Submitted:
03 January 2026
Posted:
06 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Strains, Plasmids, and Enzymes
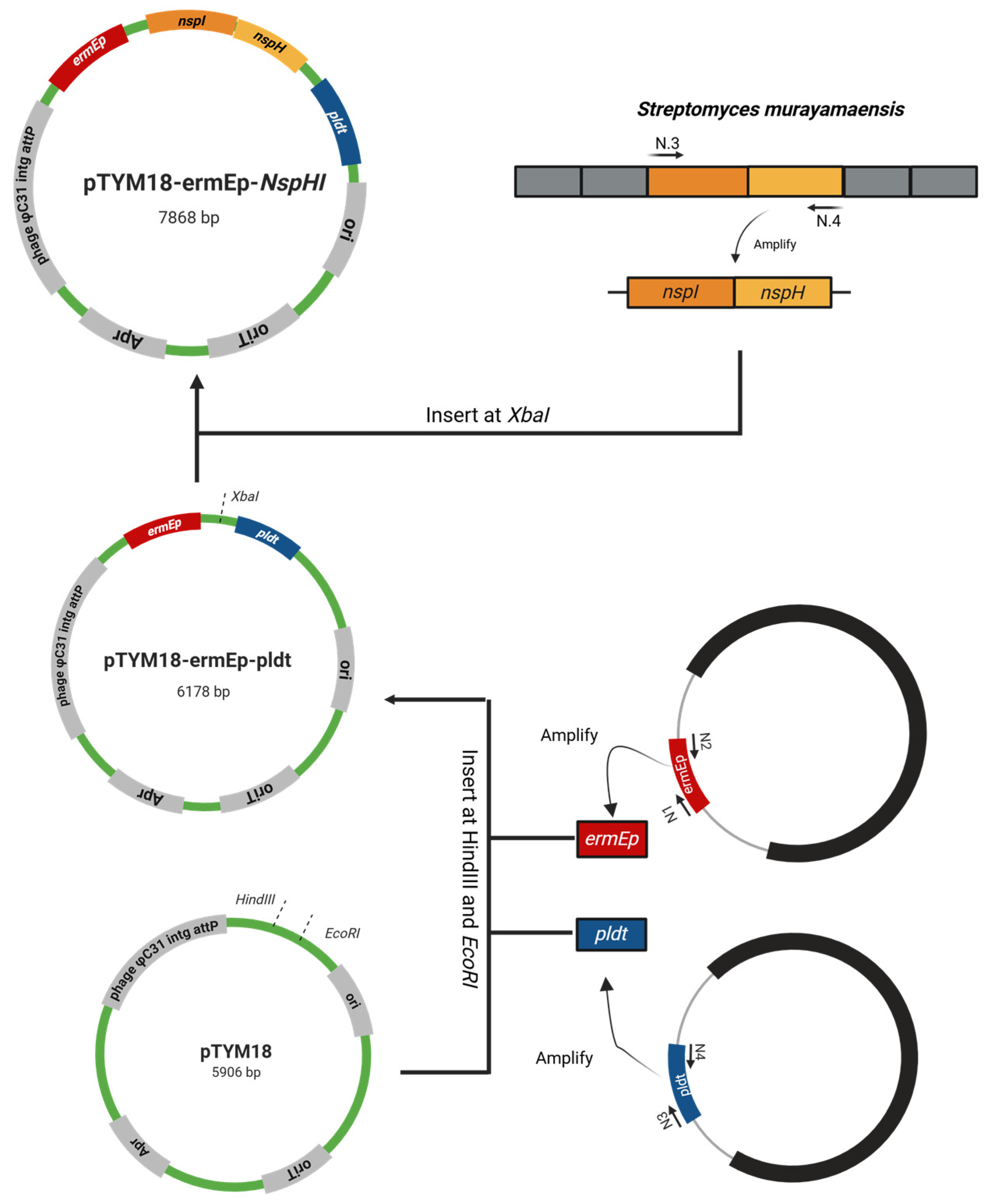
2.2. Plasmid Construction
2.3. Preparation of Transformant Strains
2.4. Fermentation and 3.4-AHBA Analysis
2.5. Model Simulations
