Submitted:
02 January 2026
Posted:
06 January 2026
You are already at the latest version
Abstract
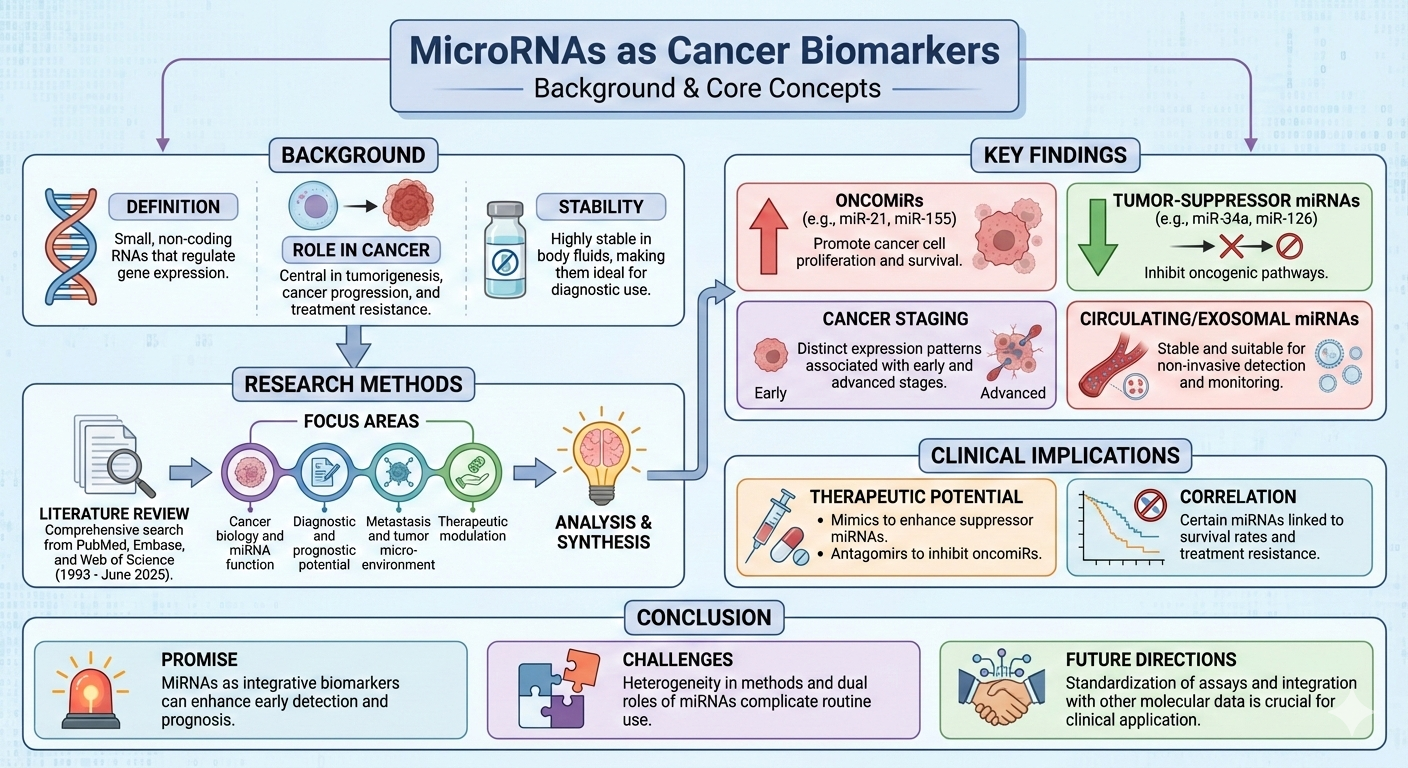
Keywords:
1. Introduction
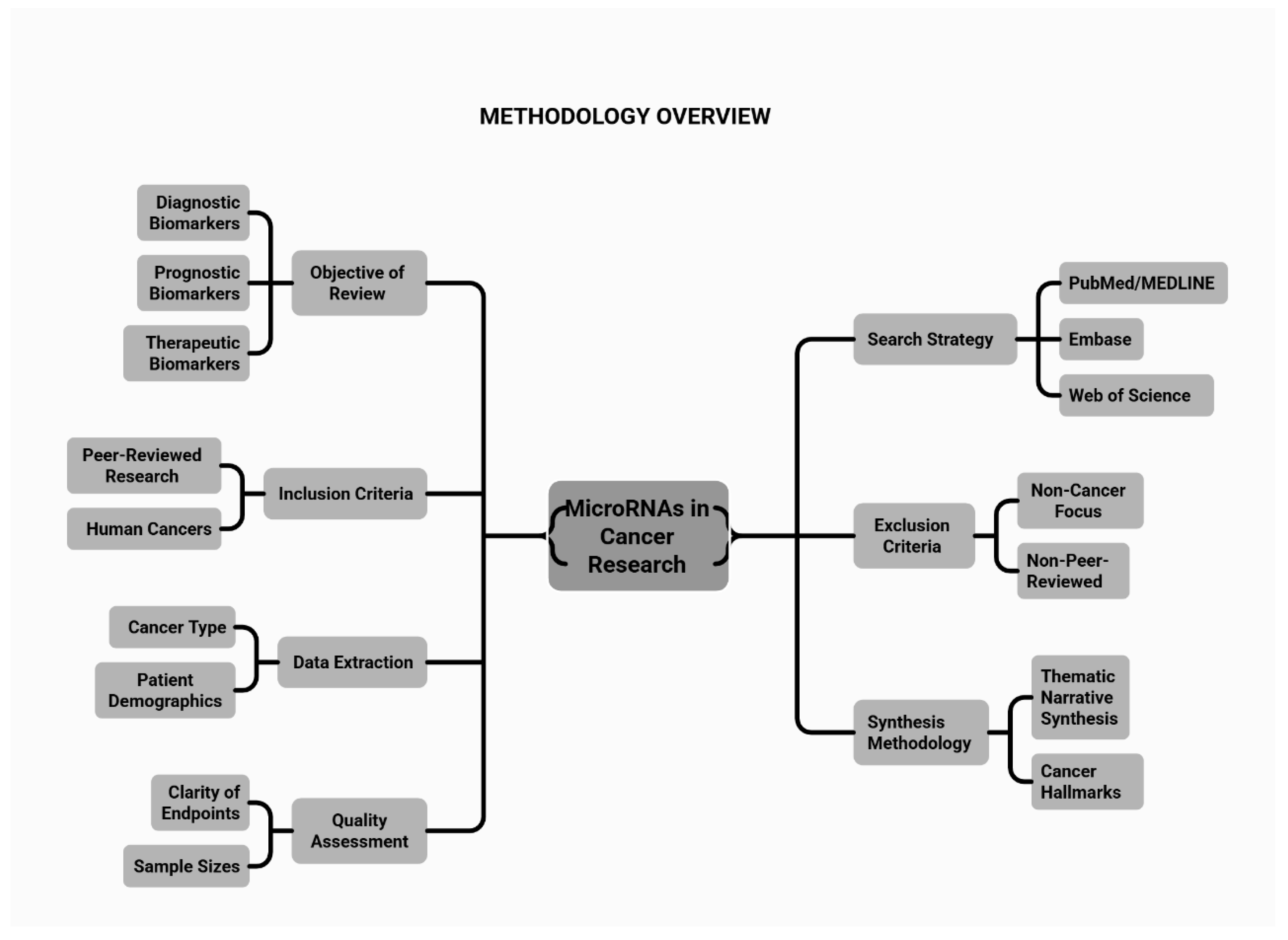
2. Methodology
2.1. Methods
2.2. Search Strategy and Selection Criteria
2.3. Inclusion and Exclusion Criteria
2.4. Data Extraction and Synthesis
2.5. Assessment of Study Quality and Limitations
3. miRNAs in Cancer Crosstalk: Tumor Suppressors versus OncomiRs
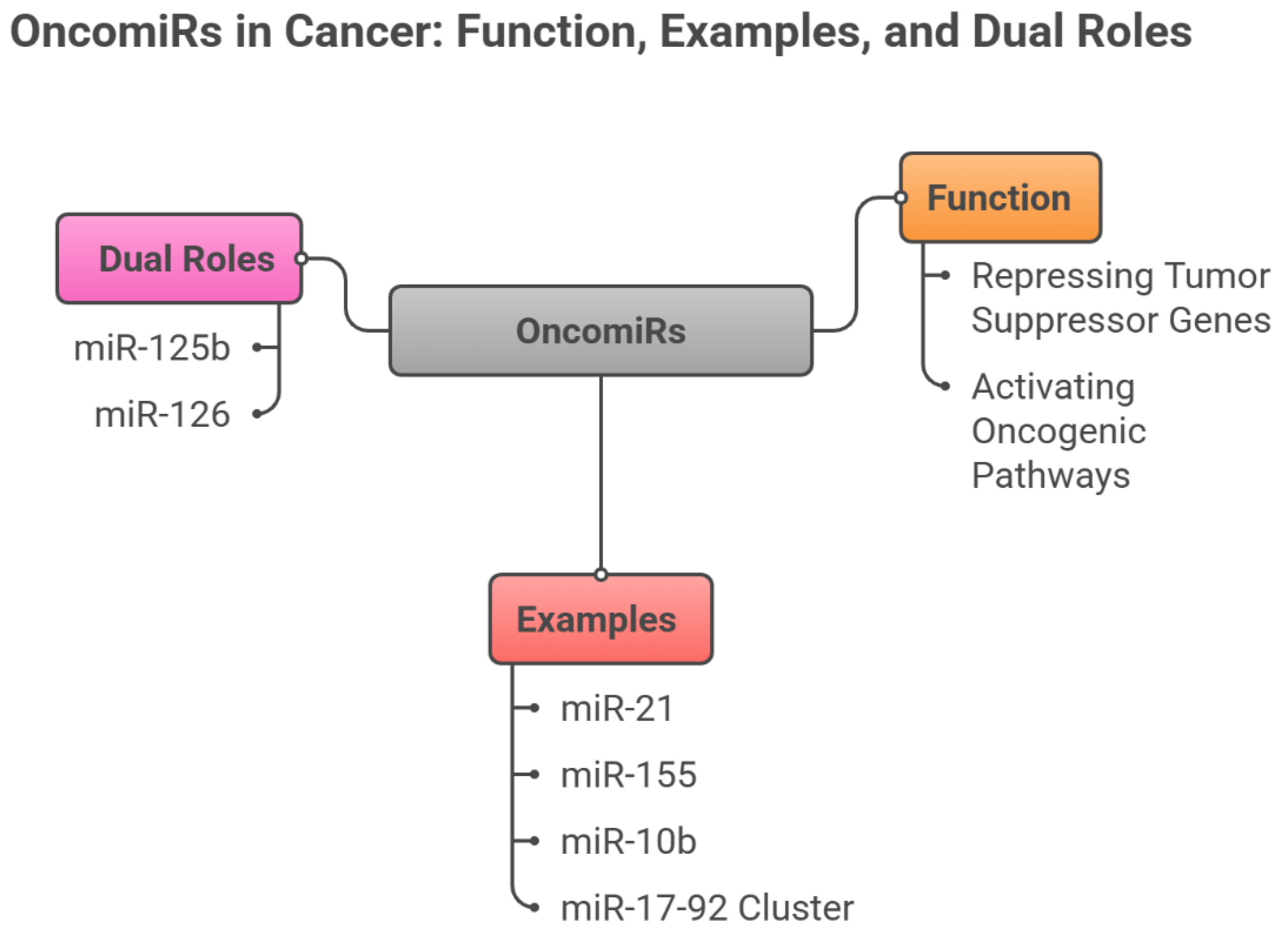
3.1. OncomiRs (Oncogenic miRNAs)
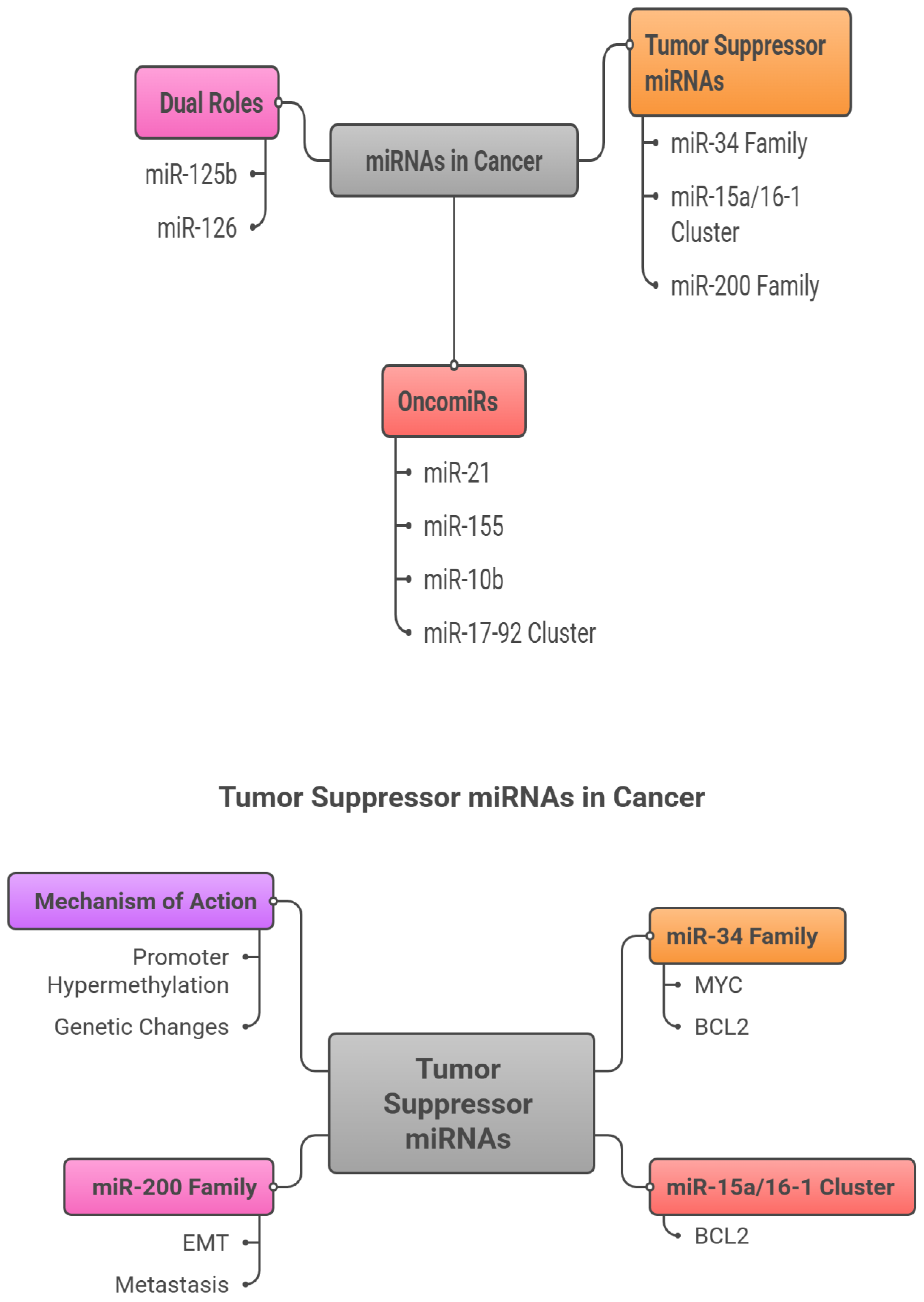
3.2. Tumor Suppressor miRNAs
4. OncomiRs as Cancer Biomarkers
4.1. Early Detection and Diagnosis
4.2. Prognostic Indicators
4.3. Metastatic Markers
5. Tumor-Suppressor miRNAs as Cancer Biomarkers
5.1. Early Detection and Diagnosis
5.2. Prognostic Indicators
5.3. Metastatic Markers
6. Frequency Patterns and Clinical Roles of Oncogenic and Tumor-Suppressive microRNAs in the Research Literature
6.1. OncomiRs: Frequency Patterns, Cancer Type, Sample Type and Clinical Roles
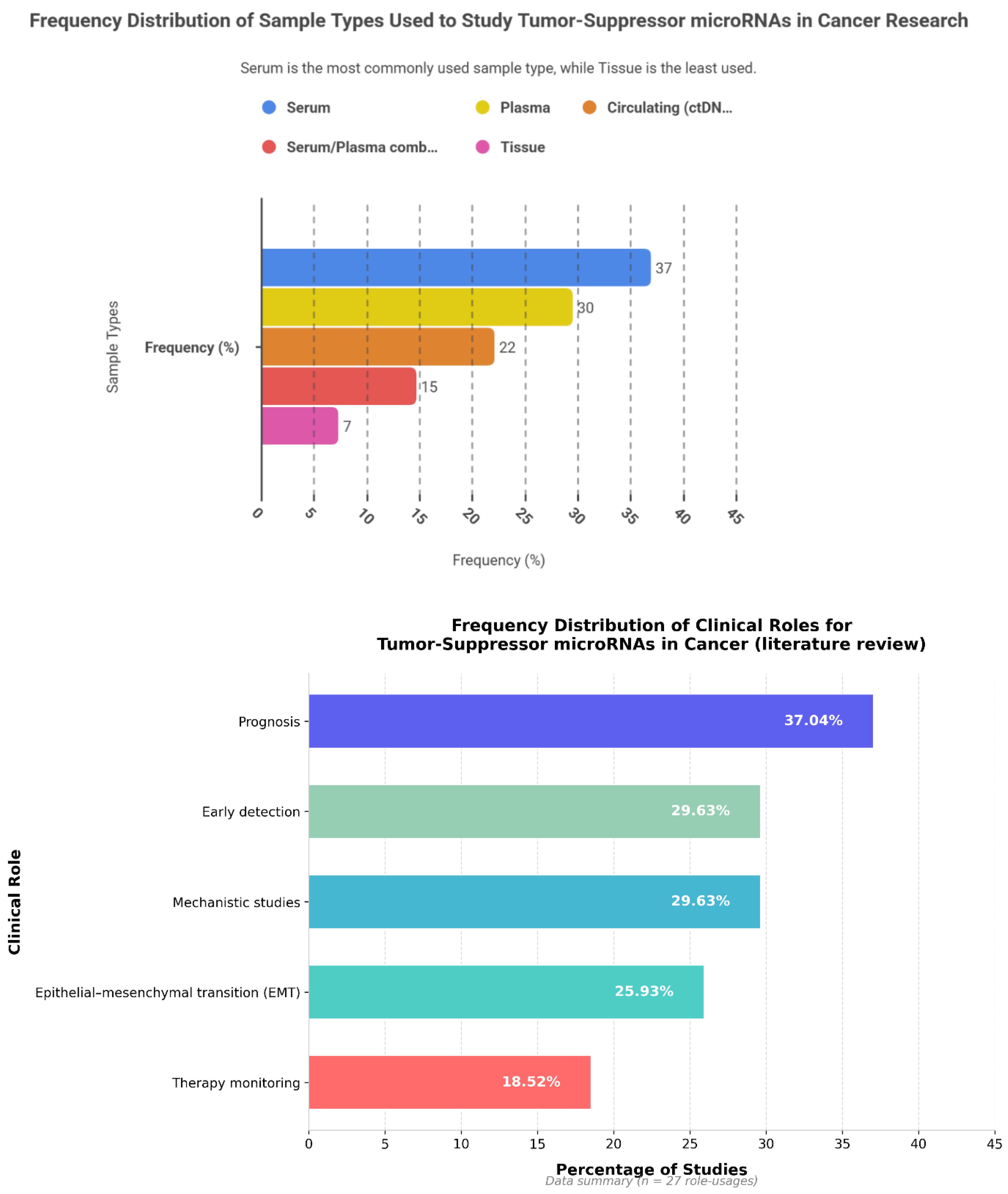
6.2. Tumor-Suppressive miRNAs: Frequency Patterns, Cancer Type, Sample Type and Clinical Roles
7. Discussion
8. Limitations
9. Future Directions / Implications
10. Conclusion
Acknowledgments
Conflicts of Interest
References
- Bautista-Sánchez, D.; Arriaga-Canon, C.; Pedroza-Torres, A.; De La Rosa-Velázquez, I.A.; González-Barrios, R.; Contreras-Espinosa, L.; Montiel-Manríquez, R.; Castro-Hernández, C.; Fragoso-Ontiveros, V.; Álvarez-Gómez, R.M. The promising role of miR-21 as a cancer biomarker and its importance in RNA-based therapeutics. Molecular therapy Nucleic acids 2020, 20, 409–420. [Google Scholar] [CrossRef]
- Billi, M.; De Marinis, E.; Gentile, M.; Nervi, C.; Grignani, F. Nuclear miRNAs: gene regulation activities. International Journal of Molecular Sciences 2024, 25(11), 6066. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nature Reviews Cancer 2006, 6(11), 857–866. [Google Scholar] [CrossRef]
- Chan, J.A.; Krichevsky, A.M.; Kosik, K.S. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer research 2005, 65(14), 6029–6033. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.-C.; Wentzel, E.A.; Kent, O.A.; Ramachandran, K.; Mullendore, M.; Lee, K.H.; Feldmann, G.; Yamakuchi, M.; Ferlito, M.; Lowenstein, C.J. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Molecular cell 2007, 26(5), 745–752. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Mao, Y.; Chen, W.; Liu, C.; Wu, H.; Zhang, J.; Wang, S.; Wang, C.; Lin, Y.; Lv, Y. Serum exosomal miR-34a as a potential biomarker for the diagnosis and prognostic of hepatocellular carcinoma. Journal of Cancer 2022, 13(5), 1410. [Google Scholar] [CrossRef]
- Chen, Z.; He, X. Application of third-generation sequencing in cancer research. Medical Review 2021, 1(2), 150–171. [Google Scholar] [CrossRef]
