1. Introduction
Antimicrobial resistance (AMR) is traditionally defined and detected through the presence of genetic determinants that confer reduced susceptibility to specific antibiotics. Advances in whole-genome sequencing (WGS) have significantly improved the resolution at which resistance genes, mutations, and mobile genetic elements can be identified, leading to widespread adoption of genomics-based AMR prediction in both research and clinical microbiology (Didelot and Maiden 2010; Ellington et al. 2017; Shahid et al. 2024). However, despite these advances, substantial discordance remains between predicted resistance genotypes and observed treatment outcomes in patients (Band and Weiss 2019; Ransom et al. 2020). Clinically, infections caused by pathogens with similar or even identical resistance gene profiles often respond differently to the same antimicrobial therapy, highlighting a persistent gap between genomic prediction and patient-specific response (Band and Weiss 2019; Su et al. 2019).
One key limitation of current AMR frameworks is their reliance on static genetic information to explain what is, in practice, a dynamic biological phenomenon. The presence of a resistance gene does not necessarily imply its expression, nor does it capture the magnitude, timing, or regulation of resistance-associated pathways during infection (Brauner et al. 2016; Andersson and Hughes 2014). Increasing evidence suggests that resistance phenotypes are shaped not only by genetic potential but also by transcriptional regulation, metabolic state, environmental cues, and host-derived pressures (Avican et al. 2021; El Meouche et al. 2024; Fang et al. 2016). These factors collectively influence whether resistance mechanisms are activated, suppressed, or bypassed under clinically relevant conditions (Brauner et al. 2016; El Meouche et al. 2024).
Precision medicine aims to tailor diagnosis and treatment based on individual variability in biological states rather than population averages (Collins and Varmus 2015). While this paradigm has gained traction in oncology and genetic disease, its integration into infectious disease management has been comparatively limited (van Belkum et al. 2020). In the context of AMR, precision medicine requires moving beyond binary classifications of “resistant” or “susceptible” toward an understanding of context-dependent resistance behavior within a specific host, at a specific time, and under specific treatment conditions (Balaban et al. 2019; Hassall et al. 2024). Current gene-centric AMR diagnostics are poorly equipped to capture this level of complexity (Su et al. 2019; Hassall et al. 2024).
Transcriptomics offers a complementary layer of information that directly reflects functional activity within microbial cells. By measuring gene expression, transcriptomic approaches can reveal which resistance pathways are actively engaged, how bacteria adapt transcriptionally to antibiotic exposure, and how stress-response programs contribute to survival without permanent genetic change (Brauner et al. 2016; Avican et al. 2021; Croucher and Thomson 2010). Importantly, transcriptomic states can vary rapidly and reversibly, allowing genetically similar pathogens to exhibit markedly different phenotypes across patients or stages of infection (Windels et al. 2019).
In this review, we argue that antimicrobial resistance should be conceptualized not solely as a genetic trait but as a dynamic resistance state emerging from regulated gene expression within specific physiological contexts (Brauner et al. 2016; El Meouche et al. 2024; Balaban et al. 2019). We synthesize current evidence demonstrating how transcriptomic data provide mechanistic insights into resistance variability that genomics alone cannot explain (Avican et al. 2021; Croucher and Thomson 2010). We further examine how integrating transcriptomics into AMR frameworks can support precision diagnostics and more individualized antimicrobial therapy, while critically addressing the technical, clinical, and translational challenges that must be overcome for real-world implementation (Hassall et al. 2024; Byron et al. 2016).
2. Resistance Genes vs Resistance States: A Conceptual Shift in Precision AMR
The dominant framework for antimicrobial resistance has historically focused on the identification of genetic determinants that reduce antibiotic susceptibility. Resistance genes, target-site mutations, and mobile genetic elements form the basis of most molecular AMR diagnostics and surveillance efforts. While this gene-centric paradigm has proven valuable for population-level monitoring and outbreak investigation, it provides an incomplete explanation for the variability observed in clinical treatment responses. Increasing experimental and clinical evidence indicates that resistance phenotypes are not solely dictated by genetic content, but also by regulated physiological states that can emerge transiently in response to environmental and host-associated pressures.
