Submitted:
04 January 2026
Posted:
06 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Construction of β-Globin Minigene Reporters
2.3. U1 snRNA Expression Construct
2.4. Extracellular Vesicle Isolation
2.5. Nanoparticle Tracking Analysis (NTA)
2.6. Western Blot Analysis of EV Markers
2.7. RNA Extraction from EVs
2.8. β-.Globin Minigene Transfection and EV Treatment
2.9. RT–PCR Splicing Analysis
2.10. Quantitative RT–PCR
2.11. Statistical Analysis
3. Results
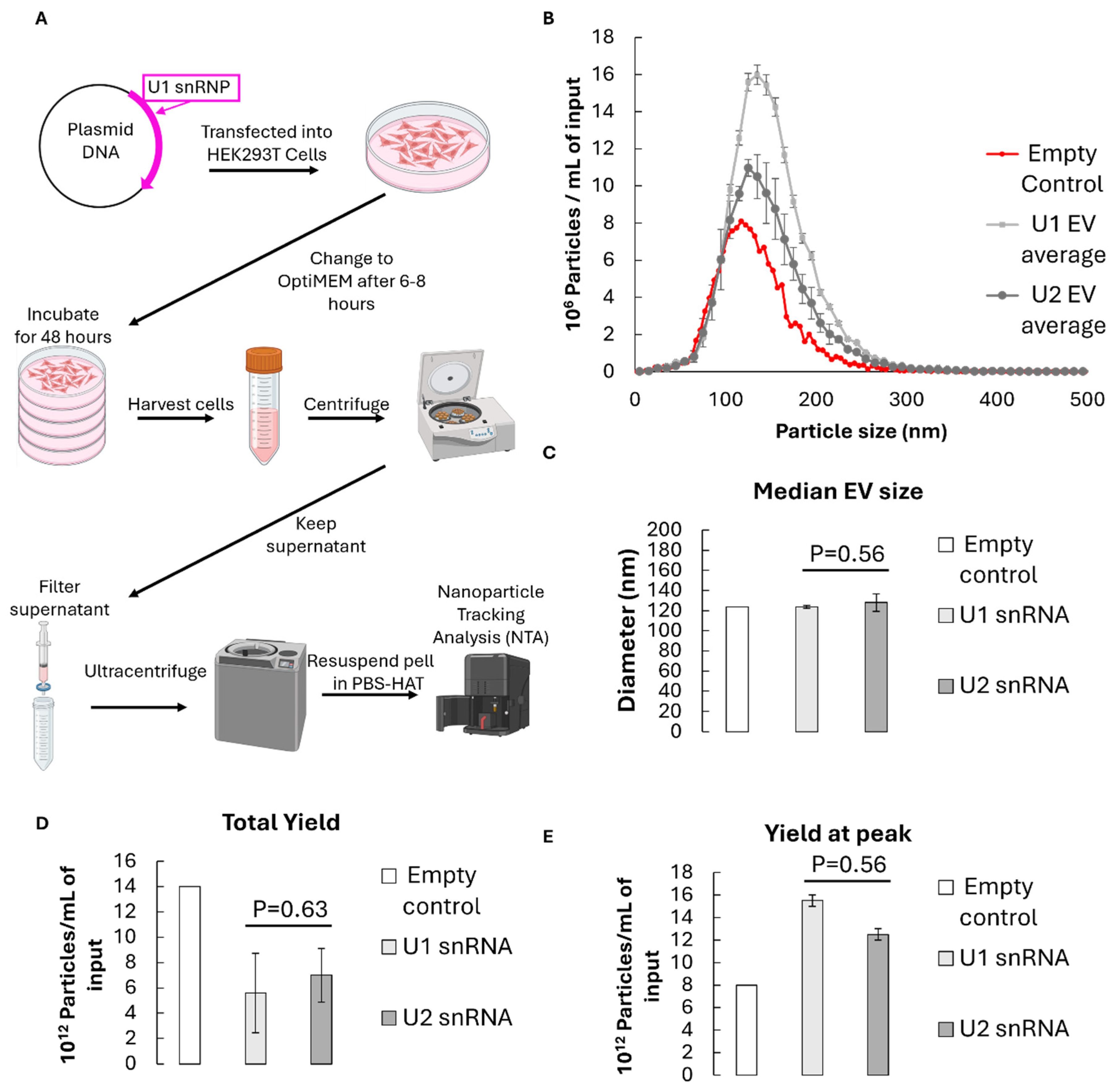
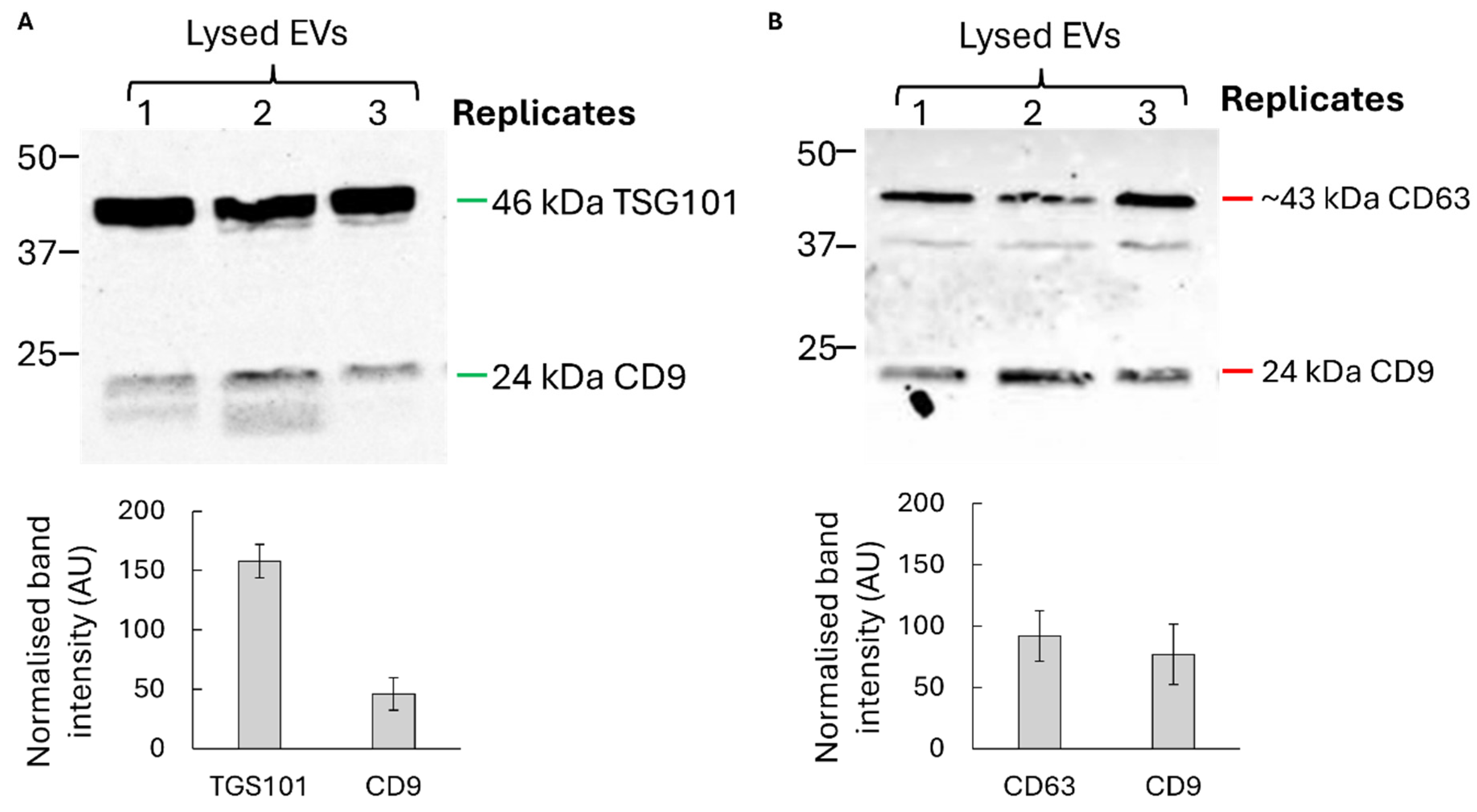
3.1. Characterisation of U1-Enriched Extracellular Vesicles
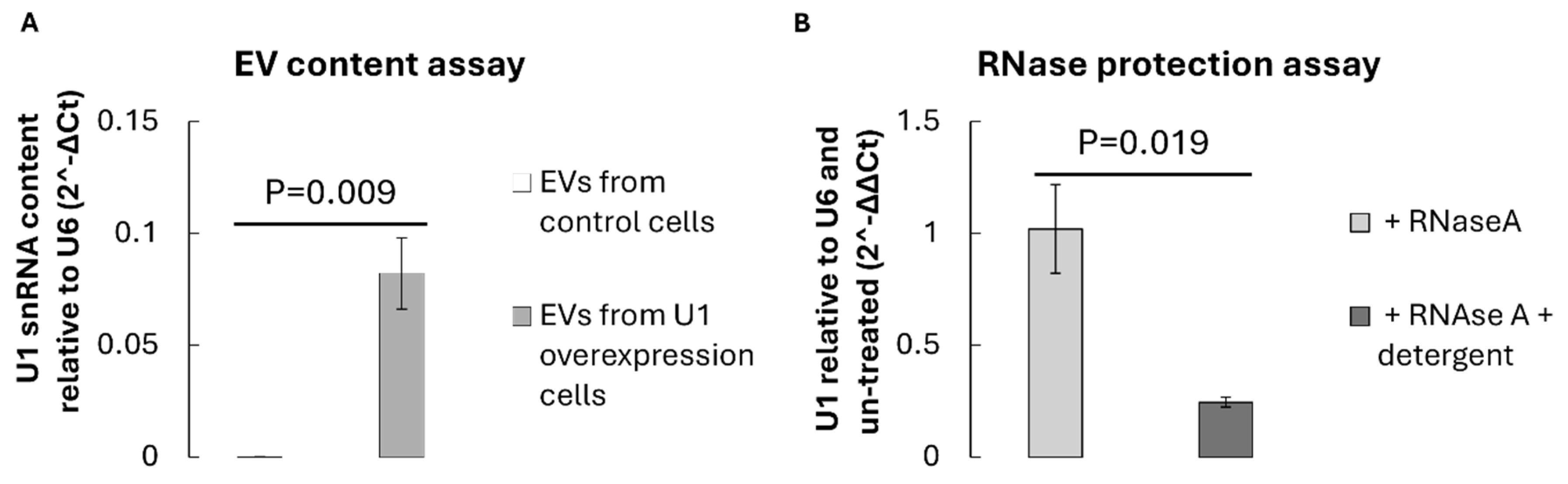
3.2. Detection of U1 snRNA Within Isolated EVs
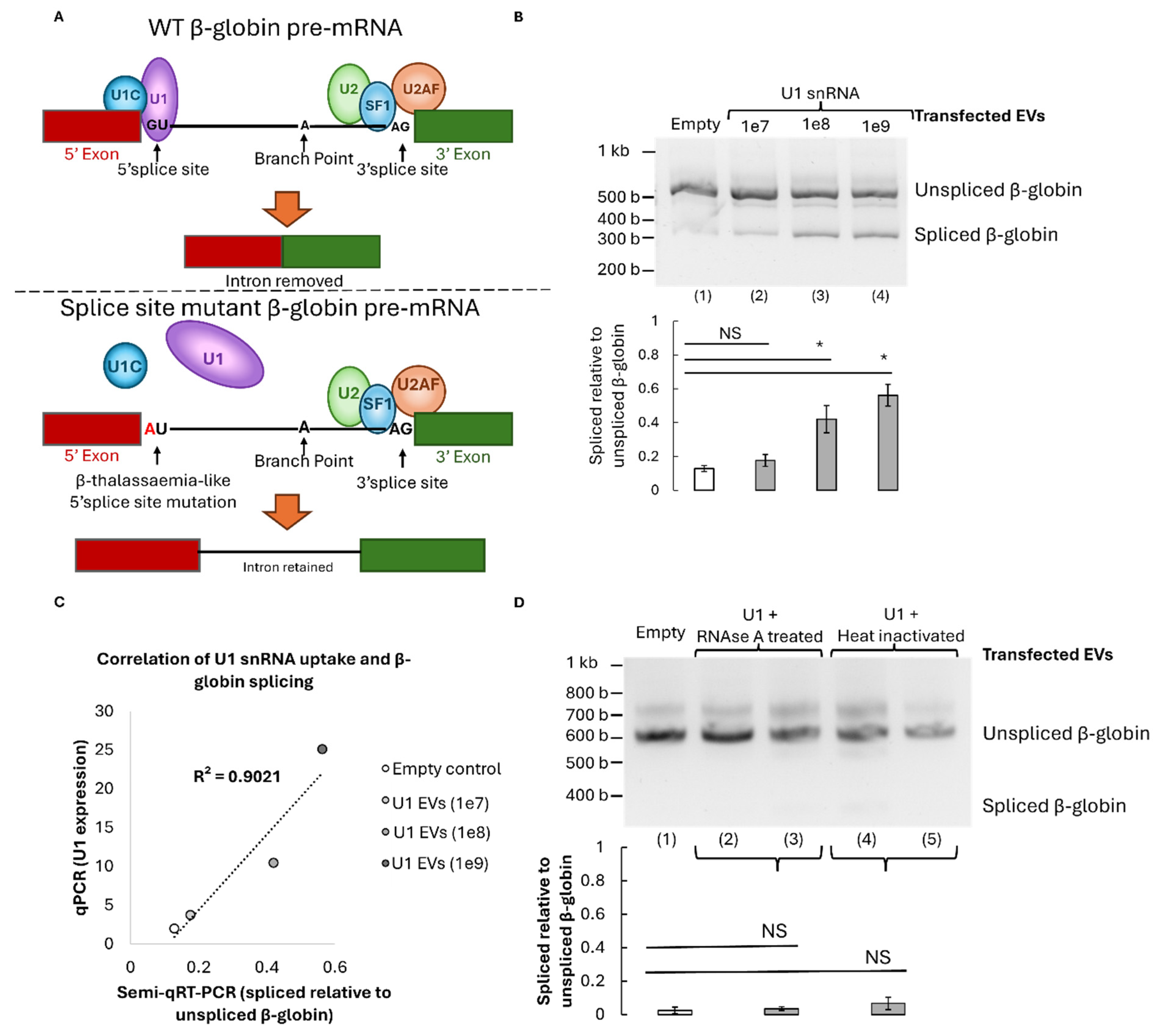
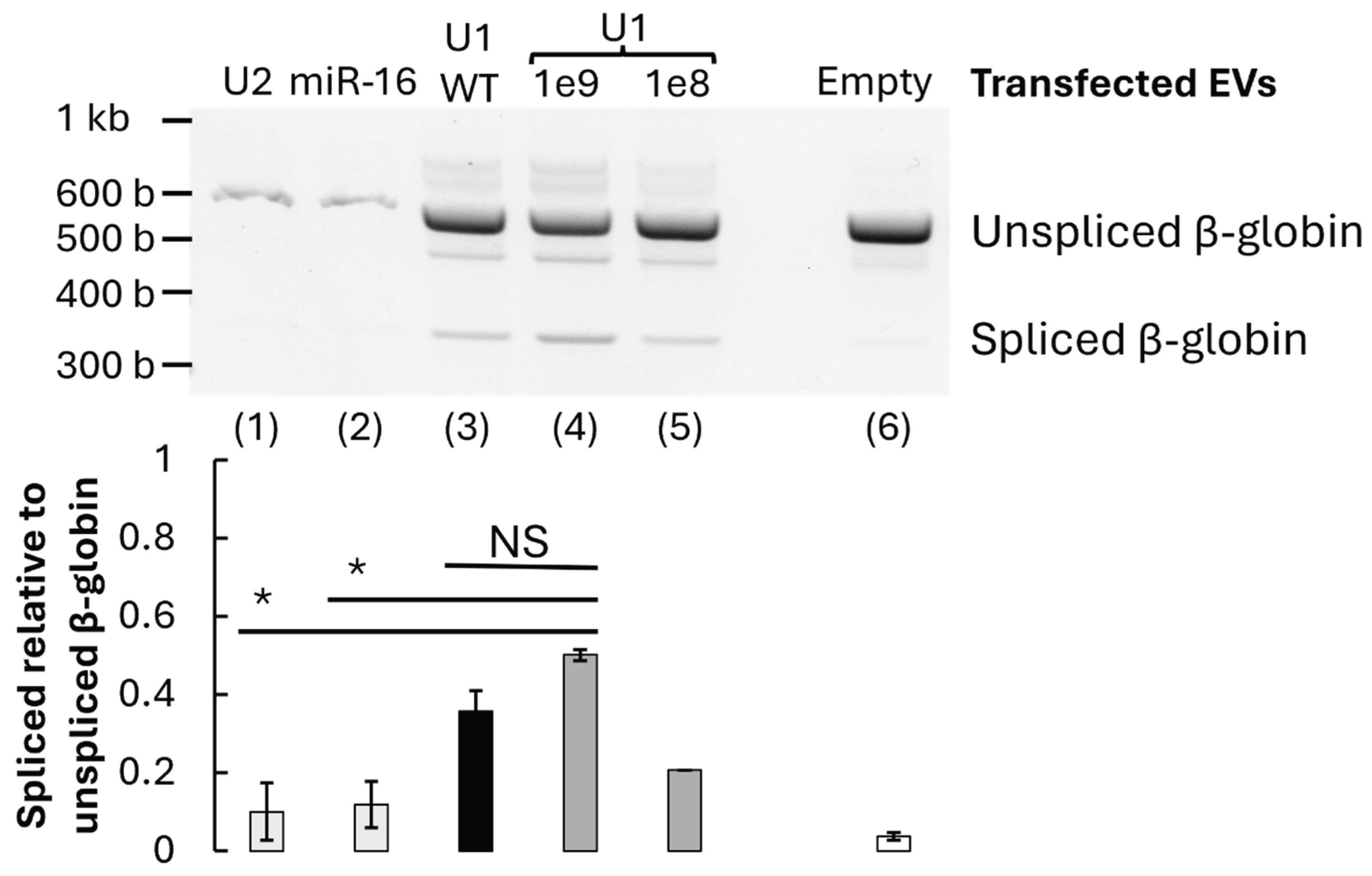
3.3. Restoration of β-Globin Pre-mRNA Splicing by U1-Containing EVs
3.4. Functional Validation and Specificity of U1-Mediated Correction
4. Discussion
4.1. EVs as Vehicles for Functional RNA Transfer
4.2. Considerations for U1-Mediated Splicing Rescue
4.3. Comparison with Existing RNA Therapeutic Approaches
5. Conclusion
6. Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Alqurashi, H.; Alsharief, M.; Perciato, M.L.; et al. Message in a bubble: the translational potential of extracellular vesicles. J. Physiol. 2023, 601, 4895–4905. [Google Scholar] [CrossRef]
- Alvarez-Erviti, L.; Seow, Y.; Yin, H.; et al. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 2011, 29, 341–345. [Google Scholar] [CrossRef]
- Andreu, Z.; Yanez-Mo, M. Tetraspanins in extracellular vesicle formation and function. Front. Immunol. 2014, 5, 442. [Google Scholar] [CrossRef]
