Submitted:
29 December 2025
Posted:
30 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Contributions and Overview
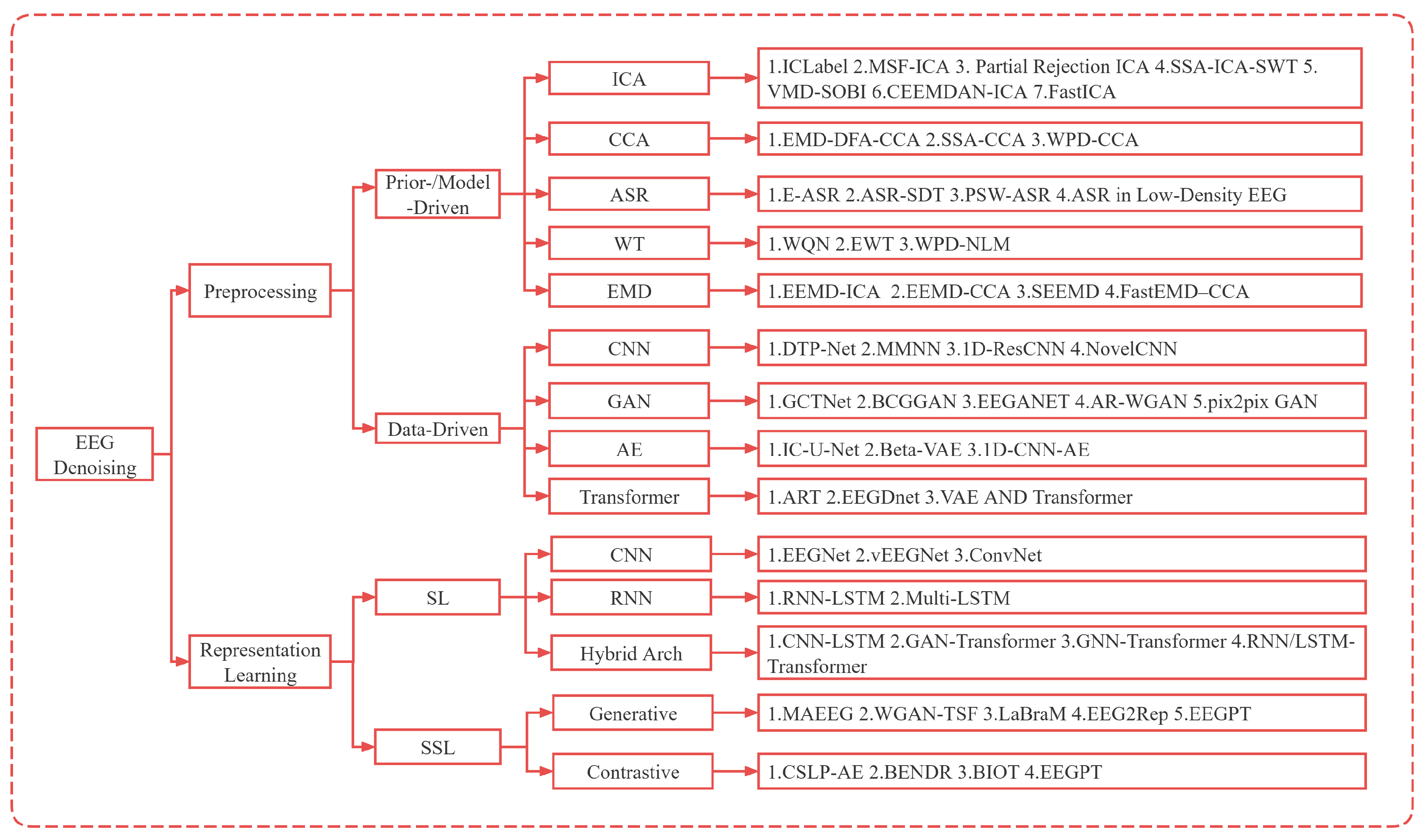
- Systematic synthesis of methods and applicability. We present a unified survey organized along two complementary routes—preprocessing for artifact mitigation and representation learning for robustness—covering prior-/model-driven signal processing, supervised data-driven models, and self-supervised learning.
- Quantified synergistic effects. Under a standardized experimental protocol, we quantify how representative traditional preprocessing modules interact with SSL encoders across downstream tasks, showing strong task dependence and providing empirical evidence for practical use.
- Practical guidelines and future directions. We provide a compact checklist to support task- and data-adaptive pipeline selection, and discuss open challenges and research directions toward hybrid frameworks that integrate signal-processing priors with self-supervised models.