- Cho, H.J.; Eun, J.W.; Baek, G.O.; Seo, C.W.; Ahn, H.R.; Kim, S.S.; Cho, S.W.; Cheong, J.Y. Serum exosomal microRNA, miR-10b-5p, as a potential diagnostic biomarker for early-stage hepatocellular carcinoma. Journal of Clinical Medicine 2020, 9(1), 281. [Google Scholar] [CrossRef]
- Cimmino, A.; Calin, G.A.; Fabbri, M.; Iorio, M.V.; Ferracin, M.; Shimizu, M.; Wojcik, S.E.; Aqeilan, R.I.; Zupo, S.; Dono, M. miR-155 and miR-16 induce apoptosis by targeting BCL2. Proceedings of the National Academy of Sciences 2005, 102(39), 13944–13949. [Google Scholar] [CrossRef]
- Cole, K.A.; Attiyeh, E.F.; Mosse, Y.P.; Laquaglia, M.J.; Diskin, S.J.; Brodeur, G.M.; Maris, J.M. A functional screen identifies miR-34a as a candidate neuroblastoma tumor suppressor gene. Molecular cancer research 2008, 6(5), 735–742. [Google Scholar] [CrossRef] [PubMed]
- Diansyah, M.N.; Prayogo, A.A.; Sedana, M.P.; Savitri, M.; Romadhon, P.Z.; Amrita, P.N.A.; Wijaya, A.Y.; Hendrata, W.M.; Bintoro, U.Y. Early detection breast cancer: role of circulating plasma miRNA-21 expression as apotential screening biomarker. Turkish Journal of Medical Sciences 2021, 51(2), 562–569. [Google Scholar] [CrossRef]
- Dwedar, F.I.; Shams-Eldin, R.S.; Nayer Mohamed, S.; Mohammed, A.F.; Gomaa, S.H. Potential value of circulatory microRNA10b gene expression and its target E-cadherin as a prognostic and metastatic prediction marker for breast cancer. Journal of clinical laboratory analysis 2021, 35(8), e23887. [Google Scholar] [CrossRef] [PubMed]
- Esquela-Kerscher, A.; Slack, F.J. Oncomirs—microRNAs with a role in cancer. Nature Reviews Cancer 2006, 6(4), 259–269. [Google Scholar] [CrossRef] [PubMed]
- Frixa, T.; Donzelli, S.; Blandino, G. Oncogenic MicroRNAs: key players in malignant transformation. Cancers 2015, 7(4), 2466–2485. [Google Scholar] [CrossRef]
- Garzon, R.; Calin, G.A.; Croce, C.M. MicroRNAs in cancer. Annual review of medicine 2009, 60(1), 167–179. [Google Scholar] [CrossRef]
- Gregory, P.A.; Bert, A.G.; Paterson, E.L.; Barry, S.C.; Tsykin, A.; Farshid, G.; Vadas, M.A.; Khew-Goodall, Y.; Goodall, G.J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nature cell biology 2008, 10(5), 593–601. [Google Scholar] [CrossRef]
- Grimaldi, A.M.; Nuzzo, S.; Condorelli, G.; Salvatore, M.; Incoronato, M. Prognostic and clinicopathological significance of MiR-155 in breast cancer: a systematic review. International Journal of Molecular Sciences 2020, 21(16), 5834. [Google Scholar] [CrossRef]
- Hassanein, S.S.; Ibrahim, S.A.; Abdel-Mawgood, A.L. Cell behavior of non-small cell lung cancer is at EGFR and microRNAs hands. International Journal of Molecular Sciences 2021, 22(22), 12496. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Hannon, G.J. MicroRNAs: small RNAs with a big role in gene regulation. Nature reviews genetics 2004, 5(7), 522–531. [Google Scholar] [CrossRef]
- He, L.; He, X.; Lim, L.P.; De Stanchina, E.; Xuan, Z.; Liang, Y.; Xue, W.; Zender, L.; Magnus, J.; Ridzon, D. A microRNA component of the p53 tumour suppressor network. Nature 2007, 447(7148), 1130–1134. [Google Scholar] [CrossRef]