- Balestra, D.; Faella, A.; Margaritis, P.; et al. An engineered U1 small nuclear RNA rescues splicing-defective coagulation F7 gene expression in mice. J. Thromb. Haemost. 2014, 12, 177–185. [Google Scholar] [CrossRef] [PubMed]
- Bradley, R.K.; Anczuków, O. RNA splicing dysregulation and the hallmarks of cancer. Nat. Rev. Cancer 2023, 23, 135–155. [Google Scholar] [CrossRef]
- Breuel, S.; Vorm, M.; Bräuer, A.U.; et al. Combining engineered U1 snRNA and antisense oligonucleotides to improve the treatment of a BBS1 splice site mutation. Mol. Ther. Nucleic Acids 2019, 18, 123–130. [Google Scholar] [CrossRef]
- Cebulla, G.; Hai, L.; Warnken, U.; et al. Long-term CSF responses in adult patients with spinal muscular atrophy type 2 or 3 on treatment with nusinersen. J. Neurol. 2025, 272, 270. [Google Scholar] [CrossRef] [PubMed]
- Cecchin, R.; Troyer, Z.; Witwer, K.; et al. Extracellular vesicles: the next generation in gene therapy delivery. Mol. Ther. 2023, 31, 1225–1230. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Heendeniya, S.N.; Le, B.T.; et al. Splice-modulating antisense oligonucleotides as therapeutics for inherited metabolic diseases. BioDrugs 2024, 38, 177–203. [Google Scholar] [CrossRef]
- Deutsch, H.M.; Song, Y.; Li, D. Spliceosome complex and neurodevelopmental disorders. Curr. Opin. Genet. Dev. 2025, 93, 102358. [Google Scholar] [CrossRef]
- Di Ianni, E.; Obuchi, W.; Breyne, K.; et al. Extracellular vesicles for the delivery of gene therapy. Nat. Rev. Bioeng. 2025, 3, 360–373. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Kantor, B. Lentiviral vectors for delivery of gene-editing systems based on CRISPR/Cas: current state and perspectives. Viruses 2021, 13, 1288. [Google Scholar] [CrossRef] [PubMed]
- El Andaloussi, S.; Mäger, I.; Breakefield, X.O.; et al. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef]
