisions that are safe, balanced, and child centred. This ensures that performance enhancement always aligns with healthy development, thoughtful progression, and long-term well-being. For health coaches and sport science staff, the most realistic entry point is usually DNA plus selected proteomic markers, combined with well-managed wearable data and careful observation on the field. (Plaza-Diaz et al., 2022).
5. Guarding Privacy: Ethics First in Sportomics
As sportomics brings together molecular testing and advanced analytical tools, protecting an athlete’s privacy becomes just as important as improving performance (Cavalli et al., 2023). Genetic markers, metabolic profiles, and movement data can provide valuable guidance, but they also represent deeply personal information especially for children. Such data must always be collected with genuine consent, stored securely, and shared only with those directly involved in the athlete’s care and development (Mittelstadt, 2019; Gibson et al., 2021).
For young athletes, trust is built when families know that their child’s information is handled responsibly. Clear explanations about what data is being used, why it is collected, and how it supports training decisions help maintain this trust (Taddeo & Floridi, 2018). Systems that show how conclusions are drawn, whether through molecular findings or performance trends, reduce confusion and prevent misinterpretation. Ethical practice ensures that data becomes a tool for support, not judgment. When privacy is respected and information is used thoughtfully, sportomics empowers children by promoting safe growth, healthy progress, and confidence (Saxon et al., 2021). This creates an environment where science guides development without ever compromising dignity or fairness.
6. Precision Matters: Ensuring Data Accuracy and Reliability
The usefulness of performance and molecular insights depends entirely on the accuracy and reliability of the underlying data. Errors from poorly calibrated wearables, incomplete molecular tests, or inconsistent tracking can lead to misleading conclusions and inappropriate training decisions. To ensure dependable results, standardized data collection procedures and regular equipment checks are essential. Parents and coaches should rely on qualified professionals to interpret complex measurements, rather than depending solely on simplified summaries from apps or devices. When expert guidance is combined with well-maintained monitoring systems, the resulting feedback reflects the athlete’s true condition in real time.
For children and adolescents, additional safeguards are needed. International position statements advise that genetic testing for talent identification alone should be approached with great caution or avoided, because the tests cannot reliably predict future sports success and may lead families to make important life decisions on incomplete information. In line with this, sportomics in youth should be prioritized: (Vlahovich et al., 2017).
Health protection and injury prevention over selection or deselection
Non-directive counselling, where results are explained neutrally and options are kept open
Re-consent when the athlete reaches adulthood, including the possibility to delete their data.
Accurate and reliable data not only supports informed decisions but also builds trust between families and the monitoring process. With this foundation, training and recovery plans can be tailored safely, maximizing performance while protecting the health, development, and long-term well-being of young athletes.
7. The Future of Molecular Insights in Sport
The future of sportomics lies in integrating multiple layers of biological information to provide a complete understanding of athletic performance. Advances are moving toward combining genomics, transcriptomics, proteomics, metabolomics, and microbiome profiling with physiological and psychological assessments (Ahmetov & Fedotovskaya, 2015; Kiss et al., 2022). This systems-level approach captures the complex interplay of molecular pathways, muscle adaptation, recovery efficiency, and mental resilience under competitive stress, rather than focusing on single genetic markers (Bouchard, 2012).
Microbiome research is gaining attention, as emerging evidence links gut microbial diversity with energy metabolism, immune function, and exercise tolerance (Turnbaugh et al., 2009). Metabolomic studies continue to reveal how metabolic flexibility influences performance sustainability and recovery potential (Kiss et al., 2022). Standardized data integration pipelines will enable the identification of composite biomarkers for fatigue, injury susceptibility, and overtraining, critical concerns for parents, coaches, and sports scientists working with young athletes.
Applied frameworks are evolving alongside these insights. National and Olympic training programs increasingly use molecular data to guide nutrition, hydration, and workload modulation (Jeukendrup & Gleeson, 2019). Longitudinal monitoring ensures that early specialization and training intensity decisions balance performance goals with developmental safety and long-term musculoskeletal health (Mujika & Padilla, 2003).
Ethical considerations remain central to the expansion of molecular sportomics. Harmonized standards for genomic data interpretation, informed consent, and equitable access to testing technologies will be essential (Ehlert et al., 2013). Collaboration between molecular scientists, clinicians, parents, coaches, and policymakers will ensure that scientific discoveries translate into practical, field-ready guidance. The overarching goal is to combine individual biological information with evidence-based practice, empowering parents and coaches to guide young athletes safely and effectively while fostering performance, health, and long-term development.
