Submitted:
06 January 2026
Posted:
06 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Treatments
2.2. Identification of WRKY Genes in Tetraploid C. oleifera Genome
2.3. Characterization and Classification of WRKY Genes
2.4. Chromosomal Localization and Phylogenetic Analysis
2.5. RNA-Seq and Differential Expression Analysis
2.6. Functional Annotation and Network Analysis
3. Results
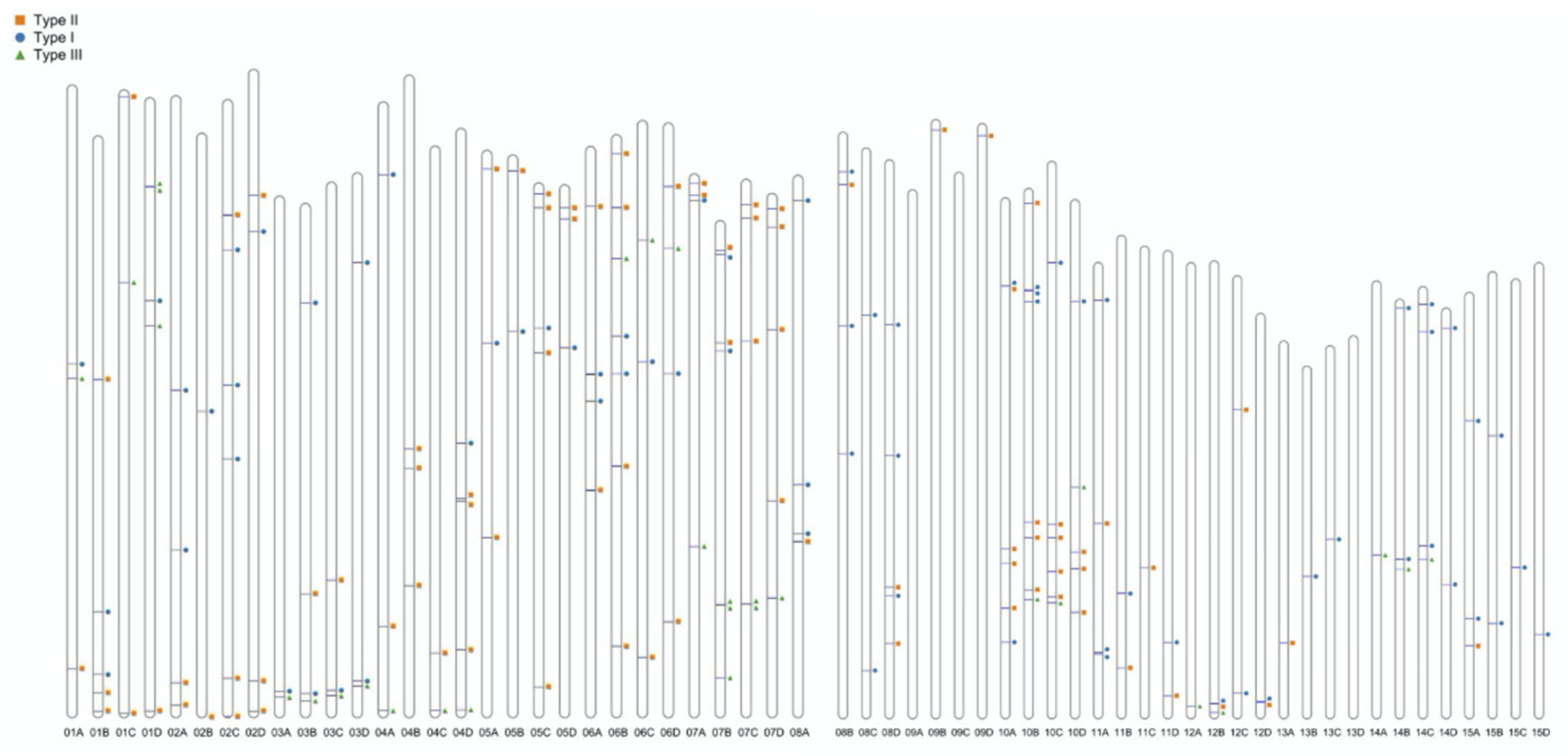
3.1. Genome-Wide Identification and Chromosomal Distribution of WRKY Genes
3.2. Conserved Domain and Gene Structure Analysis
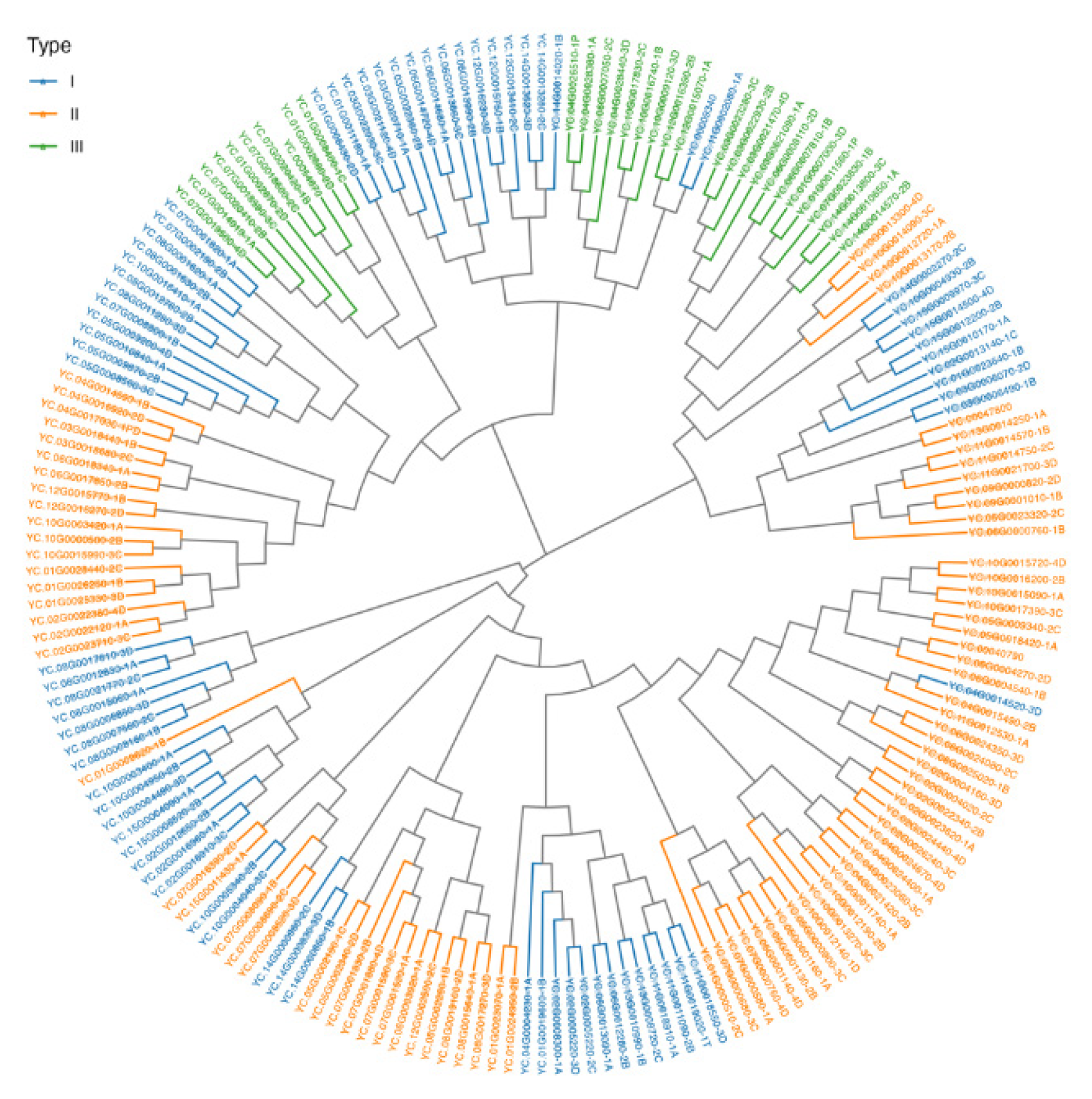
3.3. Phylogenetic Relationship of the WRKY Genes
3.4. Gene Expression Pattern of WRKY and Response to Anthracnose Infection
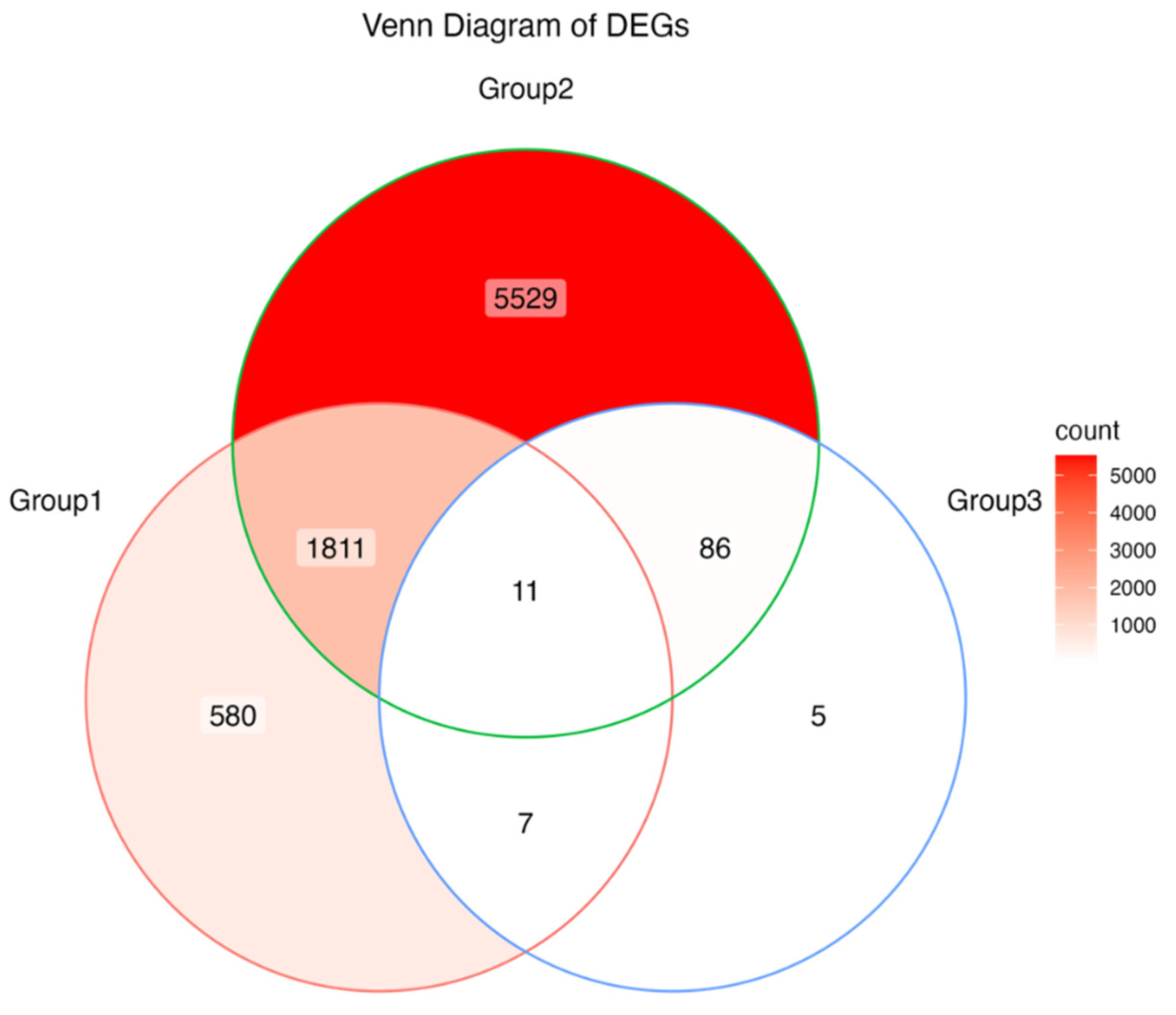
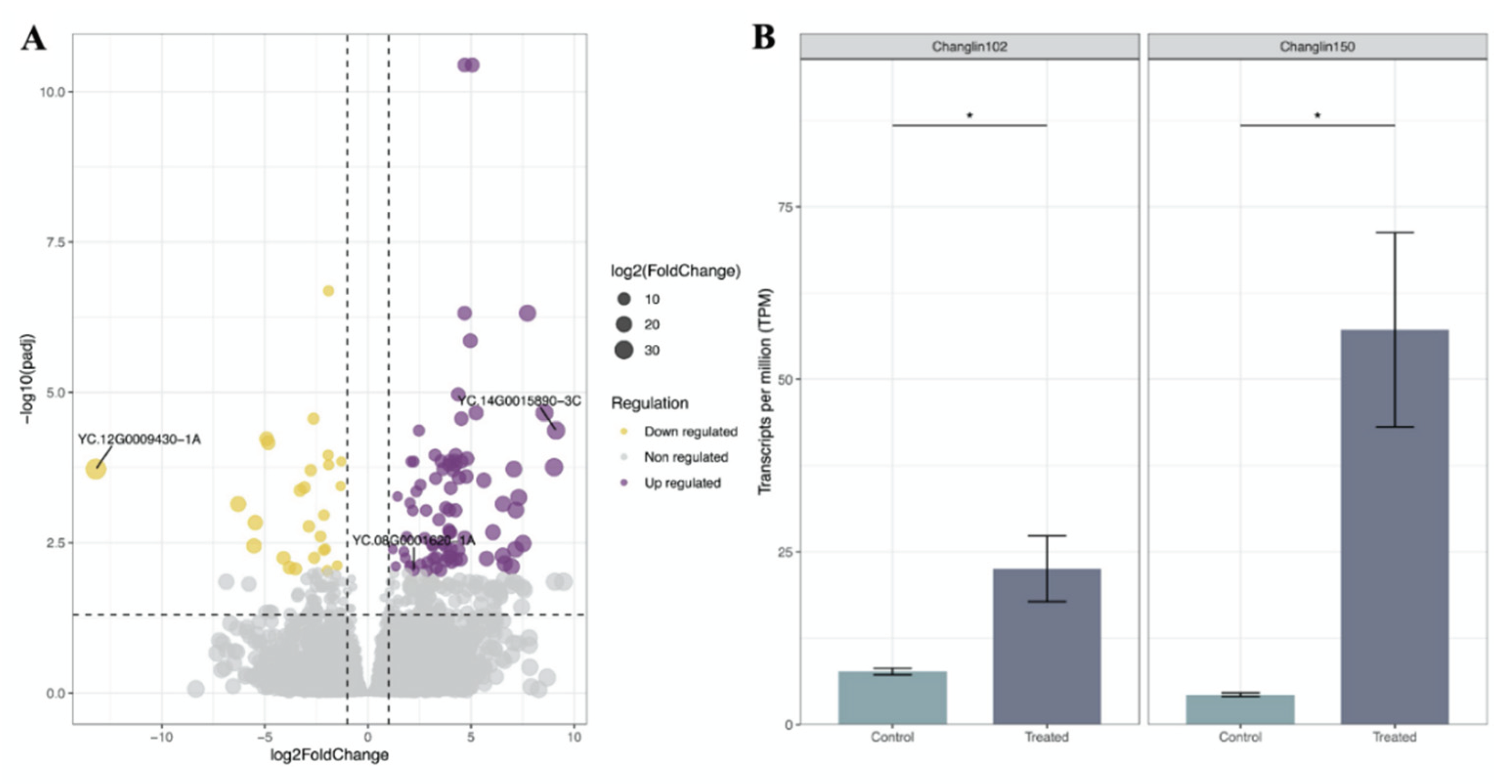
3.5. Differentially Expressed Genes (DEGs) Under Anthracnose Stress
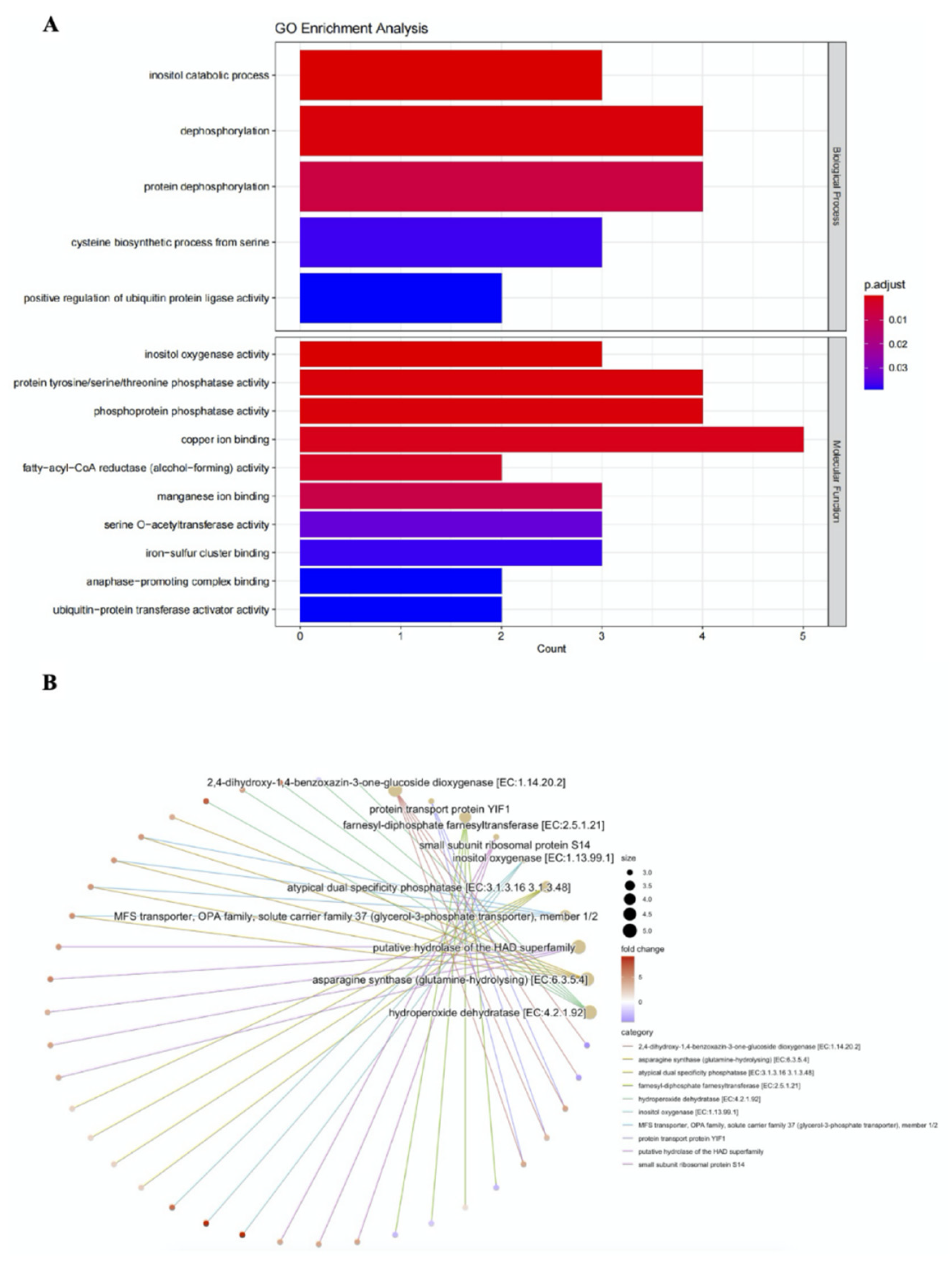
3.6. A Candidate WRKY Gene Involved into Resistance to Anthracnose
4. Discussion
4.1. Expansion and Evolutionary Dynamics of the WRKY Family
4.2. Expression Dynamics on Anthracnose Resistance
4.3. Candidate Genes and Process to Anthracnose Resistance
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Song, H.; Cao, Y.; Zhao, L.; Zhang, J.; Li, S. Review: WRKY transcription factors: Understanding the functional divergence. Plant Sci. 2023, 334, 111770. [Google Scholar] [CrossRef] [PubMed]