2.5.1. Flux balance Analysis (FBA)
2.5.2. Construction of the Metabolic Model
2.5.3. Simulation Settings and Parameter Constraints
2.5.4. Introduction of the 3,4-AHBA Biosynthetic Pathway
3. Results and Discussion
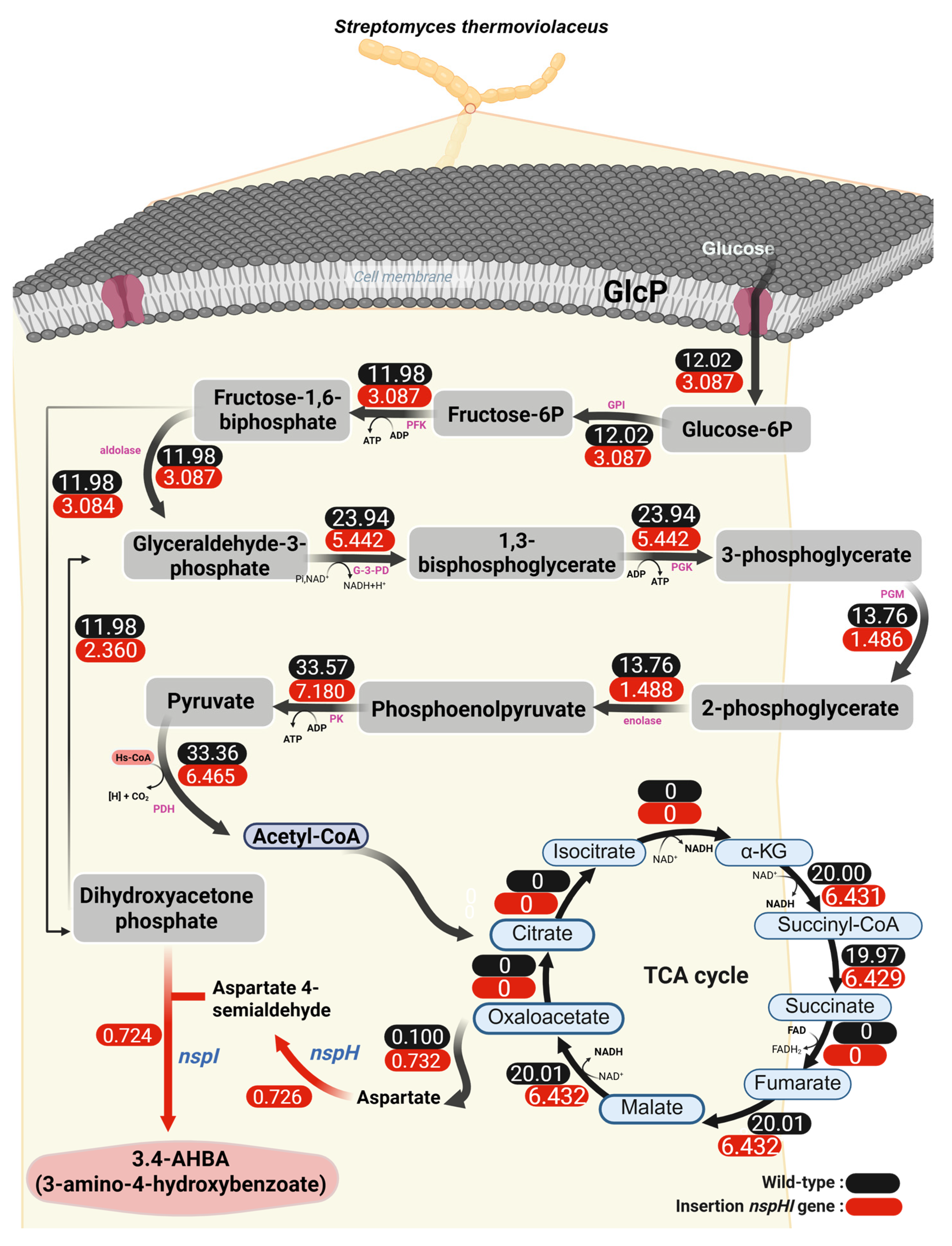
3.1. In Silico Simulation of 3.4-AHBA Production in S. thermoviolaceus Strains