- He, L.; He, X.; Lowe, S.W.; Hannon, G.J. microRNAs join the p53 network—another piece in the tumour-suppression puzzle. Nature Reviews Cancer 2007, 7(11), 819–822. [Google Scholar] [CrossRef]
- He, L.; Thomson, J.M.; Hemann, M.T.; Hernando-Monge, E.; Mu, D.; Goodson, S.; Powers, S.; Cordon-Cardo, C.; Lowe, S.W.; Hannon, G.J. A microRNA polycistron as a potential human oncogene. Nature 2005, 435(7043), 828–833. [Google Scholar] [CrossRef] [PubMed]
- Heikkinen, L. Computational analysis of small non-coding RNAs in model systems Itä-Suomen yliopisto. 2014. [Google Scholar]
- Hu, C.-B.; Li, Q.-L.; Hu, J.-F.; Zhang, Q.; Xie, J.-P.; Deng, L. miR-124 inhibits growth and invasion of gastric cancer by targeting ROCK1. Asian Pacific Journal of Cancer Prevention 2014, 15(16), 6543–6546. [Google Scholar] [CrossRef] [PubMed]
- Ji, D.; Chen, Z.; Li, M.; Zhan, T.; Yao, Y.; Zhang, Z.; Xi, J.; Yan, L.; Gu, J. MicroRNA-181a promotes tumor growth and liver metastasis in colorectal cancer by targeting the tumor suppressor WIF-1. Molecular cancer 2014, 13(1), 86. [Google Scholar] [CrossRef]
- Kalluri, R.; LeBleu, V.S. The biology, function, and biomedical applications of exosomes. Science 2020, 367(6478), eaau6977. [Google Scholar] [CrossRef]
- Koumpis, E.; Georgoulis, V.; Papathanasiou, K.; Papoudou-Bai, A.; Kanavaros, P.; Kolettas, E.; Hatzimichael, E. The Role of microRNA-155 as a Biomarker in Diffuse Large B-Cell Lymphoma. Biomedicines 2024, 12(12), 2658. [Google Scholar] [CrossRef]
- Lawrie, C.H.; Gal, S.; Dunlop, H.M.; Pushkaran, B.; Liggins, A.P.; Pulford, K.; Banham, A.H.; Pezzella, F.; Boultwood, J.; Wainscoat, J.S. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. British journal of haematology 2008, 141(5), 672–675. [Google Scholar] [CrossRef]
- Le, M.T.; Teh, C.; Shyh-Chang, N.; Xie, H.; Zhou, B.; Korzh, V.; Lodish, H.F.; Lim, B. MicroRNA-125b is a novel negative regulator of p53. Genes & development 2009, 23(7), 862–876. [Google Scholar]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75(5), 843–854. [Google Scholar] [CrossRef]
- Liao, L.; Tang, Y.; Zhou, Y.; Meng, X.; Li, B.; Zhang, X. MicroRNA-126 (MiR-126): key roles in related diseases. Journal of physiology and biochemistry 2024, 80(2), 277–286. [Google Scholar] [CrossRef]
- Lou, W.; Liu, J.; Gao, Y.; Zhong, G.; Chen, D.; Shen, J.; Bao, C.; Xu, L.; Pan, J.; Cheng, J. MicroRNAs in cancer metastasis and angiogenesis. Oncotarget 2017, 8(70), 115787. [Google Scholar] [CrossRef]
- Luo, X.; Wen, W. MicroRNA in prostate cancer: from biogenesis to applicative potential. BMC urology 2024, 24(1), 244. [Google Scholar] [CrossRef]
- Ma, L. Role of miR-10b in breast cancer metastasis. Breast cancer research 2010, 12(5), 210. [Google Scholar] [CrossRef]
- Ma, L.; Teruya-Feldstein, J.; Weinberg, R.A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 2007, 449(7163), 682–688. [Google Scholar] [CrossRef] [PubMed]