- Eulgem, T.; Rushton, P.J.; Robatzek, S.; Somssich, I.E. The WRKY superfamily of plant transcription factors. Trends Plant Sci. 2000, 5, 199–206. [Google Scholar] [CrossRef] [PubMed]
- Rushton, P.J.; et al. WRKY transcription factors. Trends Plant Sci 2010, 15, 247–58. [Google Scholar] [CrossRef] [PubMed]
- Yin, W.; Wang, X.; Liu, H.; Wang, Y.; van Nocker, S.; Tu, M.; Fang, J.; Guo, J.; Li, Z.; Wang, X. Overexpression of VqWRKY31 enhances powdery mildew resistance in grapevine by promoting salicylic acid signaling and specific metabolite synthesis. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef]
- Liu, Y.; et al. MdERF114 enhances the resistance of apple roots to Fusarium solani by regulating the transcription of MdPRX63. Plant Physiol 2023, 192, 2015–2029. [Google Scholar] [CrossRef]
- Wang, Y.; et al. VqWRKY56 interacts with VqbZIPC22 in grapevine to promote proanthocyanidin biosynthesis and increase resistance to powdery mildew. New Phytol 2023, 237, 1856–1875. [Google Scholar]
- Mu, Y.; Dong, Y.; Li, X.; Gong, A.; Yu, H.; Wang, C.; Liu, J.; Liang, Q.; Yang, K.; Fang, H. JrPHL8-JrWRKY4-JrSTH2L module regulates resistance to Colletotrichum gloeosporioides in walnut. Hortic. Res. 2024, 11. [Google Scholar] [CrossRef]
- Yan, H.; Jia, H.; Chen, X.; Hao, L.; An, H.; Guo, X. The Cotton WRKY Transcription Factor GhWRKY17 Functions in Drought and Salt Stress in Transgenic Nicotiana benthamiana Through ABA Signaling and the Modulation of Reactive Oxygen Species Production. Plant Cell Physiol. 2014, 55, 2060–2076. [Google Scholar] [CrossRef]