1.2. Paper Organization
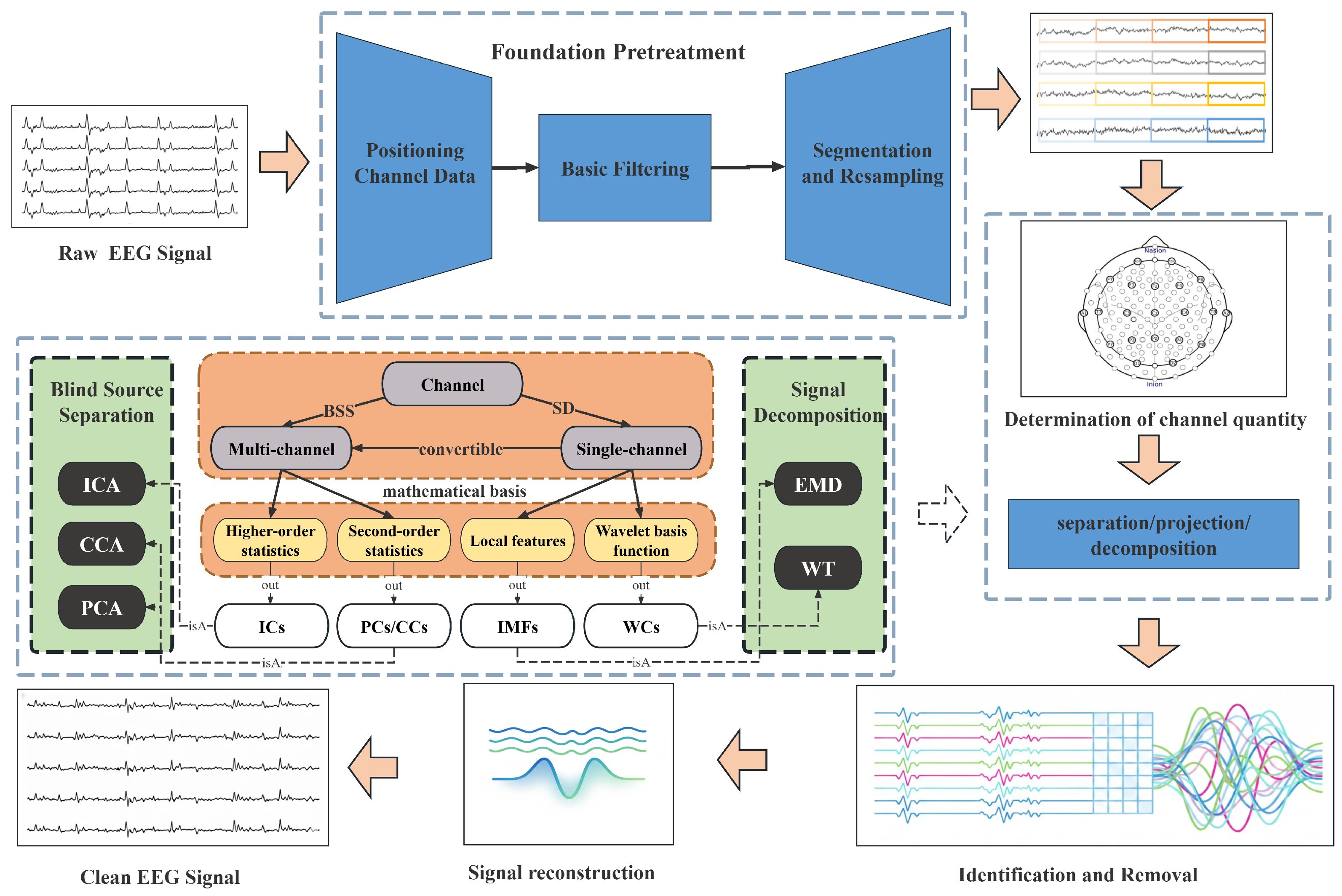
2. Preprocessing-Centric Artifact Mitigation
2.1. Prior-/Model-Driven Preprocessing Methods
2.1.1. Blind Source Separation
2.1.2. Signal Decomposition
2.2. Data-Driven Preprocessing Methods
2.2.1. CNN-Dominated Denoising Models
2.2.2. GAN-Based Denoising Models
| Model | Year | Structure | Artifacts | Dataset | Evaluation | Code Link |
|---|---|---|---|---|---|---|
| 1D-ResCNN [36] | 2020 | CNN+ResNet | Multiple Artifacts | PhysioNet | SNR: 22.944, RMSE: 0.0858 | None |
| EEGANet [40] | 2021 | GAN | EOG | Multiple Real Datasets | SNR: 3.247, RMSE: 3.6350, CC: 0.945 | EEGANet |
| NovelCNN [37] | 2021 | CNN | EMG | EEGdenoiseNet | RRMSE: 0.4480, CC: 0.863 | NovelCNN |
| IC-U-Net [41] | 2022 | U-Net+ICA | Multiple Artifacts | Simulated Data | SNR: 24.880, RMSE: 0.6320 | IC-U-Net |
| MMNN [38] | 2022 | CNN | Multiple Artifacts | EEGdenoiseNet | RRMSE: 0.3295, CC: 0.927 | MMNN |
| FrComp-CAE [42] | 2022 | 1D-CAE+TM+FrBP | EMG | Mendeley, Bonn | SNR: 8.680, RMSE: 0.0900, CC: 0.930 | None |
| AR-WGAN [43] | 2023 | WGAN | EMG, EOG | EEGdenoiseNet | RRMSE: 0.1760, CC: 0.726 | None |
| ImprovedGAN [44] | 2022 | GAN | EOG, EMG | EEGdenoiseNet | RRMSE: 0.1590, CC: 0.915 | None |
| GCTNet [45] | 2023 | GAN | Multiple Artifacts | EEGdenoiseNet | SNR: 12.141, RRMSE: 0.3080, CC: 0.925 | None |
| BCGGAN [46] | 2024 | GAN | BCG | Self-collected Data | PTPR:1.203 | None |
| Beta-VAE [47] | 2024 | VAE | Extreme Artifacts | DEAP | Gaussian/Popped MSE:0.0440 | None |
| DCAE [48] | 2024 | CAE | Antagonistic noise | ERN, MI | - | None |
| DTP-Net [39] | 2024 | CNN+TRM | Multiple Artifacts | EEGdenoiseNet | SNR: 14.312, RRMSE: 0.2342, CC: 0.981 | DTP-Net |