- Fernandez Alanis, E.; Pinotti, M.; Dal Mas, A.; et al. An exon-specific U1 small nuclear RNA (snRNA) strategy to correct splicing defects. Hum. Mol. Genet. 2012, 21, 2389–2398. [Google Scholar] [CrossRef]
- Finkel, R.S.; Mercuri, E.; Darras, B.T.; et al. Nusinersen versus sham control in infantile-onset spinal muscular atrophy. N. Engl. J. Med. 2017, 377, 1723–1732. [Google Scholar] [CrossRef]
- Furlong, R. Refining the splice region. Nat. Rev. Genet. 2018, 19, 470–471. [Google Scholar] [CrossRef]
- Gagliardi, M.; Ashizawa, A.T. The challenges and strategies of antisense oligonucleotide drug delivery. Biomedicines 2021, 9, 433. [Google Scholar] [CrossRef]
- Geng, G.; Xu, Y.; Hu, Z.; et al. Viral and non-viral vectors in gene therapy: current state and clinical perspectives. EBioMedicine 2025, 118, 105834. [Google Scholar] [CrossRef]
- Gonçalves, M.; Santos, J.I.; Coutinho, M.F.; et al. Development of engineered-U1 snRNA therapies: current status. Int. J. Mol. Sci. 2023, 24, 14617. [Google Scholar] [CrossRef]
- Goo, J.; Lee, Y.; Lee, J.; et al. Extracellular vesicles in therapeutics: a comprehensive review on applications, challenges, and clinical progress. Pharmaceutics 2024, 16, 311. [Google Scholar] [CrossRef] [PubMed]
- Guo, A.; Cao, Q.; Fang, H.; et al. Recent advances and challenges of injectable hydrogels in drug delivery. J. Control. Release 2025, 385, 114021. [Google Scholar] [CrossRef]
- Hammond, S.M.; Aartsma-Rus, A.; Alves, S.; et al. Delivery of oligonucleotide-based therapeutics: challenges and opportunities. EMBO Mol. Med. 2021, 13, e13243. [Google Scholar] [CrossRef]
- Hatch, S.T.; Smargon, A.A.; Yeo, G.W. Engineered U1 snRNAs to modulate alternatively spliced exons. Methods 2022, 205, 140–148. [Google Scholar] [CrossRef]