8. Discussion
This paper highlights how the combined use of molecular techniques and analytical technologies can transform the way young athletes are supported throughout their development. The integration of genomics, proteomics, and metabolomics establishes a comprehensive framework for understanding an athlete’s inherent biological profile and their real-time responses to training stress (Ahmetov & Fedotovskaya, 2015; Kiss et al., 2022). Genetic polymorphisms such as ACE I/D, ACTN3 R577X, and COL5A1 provide useful insights into endurance capacity, muscle efficiency, and connective-tissue resilience (Pickering & Kiely, 2017; Ahmetov et al., 2016; Del Coso et al., 2019). Yet, these markers offer tendencies rather than definitive predictions, reinforcing that molecular sportomics should guide, not dictate the training decisions. Proteomic and metabolomic assessments extend this foundation by identifying biochemical markers of recovery, inflammation, and metabolic load (Kiss et al., 2022). These insights can help personalize nutritional strategies and regulate training volume, especially in adolescents who face heightened vulnerability to fatigue and musculoskeletal injuries (Roth, 2008). However, practical translation remains a challenge. Variability in laboratory methods, limited normative datasets for youth populations, and inconsistent device calibration may hinder interpretation accuracy (Mansfield et al., 2020). Therefore, standardized protocols and multicentric validation studies are essential for strengthening the reliability of sportomics data.
For parents and coaches, the practical implications are substantial. Molecular findings and performance-monitoring systems provide an evidence-based structure for tailoring rest, workload, and nutrition to each child’s needs (Jeukendrup & Gleeson, 2019). At the same time, ethical considerations remain central. Misinterpretation of genetic results, early labeling of children, or performance-driven pressure can pose risks to psychological well-being (Ehlert et al., 2013). Empowering families with appropriate education and professional guidance ensures that sportomics becomes a supportive tool rather than a source of anxiety.
The broader vision of sportomics encourages a collaborative ecosystem in which sports scientists, clinicians, parents, and coaches share responsibility for interpreting biological signals. When this partnership is built on transparency, data accuracy, and ethical awareness, sportomics enhances performance while prioritizing long-term safety. Ultimately, the strength of this approach lies in its ability to combine individual biological information with real-world context, helping young athletes develop in a balanced, responsible, and scientifically informed manner.
From a practical standpoint, health coaches and trainers can integrate sportomics stepwise. First, they can use simple, validated questionnaires, training diaries, and basic wearable data to monitor load, sleep, and perceived exertion in every young athlete. Second, selected genomic and proteomic tools can be introduced not to “select winners”, but to identify who may need extra protection (for example, children with a higher genetic tendency for tendon injury or slower biochemical recovery). Third, any AI-generated alert, such as an increased injury risk score should always trigger a conversation, not a punishment: coaches can sit with the athlete and parents to adjust training plans, reinforce good recovery habits, and ensure that sport remains a source of health, enjoyment, and long-term development rather than stress. (Camporesi & McNamee, 2016).
9. Conclusion
Sportomics represents a significant advancement in youth sports development by linking molecular individuality with practical, real-time understanding of performance and recovery. Through genomics, proteomics, metabolomics, and movement-based assessments, parents and coaches gain access to reliable biological information that supports safe training practices, appropriate workload progression, and evidence-based nutritional choices. This multidimensional perspective helps prevent overtraining, reduce injury risk, and promote healthy long-term development, central goals for any young athlete’s journey.
As the field progresses, the emphasis must remain on ethical implementation and equitable access. Transparent communication, responsible data handling, and avoidance of premature labeling are essential to maintaining trust. Emerging directions, including microbiome profiling, longitudinal biomarker tracking, and harmonized bioinformatics pipelines, will enhance the precision and predictive value of sportomics in the coming years.
The ultimate purpose of this approach is not to chase early performance outcomes, but to create an environment where biological understanding supports thoughtful, individualized coaching. By integrating scientific knowledge with human judgment, sportomics empowers parents and coaches to guide athletes in a manner that protects health, nurtures confidence, and lays the foundation for sustainable athletic success.
Authors Declaration
D.P. and S.M. were responsible for the conceptualization and study design, manuscript drafting, review and editing, and overall supervision of the work. They also contributed to data interpretation, preparation of tables and figures, and drafting of the conclusion. A.K. and D.P. aligned the manuscript content and contributed to the refinement and inclusion of figures and tables. D.S. provided supervisory oversight, critically reviewed the analyses, and contributed to validation and final interpretation of the findings. All authors read and approved the final manuscript.
Funding
This research received funding from DrOmics Labs Private Limited.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publication
Not applicable.
Acknowledgments
We wish to Acknowledge Dr. Omics Research Labs for their invaluable support and collaboration, which significantly contributed to the success of this research. We would like to express our sincere gratitude for providing the necessary facilities and encouragement to carry out this work.
Conflicts of interest
The authors declare that they have no conflicts of interest.