| pix2pixGAN [49] | 2024 | GAN | EMG | EEGdenoiseNet | RRMSE: 0.5797, CC: 0.865 | None |
| AT-AT [50] | 2025 | AT+TRM | EMG | EEGdenoiseNet | RRMSE: 0.5400, CC: 0.825 | None |
2.2.3. Autoencoder and Transformer-Based Denoising Models
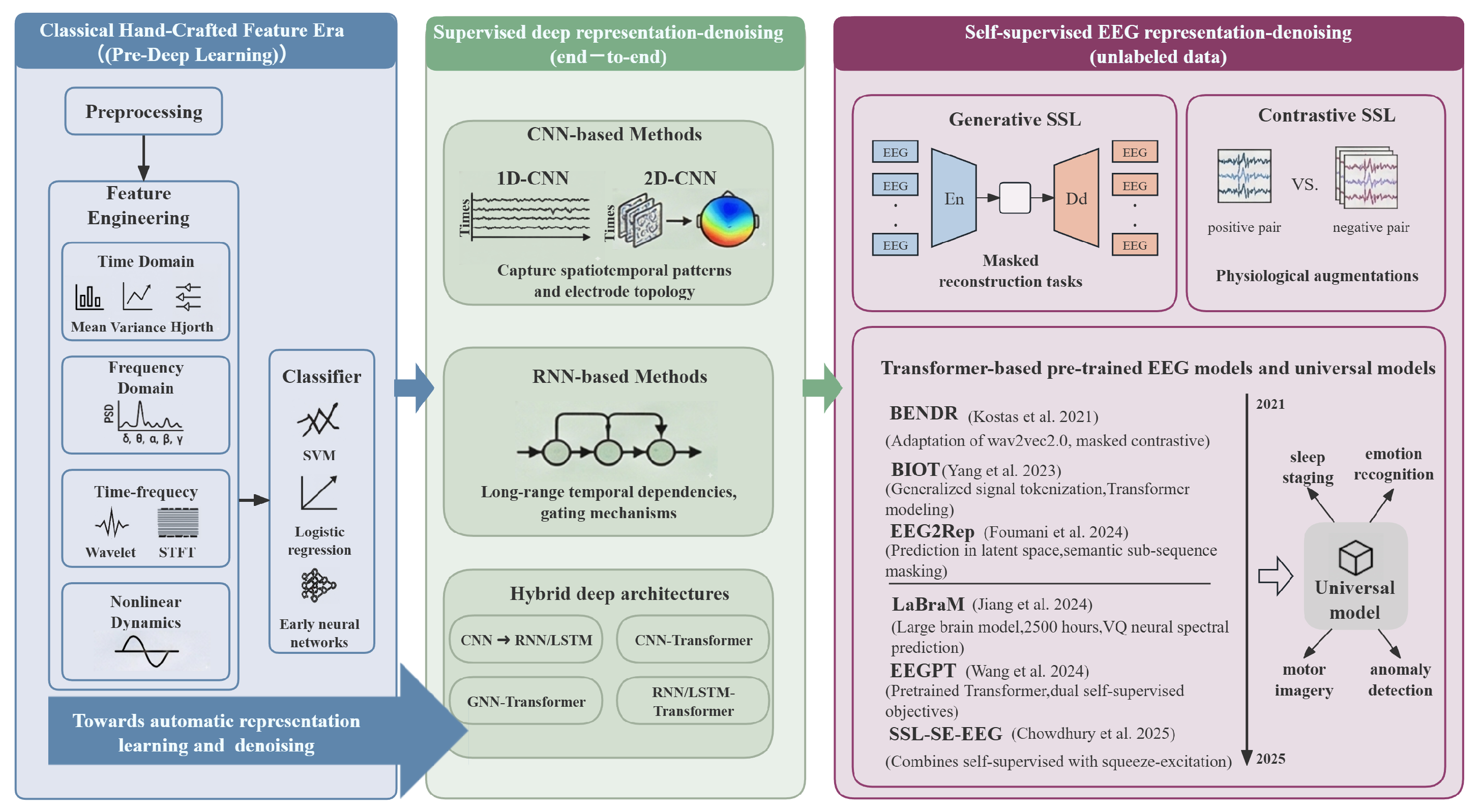
3. Representation-Learning-Centric Methods
3.1. Supervised Representation Learning
3.2. Self-Supervised Representation Learning
4. Synergy of Preprocessing and SSL
4.1. Datasets and Preprocessing
4.2. Implementation and Setup
| Datasets | Methods | Balanced Accuracy | Weighted F1 / AUROC | Cohen’s Kappa |
|---|---|---|---|---|
| BCIC-2A | Baseline | |||
| ICA+ICLabel | ||||
| EEMD | ||||
| Wavelet | ||||
| BCIC-2B | Baseline | |||
| ICA | ||||
| EEMD | ||||
| Wavelet | ||||
| PhysioP300 | Baseline | |||
| ICA+ICLabel | ||||
| EEMD | ||||
| Wavelet | ||||
| TUAB | Baseline | – | ||
| ICA+ICLabel | – | |||
| EEMD | – | |||
| Wavelet | – |
4.3. Results
5. Discussion
5.1. Challenges
5.1.1. Data Dilemma
5.1.2. Lack of Efficient Synergy Between Prior-Driven Preprocessing and Modern Representation Learning
5.1.3. Challenges in Practical Deployment
5.2. Future Research Directions
5.2.1. Prior-Guided Self-Supervision and Weak Supervision
5.2.2. Multimodal Fusion
5.2.3. Fine-Grained Subject- and Session-Aware Modeling
5.3. Practical Guidelines
Task–data–constraint checklist.
Actionable rules
- Use downstream metrics (Acc/F1/AUC/Kappa) as primary criteria; SNR/RMSE can be misleading.
- Start from the baseline SSL pipeline and add preprocessing only when necessary; treat it as tunable.
- Watch for distribution shift: strong preprocessing may reduce the usefulness of pretrained invariances.