- Havens, M.A.; Hastings, M.L. Splice-switching antisense oligonucleotides as therapeutic drugs. Nucleic Acids Res. 2016, 44, 6549–6563. [Google Scholar] [CrossRef]
- Joshi, B.S.; de Beer, M.A.; Giepmans, B.N.G.; et al. Endocytosis of extracellular vesicles and release of their cargo from endosomes. ACS Nano 2020, 14, 4444–4455. [Google Scholar] [CrossRef]
- Kalluri, R.; LeBleu, V.S. The biology, function, and biomedical applications of exosomes. Science 2020, 367, eaau6977. [Google Scholar] [CrossRef]
- Kojima, R.; Bojar, D.; Rizzi, G.; et al. Designer exosomes produced by implanted cells intracerebrally deliver therapeutic cargo for Parkinson’s disease treatment. Nat. Commun. 2018, 9, 1305. [Google Scholar] [CrossRef]
- Kondo, Y.; Oubridge, C.; van Roon, A.-M.M.; et al. Crystal structure of human U1 snRNP reveals the mechanism of 5′ splice site recognition. eLife 2015, 4, e04986. [Google Scholar] [CrossRef]
- Krawczak, M.; Reiss, J.; Cooper, D.N. The mutational spectrum of single base-pair substitutions in mRNA splice junctions of human genes: causes and consequences. Hum. Genet. 1992, 90, 55–74. [Google Scholar] [CrossRef]
- Krawczak, M.; Thomas, N.S.T.; Hundrieser, B.; et al. Single base-pair substitutions in exon-intron junctions of human genes: nature, distribution, and consequences for mRNA splicing. Hum. Mutat. 2007, 28, 150–158. [Google Scholar] [CrossRef]
- Lauffer, M.C.; van Roon-Mom, W.; Aartsma-Rus, A. Possibilities and limitations of antisense oligonucleotide therapies for the treatment of monogenic disorders. Commun. Med. 2024, 4, 6. [Google Scholar] [CrossRef]
- Lerner, M.R.; Boyle, J.A.; Mount, S.M.; et al. Are snRNPs involved in splicing? Nature 1980, 283, 220–224. [Google Scholar] [CrossRef]