- McDougall, A.R.; Hooper, S.B.; Zahra, V.A.; Sozo, F.; Lo, C.Y.; Cole, T.J.; Doran, T.; Wallace, M.J. The oncogene Trop2 regulates fetal lung cell proliferation. American Journal of Physiology-Lung Cellular and Molecular Physiology 2011, 301(4), L478–L489. [Google Scholar] [CrossRef]
- Meister, J.; Schmidt, M.H. miR-126 and miR-126: new players in cancer. The Scientific World Journal 2010, 10(1), 2090–2100. [Google Scholar] [CrossRef]
- Metanat, Y.; Sviridova, M.; Al-Nuaimi, B.N.; Janbazi, F.; Jalali, M.; Ghalamkarpour, N.; Khodabandehloo, E.; Ahmadi, E. The role of non-coding RNAs in the regulation of cell death pathways in melanoma. Discover Oncology 2025, 16(1), 1063. [Google Scholar] [CrossRef] [PubMed]
- Mogilyansky, E.; Rigoutsos, I. The miR-17/92 cluster: a comprehensive update on its genomics, genetics, functions and increasingly important and numerous roles in health and disease. Cell Death & Differentiation 2013, 20(12), 1603–1614. [Google Scholar] [CrossRef] [PubMed]
- Ng, E.K.; Li, R.; Shin, V.Y.; Jin, H.C.; Leung, C.P.; Ma, E.S.; Pang, R.; Chua, D.; Chu, K.-M.; Law, W. Circulating microRNAs as specific biomarkers for breast cancer detection. PloS one 2013, 8(1), e53141. [Google Scholar] [CrossRef]
- O'connell, R.M.; Rao, D.S.; Chaudhuri, A.A.; Baltimore, D. Physiological and pathological roles for microRNAs in the immune system. Nature Reviews Immunology 2010, 10(2), 111–122. [Google Scholar] [CrossRef]
- Otmani, K.; Rouas, R.; Lewalle, P. OncomiRs as noncoding RNAs having functions in cancer: Their role in immune suppression and clinical implications. Frontiers in immunology 2022, 13, 913951. [Google Scholar] [CrossRef]
- Pan, Y.; Li, K.; Tao, X.; Zhao, Y.; Chen, Q.; Li, N.; Liu, J.; Go, V.L.W.; Guo, J.; Gao, G. MicroRNA-34a alleviates gemcitabine resistance in pancreatic cancer by repression of cancer stem cell renewal. Pancreas 2021, 50(9), 1260–1266. [Google Scholar] [CrossRef]
- Parrella, P.; Barbano, R.; Pasculli, B.; Fontana, A.; Copetti, M.; Valori, V.M.; Poeta, M.L.; Perrone, G.; Righi, D.; Castelvetere, M. Evaluation of microRNA-10b prognostic significance in a prospective cohort of breast cancer patients. Molecular cancer 2014, 13(1), 142. [Google Scholar] [CrossRef]
- Sachdeva, M.; Zhu, S.; Wu, F.; Wu, H.; Walia, V.; Kumar, S.; Elble, R.; Watabe, K.; Mo, Y.-Y. p53 represses c-Myc through induction of the tumor suppressor miR-145. Proceedings of the National Academy of Sciences 2009, 106(9), 3207–3212. [Google Scholar] [CrossRef] [PubMed]
- Sasahira, T.; Kurihara, M.; Bhawal, U.; Ueda, N.; Shimomoto, T.; Yamamoto, K.; Kirita, T.; Kuniyasu, H. Downregulation of miR-126 induces angiogenesis and lymphangiogenesis by activation of VEGF-A in oral cancer. British journal of cancer 2012, 107(4), 700–706. [Google Scholar] [CrossRef]