- For MI, reduce denoising strength if performance drops (risk of over-denoising task-relevant rhythms).
- For ERP, prioritize temporal fidelity; avoid operations that alter waveform morphology/latency.
- For abnormal detection, moderate stabilization can help, but report efficiency and generalization (subject/session/site).
6. Conclusions
References
- Yi, Z.; Pan, J.; Chen, Z.; Lu, D.; Cai, H.; Li, J.; Xie, Q. A hybrid BCI integrating EEG and Eye-Tracking for assisting clinical communication in patients with disorders of consciousness. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2024. [Google Scholar]
- Shen, L.; Wu, Z.; Yue, Z.; Li, B.; Chen, Q.; Han, B. Prior Knowledge Uses Prestimulus Alpha Band Oscillations and Persistent Poststimulus Neural Templates for Conscious Perception. Journal of Neuroscience 2023, 43, 6164–6175. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Zhang, X.; Huang, X.; Wu, M.; Gao, J.; Lu, D.; Ding, Z.; Hu, B. An optimal channel selection for EEG-based depression detection via kernel-target alignment. IEEE Journal of Biomedical and Health Informatics 2020, 25, 2545–2556. [Google Scholar] [CrossRef] [PubMed]
- Muthukumaraswamy, S.D. High-frequency brain activity and muscle artifacts in MEG/EEG: a review and recommendations. Frontiers in human neuroscience 2013, 7, 138. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Deng, F.; Jiang, P. EEGDiR: Electroencephalogram denoising network for temporal information storage and global modeling through Retentive Network. Computers in Biology and Medicine 2024, 177, 108626. [Google Scholar] [CrossRef]
- Craik, A.; He, Y.; Contreras-Vidal, J. Deep Learning for EEG-Based Brain-Computer Interfaces: A Review. IEEE Transactions on Neural Networks and Learning Systems 2019, 30, 4–22. [Google Scholar]
- Chien, H.Y.S.; Goh, H.; Sandino, C.M.; Cheng, J.Y. Maeeg: Masked auto-encoder for eeg representation learning. arXiv 2022, arXiv:2211.02625. [Google Scholar] [CrossRef]
- Huang, Y.; Chen, Y.; Xu, S.; Wu, D.; Wu, X. Self-Supervised Learning with Adaptive Frequency-Time Attention Transformer for Seizure Prediction and Classification. Brain Sciences 2025, 15, 382. [Google Scholar]
- Zhang, X.; Zhang, X.; Huang, Q.; Chen, F. A review of epilepsy detection and prediction methods based on EEG signal processing and deep learning. Frontiers in Neuroscience 2024, 18, 1468967. [Google Scholar] [CrossRef]
- Del Pup, F.; Zanola, A.; Tshimanga, L.F.; Bertoldo, A.; Atzori, M. The more, the better? Evaluating the role of EEG preprocessing for deep learning applications. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2025. [Google Scholar]
- Yuan, Y.; Xun, G.; Jia, K.; Zhang, A. A multi-view deep learning framework for EEG seizure detection. IEEE journal of biomedical and health informatics 2018, 23, 83–94. [Google Scholar] [CrossRef]
- Xiong, W.; Li, J.; Li, J.; Zhu, K. Eeg-fm-bench: A comprehensive benchmark for the systematic evaluation of eeg foundation models. arXiv 2025, arXiv:2508.17742. [Google Scholar]
- Vigário, R.N. Extraction of ocular artefacts from EEG using independent component analysis. Electroencephalography and clinical neurophysiology 1997, 103, 395–404. [Google Scholar] [CrossRef] [PubMed]
- Mognon, A.; Jovicich, J.; Bruzzone, L.; Buiatti, M. ADJUST: An automatic EEG artifact detector based on the joint use of spatial and temporal features. Psychophysiology 2011, 48, 229–240. [Google Scholar] [CrossRef] [PubMed]
- Tamburro, G.; Fiedler, P.; Stone, D.; Haueisen, J.; Comani, S. A new ICA-based fingerprint method for the automatic removal of physiological artifacts from EEG recordings. PeerJ 2018, 6, e4380. [Google Scholar] [CrossRef] [PubMed]
- Pion-Tonachini, L.; Kreutz-Delgado, K.; Makeig, S. ICLabel: An automated electroencephalographic independent component classifier, dataset, and website. NeuroImage 2019, 198, 181–197. [Google Scholar] [CrossRef]
- Molina-Molina, M.; Tardón, L.J.; Barbancho, A.M.; Barbancho, I. Implementation of tools for lessening the influence of artifacts in EEG signal analysis. Applied Sciences 2024, 14, 971. [Google Scholar] [CrossRef]
- Villena, A.; Tardón, L.J.; Barbancho, I.; Barbancho, A.M.; Brattico, E.; Haumann, N.T. Preprocessing for lessening the influence of eye artifacts in EEG analysis. Applied Sciences 2019, 9, 1757. [Google Scholar] [CrossRef]
- Noorbasha, S.K.; Sudha, G.F. Removal of EOG artifacts and separation of different cerebral activity components from single channel EEG—An efficient approach combining SSA–ICA with wavelet thresholding for BCI applications. Biomedical Signal Processing and Control 2021, 63, 102168. [Google Scholar] [CrossRef]
- Liu, C.; Zhang, C. Remove artifacts from a single-channel EEG based on VMD and SOBI. Sensors 2022, 22, 6698. [Google Scholar] [CrossRef]