- Liang, X.; Gupta, D.; Xie, J.; et al. Engineering of extracellular vesicles for efficient intracellular delivery of multimodal therapeutics including genome editors. Nat. Commun. 2025, 16, 4028. [Google Scholar] [CrossRef]
- Liu, B.; Chen, K. Advances in hydrogel-based drug delivery systems. Gels 2024, 10, 262. [Google Scholar] [CrossRef]
- Lu, D.; Gong, X.; Guo, X.; et al. Gene editing of the endogenous cryptic 3′ splice site corrects the RNA splicing defect in the β654-thalassemia mouse model. Hum. Gene Ther. 2024, 35, 825–837. [Google Scholar] [CrossRef]
- Lv, X.; Sun, X.; Gao, Y.; et al. Targeting RNA splicing modulation: new perspectives for anticancer strategy? J. Exp. Clin. Cancer Res. 2025, 44, 32. [Google Scholar] [CrossRef]
- Matera, A.G.; Wang, Z. A day in the life of the spliceosome. Nat. Rev. Mol. Cell Biol. 2014, 15, 108–121. [Google Scholar] [CrossRef]
- Milone, M.C.; O’Doherty, U. Clinical use of lentiviral vectors. Leukemia 2018, 32, 1529–1541. [Google Scholar] [CrossRef]
- Mulcahy, L.A.; Pink, R.C.; Carter, D.R.F. Routes and mechanisms of extracellular vesicle uptake. J. Extracell. Vesicles 2014, 3, 1. [Google Scholar] [CrossRef]
- Nava, C.; Cogne, B.; Santini, A.; et al. Dominant variants in major spliceosome U4 and U5 small nuclear RNA genes cause neurodevelopmental disorders through splicing disruption. Nat. Genet. 2025, 57, 1374–1388. [Google Scholar] [CrossRef]
- Nechay, M.; Kleiner, R.E. High-throughput approaches to profile RNA-protein interactions. Curr. Opin. Chem. Biol. 2020, 54, 37–44. [Google Scholar] [CrossRef]
- Nik, S.; Bowman, T.V. Splicing and neurodegeneration: insights and mechanisms. WIREs RNA 2019, 10, e1532. [Google Scholar] [CrossRef]