- Wei, W.; Liang, D.; Bian, X.; Shen, M.; Xiao, J.; Zhang, W.; Ma, B.; Lin, Q.; Lv, J.; Chen, X.; et al. GmWRKY54 improves drought tolerance through activating genes in abscisic acid and Ca2+ signaling pathways in transgenic soybean. Plant J. 2019, 100, 384–398. [Google Scholar] [CrossRef]
- Zhang, J.; Huang, D.; Zhao, X.; Zhang, M.; Wang, Q.; Hou, X.; Di, D.; Su, B.; Wang, S.; Sun, P. Drought-responsive WRKY transcription factor genes IgWRKY50 and IgWRKY32 from Iris germanica enhance drought resistance in transgenic Arabidopsis. Front. Plant Sci. 2022, 13, 983600. [Google Scholar] [CrossRef]
- Yuan, S.; Wang, Y.; Hu, D.; Zhu, W.; Xiao, D.; Liu, T.; Hou, X.; Li, Y. BcNAC056 Interacts with BcWRKY1 to Regulate Leaf Senescence in Pak Choi. Plant Cell Physiol. 2023, 64, 1091–1105. [Google Scholar] [CrossRef]
- Zhao, L.; Zhang, W.; Song, Q.; Xuan, Y.; Li, K.; Cheng, L.; Qiao, H.; Wang, G.; Zhou, C. A WRKY transcription factor, TaWRKY40-D, promotes leaf senescence associated with jasmonic acid and abscisic acid pathways in wheat. Plant Biol. 2020, 22, 1072–1085. [Google Scholar] [CrossRef]
- Singh, D.; Debnath, P.; Sane, A.P.; Sane, V.A. Tomato (Solanum lycopersicum) WRKY23 enhances salt and osmotic stress tolerance by modulating the ethylene and auxin pathways in transgenic Arabidopsis. Plant Physiol. Biochem. 2023, 195, 330–340. [Google Scholar] [CrossRef]