2.1. Defining Resistance States
A resistance state can be defined as a condition in which bacterial survival under antibiotic exposure is mediated by context-dependent activation of molecular pathways, rather than by constitutive genetic resistance alone. These states are often characterized by reversible transcriptional programs that alter cellular processes such as metabolism, membrane permeability, stress response, and growth rate. Importantly, resistance states may arise even in the absence of classical resistance genes, and conversely, the presence of resistance genes does not guarantee their functional engagement.
Several well-described phenomena fall within this conceptual framework. Antibiotic tolerance, for example, refers to the ability of bacteria to survive transient antibiotic exposure without a change in minimum inhibitory concentration (MIC), often through growth arrest or stress-response activation rather than genetic resistance (Brauner et al. 2016; Levin-Reisman et al. 2019). Similarly, persister cells represent a subpopulation that enters a dormant or low-metabolic state, enabling survival under high antibiotic concentrations despite genetic susceptibility (Lewis 2010). These phenotypes are increasingly recognized as transcriptionally regulated and environmentally triggered, rather than genetically fixed.
In addition, heteroresistance (the coexistence of subpopulations with differing susceptibilities within an isogenic bacterial population) illustrates how phenotypic resistance can arise from differential gene expression and regulatory noise (Band and Weiss 2019). Such heterogeneity can lead to treatment failure even when standard susceptibility testing indicates sensitivity. Together, these examples demonstrate that resistance is not a binary property encoded solely in the genome, but a spectrum of adaptive states shaped by regulatory and physiological processes. To clarify the conceptual distinction between gene-centric and state-based views of antimicrobial resistance, key defining features of resistance genes and resistance states are summarized in
Table 1.
2.2. Determinants of Resistance States
Resistance states are influenced by multiple interacting factors that vary across patients and infection contexts. Antibiotic exposure itself is a major driver, as sub-inhibitory or fluctuating drug concentrations can induce specific transcriptional responses that promote survival without selecting for stable genetic resistance (Andersson and Hughes 2014). These responses often involve the upregulation of efflux pumps, modulation of cell envelope composition, and activation of global stress regulators.
Host-derived pressures further shape bacterial transcriptional states during infection. Immune-mediated stresses such as oxidative bursts, nutrient limitation, and antimicrobial peptides have been shown to induce protective gene expression programs that overlap with antibiotic tolerance pathways (Fang et al. 2016). As a result, bacterial behavior observed under standard in vitro conditions may poorly reflect in vivo resistance dynamics.
The local infection microenvironment also plays a critical role. Oxygen availability, pH, nutrient gradients, and biofilm formation can profoundly alter bacterial transcriptional profiles, leading to resistance-associated phenotypes that are not predicted by genotype alone (Stewart and Franklin 2008). Biofilm-associated cells exhibit distinct transcriptional states associated with reduced growth and increased stress tolerance, contributing to persistent infections.
Finally, the microbiome context can modulate resistance states indirectly through metabolic interactions and signaling molecules. Community-level effects have been shown to influence antibiotic efficacy and pathogen survival, although the transcriptional mechanisms underlying these interactions remain incompletely understood (Buffie and Pamer 2013). These observations further reinforce the need for resistance frameworks that incorporate functional and contextual information. This conceptual shift from static, gene-centric resistance prediction toward a dynamic, context-dependent understanding of antimicrobial resistance is illustrated in
Figure 1.
2.3. Implications for Precision Medicine
Reframing antimicrobial resistance as a dynamic resistance state has important implications for precision medicine. It highlights why static detection of resistance genes may be insufficient to predict treatment outcomes at the individual patient level. More importantly, it provides a conceptual basis for integrating transcriptomic data into AMR diagnostics, enabling assessment of whether resistance pathways are actively engaged under clinically relevant conditions.