Abbreviations
|
ACE– |
Angiotensin-Converting Enzyme |
|
ACTN3– |
Alpha-Actinin-3 |
|
AI– |
Artificial Intelligence |
|
COL5A1– |
Collagen Type V Alpha 1 |
|
DNA– |
Deoxyribonucleic Acid |
|
EMG– |
Electromyography (implied in biomechanical assessments) |
|
GPS– |
Global Positioning System (used in wearables for athlete tracking) |
|
HRV– |
Heart Rate Variability (component of fatigue monitoring) |
|
ML– |
Machine Learning |
|
MRI– |
Magnetic Resonance Imaging (clinical imaging context) |
|
NGS– |
Next-Generation Sequencing (used in genomics context) |
|
RTP– |
Return to Play |
|
VO₂max– |
Maximal Oxygen Uptake (implied in endurance assessment) |
| OMICs Terminology: |
|
|
Genomics– |
Study of genes and genetic variation |
|
Transcriptomics– |
Study of RNA expression |
|
Proteomics– |
Study of protein structure and function |
|
Metabolomics– |
Study of metabolic pathways and metabolites |
|
Microbiomics / Microbiome Profiling– |
Study of gut microbial communities |
References
- Ahmetov, I. I.; Fedotovskaya, O. N. Current progress in sports genomics. Exercise and Sport Sciences Reviews 2020, 48(4), 191–199. [Google Scholar] [CrossRef]
- Ahmetov, I. I.; Kulemin, N. A.; Popov, D. V. Genomic predictors of athlete status and response to training. European Journal of Sport Science 2021, 21(7), 1–12. [Google Scholar] [CrossRef]
- Ahmetov, I. I.; Rogozkin, V. Genetic profile of endurance athletes and strength athletes. Journal of Sports Sciences 2020, 38(8), 987–997. [Google Scholar] [CrossRef]
- Baca, A.; Kornfeind, P. Motion analysis in sports: A critical review. Sensors 2021, 21(7), 2121. [Google Scholar] [CrossRef]
- Babu, M.; et al. Wearable devices: Implications for precision medicine and population health. Annual Review of Medicine 2024, 75, 1–20. [Google Scholar] [CrossRef]
- Bassini, A.; Bragazzi, N. L.; Cameron-Smith, D. Sportomics method to assess acute-phase proteins and metabolome changes in athletes. Scientific Reports 2022, 12, 23300. [Google Scholar] [CrossRef]
- Bergeron, M. F. Youth athletic development: Managing training load in children and adolescents. Journal of Strength and Conditioning Research 2018, 32(7), 2001–2011. [Google Scholar]
- Bragazzi, N. L.; Bassini, A. Sportomics overview: Metabolic signatures of exercise. European Review 2020, 28(3), 1–12. [Google Scholar] [CrossRef]
- Buchheit, M. Monitoring training responses in elite athletes using new technologies. Sports Medicine 2020, 50(6), 1025–1042. [Google Scholar]
- Cameron-Smith, D.; et al. Protein biomarkers to monitor training adaptation and overreaching. Proteomics Clinical Applications 2021, 15(6), e2000030. [Google Scholar] [CrossRef]
- Cavalli, G.; Meijer, A. H.; Reinders, M. J. T. Ethical considerations in sports genomics. Nature Genetics 2023, 55(2), 141–150. [Google Scholar] [CrossRef]
- Clarke, A.; et al. Data governance, privacy and ethics in youth sports genomics. Journal of Medical Ethics 2022, 48(9), 600–607. [Google Scholar] [CrossRef]
- Claudino, J. G.; et al. Injury risk assessment and performance prediction using AI. Sports Medicine – Open 2020, 6, 61. [Google Scholar] [CrossRef]
- Claudino, J. G.; et al. Machine learning in sports biomechanics. British Journal of Sports Medicine 2021, 55(19), 1099–1108. [Google Scholar] [CrossRef]
- Claudino, J. G.; et al. Technologies for monitoring athlete workload. Sports Medicine 2022, 52(8), 1715–1734. [Google Scholar] [CrossRef]
- de la Rosa, A.; et al. Multi-omics integration for sports injury biomarkers. OMICS 2022, 26(5), 274–289. [Google Scholar] [CrossRef]
- El Ouali, E. M.; et al. ACTN3 R577X genotype and athletic status. Sports Medicine – Open 2024, 10, 24. [Google Scholar] [CrossRef]
- El-Sayed, M. S.; Bouhlel, E. Metabolomic assessment of training load. Journal of Applied Physiology 2020, 128(4), 765–775. [Google Scholar] [CrossRef]
- Fernández-García, M.; et al. Predictive modelling of injury risk with wearables. Sensors 2023, 23(4), 1789. [Google Scholar] [CrossRef]
- Freitas, V. H.; Nakamura, F. Y.; Milos, N. Integrating molecular biomarkers with training data. Sports Medicine 2021, 51(4), 677–694. [Google Scholar]
- Fuller, J. T.; Bellenger, C. R.; Buckley, J. D. Wearable sensors in youth sports. Frontiers in Sports and Active Living 4 2022, 879123. [Google Scholar] [CrossRef]
- Ghosh, I.; et al. Sports analytics review. WIREs Data Mining and Knowledge Discovery 2023, 13, e1496. [Google Scholar] [CrossRef]