- Luo, Z.; Yan, Z.; Fu, W. Electroencephalogram artifact filtering method of single channel EEG based on CEEMDAN-ICA. Chin. J. Sens. Actuators 2018, 31, 1211–1216. [Google Scholar]
- Hossain, M.S.; Chowdhury, M.E.; Reaz, M.B.I.; Ali, S.H.M.; Bakar, A.A.A.; Kiranyaz, S.; Khandakar, A.; Alhatou, M.; Habib, R.; Hossain, M.M. Motion artifacts correction from single-channel EEG and fNIRS signals using novel wavelet packet decomposition in combination with canonical correlation analysis. Sensors 2022, 22, 3169. [Google Scholar]
- Feng, Y.; Liu, Q.; Liu, A.; Qian, R.; Chen, X. A novel SSA-CCA framework for muscle artifact removal from ambulatory EEG. Virtual Reality & Intelligent Hardware 2022, 4, 1–21. [Google Scholar] [CrossRef]
- Dhull, S.K.; Singh, K.K.; et al. EEG artifact removal using canonical correlation analysis and EMD-DFA based hybrid denoising approach. Procedia Computer Science 2023, 218, 2081–2090. [Google Scholar] [CrossRef]
- Kaongoen, N.; Jo, S. Adapting Artifact Subspace Reconstruction Method for SingleChannel EEG using Signal Decomposition Techniques. In Proceedings of the 2023 45th Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), 2023; IEEE; pp. 1–4. [Google Scholar]
- Cataldo, A.; Criscuolo, S.; De Benedetto, E.; Masciullo, A.; Pesola, M.; Schiavoni, R.; Invitto, S. A method for optimizing the artifact subspace reconstruction performance in low-density EEG. IEEE Sensors Journal 2022, 22, 21257–21265. [Google Scholar] [CrossRef]
- Tsai, B.Y.; Diddi, S.V.S.; Ko, L.W.; Wang, S.J.; Chang, C.Y.; Jung, T.P. Development of an adaptive artifact subspace reconstruction based on Hebbian/anti-Hebbian learning networks for enhancing BCI performance. IEEE Transactions on Neural Networks and Learning Systems 2022, 35, 348–361. [Google Scholar] [CrossRef]
- Hazarika, D.; Vishnu, K.; Ransing, R.; Gupta, C.N. Dynamical Embedding of Single-Channel Electroencephalogram for Artifact Subspace Reconstruction. Sensors 2024, 24, 6734. [Google Scholar] [CrossRef]
- Zeng, K.; Chen, D.; Ouyang, G.; Wang, L.; Liu, X.; Li, X. An EEMD-ICA approach to enhancing artifact rejection for noisy multivariate neural data. IEEE transactions on neural systems and rehabilitation engineering 2015, 24, 630–638. [Google Scholar] [CrossRef]
- Chen, X.; Chen, Q.; Zhang, Y.; Wang, Z.J. A novel EEMD-CCA approach to removing muscle artifacts for pervasive EEG. IEEE Sensors Journal 2018, 19, 8420–8431. [Google Scholar] [CrossRef]
- Dai, Y.; Duan, F.; Feng, F.; Sun, Z.; Zhang, Y.; Caiafa, C.F.; Marti-Puig, P.; Solé-Casals, J. A fast approach to removing muscle artifacts for EEG with signal serialization based ensemble empirical mode decomposition. Entropy 2021, 23, 1170. [Google Scholar] [CrossRef]
- Egambaram, A.; Badruddin, N.; Asirvadam, V.S.; Begum, T.; Fauvet, E.; Stolz, C. FastEMD–CCA algorithm for unsupervised and fast removal of eyeblink artifacts from electroencephalogram. Biomedical Signal Processing and Control 2020, 57, 101692. [Google Scholar] [CrossRef]
- Nayak, A.B.; Shah, A.; Maheshwari, S.; Anand, V.; Chakraborty, S.; Kumar, T.S. An empirical wavelet transform-based approach for motion artifact removal in electroencephalogram signals. Decision Analytics Journal 2024. [Google Scholar] [CrossRef]
- Phadikar, S.; Sinha, N.; Ghosh, R.; Ghaderpour, E. Automatic muscle artifacts identification and removal from single-channel eeg using wavelet transform with meta-heuristically optimized non-local means filter. Sensors 2022, 22, 2948. [Google Scholar] [CrossRef]
- Dora, M.; Holcman, D. Adaptive single-channel EEG artifact removal with applications to clinical monitoring. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2022, 30, 286–295. [Google Scholar] [CrossRef]
- Sun, W.; Su, Y.; Wu, X.; Wu, X. A novel end-to-end 1D-ResCNN model to remove artifact from EEG signals. Neurocomputing 2020, 404, 108–121. [Google Scholar] [CrossRef]
- Zhang, H.; Wei, C.; Zhao, M.; Liu, Q.; Wu, H. A novel convolutional neural network model to remove muscle artifacts from EEG. In Proceedings of the ICASSP 2021-2021 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), 2021; IEEE; pp. 1265–1269. [Google Scholar]
- Zhang, Z.; Yu, X.; Rong, X.; Iwata, M. A novel multimodule neural network for eeg denoising. IEEE Access 2022, 10, 49528–49541. [Google Scholar] [CrossRef]
- Pei, Y.; Xu, J.; Chen, Q.; Wang, C.; Yu, F.; Zhang, L.; Luo, W. DTP-Net: learning to reconstruct EEG signals in time-frequency domain by multi-scale feature reuse. IEEE Journal of Biomedical and Health Informatics 2024, 28, 2662–2673. [Google Scholar] [CrossRef] [PubMed]
- Sawangjai, P.; Trakulruangroj, M.; Boonnag, C.; Piriyajitakonkij, M.; Tripathy, R.K.; Sudhawiyangkul, T.; Wilaiprasitporn, T. EEGANet: Removal of ocular artifacts from the EEG signal using generative adversarial networks. IEEE Journal of Biomedical and Health Informatics 2021, 26, 4913–4924. [Google Scholar] [CrossRef]
