Submitted:
31 December 2025
Posted:
02 January 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Materials and Methods
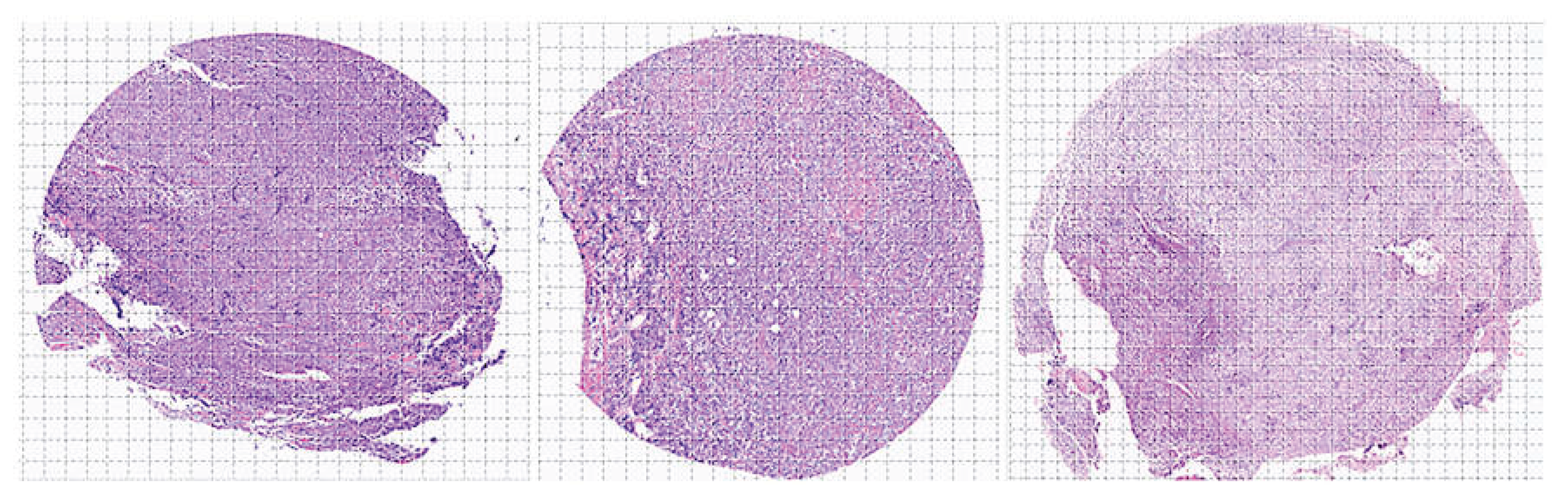
2.1. Patients and samples
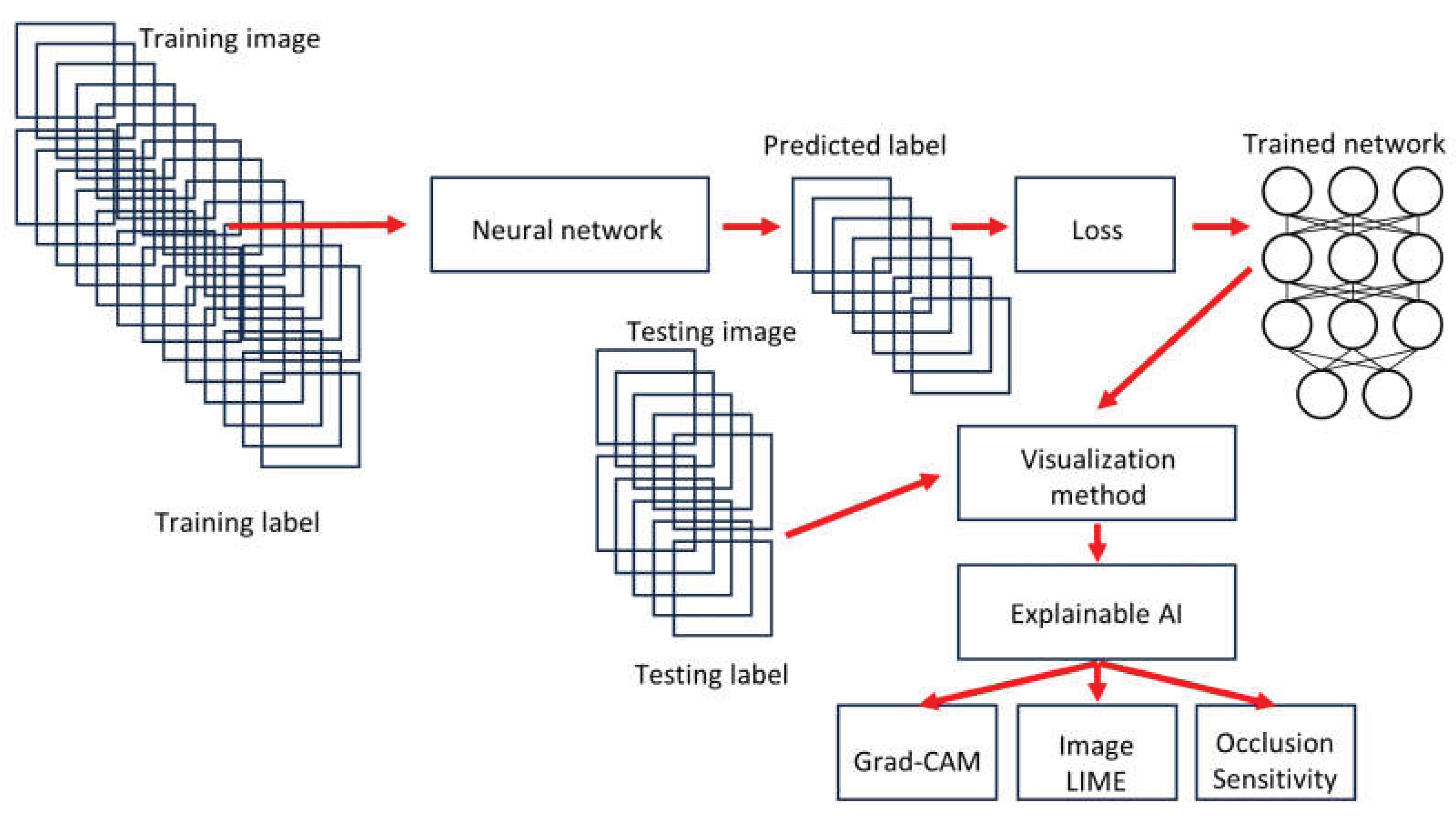
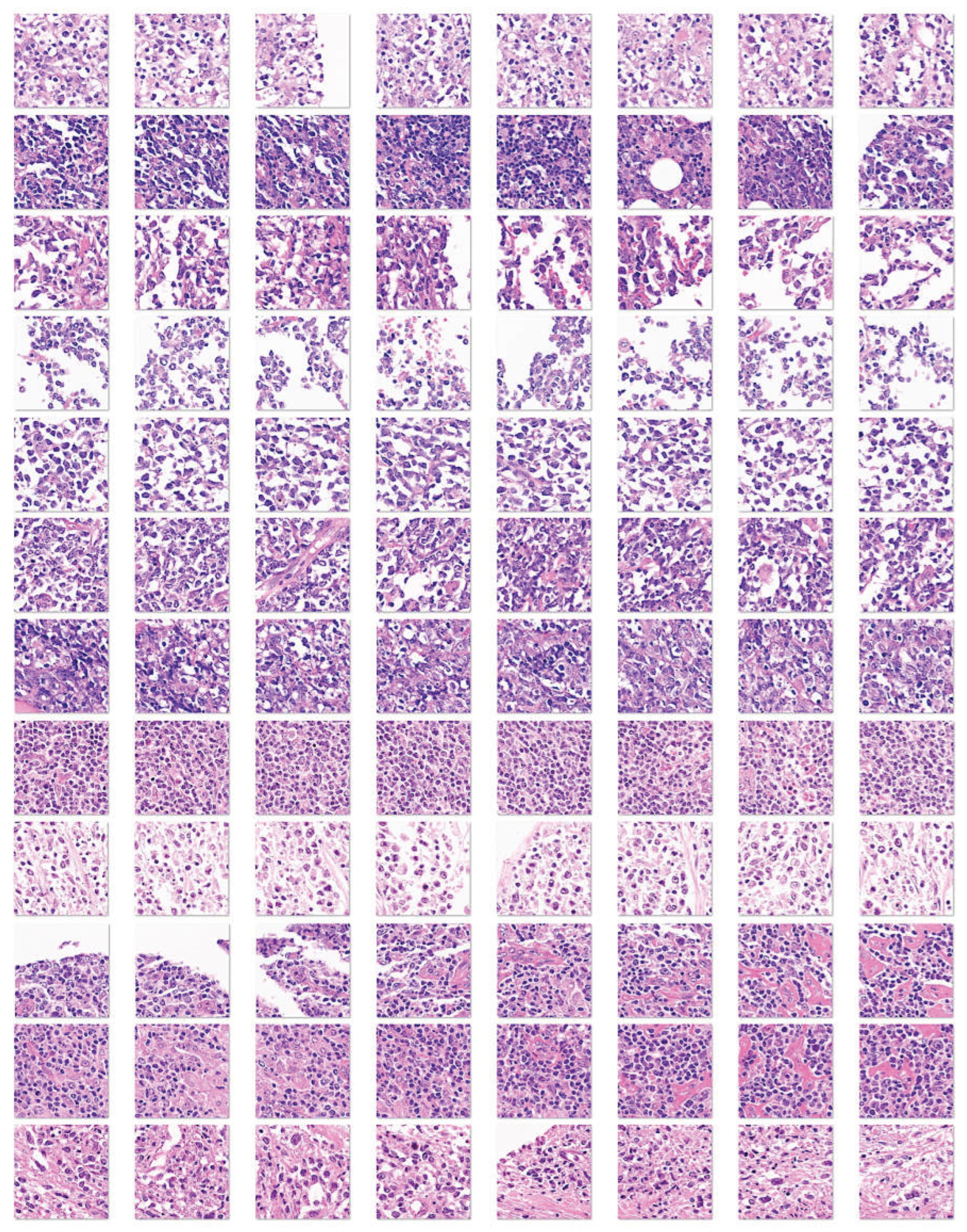
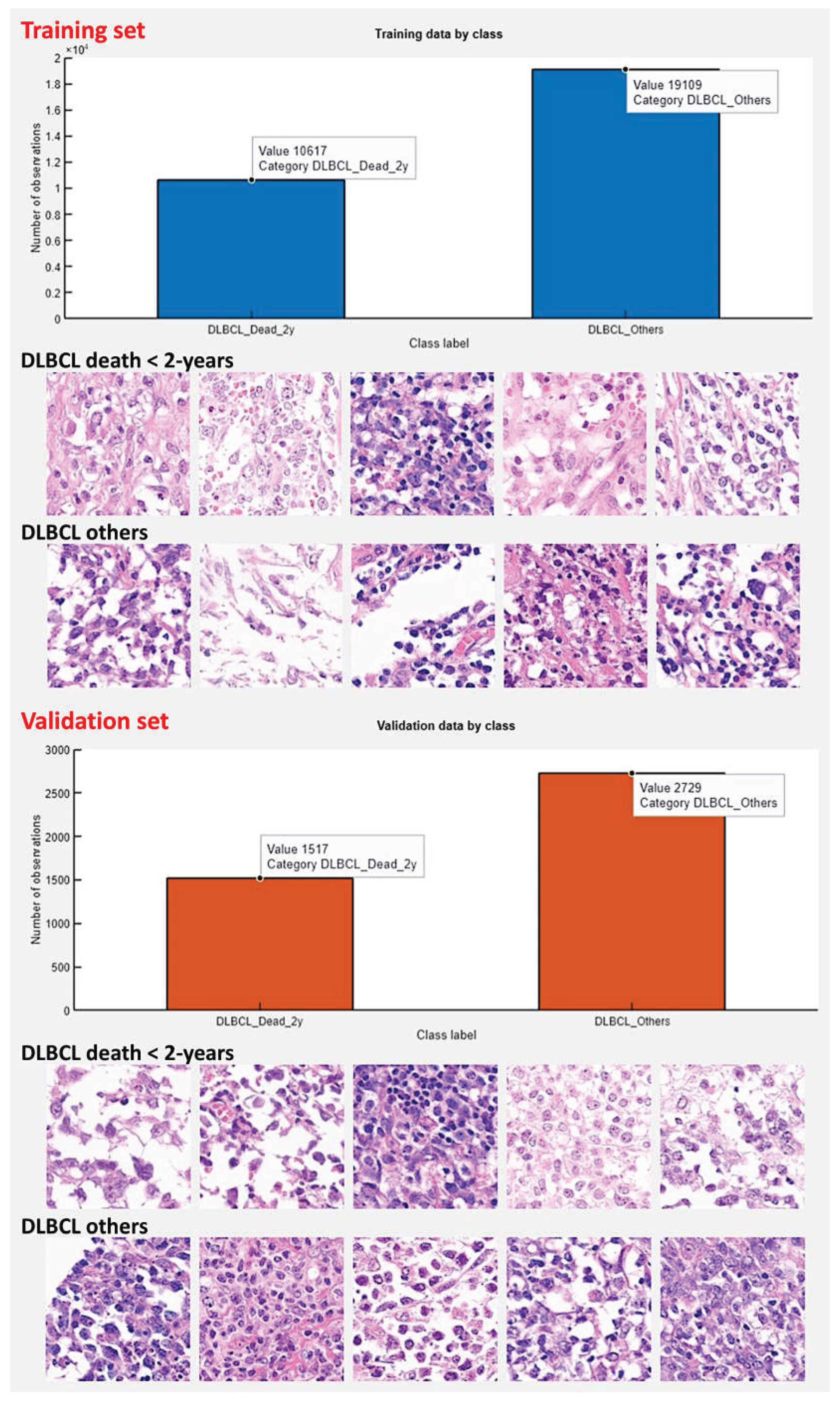
2.2. Deep learning image classification
3. Results
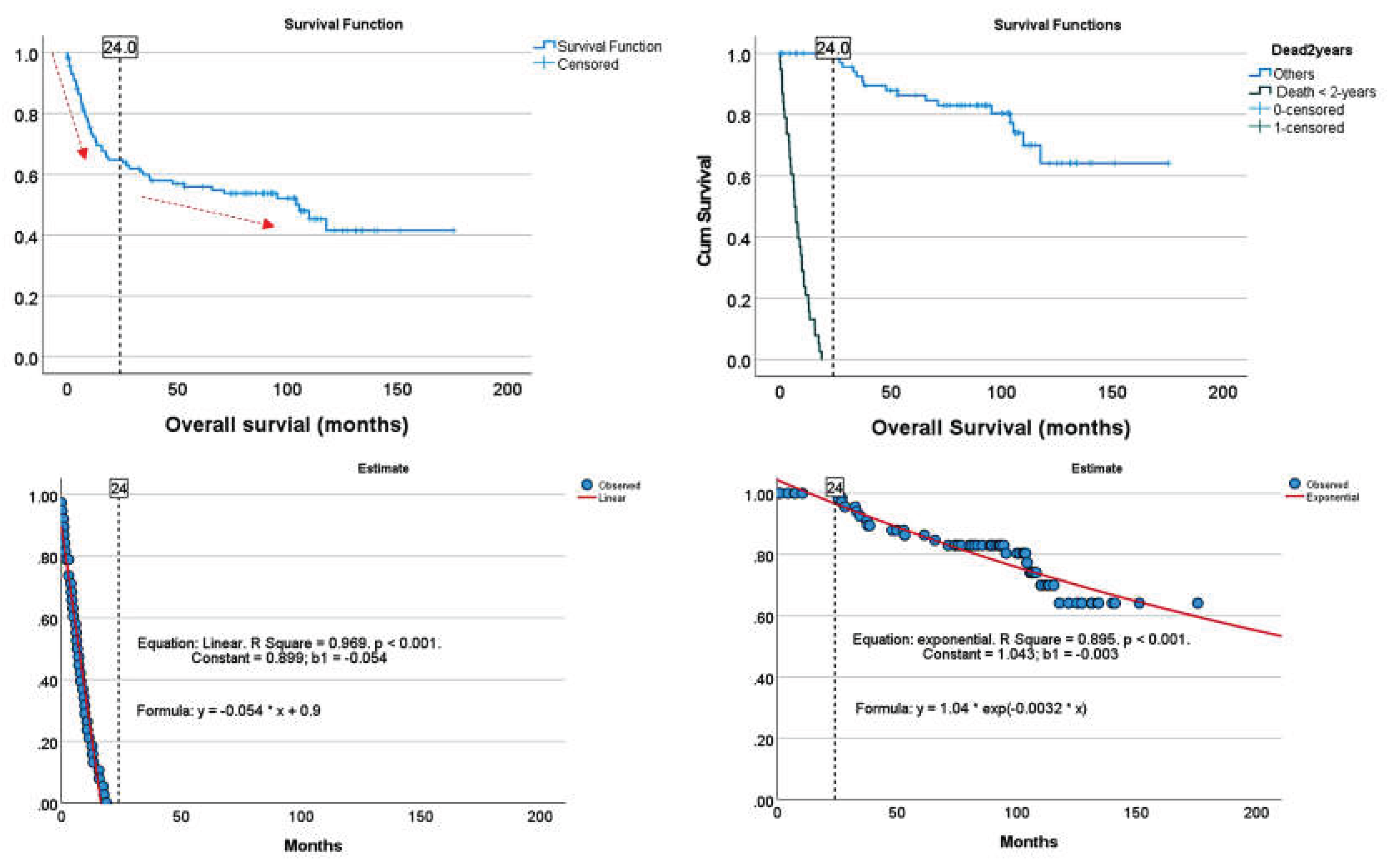
3.1. Image patch-based CNN classification of the 2-years cut-off OS groups
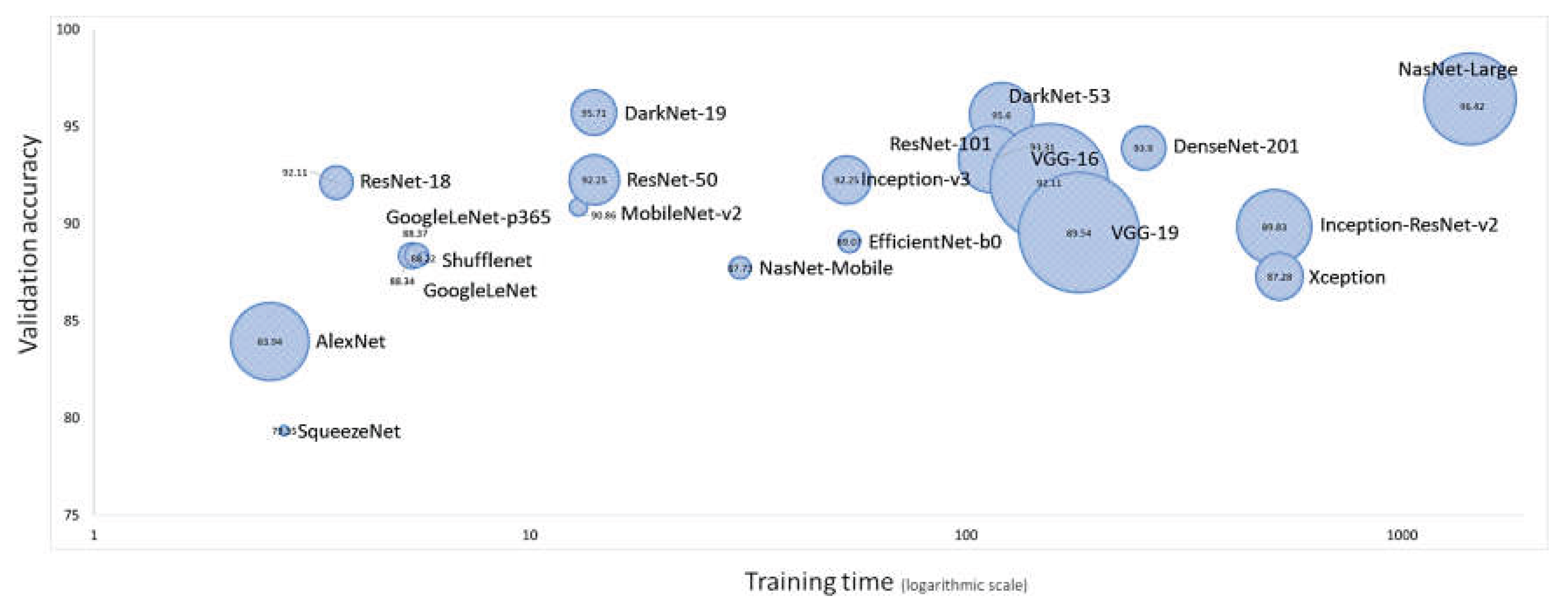
| CNN model | Learnables | Layers | Connections | Image Input | Training time | Validation accuracy (%) | Efficiency | Relative time |
|---|---|---|---|---|---|---|---|---|
| NasNet-Large | 84.9M | 1243 | 1462 | 331×331×3 | 1429 min 12 sec | 96.42 | 0.001 | 564.2 |
| DarkNet-19 | 20.8M | 64 | 63 | 256×256×3 | 14 min 1 sec | 95.71 | 0.114 | 5.5 |
| DarkNet-53 | 41.6M | 184 | 206 | 256×256×3 | 120 min 35 sec | 95.6 | 0.013 | 47.6 |
| DenseNet-201 | 20M | 708 | 805 | 224×224×3 | 255 min 9 sec | 93.9 | 0.006 | 100.7 |
| ResNet-101 | 44.6M | 347 | 379 | 224×224×3 | 114 min 21 sec | 93.31 | 0.014 | 45.1 |
| Inception-v3 | 23.8M | 315 | 349 | 299×299×3 | 53 min 15 sec | 92.25 | 0.029 | 21.0 |
| ResNet-50 | 25.5M | 177 | 192 | 224×224×3 | 14 min 3 sec | 92.25 | 0.109 | 5.5 |
| ResNet-18 | 11.6M | 71 | 78 | 224×224×3 | 3 min 36 sec | 92.11 | 0.426 | 1.4 |
| VGG-16 | 138.3M | 41 | 40 | 224×224×3 | 155 min 10 sec | 92.11 | 0.009 | 61.3 |
| MobileNet-v2 | 3.5M | 154 | 163 | 224×224×3 | 12 min 55 sec | 90.86 | 0.117 | 5.1 |
| Inception-ResNet-v2 | 55.8M | 824 | 921 | 299×299×3 | 509 min 8 sec | 89.83 | 0.003 | 201.0 |
| VGG-19 | 143.6M | 47 | 46 | 224×224×3 | 181 min 17 sec | 89.54 | 0.008 | 71.6 |
| EfficientNet-b0 | 5.3M | 290 | 363 | 224×224×3 | 54 min 0 sec | 89.07 | 0.075 | 21.3 |