- Chen, J.; Wang, H.; Li, Y.; Pan, J.; Hu, Y.; Yu, D. Arabidopsis VQ10 interacts with WRKY8 to modulate basal defense against Botrytis cinerea. J. Integr. Plant Biol. 2018, 60, 956–969. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Zhao, W.; Li, C.; Qiao, H.; Song, S.; Yang, R.; Sun, L.; Ma, J.; Ma, X.; Wang, S. SlVQ15 interacts with jasmonate-ZIM domain proteins and SlWRKY31 to regulate defense response in tomato. Plant Physiol. 2022, 190, 828–842. [Google Scholar] [CrossRef]
- Yang, C.; Liu, X.; Chen, Z.; Lin, Y.; Wang, S. Comparison of Oil Content and Fatty Acid Profile of Ten NewCamellia oleiferaCultivars. J. Lipids 2016, 2016, 3982486. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, J.-Y.; Xu, X.; Cheng, J.; Zheng, L.; Huang, J.; Li, D.-W. Identification and Characterization ofColletotrichumSpecies Associated with Anthracnose Disease ofCamellia oleiferain China. Plant Dis. 2020, 104, 474–482. [Google Scholar] [CrossRef]
- Zhang, L.; et al. The tetraploid Camellia oleifera genome provides insights into evolution, agronomic traits, and genetic architecture of oil Camellia plants. Cell Rep 2024, 43, 115032. [Google Scholar] [CrossRef]
- Zhu, H.; Wang, F.; Xu, Z.; Wang, G.; Hu, L.; Cheng, J.; Ge, X.; Liu, J.; Chen, W.; Li, Q.; et al. The complex hexaploid oil-Camellia genome traces back its phylogenomic history and multi-omics analysis of Camellia oil biosynthesis. Plant Biotechnol. J. 2024, 22, 2890–2906. [Google Scholar] [CrossRef]