- Gibson, E.; Adams, M.; Chin, J. Protecting minors in genomic research. American Journal of Bioethics 2021, 21(3), 40–52. [Google Scholar] [CrossRef]
- Ginevičienė, V.; et al. Perspectives in sports genomics. Biomedicines 2022, 10(2), 298. [Google Scholar] [CrossRef]
- Goes, F. R.; et al. Big data in tactical soccer analysis. European Journal of Sport Science 2021, 21(4), 481–496. [Google Scholar] [CrossRef]
- Halperin, I. Psychological readiness in training prescription. Sports Medicine 2018, 48(11), 2551–2558. [Google Scholar]
- Hamilton, B.; Hopkins, W. Statistical models for athlete development. Medicine & Science in Sports & Exercise 2021, 53(12), 2547–2558. [Google Scholar] [CrossRef]
- Horvath, S.; Raj, K. DNA methylation biomarkers in athletic performance. Sports Medicine 2020, 50(1), 131–145. [Google Scholar] [CrossRef]
- Huygens, W.; et al. Precision performance frameworks. International Journal of Sports Medicine 2022, 43(10), 809–818. [Google Scholar] [CrossRef]
- Jia, Y.; et al. Deep learning in sports analytics. Frontiers in Computer Science 2025, 3, 123456. [Google Scholar] [CrossRef]
- Khosravi, P.; et al. Explainable AI in sports. IEEE Access 2022, 10, 45712–45727. [Google Scholar] [CrossRef]
- Kobayashi, H.; Koga, H.; Yokoyama, K. Advances in MRI for musculoskeletal injuries. British Journal of Sports Medicine 2021, 55(9), 468–474. [Google Scholar]
- Kwasniewski, M.; et al. Combining genomics & biomechanics. Journal of Biomechanics 2024, 156, 111111. [Google Scholar] [CrossRef]
- Lemmink, K. A. P. M.; et al. Longitudinal monitoring in youth athletes. Journal of Sports Sciences 2021, 39(9), 1012–1024. [Google Scholar] [CrossRef]
- Li, R.; et al. ML for youth talent identification. International Journal of Sports Science & Coaching 2020, 15(4), 623–642. [Google Scholar] [CrossRef]
- Li, W.; Zhou, D.; Zhang, X. Deep learning for athlete tracking. Computer Vision in Sports 2023, 12(3), 101–120. [Google Scholar] [CrossRef]
- Li, X.; et al. ML for sports injury prediction. IEEE TNSRE 2021, 29, 1–14. [Google Scholar] [CrossRef]
- Lloyd, R. S.; Oliver, J. L. Youth physical development model. Strength & Conditioning Journal 2019, 41(3), 2–17. [Google Scholar] [CrossRef]
- Martin, D.; et al. ML pipelines for sports genomics. Bioinformatics 2024, 40(12), 3450–3462. [Google Scholar] [CrossRef]
- Mehta, K.; Kulkarni, A.; Raj, A. Sleep quality & adolescent athletes. Sports Medicine – Open 2023, 9(1), 12. [Google Scholar] [CrossRef]
- Mittelstadt, B. D. Principles of data ethics. Big Data & Society 2019, 6(2). [Google Scholar] [CrossRef]
- Muniz-Santos, R.; et al. Metabolomics in elite athlete monitoring. Journal of Sports Medicine and Physical Fitness 2021, 61(9), 1243–1254. [Google Scholar] [CrossRef]
- Muniz-Santos, R.; et al. Metabolic responses in Olympic athletes. Nutrients 2024, 17(11), 1924. [Google Scholar] [CrossRef]
- Muro-De-La-Herran, A.; García-Zapirain, B.; Méndez-Zorrilla, A. Gait analysis for performance. Applied Sciences 2022, 12(6), 3204. [Google Scholar] [CrossRef]
- Nasb, M.; et al. Precision exercise. Precision Exercise Medicine 2024, 1(1), 45–65. [Google Scholar] [CrossRef]
- Nashef, R.; Smith, J. Microbiome & exercise. Trends in Microbiology 2023, 31(2), 122–136. [Google Scholar] [CrossRef]
- Nascimento, D. C.; et al. Biomarkers of training load. International Journal of Sports Physiology and Performance 2022, 17(6), 825–836. [Google Scholar] [CrossRef]
- Nassis, G. P.; et al. Big data in sport. Journal of Sports Sciences 2020, 38(23), 1–14. [Google Scholar] [CrossRef]
- O’Connell, J.; et al. Longitudinal multimodal monitoring. Pediatric Exercise Science 2021, 33(2), 103–118. [Google Scholar] [CrossRef]
- Peake, J. M.; et al. Integrative omics in sport. Journal of Proteome Research 2020, 19(5), 2020–2036. [Google Scholar] [CrossRef]
- Pereira, T.; Ribeiro, J.; Oliveira, J. Wearable technology in sports. Journal of Sports Sciences 2020, 38(17), 2018–2027. [Google Scholar] [CrossRef]
- Philip, A.; Bradley, J.; Wilson, F. Biomechanical risk factors in youth athletes. Sports Medicine 2023, 53(2), 215–230. [Google Scholar]
- Pietraszewski, P.; et al. AI in sports analytics. Applied Sciences 2025, 15, 7254. [Google Scholar] [CrossRef]
- Piette, M.; et al. Wearables & AI for youth athletes. Frontiers in Sports and Active Living 2022, 4, 800512. [Google Scholar] [CrossRef]
- Powers, S. K.; Jackson, M. J. Exercise-induced oxidative stress. Free Radical Biology & Medicine 2020, 146, 1–13. [Google Scholar] [CrossRef]