| GoogleLeNet-places365 | 5.9M | 144 | 170 | 224×224×3 | 5 min 30 sec | 88.37 | 0.268 | 2.2 |
| GoogleLeNet | 6.9M | 144 | 170 | 224×224×3 | 5 min 21 sec | 88.34 | 0.275 | 2.1 |
| Shufflenet | 1.4M | 172 | 187 | 224×224×3 | 5 min 42 sec | 88.22 | 0.258 | 2.3 |
| NasNet-Mobile | 5.3M | 913 | 1072 | 224×224×3 | 30 min 19 sec | 87.73 | 0.048 | 12.0 |
| Xception | 22.9M | 170 | 181 | 299×299×3 | 522 min 52 sec | 87.28 | 0.003 | 206.4 |
| AlexNet | 60.9M | 25 | 24 | 227×227×3 | 2 min 32 sec | 83.94 | 0.552 | 1.0 |
| SqueezeNet | 1.2M | 68 | 75 | 227×227×3 | 2 min 44 sec | 79.35 | 0.484 | 1.1 |
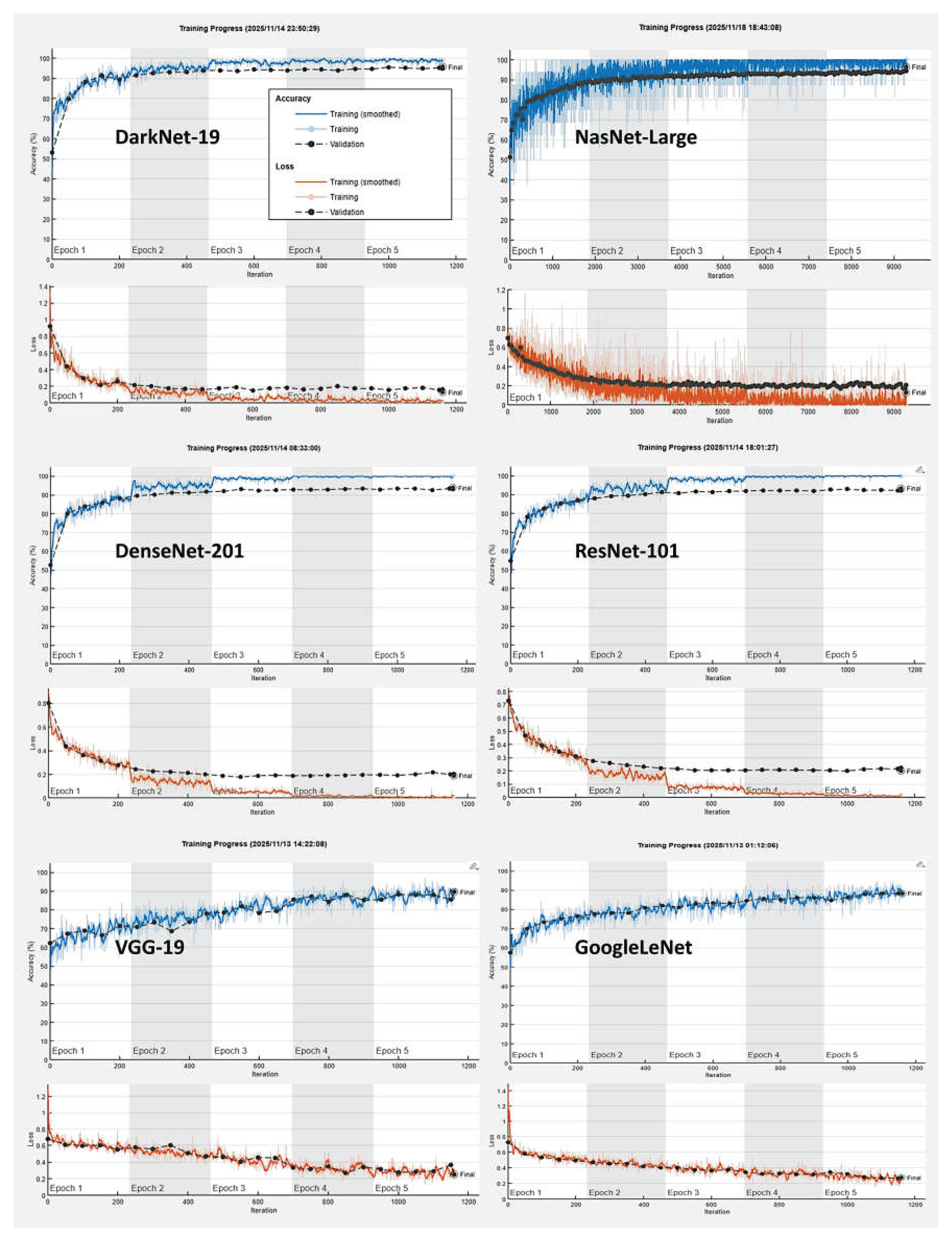
3.3. Clinicopathological Characteristics
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CNIO | Spanish National Cancer Research Center |
| CNN | Convolutional neural network |
| DLBCL | Diffuse large B-cell lymphoma |
| H&E | Hematoxylin and eosin |
| NHL | Non-Hodgkin lymphoma |
| XAI | Explainable artificial intelligence |
Appendix A
Appendix B
| Marker | Target | Clone | Company |
|---|---|---|---|
| CD3 | T lymphocytes | LN10 | Novocastra (Leica) |
| CD20 | B lymphocytes | L26 | Novocastra (Leica) |
| CD5 | T lymphocytes | 4C7 | Novocastra (Leica) |
| CD10 | Germinal center | 56C6 | Novocastra (Leica) |
| BCL6 | Germinal center | LN22 | Novocastra (Leica) |
| MUM1 / IRF4 | Plasma cell differentiation | EAU32 | Novocastra (Leica) |
| BCL2 | Apoptosis | Bcl2/10/D5 | Novocastra (Leica) |
| EBER | EBV-encoded mRNA | BP0589/AR0833 | Novocastra (Leica) |
| Ki67 | Cell proliferation | MM1 | Novocastra (Leica) |
| IL10 | Immuno-oncology | LS-B7432 | Lifespan Bioscience |
| PD-L1 (CD274) | Immuno-oncology | E1J2 | Cell Signaling |
| CSF1R | Immuno-oncology | FER216 | CNIO |
| CD163 | Tumor-associated macrophages | 10D6 | Novocastra (Leica) |
| CASP8 | Active subunit p18 | 11B6 | Novocastra |