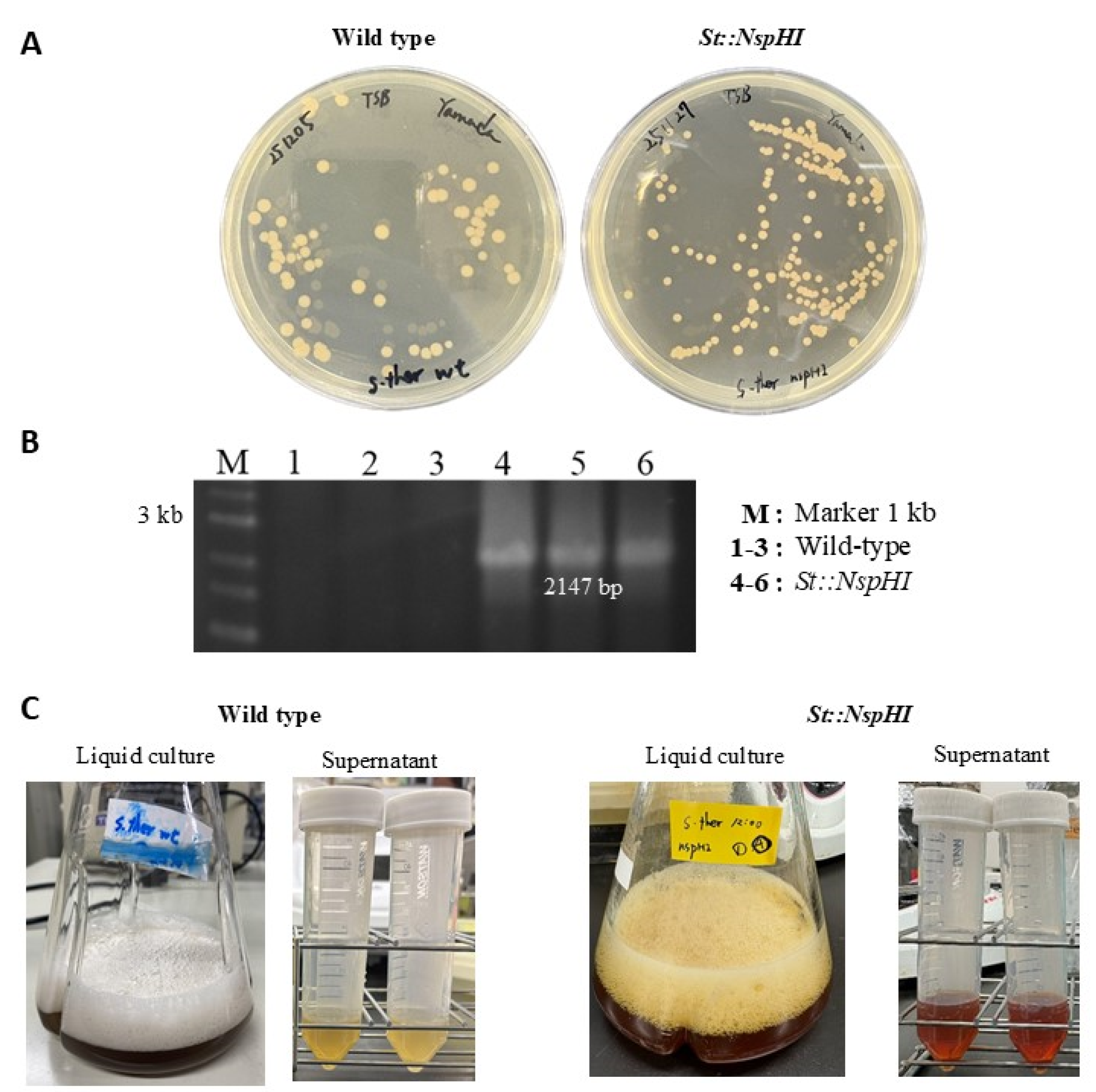
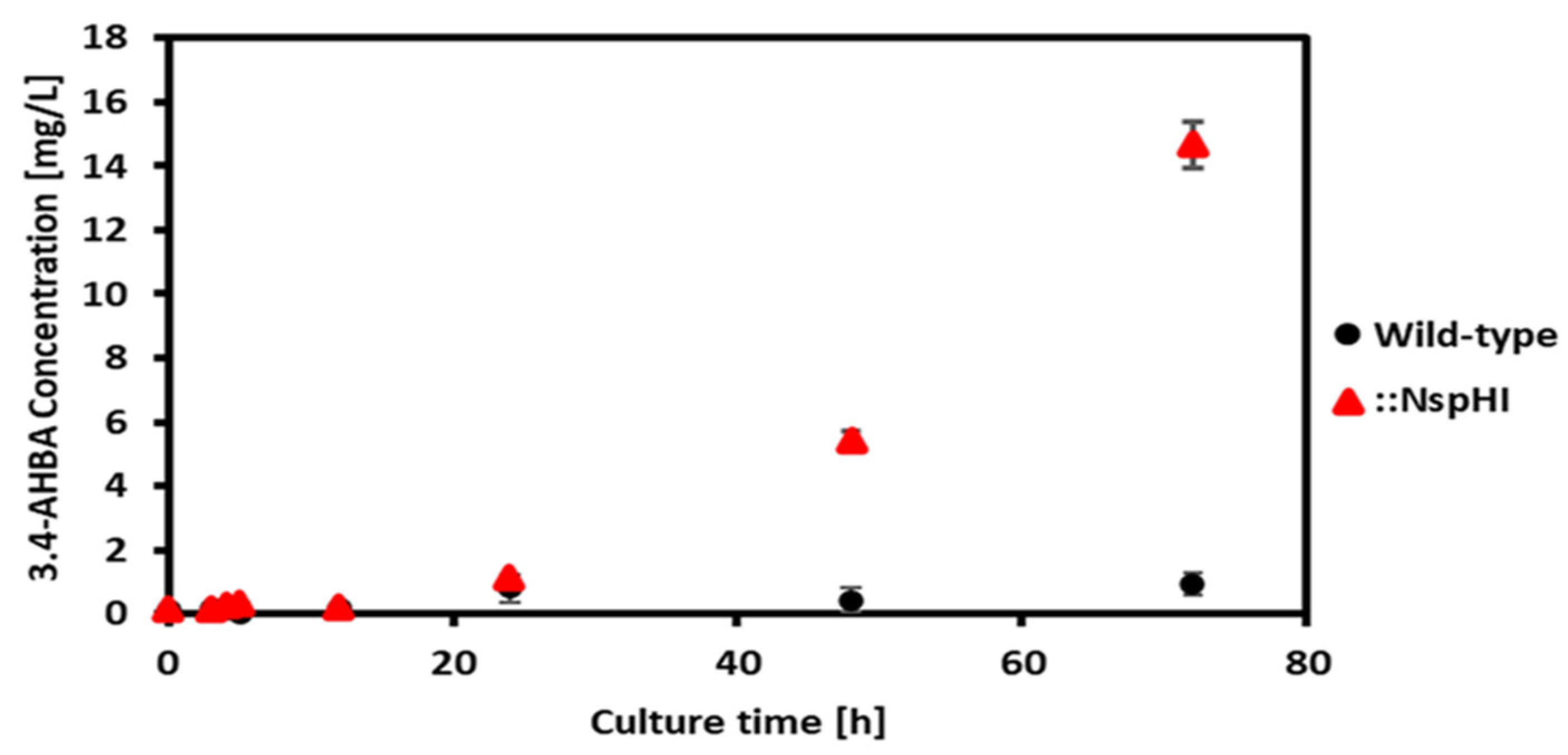
3.2. Effect of nspH–nspI Insertion on 3,4-AHBA Production
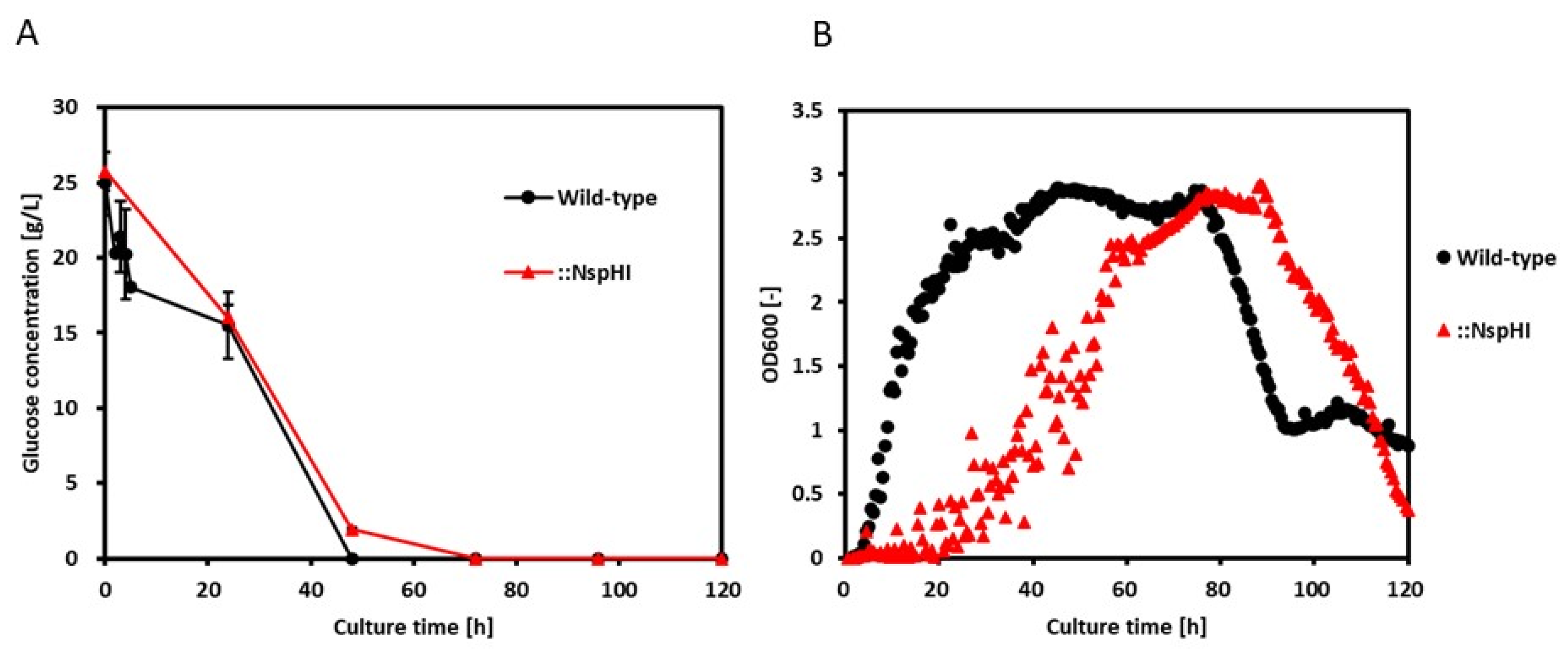
3.3. Comparison of FBA Predictions with Experimental Data
4. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviations
| 3.4-AHBA | 3-amino-4-hydroxybenzoic acid |
| GSM | Genome-Scale Metabolic modeling |
| GEM | Genome-scale metabolic model |
| FBA | Flux Balance Analysis |
| S. thermoviolaceus | Streptomyces thermoviolaceus |
| GAM | Growth-associated maintenance |
| NGAM | Non-growth-associated maintenance |
| NBRC | NITE Biological Resource Center |
| NCBI | National Center for Biotechnology Information |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
References
- Osterlund, T.; Nookaew, I.; Bordel, S.; Nielsen, J. Mapping Condition-Dependent Regulation of Metabolism in Yeast through Genome-Scale Modeling. 2013.