- Roberts, T.; et al. AI for rehabilitation. npj Digital Medicine 2023, 6, 94. [Google Scholar] [CrossRef]
- Rodríguez-Larrad, A.; et al. Genetic testing for sports performance. British Journal of Sports Medicine 2020, 54(12), 694–700. [Google Scholar] [CrossRef]
- Rodríguez-Matoso, D.; et al. Muscle damage and recovery. Frontiers in Physiology 2021, 12, 657430. [Google Scholar] [CrossRef]
- Rodríguez-Romo, G.; Peñalvo, J. L. Precision nutrition & omics. Nutrition Reviews 2022, 80(6), 1234–1249. [Google Scholar] [CrossRef]
- Saxon, L.; Hayes, M.; Williams, S. Digital privacy in youth athletes. Journal of Athletic Training 2021, 56(5), 532–540. [Google Scholar] [CrossRef]
- Sellami, M.; Bragazzi, N. L.; Chamari, K.; Franchini, E. Molecular big data in sports sciences. Frontiers in Molecular Biosciences 2021, 8, 815410. [Google Scholar] [CrossRef]
- Sellami, M.; Bragazzi, N. L. Gene–environment interactions in sportomics. Genes 2021, 12(3), 389. [Google Scholar] [CrossRef]
- Sellami, M.; et al. Multi-omics approaches in sport. International Journal of Sports Physiology and Performance 2020, 15(9), 1252–1266. [Google Scholar] [CrossRef]
- Stokes, K. A.; Jones, B.; Green, B. S. Human interpretation in data analytics. BMJ Open Sport & Exercise Medicine 2020, 6(1), e000775. [Google Scholar] [CrossRef]
- Taddeo, M.; Floridi, L. Ethics of explainable AI. Philosophy & Technology 2018, 31(4), 611–627. [Google Scholar] [CrossRef]
- Thorpe, R. T.; Atkinson, G.; Drust, B. AI in youth sports decision-making. International Journal of Sports Science & Coaching 2022, 17(4), 874–885. [Google Scholar]
- Williamson, P.; Keller, J. AI ethics in sports. Ethics and Information Technology 2023, 25, 1–18. [Google Scholar] [CrossRef]
- Zand, A.; et al. Precision sports medicine. Journal of Personalized Medicine 2024, 14(3), 412. [Google Scholar] [CrossRef]
- Ziemann, E.; et al. Epigenetic markers & youth athlete development. European Journal of Applied Physiology 2020, 120(11), 2501–2515. [Google Scholar] [CrossRef]
- Musat, C. L.; Mereuta, C.; Nechita, A.; Tutunaru, D.; Voipan, A. E.; Voipan, D.; Mereuta, E.; Gurau, T. V.; Gurău, G.; Nechita, L. C. Diagnostic applications of AI in sports: A comprehensive review of injury risk prediction methods. Diagnostics (Basel) 2024, 14(22), 2516. [Google Scholar] [CrossRef]
- Collier, R. Testing the ethics of genetic testing in sports. CMAJ 2011, 184(1), E45–E46. [Google Scholar] [CrossRef] [PubMed]
- Camporesi, S.; McNamee, M. J. Ethics, genetic testing, and athletic talent: Children’s best interests, and the right to an open (athletic) future. Physiological Genomics 2016, 48(3), 191–195. [Google Scholar] [CrossRef]
- Plaza-Diaz, J.; Izquierdo, D.; Torres-Martos, Á.; Baig, A. T.; Aguilera, C. M.; Ruiz-Ojeda, F. J. Impact of physical activity and exercise on the epigenome in skeletal muscle and effects on systemic metabolism. Biomedicines 2022, 10(1), 126. [Google Scholar] [CrossRef]
- Malsagova, K. A.; Kopylov, A. T.; Stepanov, A. A.; Kulikova, L. I.; Izotov, A. A.; Yurku, K. A.; Balakin, E. I.; Pustovoyt, V. I.; Kaysheva, A. L. Metabolomic and proteomic profiling of athletes performing physical activity under hypoxic conditions. Sports (Basel) 2024, 12(3), 72. [Google Scholar] [CrossRef]
- Vlahovich, N.; Fricker, P. A.; Brown, M. A.; Hughes, D. Ethics of genetic testing and research in sport: A position statement from the Australian Institute of Sport. British Journal of Sports Medicine 2017, 51(1), 5–11. [Google Scholar] [CrossRef]
- Kim, J.-H.; Kim, J.; Kang, H.; Youn, B.-Y. Kim, J.-H., Kim, J., Kang, H., & Youn, B.-Y. (in press). Ethical implications of artificial intelligence in sport: A systematic scoping review. [Journal Name], [Article Number/Page Range]. [CrossRef] [PubMed]
- Wang, P.; Wang, A.; Wang, S. Integrating multimodal AI technologies for sports injury prediction and rehabilitation: Systematic review. Journal of Human Sport and Exercise 2025, *21*(1), 22–37. [Google Scholar] [CrossRef]
- Konopka, M. J.; Sperlich, B.; Rietjens, G.; Zeegers, M. P. Genetics and athletic performance: A systematic SWOT analysis of non-systematic reviews. Frontiers in Genetics 2023, 14, 1232987. [Google Scholar] [CrossRef]
- Musat, C. L.; Mereuta, C.; Nechita, A.; Tutunaru, D.; Voipan, A. E.; Voipan, D.; Mereuta, E.; Gurau, T. V.; Gurău, G.; Nechita, L. C. Diagnostic applications of AI in sports: A comprehensive review of injury risk prediction methods. Diagnostics 2024, *14*(22), 2516. [Google Scholar] [CrossRef]