| TNFAIP8 | Apoptosis | 14559-MM0 | Sino Biological |
| LMO2 | Hematopoietic development | 299B | CNIO |
| MYC | Proto-oncogene | Y69 | Abcam |
| MDM2 | Proto-oncogene | IF2 | Invitrogen |
| CDK6 | Cell cycle | 98D | CNIO |
| E2F1 | Cell cycle | Agro368V | CNIO |
| TP53 | Cell regulation | DO-7 | Novocastra (Leica) |
References
- Brown JR, Aster JC, Lister A, Rosmarin A. Pathobiology of diffuse large B cell lymphoma and primary mediastinal large B cell lymphoma. In UpToDate. Website: www.uptodate.com (last accessed on 19th November 2025).
- Alaggio, R.; Amador, C.; Anagnostopoulos, I.; Attygalle, A.D.; Araujo, I.B.O.; Berti, E.; Bhagat, G.; Borges, A.M.; Boyer, D.; Calaminici, M.; et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms. Leukemia 2022, 36, 1720-1748. [CrossRef]
- Arber, D.A.; Campo, E.; Jaffe, E.S. Advances in the Classification of Myeloid and Lymphoid Neoplasms. Virchows Arch 2023, 482, 1-9. [CrossRef]
- Campo, E. The 2022 classifications of lymphoid neoplasms : Keynote. Pathologie (Heidelb) 2023, 44, 121-127. [CrossRef]
- Campo, E.; Jaffe, E.S.; Cook, J.R.; Quintanilla-Martinez, L.; Swerdlow, S.H.; Anderson, K.C.; Brousset, P.; Cerroni, L.; de Leval, L.; Dirnhofer, S.; et al. The International Consensus Classification of Mature Lymphoid Neoplasms: a report from the Clinical Advisory Committee. Blood 2022, 140, 1229-1253. [CrossRef]
- de Leval, L.; Alizadeh, A.A.; Bergsagel, P.L.; Campo, E.; Davies, A.; Dogan, A.; Fitzgibbon, J.; Horwitz, S.M.; Melnick, A.M.; Morice, W.G.; et al. Genomic profiling for clinical decision making in lymphoid neoplasms. Blood 2022, 140, 2193-2227. [CrossRef]
- Song, J.Y.; Dirnhofer, S.; Piris, M.A.; Quintanilla-Martinez, L.; Pileri, S.; Campo, E. Diffuse large B-cell lymphomas, not otherwise specified, and emerging entities. Virchows Arch 2023, 482, 179-192. [CrossRef]