- Ottesen, E.W.; Singh, N.N.; Seo, J.; et al. U1 snRNA interactions with deep intronic sequences regulate splicing of multiple exons of spinal muscular atrophy genes. Front. Neurosci. 2024, 18, 1412893. [Google Scholar] [CrossRef]
- Pan, Q.; Shai, O.; Lee, L.; et al. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 2008, 40, 1413–1415. [Google Scholar] [CrossRef]
- Pasteris, M.; Cakir, S.; Bellizzi, A.; et al. Alternative splicing in Alzheimer’s disease: mechanisms, therapeutic implications, and 3D modeling approaches. J. Alzheimers Dis. 2025, 107, 5–14. [Google Scholar] [CrossRef]
- Peruzzo, P.; Bergamin, N.; Bon, M.; et al. Rescue of common and rare exon 2 skipping variants of the GAA gene using modified U1 snRNA. Mol. Med. 2025, 31, 45. [Google Scholar] [CrossRef]
- Pokorná, P.; Aupič, J.; Fica, S.; et al. Decoding spliceosome dynamics through computation and experiment. Chem. Rev. 2025, 125, 9807–9833. [Google Scholar] [CrossRef]
- Qiu, J.; Wu, L.; Qu, R.; et al. History of development of the life-saving drug “Nusinersen” in spinal muscular atrophy. Front. Cell. Neurosci. 2022, 16, 942976. [Google Scholar] [CrossRef]
- Raguraman, P.; Balachandran, A.A.; Chen, S.; et al. Antisense oligonucleotide-mediated splice switching: potential therapeutic approach for cancer mitigation. Cancers 2021, 13, 5555. [Google Scholar] [CrossRef]
- Sabrina Haque, U.; Kohut, M.; Yokota, T. Comprehensive review of adverse reactions and toxicology in ASO-based therapies for Duchenne muscular dystrophy: from FDA-approved drugs to peptide-conjugated ASO. Curr. Res. Toxicol. 2024, 7, 100182. [Google Scholar] [CrossRef]
- Sakharkar, M.K.; Chow, V.T.K.; Kangueane, P. Distributions of exons and introns in the human genome. In Silico Biol. 2004, 4, 387–393. [Google Scholar] [CrossRef]