- Wang, R.; Li, W.; He, Z.; Lyu, H.; Wang, X.; Ye, C.; Xun, C.; Xiao, G.; Zhang, Y.; Zhang, Z.; et al. Haplotype-resolved genome assembly of the tetraploid Youcha tree Camellia meiocarpa Hu. Sci. Data 2025, 12, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Dou, L.; Zhang, X.; Pang, C.; Song, M.; Wei, H.; Fan, S.; Yu, S. Genome-wide analysis of the WRKY gene family in cotton. Mol. Genet. Genom. 2014, 289, 1103–1121. [Google Scholar] [CrossRef]
- Yu, Y.; Wang, N.; Hu, R.; Xiang, F. Genome-wide identification of soybean WRKY transcription factors in response to salt stress. SpringerPlus 2016, 5, 92. [Google Scholar] [CrossRef]
- Yang, C.; Wu, P.; Yao, X.; Sheng, Y.; Zhang, C.; Lin, P.; Wang, K. Integrated Transcriptome and Metabolome Analysis Reveals Key Metabolites Involved in Camellia oleifera Defense against Anthracnose. Int. J. Mol. Sci. 2022, 23, 536. [Google Scholar] [CrossRef]
- Eddy, S.R. Accelerated Profile HMM Searches. PLOS Comput. Biol. 2011, 7, e1002195. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Hao, Z.; Lv, D.; Ge, Y.; Shi, J.; Weijers, D.; Yu, G.; Chen, J. RIdeogram: drawing SVG graphics to visualize and map genome-wide data on the idiograms. PeerJ Comput. Sci. 2020, 6, e251. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Stecher, G.; Tamura, K.; Kumar, S. Molecular Evolutionary Genetics Analysis (MEGA) for macOS. Mol. Biol. Evol. 2020, 37, 1237–1239. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows—Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Smyth, G.K.; Shi, W. feature Counts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014, 15, 550. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef]