- Li, S.; Young, K.H.; Medeiros, L.J. Diffuse large B-cell lymphoma. Pathology 2018, 50, 74-87. [CrossRef]
- Crump, M. Management of Relapsed Diffuse Large B-cell Lymphoma. Hematol Oncol Clin North Am 2016, 30, 1195-1213. [CrossRef]
- Swerdlow, S.H.; Campo, E.; Pileri, S.A.; Harris, N.L.; Stein, H.; Siebert, R.; Advani, R.; Ghielmini, M.; Salles, G.A.; Zelenetz, A.D.; et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 2016, 127, 2375-2390. [CrossRef]
- Xie, Y.; Pittaluga, S.; Jaffe, E.S. The histological classification of diffuse large B-cell lymphomas. Semin Hematol 2015, 52, 57-66. [CrossRef]
- de Leval, L.; Jaffe, E.S. Lymphoma Classification. Cancer J 2020, 26, 176-185. [CrossRef]
- Willemze, R.; Cerroni, L.; Kempf, W.; Berti, E.; Facchetti, F.; Swerdlow, S.H.; Jaffe, E.S. The 2018 update of the WHO-EORTC classification for primary cutaneous lymphomas. Blood 2019, 133, 1703-1714. [CrossRef]
- Jaffe, E.S. Diagnosis and classification of lymphoma: Impact of technical advances. Semin Hematol 2019, 56, 30-36. [CrossRef]
- Quintanilla-Martinez, L.; Swerdlow, S.H.; Tousseyn, T.; Barrionuevo, C.; Nakamura, S.; Jaffe, E.S. New concepts in EBV-associated B, T, and NK cell lymphoproliferative disorders. Virchows Arch 2023, 482, 227-244. [CrossRef]
- Hans, C.P.; Weisenburger, D.D.; Greiner, T.C.; Gascoyne, R.D.; Delabie, J.; Ott, G.; Muller-Hermelink, H.K.; Campo, E.; Braziel, R.M.; Jaffe, E.S.; et al. Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood 2004, 103, 275-282. [CrossRef]
- Collinge, B.; Hilton, L.K.; Wong, J.; Alduaij, W.; Ben-Neriah, S.; Slack, G.W.; Farinha, P.; Boyle, M.; Meissner, B.; Cook, J.R.; et al. High-grade B-cell lymphoma, not otherwise specified: an LLMPP study. Blood Adv 2025, 9, 5409-5422. [CrossRef]
- Herlevic, V.; Reynolds, S.B.; Morris, J.D. Gastric Lymphoma. In StatPearls; Treasure Island (FL), 2025.