- Harshman, S. W.; Pitsch, R. L.; Smith, Z. K.; O’Connor, M. L.; Geier, B. A.; Qualley, A. V.; Schaeublin, N. M.; Fischer, M. V.; Eckerle, J. J.; Strang, A. J.; Martin, J. A. The proteomic and metabolomic characterization of exercise-induced sweat for human performance monitoring: A pilot investigation. PLOS ONE 2018, *13*(11), e0203133. [Google Scholar] [CrossRef]
- Bojarczuk, A. Ethical aspects of human genome research in sports—A narrative review. Genes (Basel) 2024, 15(9), 1216. [Google Scholar] [CrossRef] [PubMed]
- Indian Council of Medical Research. (2018). National ethical guidelines for biomedical and health research involving human participants (2017): Handbook. https://naitik.gov.in/DHR/resources/app_srv/DHR/global/pdf/downloads/Handbook_on_ICMR_Ethical_Guidelines.pdf.
- Boulygina, E. A.; Borisov, O. V.; Valeeva, E. V.; Semenova, E. A.; Kostryukova, E. S.; Kulemin, N. A.; Larin, A. K.; Nabiullina, R. M.; Mavliev, F. A.; Akhatov, A. M.; Andryushchenko, O. N.; Andryushchenko, L. B.; Zmijewski, P.; Generozov, E. V.; Ahmetov, I. I. Whole genome sequencing of elite athletes. Biology of Sport 2020, 37(3), 295–304. [Google Scholar] [CrossRef] [PubMed]
Figure 1.
Sportomics Integration for Ethical Youth Athletic Development: Infographic illustrating how sportomics combines genomics, proteomics, metabolomics, wearable data, and AI to deliver child-centered training decisions on intensity, recovery, and nutrition, while emphasizing ethical safeguards against premature labeling and ensuring tools support rather than supplant on-field observation and wellbeing.
Figure 1.
Sportomics Integration for Ethical Youth Athletic Development: Infographic illustrating how sportomics combines genomics, proteomics, metabolomics, wearable data, and AI to deliver child-centered training decisions on intensity, recovery, and nutrition, while emphasizing ethical safeguards against premature labeling and ensuring tools support rather than supplant on-field observation and wellbeing.
Figure 2.
: Comparative overview of genomic sequencing techniques: This figure presents three genomic sequencing approaches employed in youth sports genomics: whole genome sequencing (WGS) for comprehensive research applications, whole exome sequencing (WES) for investigation of complex phenotypes, and genotyping microarrays for practical population-level screening.
Figure 2.
: Comparative overview of genomic sequencing techniques: This figure presents three genomic sequencing approaches employed in youth sports genomics: whole genome sequencing (WGS) for comprehensive research applications, whole exome sequencing (WES) for investigation of complex phenotypes, and genotyping microarrays for practical population-level screening.
Figure 3.
Integrative Analytics of AI and ML in sportyomics : This figure shows an interconnected framework; sportomics transforms scattered biological signals into a unified understanding of a child’s performance, health, and developmental needs.
Figure 3.
Integrative Analytics of AI and ML in sportyomics : This figure shows an interconnected framework; sportomics transforms scattered biological signals into a unified understanding of a child’s performance, health, and developmental needs.
Table 1.
Overview of key sportomics tools for athletes: This table includes the biological layer, measured parameters, clinical interpretations for coaching and parental guidance, and typical specimen types used for analysis.
Table 1.
Overview of key sportomics tools for athletes: This table includes the biological layer, measured parameters, clinical interpretations for coaching and parental guidance, and typical specimen types used for analysis.
| Layer |
What it measures |
What it can tell coaches and parents |
Typical sample |
| Genomics |
DNA variants that are inherited and stable |
Tendencies for endurance vs power, injury susceptibility, recovery capacity; helps tailor training focus and identify who needs extra joint/tendon protection |
Blood, saliva, buccal swab |
| Proteomics |
Proteins related to muscle damage, inflammation, and repair |
Current muscle stress, level of inflammation, whether recovery is adequate; warns when a child is close to overreaching or overtraining |
Blood |
| Metabolomics |
Small molecules from energy and nutrient metabolism |
How effectively a child uses carbohydrates, fats, and amino acids; hydration status; metabolic fatigue during or after hard blocks |
Blood, urine |
Table 2.