- Schwarzenbach, H.; Nishida, N.; Calin, G.A.; Pantel, K. Clinical relevance of circulating cell-free microRNAs in cancer. Nature reviews Clinical oncology 2014, 11(3), 145–156. [Google Scholar] [CrossRef] [PubMed]
- Sheedy, P.; Medarova, Z. The fundamental role of miR-10b in metastatic cancer. American journal of cancer research 2018, 8(9), 1674. [Google Scholar]
- Vasudevan, S. Posttranscriptional upregulation by microRNAs. Wiley Interdisciplinary Reviews: RNA 2012, 3(3), 311–330. [Google Scholar] [CrossRef]
- Wang, W.; Li, X.; Liu, C.; Zhang, X.; Wu, Y.; Diao, M.; Tan, S.; Huang, S.; Cheng, Y.; You, T. MicroRNA-21 as a diagnostic and prognostic biomarker of lung cancer: a systematic review and meta-analysis. Bioscience Reports 2022, 42(5), BSR20211653. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-Y.; Li, L.; Ye, Z.-Y.; Zhao, Z.-S.; Yan, Z.-L. MicroRNA-10b promotes migration and invasion through Hoxd10 in human gastric cancer. World journal of surgical oncology 2015, 13(1), 259. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.; Gao, W.; Zhu, C.-J.; Liu, Y.-Q.; Mei, Z.; Cheng, T.; Shu, Y.-Q. Identification of plasma microRNA-21 as a biomarker for early detection and chemosensitivity of non–small cell lung cancer. Chinese journal of cancer 2011, 30(6), 407. [Google Scholar] [CrossRef]
- Wu, Y.; Hong, Q.; Lu, F.; Zhang, Z.; Li, J.; Nie, Z.; He, B. The diagnostic and prognostic value of miR-155 in cancers: an updated meta-analysis. Molecular Diagnosis & Therapy 2023, 27(3), 283–301. [Google Scholar] [CrossRef]
- Zhang, C.; Zhao, Y.; Wang, Q.; Qin, J.; Ye, B.; Xu, C.; Yu, G. Overexpression of miR-125a-5p inhibits hepatocyte proliferation through the STAT3 regulation in vivo and in vitro. International Journal of Molecular Sciences 2022, 23(15), 8661. [Google Scholar] [CrossRef]
- Zhang, H.; Zhou, Y.; Wen, D.; Wang, J. Noncoding RNAs: master regulator of fibroblast to myofibroblast transition in fibrosis. International Journal of Molecular Sciences 2023, 24(2), 1801. [Google Scholar] [CrossRef]
- Zhao, K.; Cheng, J.; Chen, B.; Liu, Q.; Xu, D.; Zhang, Y. Circulating microRNA-34 family low expression correlates with poor prognosis in patients with non-small cell lung cancer. Journal of thoracic disease 2017, 9(10), 3735. [Google Scholar] [CrossRef]
- Zhao, S.; Wu, Q.; Gao, F.; Zhang, C.; Yang, X. Serum microRNA-155 as a potential biomarker for breast cancer screening. Chinese Science Bulletin 2012, 57(26), 3466–3468. [Google Scholar] [CrossRef]
- Zhou, Z.; Lv, J.; Wang, J.; Yu, H.; Lu, H.; Yuan, B.; Han, J.; Zhou, R.; Zhang, X.; Yang, X. Role of microRNA-124 as a prognostic factor in multiple neoplasms: a meta-analysis. Disease markers 2019, 2019(1), 1654780. [Google Scholar] [CrossRef] [PubMed]
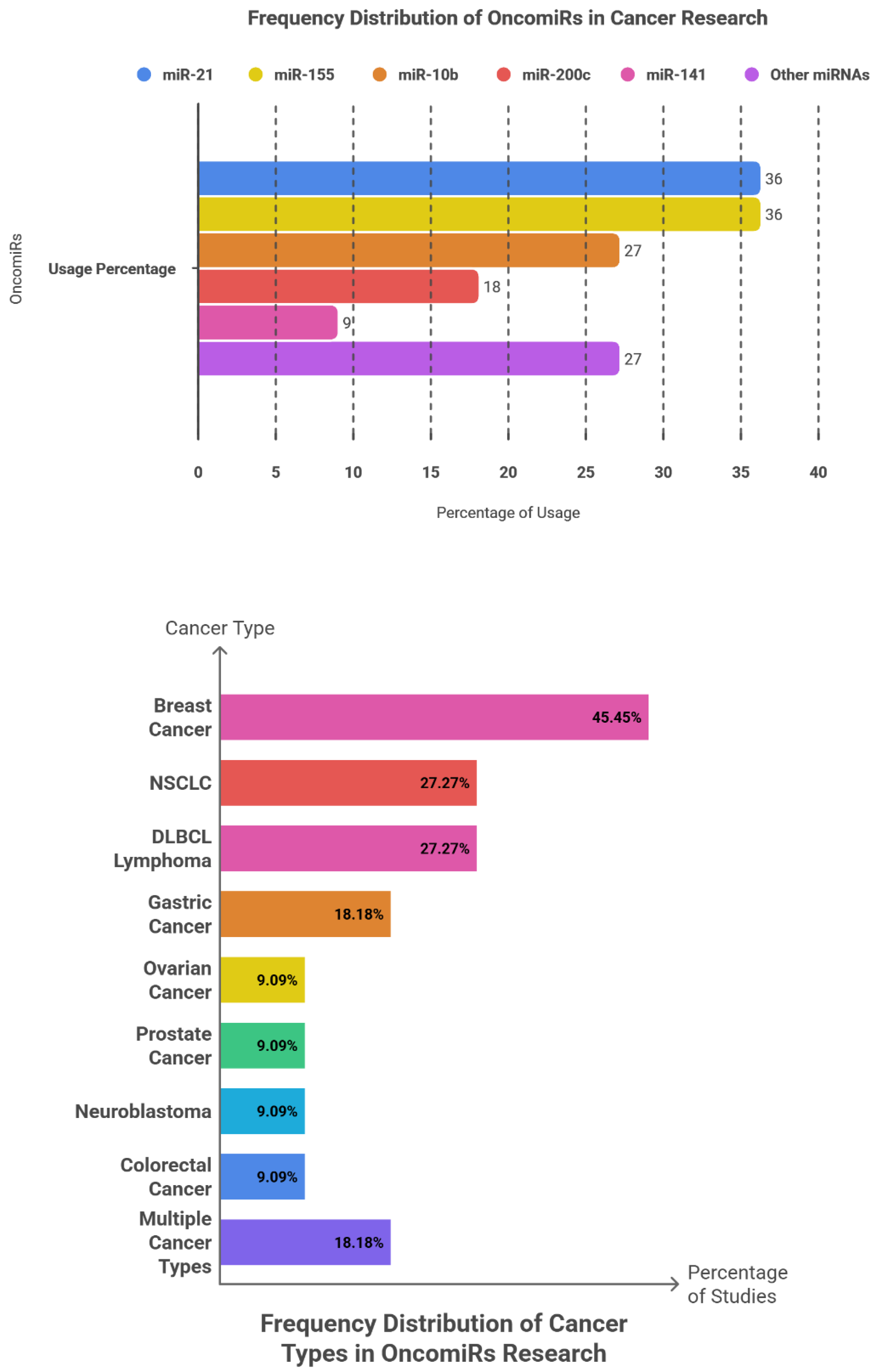
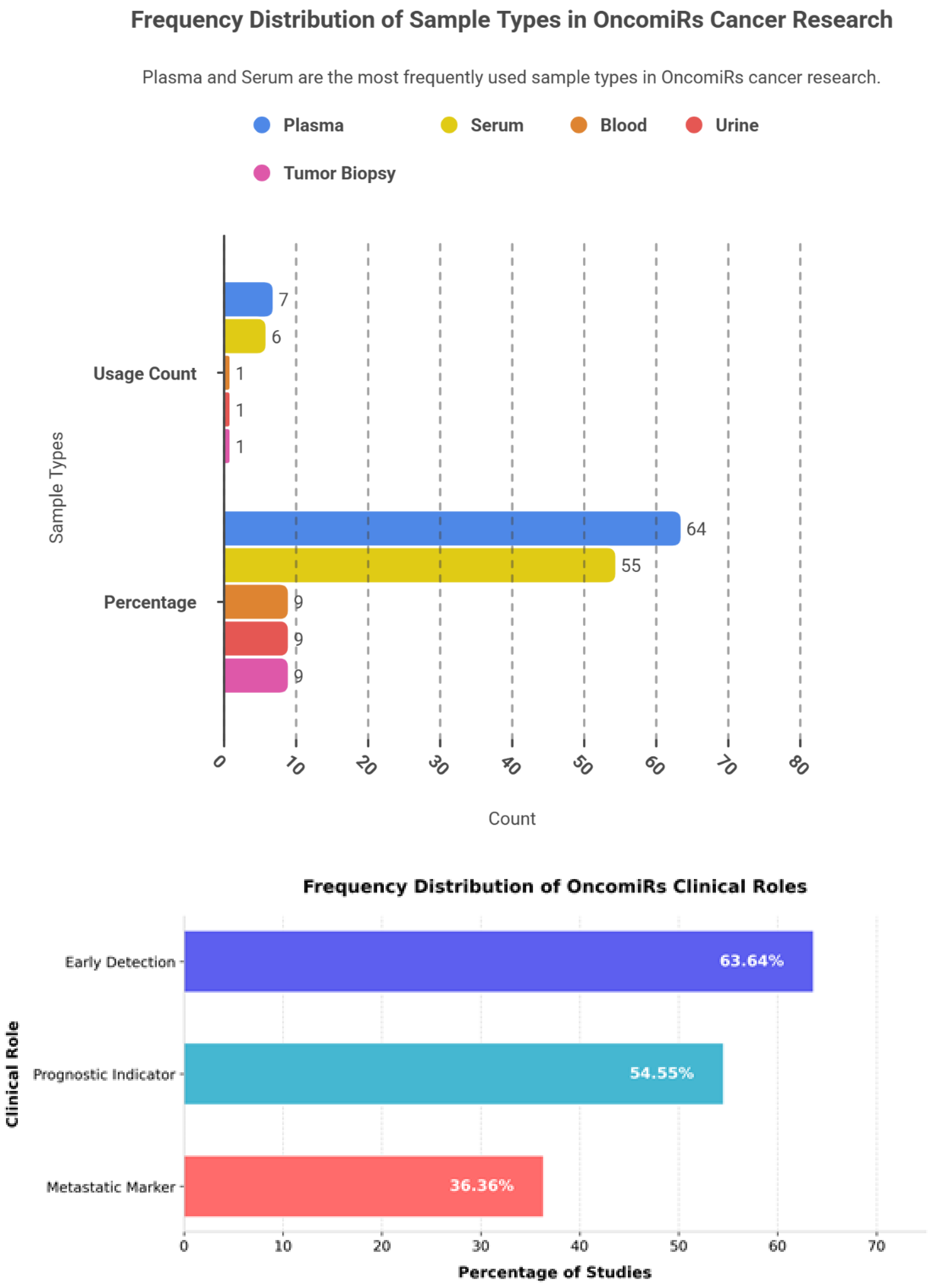
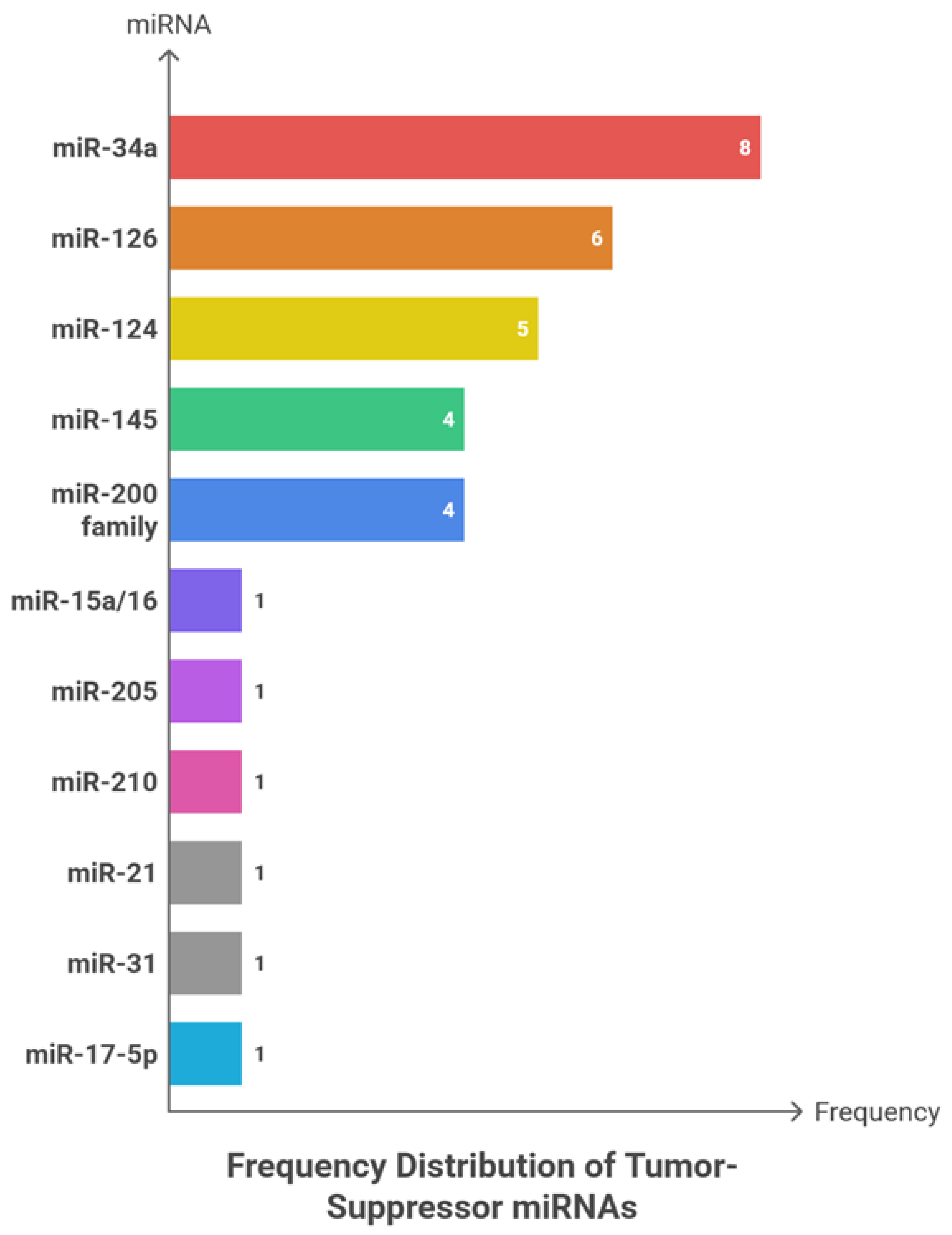
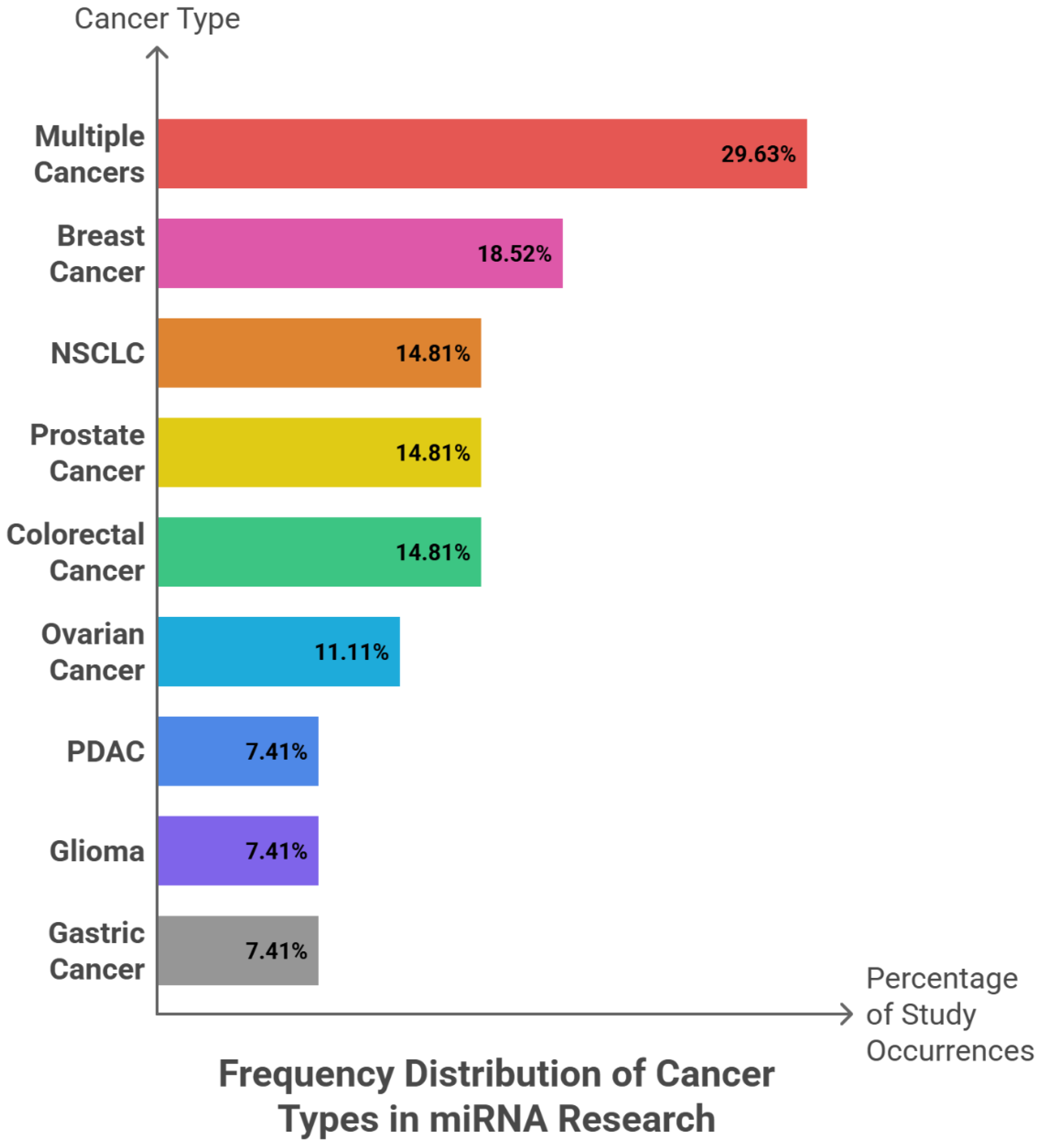
| miRNA | Clinical Role | Cancer Type | Sample Type | Findings and Key Results |
(Reference) |
|---|---|---|---|---|---|