- Grainger, B.T.; Cheah, C.Y. Primary testicular lymphoma. Cancer Treat Rev 2025, 136, 102927. [CrossRef]
- Chen, S.Y.; Xu, P.P.; Feng, R.; Cui, G.H.; Wang, L.; Cheng, S.; Mu, R.J.; Zhang, H.L.; Wei, X.L.; Song, Y.P.; et al. Extranodal diffuse large B-cell lymphoma: Clinical and molecular insights with survival outcomes from the multicenter EXPECT study. Cancer Commun (Lond) 2025, 45, 919-935. [CrossRef]
- International Non-Hodgkin’s Lymphoma Prognostic Factors, P. A predictive model for aggressive non-Hodgkin’s lymphoma. N Engl J Med 1993, 329, 987-994. [CrossRef]
- Scott, D.W. Cell-of-Origin in Diffuse Large B-Cell Lymphoma: Are the Assays Ready for the Clinic? Am Soc Clin Oncol Educ Book 2015, e458-466. [CrossRef]
- Choi, W.W.; Weisenburger, D.D.; Greiner, T.C.; Piris, M.A.; Banham, A.H.; Delabie, J.; Braziel, R.M.; Geng, H.; Iqbal, J.; Lenz, G.; et al. A new immunostain algorithm classifies diffuse large B-cell lymphoma into molecular subtypes with high accuracy. Clin Cancer Res 2009, 15, 5494-5502. [CrossRef]
- Alizadeh, A.A.; Eisen, M.B.; Davis, R.E.; Ma, C.; Lossos, I.S.; Rosenwald, A.; Boldrick, J.C.; Sabet, H.; Tran, T.; Yu, X.; et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 2000, 403, 503-511. [CrossRef]
- Rosenwald, A.; Wright, G.; Chan, W.C.; Connors, J.M.; Campo, E.; Fisher, R.I.; Gascoyne, R.D.; Muller-Hermelink, H.K.; Smeland, E.B.; Giltnane, J.M.; et al. The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N Engl J Med 2002, 346, 1937-1947. [CrossRef]
- Wright, G.; Tan, B.; Rosenwald, A.; Hurt, E.H.; Wiestner, A.; Staudt, L.M. A gene expression-based method to diagnose clinically distinct subgroups of diffuse large B cell lymphoma. Proc Natl Acad Sci U S A 2003, 100, 9991-9996. [CrossRef]
- Berglund, M.; Thunberg, U.; Amini, R.M.; Book, M.; Roos, G.; Erlanson, M.; Linderoth, J.; Dictor, M.; Jerkeman, M.; Cavallin-Stahl, E.; et al. Evaluation of immunophenotype in diffuse large B-cell lymphoma and its impact on prognosis. Mod Pathol 2005, 18, 1113-1120. [CrossRef]
- Aster JC, Abramson JS, Lister A, Rosmarin AG. Prognosis of diffuse large B cell lymphoma. In UpToDate (last accessed on 20th November, 2025).
- Horn, H.; Ziepert, M.; Becher, C.; Barth, T.F.; Bernd, H.W.; Feller, A.C.; Klapper, W.; Hummel, M.; Stein, H.; Hansmann, M.L.; et al. MYC status in concert with BCL2 and BCL6 expression predicts outcome in diffuse large B-cell lymphoma. Blood 2013, 121, 2253-2263. [CrossRef]