- Vikromvarasiri, N.; Shirai, T.; Kondo, A. Metabolic Engineering Design to Enhance (R,R)-2,3-Butanediol Production from Glycerol in Bacillus Subtilis Based on Flux Balance Analysis. Microb Cell Fact 2021, 20, 196. [CrossRef]
- Feist, A.M.; Henry, C.S.; Reed, J.L.; Krummenacker, M.; Joyce, A.R.; Karp, P.D.; Broadbelt, L.J.; Hatzimanikatis, V.; Palsson, B.O. A Genome-scale Metabolic Reconstruction for Escherichia Coli K-12 MG1655 That Accounts for 1260 ORFs and Thermodynamic Information. Mol Syst Biol 2007, 3. [CrossRef]
- Orth, J.D.; Thiele, I.; Palsson, B.Ø. What Is Flux Balance Analysis? Nat Biotechnol 2010, 28, 245–248. [CrossRef]
- Simensen, V.; Schulz, C.; Karlsen, E.; Bråtelund, S.; Burgos, I.; Thorfinnsdottir, L.B.; García-Calvo, L.; Bruheim, P.; Almaas, E. Experimental Determination of Escherichia Coli Biomass Composition for Constraint-Based Metabolic Modeling. PLoS One 2022, 17, e0262450. [CrossRef]
- Tokuyama, K.; Ohno, S.; Yoshikawa, K.; Hirasawa, T.; Tanaka, S.; Furusawa, C.; Shimizu, H. Increased 3-Hydroxypropionic Acid Production from Glycerol, by Modification of Central Metabolism in Escherichia Coli. Microb Cell Fact 2014, 13, 64. [CrossRef]
- Bentley, S.D.; Chater, K.F.; Cerdeño-Tárraga, A.M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete Genome Sequence of the Model Actinomycete Streptomyces Coelicolor A3(2). Nature 2002, 417, 141–147. [CrossRef]
- Kashiwagi, N.; Ogino, C.; Kondo, A. Production of Chemicals and Proteins Using Biomass-Derived Substrates from a Streptomyces Host. Bioresour Technol 2017, 245, 1655–1663. [CrossRef]
- Lee, N.; Hwang, S.; Kim, W.; Lee, Y.; Kim, J.H.; Cho, S.; Kim, H.U.; Yoon, Y.J.; Oh, M.-K.; Palsson, B.O.; et al. Systems and Synthetic Biology to Elucidate Secondary Metabolite Biosynthetic Gene Clusters Encoded in Streptomyces Genomes. Nat Prod Rep 2021, 38, 1330–1361. [CrossRef]
- Tsujibo, H.; Hatano, N.; Endo, H.; Miyamoto, K.; Inamori, Y. Purification and Characterization of a Thermostable Chitinase from Streptomyces Thermoviolaceus OPC-520 and Cloning of the Encoding Gene. Biosci Biotechnol Biochem 2000, 64, 96–102. [CrossRef]
- Kawaguchi, H.; Sasaki, K.; Uematsu, K.; Tsuge, Y.; Teramura, H.; Okai, N.; Nakamura-Tsuruta, S.; Katsuyama, Y.; Sugai, Y.; Ohnishi, Y.; et al. 3-Amino-4-Hydroxybenzoic Acid Production from Sweet Sorghum Juice by Recombinant Corynebacterium Glutamicum. Bioresour Technol 2015, 198, 410–417. [CrossRef]