- Abdullah-Zawawi, M.-R.; Ahmad-Nizammuddin, N.-F.; Govender, N.; Harun, S.; Mohd-Assaad, N.; Mohamed-Hussein, Z.-A. Comparative genome-wide analysis of WRKY, MADS-box and MYB transcription factor families in Arabidopsis and rice. Sci. Rep. 2021, 11, 1–18. [Google Scholar] [CrossRef]
- Du, C.; Yu, H.; Hu, H.; Sun, E.; Cai, M.; Dou, Z.; Dong, H.; Zuo, C. Evolutional, expressional and functional analysis of WRKY gene family reveals that PbeWRKY16 and PbeWRKY31 contribute to the Valsa canker resistance in Pyrus betulifolia. Plant Physiol. Biochem. 2025, 222, 109719. [Google Scholar] [CrossRef]
- Wu, W.; Zhu, S.; Xu, L.; Zhu, L.; Wang, D.; Liu, Y.; Liu, S.; Hao, Z.; Lu, Y.; Yang, L.; et al. Genome-wide identification of the Liriodendron chinense WRKY gene family and its diverse roles in response to multiple abiotic stress. BMC Plant Biol. 2022, 22, 1–27. [Google Scholar] [CrossRef] [PubMed]
- Ryu, H.-S.; Han, M.; Lee, S.-K.; Cho, J.-I.; Ryoo, N.; Heu, S.; Lee, Y.-H.; Bhoo, S.H.; Wang, G.-L.; Hahn, T.-R.; et al. A comprehensive expression analysis of the WRKY gene superfamily in rice plants during defense response. Plant Cell Rep. 2006, 25, 836–847. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Kong, X.; Yang, L.; Fu, M.; Zhang, S. Genome-Wide Identification of WRKY Family Genes and the Expression Profiles in Response to Nitrogen Deficiency in Poplar. Genes 2022, 13, 2324. [Google Scholar] [CrossRef] [PubMed]