- Johnson, N.A.; Slack, G.W.; Savage, K.J.; Connors, J.M.; Ben-Neriah, S.; Rogic, S.; Scott, D.W.; Tan, K.L.; Steidl, C.; Sehn, L.H.; et al. Concurrent expression of MYC and BCL2 in diffuse large B-cell lymphoma treated with rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone. J Clin Oncol 2012, 30, 3452-3459. [CrossRef]
- Green, T.M.; Young, K.H.; Visco, C.; Xu-Monette, Z.Y.; Orazi, A.; Go, R.S.; Nielsen, O.; Gadeberg, O.V.; Mourits-Andersen, T.; Frederiksen, M.; et al. Immunohistochemical double-hit score is a strong predictor of outcome in patients with diffuse large B-cell lymphoma treated with rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone. J Clin Oncol 2012, 30, 3460-3467. [CrossRef]
- Hu, S.; Xu-Monette, Z.Y.; Tzankov, A.; Green, T.; Wu, L.; Balasubramanyam, A.; Liu, W.M.; Visco, C.; Li, Y.; Miranda, R.N.; et al. MYC/BCL2 protein coexpression contributes to the inferior survival of activated B-cell subtype of diffuse large B-cell lymphoma and demonstrates high-risk gene expression signatures: a report from The International DLBCL Rituximab-CHOP Consortium Program. Blood 2013, 121, 4021-4031; quiz 4250. [CrossRef]
- Somasundaram, E.; Abramson, J.S. Double hit lymphoma: contemporary understanding and practices. Leuk Lymphoma 2025, 66, 26-33. [CrossRef]
- Karmali, R.; Shouse, G.; Torka, P.; Moyo, T.K.; Romancik, J.; Barta, S.K.; Bhansali, R.; Cohen, J.B.; Shah, N.N.; Zurko, J.; et al. Double hit & double expressor lymphomas: a multicenter analysis of survival outcomes with CD19-directed CAR T-cell therapy. Blood Cancer J 2025, 15, 43. [CrossRef]
- Qiu, L.; Medeiros, L.J.; Li, S. High-grade B-cell lymphomas: Double hit and non-double hit. Hum Pathol 2025, 156, 105700. [CrossRef]
- Reddy, A.; Zhang, J.; Davis, N.S.; Moffitt, A.B.; Love, C.L.; Waldrop, A.; Leppa, S.; Pasanen, A.; Meriranta, L.; Karjalainen-Lindsberg, M.L.; et al. Genetic and Functional Drivers of Diffuse Large B Cell Lymphoma. Cell 2017, 171, 481-494 e415. [CrossRef]
- Schmitz, R.; Wright, G.W.; Huang, D.W.; Johnson, C.A.; Phelan, J.D.; Wang, J.Q.; Roulland, S.; Kasbekar, M.; Young, R.M.; Shaffer, A.L.; et al. Genetics and Pathogenesis of Diffuse Large B-Cell Lymphoma. N Engl J Med 2018, 378, 1396-1407. [CrossRef]
- Chapuy, B.; Stewart, C.; Dunford, A.J.; Kim, J.; Kamburov, A.; Redd, R.A.; Lawrence, M.S.; Roemer, M.G.M.; Li, A.J.; Ziepert, M.; et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat Med 2018, 24, 679-690. [CrossRef]
- Carreras, J.; Kikuti, Y.Y.; Roncador, G.; Miyaoka, M.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; Shiraiwa, S.; et al. High Expression of Caspase-8 Associated with Improved Survival in Diffuse Large B-Cell Lymphoma: Machine Learning and Artificial Neural Networks Analyses. BioMedInformatics 2021, 1, 18-46. [CrossRef]