| miR-21 |
Early Detection |
Breast Cancer |
Plasma |
Elevated miR-21 levels observed months before clinical diagnosis. |
Wang et al. (2020) |
| miR-21 |
Early Detection |
Non-Small Cell Lung Cancer (NSCLC) |
Serum |
Significant increase in serum miR-21 levels years before radiographic detection. | Zhang et al. (2022) |
| miR-21 | Early Detection |
Glioblastoma |
Serum |
Elevated serum miR-21 levels before clinical symptoms or MRI findings. | Liu et al. (2021) |
| miR-21 | Prognostic Indicator |
Lung Cancer |
Plasma |
High miR-21 levels associated with poor prognosis and reduced overall survival (OS). |
Wang et al. (2020) |
| miR-21 | Prognostic Indicator |
Gastric Cancer |
Plasma |
Increased miR-21 levels correlated with advanced disease stages and reduced survival. | Zhang et al. (2022) |
| miR155 | Early Detection | DLBCL Lymphoma |
Serum |
High miR-155 levels at diagnosis compared to healthy controls; levels decrease after successful treatment. |
Li et al. (2019) |
| miR155 | Early Detection |
Breast Cancer |
Serum | Elevated miR-155 levels across different stages, including stage I. |
Zhang et al. (2020) |
| miR155 | Prognostic Indicator |
DLBCL Lymphoma |
Serum | High miR-155 levels associated with poor prognosis. |
Li et al. (2019) |
| miR155 | Prognostic Indicator | Breast Cancer |
Serum |
Zhang et al. (2020) |
|
| miR10b | Early Detection | Breast, Colorectal, Pancreatic Cancer | Blood |
High miR-10b levels detectable before clinical metastasis. |
Ma et al. (2007); Sheedy & Medarova (2018) |
| miR10b | Prognostic Indicator |
Breast Cancer | Serum |
High miR-10b levels predictive of metastatic spread and reduced disease-free survival (DFS). |
Gómez et al. (2023) |
| miR10b | Metastatic Marker |
Breast, Pancreatic Cancer |
Plasma, Tumor Biopsy |
Elevated miR-10b levels associated with higher metastatic burden and poorer prognosis; role in invasion via HOXD10. |
Ma et al. (2007); Zhang et al. (2021); Liu et al. (2023) |
| miR-200 family (miR-200a, miR-200c) |
Early Detection |
Ovarian, Breast Cancer |
Serum |
Elevated miR-200a and miR-200c levels in early stages, correlated with tumor size and stage. |
Song et al. (2021) |
| miR-200 family |
Metastatic Marker |
Multiple Cancer Types |
Serum |
Downregulation of miR-200 associated with increased metastatic capacity via EMT inhibition (targeting ZEB1 and ZEB2). |
Bracken et al. (2010) |
| miR-141 |
Early Detection |
Prostate Cancer |
Urine |
High miR-141 levels in urine of patients compared to healthy controls; suitable for non-invasive screening. |
Liu et al. (2022) |
| miR-17-92 cluster |
Early Detection |
Neuroblastoma |
Plasma |
Detectable in early stages, correlated with tumor burden. |
Kim et al. (2019) |
| miR-9 |
Metastatic Marker |
Multiple Cancer Types |
- | Facilitates metastasis via downregulation of E-cadherin and promotion of EMT. |
Ma et al. (2010) |
| miR-181a |
Metastatic Marker |
Colorectal, Breast Cancer |