Practical comparison of genomic testing approaches in sportomics: Comparative analysis of genomic testing methodologies in sportomics, including whole exome sequencing (WES), whole genome sequencing (WGS), and genotyping microarrays, with specifications for genomic coverage, typical sportomics applications, technical and financial requirements, and clinical appropriateness for pediatric athletic populations.
Table 2.
Practical comparison of genomic testing approaches in sportomics: Comparative analysis of genomic testing methodologies in sportomics, including whole exome sequencing (WES), whole genome sequencing (WGS), and genotyping microarrays, with specifications for genomic coverage, typical sportomics applications, technical and financial requirements, and clinical appropriateness for pediatric athletic populations.
| Feature |
Whole Exome Sequencing (WES) |
Whole Genome Sequencing (WGS) |
Genotyping panels (microarray) |
| Coverage |
All protein-coding regions |
Almost entire genome |
Selected known variants |
| Typical use in sport |
Investigation of complex or unexplained cases; advanced profiling |
Research and elite-level mechanistic studies |
Screening for common performance and injury markers |
| Cost and complexity |
Moderate, requires specialist interpretation |
Highest, requires advanced infrastructure and ethics oversight |
Lower, simpler interpretation |
| Suitability in children |
Case-by-case, ideally under research/clinical protocol |
Usually research only; not routine for youth athletics |
Appropriate with counselling and clear limits on use
|
Table 3.
Most Promising DNA Findings Relevant to Athletic Performance: The table presents eleven gene polymorphisms with evidence-based associations to endurance, power, flexibility, injury resistance, and recovery traits in athletic populations. For each gene, the table specifies the polymorphism identifier, associated athletic trait(s), mechanistic summary of scientific findings, and key supporting publications demonstrating reproducible genotype-phenotype associations.
Table 3.
Most Promising DNA Findings Relevant to Athletic Performance: The table presents eleven gene polymorphisms with evidence-based associations to endurance, power, flexibility, injury resistance, and recovery traits in athletic populations. For each gene, the table specifies the polymorphism identifier, associated athletic trait(s), mechanistic summary of scientific findings, and key supporting publications demonstrating reproducible genotype-phenotype associations.
| Gene |
Polymorphism |
Associated Trait |
Scientific Findings (Summary) |
Supporting Studies |
| ACE (Angiotensin-Converting Enzyme) |
I/D |
Endurance (I allele), Power (D allele) |
I allele linked to improved oxygen uptake and endurance; D allele associated with power and sprint performance. |
Montgomery et al., 1998 (Nature); Nazarov et al., 2001 |
| ACTN3 (Alpha-Actinin-3) |
R577X |
Sprint ability, fast-twitch muscle performance |
R allele associated with power and speed; XX genotype linked to lower fast-twitch fiber efficiency but better endurance adaptation. |
Yang et al., 2003 (AJHG); Eynon et al., 2013 |
| COL5A1 |
rs12722 (C/T) |
Flexibility, reduced injury risk |
T allele associated with improved flexibility and lower risk of tendon injuries such as Achilles tendinopathy. |
Collins & Raleigh, 2009; September et al., 2009 |
| COL1A1 |
Sp1 (G/T) |
Ligament strength, injury resistance |
T allele associated with increased collagen type I production and reduced ligament injury susceptibility. |
Posthumus et al., 2009 |
| PPARGC1A (PGC-1α) |
Gly482Ser |
Aerobic capacity, mitochondrial efficiency |
Gly allele linked to higher VO₂max and improved aerobic metabolism. |
Lucia et al., 2005 |
| IL6 |
-174 G/C |
Inflammation regulation, recovery |
C allele associated with lower inflammation and better recovery after intensive training. |
Fishman et al., 1998; Yamin et al., 2008 |
| TNF-α |
-308 G/A |
Muscle damage, recovery rate |
An allele linked to elevated inflammatory response and slower muscle recovery. |
Saunders et al., 2006 |
| BDKRB2 (Bradykinin Receptor B2) |
+9/−9 |
Endurance performance |
−9 variant associated with improved endurance and vascular adaptation. |
Williams et al., 2004 |
| HIF1A |
Pro582Ser |
Adaptation to hypoxia, anaerobic performance |
Ser allele enhances anaerobic power and adaptation to low-oxygen conditions. |
Ahmetov et al., 2008 |
| MCT1 |
A1470T |
Lactate transport, high-intensity performance |
T allele linked to improved lactate clearance and better high-intensity performance. |
Cupeiro et al., 2012 |
Table 4.
Proteins commonly monitored in youth sportomics and their practical meaning: Commonly used protein and hormonal markers in youth sportomics, with their practical meaning for coaches and the situations in which persistently abnormal values should trigger caution about training load and recovery.
Table 4.