- Carreras, J.; Kikuti, Y.Y.; Miyaoka, M.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; Shiraiwa, S.; Hamoudi, R.; et al. A Single Gene Expression Set Derived from Artificial Intelligence Predicted the Prognosis of Several Lymphoma Subtypes; and High Immunohistochemical Expression of TNFAIP8 Associated with Poor Prognosis in Diffuse Large B-Cell Lymphoma. AI 2020, 1, 342-360. [CrossRef]
- Carreras, J.; Kikuti, Y.Y.; Miyaoka, M.; Roncador, G.; Garcia, J.F.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; et al. Integrative Statistics, Machine Learning and Artificial Intelligence Neural Network Analysis Correlated CSF1R with the Prognosis of Diffuse Large B-Cell Lymphoma. Hemato 2021, 2, 182-206. [CrossRef]


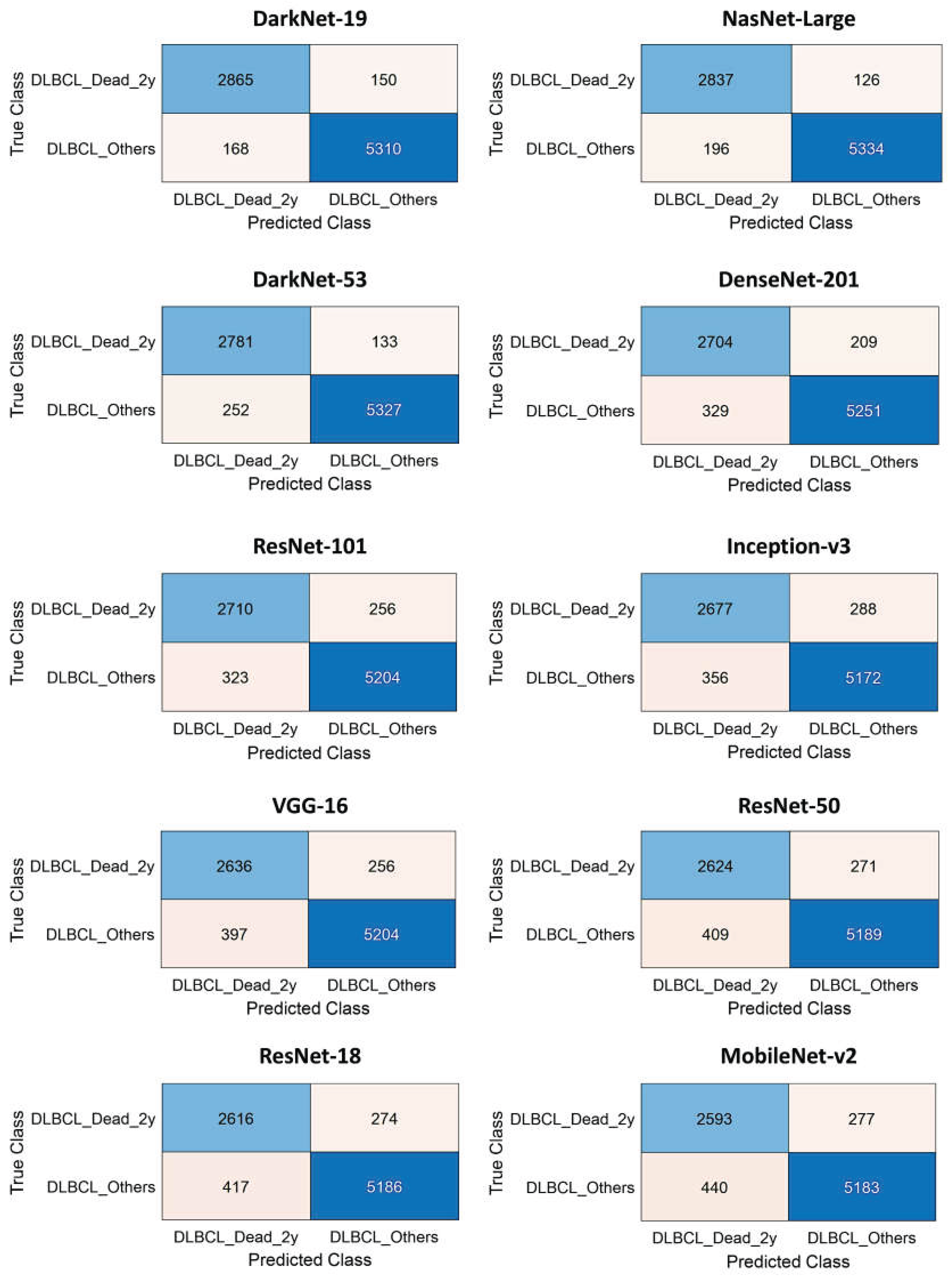
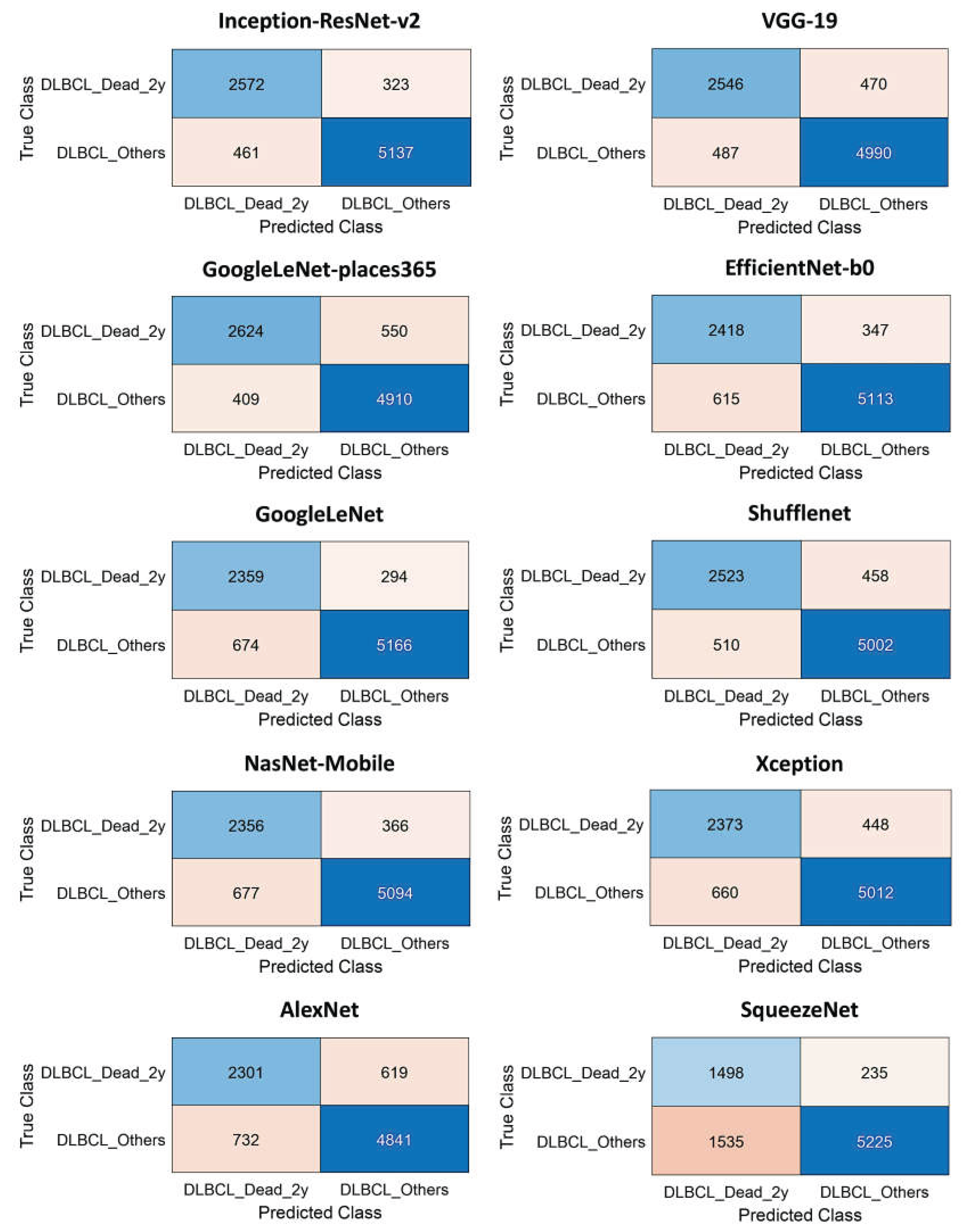
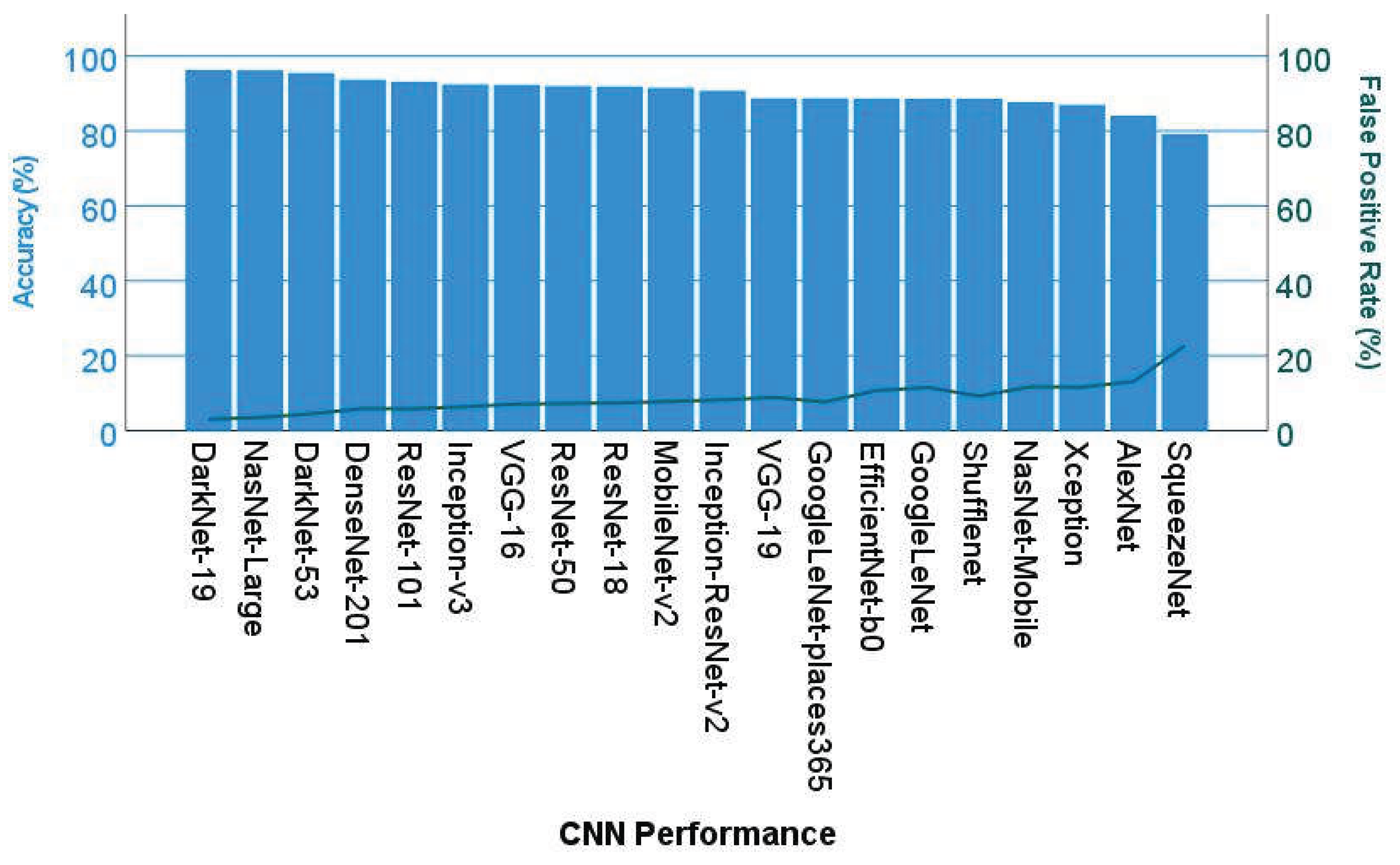
| CNN | Accuracy (%) | Precision (%) | Recall (%) | False Positive Rate (%) | Specificity (%) | F1 score (%) |
|---|---|---|---|---|---|---|
| DarkNet-19 | 96.26 | 94.46 | 95.02 | 3.07 | 96.93 | 94.74 |
| NasNet-Large | 96.21 | 93.54 | 95.75 | 3.54 | 96.46 | 94.63 |
| DarkNet-53 | 95.47 | 91.69 | 95.44 | 4.52 | 95.48 | 93.53 |
| DenseNet-201 | 93.67 | 89.15 | 92.83 | 5.9 | 94.1 | 90.95 |
| ResNet-101 | 93.18 | 89.35 | 91.37 | 5.84 | 94.16 | 90.35 |
| Inception-v3 | 92.42 | 88.26 | 90.29 | 6.44 | 93.56 | 89.26 |
| VGG-16 | 92.31 | 86.91 | 91.15 | 7.09 | 92.91 | 88.98 |
| ResNet-50 | 91.99 | 86.52 | 90.64 | 7.31 | 92.69 | 88.53 |
| ResNet-18 | 91.86 | 86.25 | 90.52 | 7.44 | 92.56 | 88.33 |
| MobileNet-v2 | 91.56 | 85.49 | 90.35 | 7.83 | 92.17 | 87.85 |
| Inception-ResNet-v2 | 90.77 | 84.8 | 88.84 | 8.24 | 91.76 | 86.77 |
| VGG-19 | 88.73 | 83.94 | 84.42 | 8.89 | 91.11 | 84.18 |
| GoogleLeNet-places365 | 88.71 | 86.52 | 82.67 | 7.69 | 92.31 | 84.55 |
| EfficientNet-b0 | 88.67 | 79.72 | 87.45 | 10.74 | 89.26 | 83.41 |
| GoogleLeNet | 88.6 | 77.78 | 88.92 | 11.54 | 88.46 | 82.98 |
| Shufflenet | 88.6 | 83.18 | 84.64 | 9.25 | 90.75 | 83.9 |
| NasNet-Mobile | 87.72 | 77.68 | 86.55 | 11.73 | 88.27 | 81.88 |
| Xception | 86.95 | 78.23 | 84.11 | 11.64 | 88.36 | 81.07 |
| AlexNet | 84.09 | 75.87 | 78.8 | 13.13 | 86.87 | 77.31 |
| SqueezeNet | 79.16 | 49.39 | 86.44 | 22.71 | 77.29 | 62.86 |
| Variable | All cases | Dead 2-years | Others | P value |
|---|---|---|---|---|
| Frequency | 114 | 38/114 (33.3%) | 76/114 (66.7%) | - |
| Clinical characteristics | ||||
| Age > 60 years | 81/114 (71.1%) | 30/38 (78.9%) | 51/76 (67.1%) | 0.273 |
| Male | 60/114 (52.6%) | 19/38 (50%) | 41/76 (53.9%) | 0.697 |