- | Increases metastatic potential via regulation of migration-related genes. |
Li et al. (2018) |
| miRNA | Clinical Role | Cancer Type | Sample Type | Findings and Key Results | Reference |
|---|---|---|---|---|---|
| miR-34a | Early detection; Prognosis; Therapy monitoring | NSCLC | Plasma/Serum | Reduced levels correlate with early-stage disease; inverse with tumor size, nodal status, and metastasis. | Li et al., 2020 |
| miR-34a | Early detection; Prognosis | PDAC | Serum | Low levels indicate early diagnosis value; associated with poor prognosis and chemoresistance. | Zhang et al., 2023 |
| miR-34a | Early detection | Colorectal cancer | Serum | Levels decline at adenoma stage; downregulation increases MET and ZEB1, promoting metastasis. | — (from text) |
| miR-34a | Prognosis | Breast, Prostate | Serum | Lower levels predict aggressive behavior and worse survival via EMT and cell-cycle regulation. | Chen et al., 2022; Zhao et al., 2017 |
| miR-34a | Mechanism; Early detection | Multiple cancers | Circulating/ctDNA; single-cell | p53 effector; targets CDK6, CCND1, BCL2, ZEB1/2, SNAIL, MET; downregulation linked to Wnt/β-catenin, PI3K/Akt activation; early monitoring feasible. | Cole et al., 2008; Zhang et al., 2025 |
| miR-34a | Prognosis; Therapy monitoring | NSCLC, PDAC, Gastric, Colorectal, Glioma | Serum/Plasma | Lower levels link to advanced stage, metastasis, resistance; restoration may enhance chemo sensitivity; serial decline signals progression/relapse. | Hassanein et al., 2021; Zhao et al., 2017; Zhang et al., 2025; Chen et al., 2025; Pan et al., 2021 |
| miR-126 | Early detection; Risk assessment | Colorectal, Lung, Breast, Gastric, Ovarian | Circulating (plasma/serum) | Decreased levels correlate with early tumorigenic changes and poorer prognosis; anti-angiogenic by targeting VEGF-A, PI3K, SPRED1. | Meister & Schmidt, 2010; Liu et al., 2021 |
| miR-126 | Mechanism; Monitoring | Multiple | Circulating; single-cell/ctDNA | Suppresses angiogenesis and invasion; links to advanced stage and metastasis; sensitive detection via single-cell/ctDNA profiling. | Sasahira et al., 2012 |
| miR-126 | Prognosis; Mechanism | Multiple | Circulating | Targets VEGF-A, PI3K/AKT (p85β), p-STAT3; additional targets SDC4, Crk/CrkL, IRS-1/2; loss promotes angiogenesis, invasion, survival. | Lou et al., 2017; Liu et al., 2023 |
| miR-126 | Prognosis; Metastasis risk | mCRC | Plasma | Lower levels associate with metastasis and worse survival; longitudinal reduction predicts higher risk. | Liao et al., 2024 |
| miR-126 | Prognosis; Chemoresistance | NSCLC | Serum/Plasma | Downregulated in metastatic lesions; low levels predict poor prognosis and platinum resistance. | — (from text) |
| miR-126 | Prognosis; Therapeutic potential | TNBC; Gastric; Pancreatic; Prostate | Circulating | Decrease linked to invasiveness and metastasis; restoration reduces metastasis; ties to EMT and PI3K/AKT activity. | Wang et al., 2024; Li et al., 2023 |