- Chuang, C.H.; Chang, K.Y.; Huang, C.S.; Jung, T.P. IC-U-Net: a U-Net-based denoising autoencoder using mixtures of independent components for automatic EEG artifact removal. NeuroImage 2022, 263, 119586. [Google Scholar] [CrossRef]
- Nagar, S.; Kumar, A. Orthogonal features based EEG signals denoising using fractional and compressed one-dimensional CNN AutoEncoder. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2022, 30, 2474–2485. [Google Scholar] [CrossRef]
- Dong, Y.; Tang, X.; Li, Q.; Wang, Y.; Jiang, N.; Tian, L.; Zheng, Y.; Li, X.; Zhao, S.; Li, G.; et al. An approach for EEG denoising based on wasserstein generative adversarial network. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2023, 31, 3524–3534. [Google Scholar] [CrossRef]
- Wang, S.; Luo, Y.; Shen, H. An improved Generative Adversarial Network for Denoising EEG signals of brain-computer interface systems. In Proceedings of the 2022 China Automation Congress (CAC). IEEE, 2022; pp. 6498–6502. [Google Scholar]
- Yin, J.; Liu, A.; Li, C.; Qian, R.; Chen, X. A GAN guided parallel CNN and transformer network for EEG denoising. IEEE Journal of Biomedical and Health Informatics 2023. [Google Scholar] [CrossRef]
- Lin, G.; Zhang, J.; Liu, Y.; Gao, T.; Kong, W.; Lei, X.; Qiu, T. Ballistocardiogram artifact removal in simultaneous EEG-fMRI using generative adversarial network. Journal of Neuroscience Methods 2022, 371, 109498. [Google Scholar] [CrossRef] [PubMed]
- Mahaseni, B.; Khan, N.M. EEG Signal Denoising Using Beta-Variational Autoencoder. In Proceedings of the 2024 46th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), 2024; IEEE; pp. 1–4. [Google Scholar]
- Ding, Y.; Li, L.; Li, Q. Adversarial Defense Based on Denoising Convolutional Autoencoder in EEG-Based Brain–Computer Interfaces. IEEE Access 2024. [Google Scholar]
- Wang, H.; Chen, X.; Yang, Y.; Zhou, K.; Lv, M.; Wang, D.; Zhang, W. EEG Signal Denoising Using pix2pix GAN: Enhancing Neurological Data Analysis. arXiv 2024, arXiv:2411.13288. [Google Scholar] [CrossRef]
- Choi, B.J. Removing neural signal artifacts with autoencoder-targeted adversarial transformers (AT-AT). arXiv 2025, arXiv:2502.05332. [Google Scholar] [CrossRef]
- Hjorth, B. EEG analysis based on time domain properties. Electroencephalography and clinical neurophysiology 1970, 29, 306–310. [Google Scholar] [CrossRef]
- Amin, H.U.; Malik, A.S.; Ahmad, R.F.; Badruddin, N.; Kamel, N.; Hussain, M.; Chooi, W.T. Feature extraction and classification for EEG signals using wavelet transform and machine learning techniques. Australasian physical & engineering sciences in medicine 2015, 38, 139–149. [Google Scholar]
- Subasi, A.; Ercelebi, E. Classification of EEG signals using neural network and logistic regression. Computer methods and programs in biomedicine 2005, 78, 87–99. [Google Scholar] [CrossRef]
- Acharya, U.R.; Sree, S.V.; Ang, P.C.A.; Yanti, R.; Suri, J.S. Application of non-linear and wavelet based features for the automated identification of epileptic EEG signals. International journal of neural systems 2012, 22, 1250002. [Google Scholar] [CrossRef]
- Schirrmeister, R.T.; Springenberg, J.T.; Fiederer, L.D.J.; Glasstetter, M.; Eggensperger, K.; Tangermann, M.; Hutter, F.; Burgard, W.; Ball, T. Deep learning with convolutional neural networks for EEG decoding and visualization. Human brain mapping 2017, 38, 5391–5420. [Google Scholar] [CrossRef]
- Pan, S.J.; Yang, Q. A survey on transfer learning. ieee transactions on knowledge and data engineering 2010, 22(10), 1345. [Google Scholar] [CrossRef]
- Zancanaro, A.; Cisotto, G.; Zoppis, I.; Manzoni, S.L. vEEGNet: learning latent representations to reconstruct EEG raw data via variational autoencoders. In Proceedings of the International Conference on Information and Communication Technologies for Ageing Well and e-Health, 2023; Springer; pp. 114–129. [Google Scholar]
- Lawhern, V.J.; Solon, A.J.; Waytowich, N.R.; Gordon, S.M.; Hung, C.P.; Lance, B.J. EEGNet: a compact convolutional neural network for EEG-based brain–computer interfaces. Journal of neural engineering 2018, 15, 056013. [Google Scholar] [CrossRef]
- Zhao, D.; Tang, F.; Si, B.; Feng, X. Learning joint space–time–frequency features for EEG decoding on small labeled data. Neural Networks 2019, 114, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, L.; Xia, P.; Wang, P.; Chen, X.; Du, L.; Fang, Z.; Du, M. EEG-based emotion recognition using a 2D CNN with different kernels. Bioengineering 2022, 9, 231. [Google Scholar] [CrossRef]
- Najafi, T.; Jaafar, R.; Remli, R.; Wan Zaidi, W.A. A classification model of EEG signals based on RNN-LSTM for diagnosing focal and generalized epilepsy. Sensors 2022, 22, 7269. [Google Scholar] [CrossRef] [PubMed]
- Falaschetti, L.; Biagetti, G.; Alessandrini, M.; Turchetti, C.; Luzzi, S.; Crippa, P. Multi-class detection of neurodegenerative diseases from EEG signals using lightweight lstm neural networks. Sensors 2024, 24, 6721. [Google Scholar] [CrossRef] [PubMed]