| Location | ||||
| Nodal (+Spleen) | 58/114 (50.9%) | 16/38 (42.1%) | 42/76 (55.3%) | 0.430 |
| Waldeyer’s ring | 11/114 (9.6%) | 3/38 (7.9%) | 8/76 (10.5%) | |
| Gastrointestinal | 13/114 (11.4%) | 5/38 (13.2%) | 8/76 (10.5%) | |
| Other extranodal | 32/114 (28.1%) | 14/38 (36.8%) | 18/76 (23.7%) | |
| Stage III-IV | 46/97 (47.4%) | 18/28 (64.3%) | 28/69 (40.6%) | 0.044 |
| IPI High+High/Intermediate | 31/91 (34.1%) | 14/27 (51.9%) | 17/64 (26.6%) | 0.029 |
| RCHOP/RCHOP-like treatment | 93/98 (94.9%) | 26/28 (92.9%) | 67/70 (95.7%) | 0.513 |
| Clinical response | 68/92 (73.9%) | 5/24 (20.8%) | 63/68 (92.5%) | < 0.001 |
| Hight sIL2R | 79/99 (79.8%) | 27/29 (93.1%) | 52/70 (74.3%) | 0.052 |
| Pathological characteristics | ||||
| CD3+ | 0/114 (0%) | 0/38 (0%) | 0/76 (0%) | 1.0 |
| CD20+ | 114/114 (100%) | 38/38 (100%) | 76/76 (100%) | 1.0 |
| CD5+ | 13/113 (11.5%) | 4/38 (10.5%) | 9/75 (12.0%) | 1.0 |
| CD10+ | 33/113 (29.2%) | 2/38 (5.3%) | 31/75 (41.3%) | < 0.001 |
| BCL6+ | 76/113 (67.3%) | 26/38 (68.4%) | 50/75 (66.7%) | 1.0 |
| MUM1+ | 93/113 (82.3%) | 33/38 (86.8%) | 60/75 (80%) | 0.442 |
| Non-GCB | 77/114 (67.5%) | 35/38 (92.1%) | 42/76 (55.3%) | < 0.001 |
| BCL2+ | 89/113 (78.8%) | 36/38 (94.7%) | 53/75 (70.7%) | 0.003 |
| MYC rearrangement | 9/98 (9.2%) | 2/29 (6.9%) | 7/69 (10.1%) | 1.0 |
| EBER+ | 28/114 (25%) | 15/37 (40.5%) | 13/75 (17.3%) | 0.011 |
| Ki67 | 16.1% +/- 14.2 | 15.3% +/- 12.2 | 16.5% +/- 14.9 | 0.959 |
| Immune microenvironment | ||||
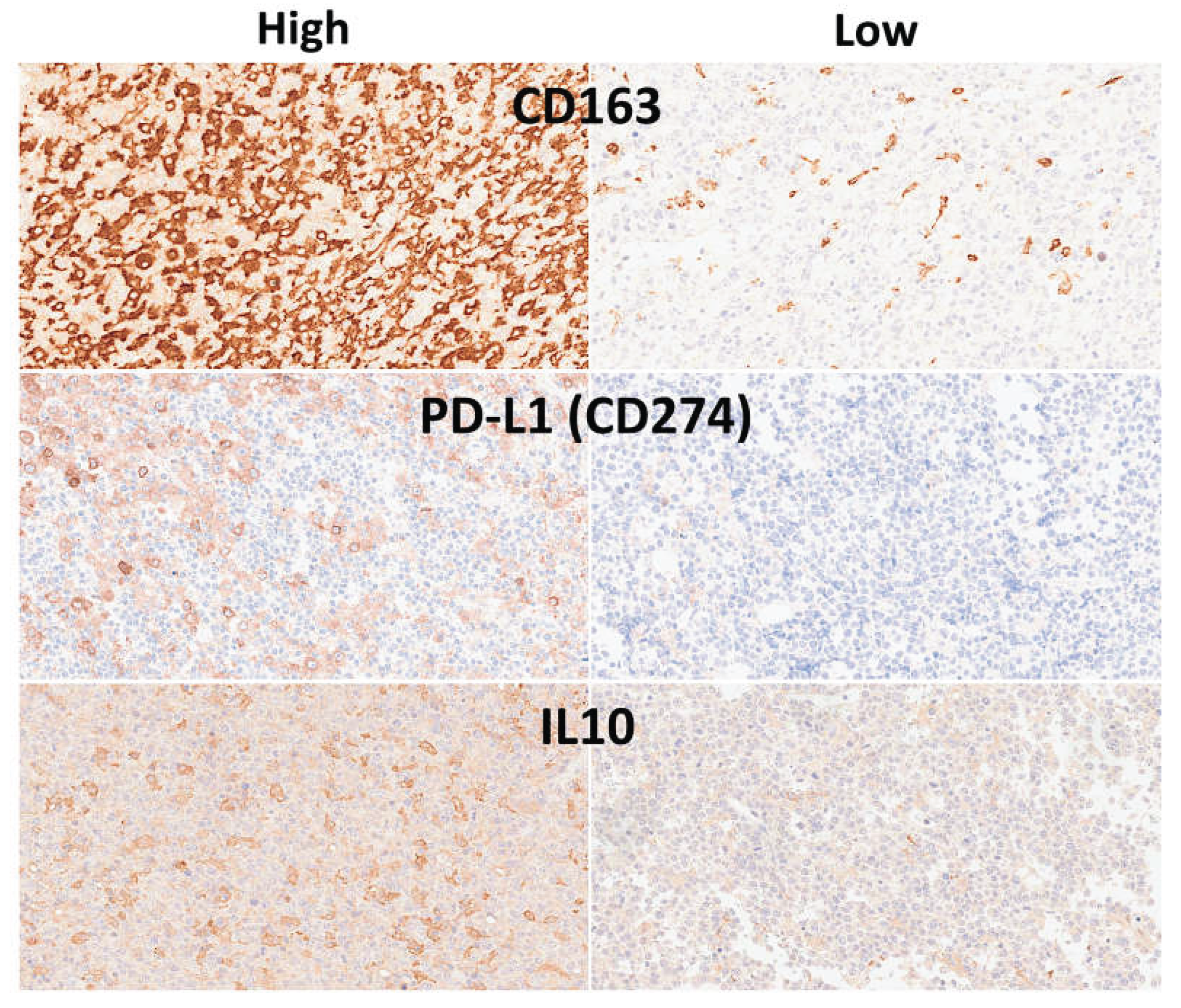
| IL10 | 12.2% +/- 15.8 (n = 102) | 18.6% +/- 19.6 | 9.2% +/- 12.8 | 0.006 |
| PD-L1 (CD274) | 12.2% +/- 15.8% (n = 102) | 18.5% +/- 19.6 | 9.1% +/- 12.8 | 0.026 |
| CSF1R | 33.5% +/- 27.5 (n = 94) | 28.7% +/- 25.4 | 35.8% +/- 28.3 | 0.247 |
| CD163 | 39.2% +/- 25.9 (n = 114) | 48.2% +/- 24.5 | 34.6% +/- 25.6 | 0.008 |
| CASP8 | 6.7% +/- 8.4 (n = 94) | 6.0% +/- 9.4 | 7.1% +/- 8.0 | 0.268 |
| TNFAIP8 | 41.3% +/- 25.6 (n =93) | 46.2 +/- 24.0 | 39.3% +/- 26.1 | 0.223 |
| Cell cycle / GC-related | ||||
| LMO2 | 2.6% +/- 3.5 (n = 92) | 2.4% +/- 3.9 | 2.7% +/- 3.4 | 0.051 |
| MYC | 5.4% +/- 5.7 (n = 93) | 6.5% +/- 6.4 | 4.9% +/- 5.5 | 0.318 |
| MDM2 | 10.8% +/- 8.1 (n = 93) | 9.7% +/- 6.1 | 11.3% +/- 8.8 | 0.594 |
| CDK6 | 5.1% +/- 7.4 (n = 93) | 3.6% +/- 5.3 | 5.7% +/- 8.1 | 0.056 |
| E2F1 | 1.8% +/- 1.8 (n = 93) | 1.2% +/- 0.9 | 2.0% +/- 1.9 | 0.020 |
| BCL2 | 6.8% +/- 9.7 (n = 93) | 3.4% +/- 4.5 | 8.1% +/- 10.9 | 0.087 |
| TP53 | 5.2% +/- 8.1 (n = 94) | 6.6% +/- 10.3 | 4.6% +/- 7.0 | 0.128 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