| miR-124 | Early detection | Glioma, HCC | Serum/Plasma | Decreased levels in early-stage disease; precede clinical detection. | Hu et al., 2014 |
| miR-124 | Prognosis | Glioma | Serum | Lower levels correlate with higher grade, invasion, and reduced survival. | Li et al., 2022 |
| miR-124 | Prognosis | HCC | Serum | Low circulating levels predict poorer overall survival and higher recurrence; complements AFP. | Zhang et al., 2025 |
| miR-124 | Prognosis; Mechanism | Multiple (incl. Cervical) | Serum | Targets SOX9, STAT3, AKT components; reduced levels link to worse outcomes; qRT-PCR supports clinical utility. | Zhou et al., 2019; Chen et al., 2024 |
| miR-145 | Prognosis; Risk stratification | Breast, Prostate | Serum | Lower levels associate with high grade, nodal involvement, and therapy resistance; predicts early relapse and EGFR therapy resistance. | Wang et al., 2021; Wu et al., 2022 |
| miR-145 | Prognosis | Multiple (incl. Gynecologic) | Plasma/Serum | Decreased levels link to advanced stage, metastasis, chemoresistance, worse survival. | Zhang et al., 2023; Chen et al., 2024 |
| miR-145 | Mechanism | Multiple | - | Targets Wnt/β-catenin axis (RGS17, KEAP1), MUC1, c-MYC, FSCN1; suppresses EMT and invasion. | — (from text) |
| miR-15a/16-1 cluster | Prognosis; EMT suppression | Leukemia, Breast, Colorectal, Gastric | Serum/Tissue | Downregulation associates with increased EMT markers, invasion, poor prognosis; restoration inhibits growth/metastasis; hallmark in CLL. | Zhang et al., 2024 |
| miR-200 family (a/b/c) | Prognosis; Metastasis classifier | Breast, Ovarian, Lung, Pancreatic, Ovarian | Serum/Plasma/Tissue | Reduced levels associate with EMT, metastasis, chemoresistance; distinguish localized vs metastatic disease; lower in metastatic cases. | Korpal et al., 2018; Song et al., 2021; Zhan et al., 2022; Wang et al., 2023; Lee et al., 2022 |
| miR-205 | Metastasis marker; EMT inhibition | Breast, Prostate | Serum | Decreased levels observed in metastatic disease; targets ZEB1/2. | Li et al., 2024 |
| miR-30a | Metastasis marker; EMT suppression | Gastric; Lung adenocarcinoma | Serum/Plasma | Downregulation correlates with lymph node metastasis (gastric) and brain metastasis (lung adenocarcinoma). | Zhou et al., 2023; — (from text) |
| miR-210 | Metastasis; Hypoxia-associated | Multiple | Plasma | Linked to hypoxia and metastatic spread. | Chen et al., 2025 |
| miR-21 | Metastasis dynamics | Multiple | Plasma | Lower plasma levels reported in metastatic cases, reflecting complex regulation. | Liu et al., 2025 |
| miR-373 | Metastasis; EMT | Ovarian, Breast, Lung | Serum/Plasma | Emerging indicator of EMT and metastatic colonization. | Wang et al., 2024 |
| miR-17-5p | Metastasis; EMT | Ovarian, Breast, Lung | Serum/Plasma | Associated with EMT and metastatic colonization across cancers. | Zhao et al., 2025 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