- Santangelo, L.; Giurato, G.; Cicchini, C.; et al. The RNA-binding protein SYNCRIP is a component of the hepatocyte exosomal machinery controlling microRNA sorting. Cell Rep. 2016, 17, 799–808. [Google Scholar] [CrossRef]
- Schmid, F.; Glaus, E.; Barthelmes, D.; et al. U1 snRNA-mediated gene therapeutic correction of splice defects caused by an exceptionally mild BBS mutation. Hum. Mutat. 2011, 32, 815–824. [Google Scholar] [CrossRef]
- Scotti, M.M.; Swanson, M.S. RNA mis-splicing in disease. Nat. Rev. Genet. 2016, 17, 19–32. [Google Scholar] [CrossRef]
- Soares, E.S.; Leal, C.B.Q.S.; Sinatti, V.V.C.; et al. Role of the U1 snRNP complex in human health and disease. WIREs RNA 2025, 16, e70026. [Google Scholar] [CrossRef]
- Sönmezler, E.; Stuani, C.; Hız Kurul, S.; et al. Characterization and engineered U1 snRNA rescue of splicing variants in a Turkish neurodevelopmental disease cohort. Hum. Mutat. 2024, 2024, 1–17. [Google Scholar] [CrossRef]
- Théry, C.; Witwer, K.W.; Aikawa, E.; et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef]
- Treisman, R.; Orkin, S.H.; Maniatis, T. Specific transcription and RNA splicing defects in five cloned β-thalassaemia genes. Nature 1983, 302, 591–596. [Google Scholar] [CrossRef]
- Urbanski, L.M.; Leclair, N.; Anczuków, O. Alternative-splicing defects in cancer: splicing regulators and their downstream targets, guiding the way to novel cancer therapeutics. WIREs RNA 2018, 9, e1476. [Google Scholar] [CrossRef]
- Valadi, H.; Ekström, K.; Bossios, A.; et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef]
- Van Delen, M.; Derdelinckx, J.; Wouters, K.; et al. A systematic review and meta-analysis of clinical trials assessing safety and efficacy of human extracellular vesicle-based therapy. J. Extracell. Vesicles 2024, 13, e12458. [Google Scholar] [CrossRef]