- Fu, M.; Wang, Y.; Chen, Z.; Li, J.; Xu, F.; Liu, X.; Hou, F. Deep learning in automatic sleep staging with a single channel electroencephalography. Frontiers in Physiology 2021, 12, 628502. [Google Scholar] [CrossRef]
- Liu, D.; Dai, W.; Zhang, H.; Jin, X.; Cao, J.; Kong, W. Brain-machine coupled learning method for facial emotion recognition. IEEE Transactions on Pattern Analysis and Machine Intelligence 2023, 45, 10703–10717. [Google Scholar] [CrossRef]
- Li, C.; Li, H.; Dong, X.; Zhong, X.; Cui, H.; Ji, D.; He, L.; Liu, G.; Zhou, W. CNN-Informer: A hybrid deep learning model for seizure detection on long-term EEG. Neural Networks 2025, 181, 106855. [Google Scholar] [CrossRef]
- Yang, C.; Westover, M.; Sun, J. Biot: Biosignal transformer for cross-data learning in the wild. Advances in Neural Information Processing Systems 2023, 36, 78240–78260. [Google Scholar]
- Yu, L.; Tang, P.; Jiang, Z.; Zhang, X. Denoise enhanced neural network with efficient data generation for automatic sleep stage classification of class imbalance. In Proceedings of the 2023 International Joint Conference on Neural Networks (IJCNN), 2023; IEEE; pp. 1–8. [Google Scholar]
- Duan, Y.; Hu, S.; Ma, X.; Tao, X. Multi-Class Image Generation from EEG Features with Conditional Generative Adversarial Networks. In Proceedings of the 2023 International Conference on Wireless Communications and Signal Processing (WCSP), 2023; IEEE; pp. 534–539. [Google Scholar]
- Zhao, X.; Guo, Y. Dual AxAtGAN: A Feature Intregrate BCI Model for Image Reconstruction. In Proceedings of the 2023 13th International Conference on Information Technology in Medicine and Education (ITME), 2023; IEEE; pp. 135–139. [Google Scholar]
- Sun, M.; Cui, W.; Yu, S.; Han, H.; Hu, B.; Li, Y. A dual-branch dynamic graph convolution based adaptive transformer feature fusion network for EEG emotion recognition. IEEE Transactions on Affective Computing 2022, 13, 2218–2228. [Google Scholar] [CrossRef]
- Wang, Y.; Cui, W.; Yu, T.; Li, X.; Liao, X.; Li, Y. Dynamic multi-graph convolution-based channel-weighted transformer feature fusion network for epileptic seizure prediction. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2023, 31, 4266–4277. [Google Scholar] [CrossRef]
- kumar Ravikanti, D.; Saravanan, S. EEGAlzheimer’sNet: Development of transformer-based attention long short term memory network for detecting Alzheimer disease using EEG signal. Biomedical Signal Processing and Control 2023, 86, 105318. [Google Scholar] [CrossRef]
- Qin, Y.; Zhang, W.; Tao, X. TBEEG: A two-branch manifold domain enhanced transformer algorithm for learning EEG decoding. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2024, 32, 1466–1476. [Google Scholar] [CrossRef] [PubMed]
- Mohsenvand, M.N.; Izadi, M.R.; Maes, P. Contrastive representation learning for electroencephalogram classification. In Proceedings of the Machine learning for health. PMLR, 2020; pp. 238–253. [Google Scholar]
- Kostas, D.; Aroca-Ouellette, S.; Rudzicz, F. BENDR: Using transformers and a contrastive self-supervised learning task to learn from massive amounts of EEG data. Frontiers in Human Neuroscience 2021, 15, 653659. [Google Scholar] [CrossRef]
- Nørskov, A.; Neergaard Zahid, A.; Mørup, M. Cslp-ae: A contrastive split-latent permutation autoencoder framework for zero-shot electroencephalography signal conversion. Advances in Neural Information Processing Systems 2023, 36, 13179–13199. [Google Scholar]
- Mohammadi Foumani, N.; Mackellar, G.; Ghane, S.; Irtza, S.; Nguyen, N.; Salehi, M. Eeg2rep: enhancing self-supervised eeg representation through informative masked inputs. In Proceedings of the Proceedings of the 30th ACM SIGKDD Conference on Knowledge Discovery and Data Mining, 2024; pp. 5544–5555. [Google Scholar]
- Jiang, W.B.; Zhao, L.M.; Lu, B.L. Large brain model for learning generic representations with tremendous EEG data in BCI. arXiv 2024. arXiv:2405.18765. [CrossRef]
- Wang, G.; Liu, W.; He, Y.; Xu, C.; Ma, L.; Li, H. Eegpt: Pretrained transformer for universal and reliable representation of eeg signals. Advances in Neural Information Processing Systems 2024, 37, 39249–39280. [Google Scholar]
- Chowdhury, M.R.; Ding, Y.; Sen, S. SSL-SE-EEG: A Framework for Robust Learning from Unlabeled EEG Data with Self-Supervised Learning and Squeeze-Excitation Networks. arXiv 2025, arXiv:2510.19829. [Google Scholar]
- Luo, T.j.; Fan, Y.; Chen, L.; Guo, G.; Zhou, C. EEG signal reconstruction using a generative adversarial network with wasserstein distance and temporal-spatial-frequency loss. Frontiers in neuroinformatics 2020, 14, 15. [Google Scholar] [CrossRef]
- Uraechu, U. Self-Supervised Contrastive Learning Using EEG Signals For Mental Stress Assessment. 2024. [Google Scholar]