Proteins commonly monitored in youth sportomics and their practical meaning: Commonly used protein and hormonal markers in youth sportomics, with their practical meaning for coaches and the situations in which persistently abnormal values should trigger caution about training load and recovery.
| Marker |
What it reflects for coaches |
When to be cautious |
| Creatine kinase (CK) |
Degree of muscle fibre disruption after training |
If CK stays elevated across multiple tests, training loads may be too high or recovery insufficient |
| Myoglobin |
Acute muscle damage |
Very high values after repeated intense sessions warrant rest and technique review |
| IL-6, TNF-α |
Systemic inflammation |
Persistent elevation suggests inadequate recovery or early overuse risk |
| C-reactive protein (CRP / hsCRP) |
General inflammatory load |
High baseline levels may signal that the athlete is not fully recovered from previous blocks |
Testosterone:
cortisol ratio |
Balance between anabolic (building) and catabolic (breaking down) processes |
A chronically reduced ratio in adolescents can indicate accumulated stress and need for modified training |
Table 5.
Non-comparative directory of sportomics testing services in India: This table consists of genomics focus; proteomics/metabolomics research/clinical.(ICMR labs recommended).
Table 5.
Non-comparative directory of sportomics testing services in India: This table consists of genomics focus; proteomics/metabolomics research/clinical.(ICMR labs recommended).
| Testing Category |
Description |
Sample Type |
Key Features |
| Genomic Panels (DNA-based athletic profiling) |
Tests for performance-related polymorphisms (e.g., ACTN3, ACE, COL5A1 variants for power/endurance/injury risk) |
Saliva, buccal swab, blood |
20-40+ markers; reports on athletic traits, recovery, nutrition guidance |
| Sports Gene Genotyping (Single/Multi-gene) |
Focused tests (e.g., ACTN3 R577X for fast-twitch muscle) |
Blood, saliva |
Single gene or small panels; quick turnaround |
| Integrated Wellness + Sports Genomics |
Combines athletic traits with nutrition, injury risk, wellness |
Saliva/blood |
40+ traits + nutrigenomics; personalized reports |
| Proteomics (Muscle/Recovery Markers) |
Blood panels for CK, LDH, IL-6 (muscle damage, inflammation) |
Blood serum/plasma |
Clinical-grade; for overtraining assessment |
| Metabolomics (Energy/Nutrition) |
Serum/urine analysis of metabolites (lactate, BCAAs) |
Blood/urine |
Training adaptation, fatigue biomarkers |
| Performance Physiology Labs (VO2max + Biochemical) |
VO2max testing + lactate/blood markers |
Blood + exercise testing |
Integrated with wearables |
Table 6.
Practical view of DNA, RNA, protein and integrated sportomics tools for youth sport: This table presents a practical framework stratifying sportomics assessment tools across biological levels: genomics provides stable, heritable information informing long-term training planning; transcriptomics enables research-level understanding of training-induced molecular adaptation; proteomics offers dynamic biomarkers indicating whether training stimulus matches athlete capacity; and integrated multi-omics provides comprehensive systems-level assessment appropriate for elite programs and research collaborations.
Table 6.
Practical view of DNA, RNA, protein and integrated sportomics tools for youth sport: This table presents a practical framework stratifying sportomics assessment tools across biological levels: genomics provides stable, heritable information informing long-term training planning; transcriptomics enables research-level understanding of training-induced molecular adaptation; proteomics offers dynamic biomarkers indicating whether training stimulus matches athlete capacity; and integrated multi-omics provides comprehensive systems-level assessment appropriate for elite programs and research collaborations.
| Level |
What it is (for coaches) |
Main benefit in practice |
Who might consider it |
Key limitations / points to remember |
| DNA (Genomics) |
Relatively fixed blueprint: variants in genes related to muscle type, cardiovascular function, connective tissue, inflammation |
Helps tailor training focus (endurance vs power), identify children who may need slower load progression or extra injury-prevention work |
Competitive youth athletes and academies seeking long-term planning; usually one-time testing with updates only if needed |
Shows tendencies, not guarantees; must not be used to label or exclude; requires robust consent and privacy safeguards |
| RNA (Transcriptomics) |
Snapshot of which genes are “switched on” in response to training or recovery |
Helps researchers understand how a particular training block is affecting muscle adaptation at a molecular level |
Mainly research settings and high-performance centres; can inform future guidelines |
Not yet routine for day-to-day coaching; invasive, costlier, and interpretation is complex |
| Protein (Proteomics) |
Markers of muscle damage, inflammation, and repair in blood |
Indicates whether a training block is within the athlete’s current capacity or causing excessive stress; helps optimize rest days and deload weeks |
Health professionals supporting youth teams; intensive training camps; return-to-sport monitoring after injury |
Values fluctuate; must always be interpreted with symptoms and training load; over-testing may cause confusion and unnecessary anxiety |
| Integrated panels (multi-omics + wearables) |
Combined picture using DNA + proteins + metabolites + heart rate, sleep, workload data |
Allows individualized training and recovery plans; supports early detection of overuse risk; can inform nutrition strategies |
Elite development programmes; research collaborations; selected high-commitment families |
More data ≠ better decisions without expert interpretation; higher cost; strong data protection and ethical oversight required, especially for minors |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).