- Villarroya-Beltri, C.; Gutiérrez-Vázquez, C.; Sánchez-Cabo, F.; et al. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat. Commun. 2013, 4, 2980. [Google Scholar] [CrossRef]
- Volodina, O.; Smirnikhina, S. The future of gene therapy: a review of in vivo and ex vivo delivery methods for genome editing-based therapies. Mol. Biotechnol. 2025, 67, 425–437. [Google Scholar] [CrossRef]
- Wai, H.A.; Svobodova, E.; Herrera, N.R.; et al. Tailored antisense oligonucleotides designed to correct aberrant splicing reveal actionable groups of mutations for rare genetic disorders. Exp. Mol. Med. 2024, 56, 1816–1825. [Google Scholar] [CrossRef]
- Wijnant, K.N.; Nadif Kasri, N.; Vissers, L.E.L.M. Systematic analysis of genetic and phenotypic characteristics reveals antisense oligonucleotide therapy potential for one-third of neurodevelopmental disorders. Genome Med. 2025, 17, 59. [Google Scholar] [CrossRef]
- Yim, N.; Ryu, S.-W.; Choi, K.; et al. Exosome engineering for efficient intracellular delivery of soluble proteins using optically reversible protein–protein interaction module. Nat. Commun. 2016, 7, 12277. [Google Scholar] [CrossRef]
- Zhang, X.; Yan, C.; Hang, J.; et al. An atomic structure of the human spliceosome. Cell 2017, 169, 918–929. [Google Scholar] [CrossRef]
- Zhuang, Y.; Weiner, A.M. A compensatory base change in U1 snRNA suppresses a 5′ splice site mutation. Cell 1986, 46, 827–835. [Google Scholar] [CrossRef]
- Lieberman, H.B.; Bernstock, J.D.; Broustas, C.G.; et al. The role of RAD9 in tumorigenesis. J. Mol. Cell Biol. 2011, 3, 39–43. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