- Kim, M.; Sang Yi, J.; Kim, J.; Kim, J.; Kim, M.W.; Kim, B. Reconstruction of a High-quality Metabolic Model Enables the Identification of Gene Overexpression Targets for Enhanced Antibiotic Production in Streptomyces Coelicolor A3(2). Biotechnol J 2014, 9, 1185–1194. [CrossRef]
- Orth, J.D.; Fleming, R.M.T.; Palsson, B.Ø. Reconstruction and Use of Microbial Metabolic Networks: The Core Escherichia Coli Metabolic Model as an Educational Guide. EcoSal Plus 2010, 4. [CrossRef]
- Becker, S.A.; Feist, A.M.; Mo, M.L.; Hannum, G.; Palsson, B.Ø.; Herrgard, M.J. Quantitative Prediction of Cellular Metabolism with Constraint-Based Models: The COBRA Toolbox. Nat Protoc 2007, 2, 727–738. [CrossRef]
- Hasibi, R.; Michoel, T.; Oyarzún, D.A. Integration of Graph Neural Networks and Genome-Scale Metabolic Models for Predicting Gene Essentiality. NPJ Syst Biol Appl 2024, 10, 1–10. [CrossRef]
- Kawaguchi, H.; Hasunuma, T.; Ohnishi, Y.; Sazuka, T.; Kondo, A.; Ogino, C. Enhanced Production of γ-Amino Acid 3-Amino-4-Hydroxybenzoic Acid by Recombinant Corynebacterium Glutamicum under Oxygen Limitation. Microb Cell Fact 2021, 20, 228. [CrossRef]
- Huang, D.; Wen, J.; Wang, G.; Yu, G.; Jia, X.; Chen, Y. In Silico Aided Metabolic Engineering of Streptomyces Roseosporus for Daptomycin Yield Improvement. Appl Microbiol Biotechnol 2012, 94, 637–649. [CrossRef]
- Rubio, A.; Razmilic, V.; Brain-Isasi, S.; Andrews, B.; Asenjo, J.A. Metabolic Engineering of Streptomyces Leeuwenhoekii C34 T to Increase Chaxamycin Production Based on the i VR1007 Genome-Scale Model. Biotechnol Bioeng 2025, 122, 3376–3392. [CrossRef]
- Suzuki, H.; Ohnishi, Y.; Furusho, Y.; Sakuda, S.; Horinouchi, S. Novel Benzene Ring Biosynthesis from C3 and C4 Primary Metabolites by Two Enzymes. Journal of Biological Chemistry 2006, 281, 36944–36951. [CrossRef]
- Niimi-Nakamura, S.; Kawaguchi, H.; Uematsu, K.; Teramura, H.; Nakamura-Tsuruta, S.; Kashiwagi, N.; Sugai, Y.; Katsuyama, Y.; Ohnishi, Y.; Ogino, C.; et al. 3-Amino-4-Hydroxybenzoic Acid Production from Glucose and/or Xylose via Recombinant Streptomyces Lividans. J Gen Appl Microbiol 2022, 68, 2022.06.001. [CrossRef]
- OHNISHI, Y.; FURUSHO, Y.; HIGASHI, T.; CHUN, H.-K.; FURIHATA, K.; SAKUDA, S.; HORINOUCHI, S. Structures of Grixazone A and B, A-Factor-Dependent Yellow Pigments Produced under Phosphate Depletion by Streptomyces Griseus. J Antibiot (Tokyo) 2004, 57, 218–223. [CrossRef]
- Wu, G.; Yan, Q.; Jones, J.A.; Tang, Y.J.; Fong, S.S.; Koffas, M.A.G. Metabolic Burden: Cornerstones in Synthetic Biology and Metabolic Engineering Applications. Trends Biotechnol 2016, 34, 652–664. [CrossRef]