- Yin, G.; Xu, H.; Xiao, S.; Qin, Y.; Li, Y.; Yan, Y.; Hu, Y. The large soybean (Glycine max) WRKY TF family expanded by segmental duplication events and subsequent divergent selection among subgroups. BMC Plant Biol. 2013, 13, 148. [Google Scholar] [CrossRef]
- Chen, H.; Lai, Z.; Shi, J.; Xiao, Y.; Chen, Z.; Xu, X. Roles of arabidopsis WRKY18, WRKY40 and WRKY60 transcription factors in plant responses to abscisic acid and abiotic stress. BMC Plant Biol. 2010, 10, 281–281. [Google Scholar] [CrossRef]
- Yang, L.; Ye, C.; Zhao, Y.; Cheng, X.; Wang, Y.; Jiang, Y.-Q.; Yang, B. An oilseed rape WRKY-type transcription factor regulates ROS accumulation and leaf senescence in Nicotiana benthamiana and Arabidopsis through modulating transcription of RbohD and RbohF. Planta 2018, 247, 1323–1338. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Wang, H.; Yu, X.; Cui, K.; Hu, Y.; Xiao, S.; Wen, Y.-Q. Transcription factors VviWRKY10 and VviWRKY30 co-regulate powdery mildew resistance in grapevine. Plant Physiol. 2024, 195, 446–461. [Google Scholar] [CrossRef] [PubMed]


| Homolog Group | Subgenome | Tgroup | |||
|---|---|---|---|---|---|
| A | B | C | D | ||
| 1 | 3 | 5 | 3 | 5 | 16 |
| 2 | 4 | 2 | 6 | 4 | 16 |
| 3 | 2 | 4 | 3 | 3 | 12 |
| 4 | 3 | 3 | 2 | 5 | 13 |