- Zhou, Y.; Zhao, S.; Wang, J.; Jiang, H.; Li, S.; Li, T.; Pan, G. SPICED: A Synaptic Homeostasis-Inspired Framework for Unsupervised Continual EEG Decoding. arXiv 2025, arXiv:2509.17439. [Google Scholar] [CrossRef]
- Chen, R.; Xie, C.; Zhang, J.; You, Q.; Pan, J. A Progressive Multi-Domain Adaptation Network With Reinforced Self-Constructed Graphs for Cross-Subject EEG-Based Emotion and Consciousness Recognition. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2025. [Google Scholar]
- Kim, M.; Bae, J.; Wang, B.; Ko, H.; Lim, J.S. Feature selection method using multi-agent reinforcement learning based on guide agents. Sensors 2022, 23, 98. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Shang, S.; Tang, L.; He, L.; Zhou, M. EEG channel selection algorithm based on Reinforcement Learning. In Proceedings of the 2022 IEEE International Conference on Networking, Sensing and Control (ICNSC), 2022; IEEE; pp. 1–6. [Google Scholar]
- Li, J.; Wu, C.; Pan, J.; Wang, F. Few-shot EEG sleep staging based on transductive prototype optimization network. Frontiers in Neuroinformatics 2023, 17, 1297874. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Chen, P.; Zhang, M.; Zhang, J.; Sun, X.; Li, M.; Yang, X.; Gao, Z. EEG-based motor imagery recognition framework via multisubject dynamic transfer and iterative self-training. IEEE Transactions on Neural Networks and Learning Systems 2023, 35, 10698–10712. [Google Scholar] [CrossRef] [PubMed]
- Mellot, A.; Collas, A.; Chevallier, S.; Engemann, D.; Gramfort, A. Physics-informed and unsupervised riemannian domain adaptation for machine learning on heterogeneous EEG datasets. In Proceedings of the 2024 32nd European Signal Processing Conference (EUSIPCO). IEEE, 2024; pp. 1367–1371. [Google Scholar]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: components of a new research resource for complex physiologic signals. circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, X.; Gao, X.; Gao, S. A benchmark dataset for SSVEP-based brain–computer interfaces. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2016, 25, 1746–1752. [Google Scholar] [CrossRef]
- Zheng, W.L.; Lu, B.L. Investigating critical frequency bands and channels for EEG-based emotion recognition with deep neural networks. IEEE Transactions on autonomous mental development 2015, 7, 162–175. [Google Scholar] [CrossRef]
- Brunner, C.; Leeb, R.; Müller-Putz, G.; Schlögl, A.; Pfurtscheller, G. BCI Competition 2008–Graz data set A. Institute for knowledge discovery (laboratory of brain-computer interfaces), Graz University of Technology 2008, 16, 34. [Google Scholar]
- Leeb, R.; Brunner, C.; Müller-Putz, G.; Schlögl, A.; Pfurtscheller, G. BCI Competition 2008–Graz data set B. Graz University of Technology, Austria 2008, 16, 1–6. [Google Scholar]
- Obeid, I.; Picone, J. The temple university hospital EEG data corpus. Frontiers in neuroscience 2016, 10, 196. [Google Scholar] [CrossRef]
| Model | Year | Structure | Core Contribution | Dataset | Downstream Tasks | Code Link |
|---|---|---|---|---|---|---|
| SeqCLR[74] | 2020 | SimCLR | Channel permutation and dataset fusion, contrastive learning | SEED,TUAB, SleepEDF et al. | subset of pre-training | None |
| BENDR [75] | 2021 | Transformer+1D-CNN | Contrastive pre-training for general EEG feature learning | TUEG | MMI,BCIC,ERN et al. | BENDR |
| CSLP-AE [76] | 2023 | Transformer+1D-CNN | Split latent space, contrastive learning and latent permutation loss | ERP PhysioNet | subset of pre-training | CSLP-AE |
| BIOT [66] | 2023 | Transformer | Biosignal tokenization, adapts to channel mismatch/missing values | SHHS,PREST et al. | CHB-MIT,TUAB,PTB-XL et al. | BIOT |
| EEG2Rep [77] | 2024 | CNN+Transformer | SSP masking; predict masked inputs in latent space | DREAMER,TUAB et al. | Emotion/mental workload/abnormal et al. | EEG2Rep |
| LaBraM [78] | 2024 | Transformer+1D-CNN | 2500h multi-dataset pre-training, channel patching and symmetric masking | BCIC,SEED | TUAB,TUEV,SEED-V et al. | LaBraM |
| EEGPT [79] | 2024 | Transformer+ LSTE | Dual self-supervision, local spatiotemporal embedding | PhysioMI,SEED et al. | BCIC-2A/2B,Sleep-EDFx et al. | EEGPT |
| SSL-SE [80] | 2025 | CNN+SE-Net | EEG-to-2D RGB; SSL+SE-Net for less labeled data, noise robustness, low power | MindBigData,TUAB | Digit/EEG abnormal/emotion et al. | None |
| Task | Typical cue | Main risk | Recommended starting |
|---|---|---|---|
| MI | subtle ERD/ERS | over-denoising task rhythms | Minimal preprocessing; add ICA/EEMD/WT only if artifacts dominate. |
| ERP | time-locked components | latency/shape distortion | Baseline is often sufficient; avoid aggressive smoothing/reconstruction. |
| Abnormal | morphology and broad spectral patterns | lower,but possible distribution shift | Try moderate EEMD/WT; validate generalization and report efficiency. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