- Asah-Asante, R.; Tang, L.; Gong, X.; Fan, S.; Yan, C.; Asante, J.O.; Zeng, Q. Exploring Pigment-Producing Streptomyces as an Alternative Source to Synthetic Pigments: Diversity, Biosynthesis, and Biotechnological Applications. A Review. World J Microbiol Biotechnol 2025, 41, 211. [CrossRef]
- Onaka, H.; Taniguchi, S.-I.; Ikeda, H.; Igarashi, Y.; Fuurumai, T. PTOYAMAcos, PTYM18, and PTYM19, Actinomycete-Escherichia Coli Integrating Vectors for Heterologous Gene Expression. J Antibiot (Tokyo) 2003, 56, 950–956. [CrossRef]
| Materials | Description | Source |
| Strains: | ||
| E. coli Nova Blue | For routine plasmid maintenance (endA1 hsdR17 (rK12– mK12+) supE44 thi-1 recA1 gyrA96 relA1 lac F′[proA+B+ lacIqZΔM15::Tn10] (TetR) | |
| E. coli HST04 | Cloning host. ((F-, ara, Δ(lacproAB) [Φ80dlacZΔM15], rpsL(str), thi, Δ(mrr-hsdRMS-mcrBC), ΔmcrA, dam, dcm)) | Takara |
| E. coli JM109 | For conjugation plasmid into Streptomyces (endA1 recA1 gyrA96 thi-1 hsdR17 (rK–, mK+) relA1 supE44 Δ(lac-proAB) / F' [traD36 proA+B+ lacIq lacZΔM15]) | Takara |
| Strptomyces thermoviolaceus NBRC 13905 | Expression host | NBRC |
| St::NspHI | Strain of Streptomyces thermoviolaceus::nspH::nspI | This study |
| Plasmids: | ||
| pTYM18 | [11] | |
| pTYM18-ermE-nspHI | Plasmid for nspHI expression; apramycin resistance marker; kasO promoter; pld terminator; rep, replication gene | This study |
| PUB307 | PΔTnAc,IncPαrepliconKmrTCr |
| Primers | Description |
| ermEp-MCS_Fw (N.1) | ggccagtgccaagcttCACGCGGTCGATCTTGACGGCTG |
| ermEp-MCS-PLDt_Rv (N.2) | ggatccggtaccCGCTGGATCCTACCAACCGGCAC- |
| PLDt_Fw | GCGggtaccggatcctctagaTGAGACGACTGAGCGCCCGGAC |
| PLDt_Rv | acatgattacgaattcATTTCCTCGCTGGTCGGTTC |
| nspHI-tandem1-Hind3-Fw (N.3) | gccagtgccaagcttCACGCGGTCGATCTTGACG |
| nspHI-tandem2-Xba1-Rv (N.4) | GTCGTCTCAtctagactacttctccaggcagaactc |
| Wild-type | St::NspHI | ||||
| Glucose uptake rate [mmol/gDCW/h] | Experimental growth [/h] |
In silico growth [/h] | Glucose uptake rate [mmol/gDCW/h] | Experimental growth [/h] |
In silico growth [/h] |
| -12.02 | 0.916 | 0.874 | -3.087 | 0.0736 | 0.336 |
| Parameter (mmol/gDCW/h) | S. thermoviolaceus | |
| Wild type | St::NspHI | |
| *Glucose consumption | -12.02 | -3.087 |
| *Biomass (Growth rate) [/h] | 0.916 | 0.0377 |
| Oxygen consumption | -10 | -10 |
| DHAP exchange | 11.98 | 3.084 |
| ASA exchange | 0.0257 | 0.726 |
| 3.4-AHBA exchange | 0 | 0.724 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