| 5 | 3 | 2 | 5 | 3 | 13 |
| 6 | 4 | 7 | 3 | 4 | 18 |
| 7 | 4 | 7 | 5 | 5 | 21 |
| 8 | 4 | 4 | 2 | 5 | 15 |
| 9 | 0 | 1 | 0 | 1 | 2 |
| 10 | 6 | 8 | 6 | 5 | 25 |
| 11 | 4 | 2 | 1 | 2 | 9 |
| 12 | 1 | 3 | 2 | 2 | 8 |
| 13 | 1 | 1 | 1 | 0 | 3 |
| 14 | 1 | 3 | 4 | 2 | 10 |
| 15 | 3 | 2 | 1 | 1 | 7 |
| Tsubgenome | 43 | 54 | 44 | 47 | 188 |
| Gene ID | Fold Change | p.adj Value | Function Description |
|---|---|---|---|
| YC.01G0018630-2C | 6.60 | 7.03E-03 | manganese ion binding |
| YC.02G0003640-2D | 4.38 | 5.20E-03 | acyltransferase activity, transferring groups other than amino-acyl groups |
| YC.02G0003770-1C | 3.95 | 1.39E-04 | acyltransferase activity, transferring groups other than amino-acyl groups |
| YC.11G0011070-2C | 8.56 | 2.19E-05 | alcohol-forming very long-chain fatty acyl-CoA reductase activity |
| YC.11G0010960-1B | 7.74 | 4.82E-07 | alcohol-forming very long-chain fatty acyl-CoA reductase activity |
| YC.05G0019060-3C | 6.54 | 5.20E-03 | unknown |
| YC.08G0001620-1A | 2.23 | 9.10E-03 | DNA-binding transcription factor activity |
| YC.09G0021520-3C | 7.07 | 1.89E-04 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen |
| YC.09G0020800-2B | 4.37 | 4.23E-03 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen |
| YC.09G0022420-1A | 5.75 | 5.80E-03 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen |
| YC.15G0010630-1C | -2.87 | 1.69E-03 | oxidoreductase activity |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