By focusing on resistance states rather than genes alone, precision medicine approaches can begin to account for temporal variability, host-pathogen interactions, and environmental influences that shape antimicrobial response. This shift does not negate the importance of genomics but rather positions it as one component within a broader, functionally informed framework.
3. What Pathogen Transcriptomics Adds Beyond Genomics
Genomic methods (principally WGS) have transformed our ability to catalogue resistance loci, mobile elements and insertion sequences that underlie classical AMR mechanisms. However, genomes encode potential, not current activity. Transcriptomics directly measures gene expression and therefore provides a time-resolved readout of which pathways are engaged under particular conditions - information that is essential for understanding dynamic resistance behaviours such as tolerance, persistence, inducible efflux, and stress-response activation.
3.1. Mechanistic Insights Uniquely Revealed by Transcriptomics
Transcriptome profiling reveals regulatory and physiological responses that cannot be inferred from static sequence data alone. Antibiotic exposure frequently induces rapid transcriptional activation of efflux systems, modifying enzymes, and global stress-response regulators that mediate transient survival without stable genetic change (Martínez and Rojo 2011; Bie et al. 2023). Large-scale stress-response atlases across human bacterial pathogens demonstrate conserved transcriptional programs activated under host-like stresses, highlighting how transcriptional reprogramming shapes survival strategies in clinically relevant contexts (Avican et al. 2021).
RNA-level signatures further capture metabolic rerouting, including changes in ATP production and central carbon flux, modulation of ribosomal and translational machinery, and the expression of regulatory small RNAs that fine-tune stress responses. These transcriptionally regulated processes can substantially alter antimicrobial susceptibility independent of genotype, helping to explain divergent phenotypes among genetically similar isolates during infection (Martínez and Rojo 2011; El Meouche et al. 2024).
Beyond canonical resistance mechanisms, transcriptomics uncovers conditional activation of pathways that compensate for antibiotic action. For example, β-lactam exposure induces transcriptional programs associated with cell-wall remodeling, whereas fluoroquinolone treatment activates oxidative stress defense pathways that promote survival (Martínez and Rojo 2011; Bie et al. 2023). Importantly, time-series transcriptomic studies demonstrate that such adaptive expression programs can arise before the fixation of stable resistance mutations, suggesting that early transcriptional states may define reversible phases of resistance evolution (Kirchner et al. 2024). These adaptive responses are frequently coordinated by global regulators whose transient activation would not be detectable through genomic analysis alone. Such transcriptionally regulated resistance programs do not operate in isolation but form overlapping and coordinated physiological states within bacterial cells. The major transcriptional architectures underlying inducible resistance, tolerance, and persistence are summarized schematically in
Figure 2.
3.2. Transcriptomic Signatures Under Antibiotic Stress: Empirical Evidence
Comparative RNA-sequencing studies across antibiotic classes and bacterial species reveal both drug-specific and shared transcriptional responses. In Escherichia coli and other Enterobacterales, antibiotic exposure drives differential expression of genes involved in efflux, redox balance, and central metabolism, with these transcriptional changes correlating with phenotypic tolerance under laboratory conditions (Bie et al. 2023). Such findings indicate that transcriptional remodeling contributes substantially to resistance-associated phenotypes beyond the presence or absence of resistance genes. Transcriptomic atlases generated under host-mimicking stressors (including nutrient limitation, oxygen restriction, and pH variation) show partial overlap with in vivo expression profiles, supporting the relevance of carefully designed in vitro models for studying infection-associated resistance states (Avican et al. 2021). However, these studies also reveal context-specific expression programs that are absent under standard laboratory growth conditions, underscoring the importance of physiological relevance in transcriptomic analyses.
Integrated transcriptomic-metabolomic and transcriptomic-proteomic studies further demonstrate coordinated rewiring across molecular layers in response to antibiotic exposure. These analyses consistently identify metabolic shifts, such as altered glycolytic and tricarboxylic acid cycle fluxes, that accompany transcriptional programs associated with survival under drug pressure (Liao et al. 2024). Such coordinated changes are mechanistically informative because they point to metabolic bottlenecks and co-dependencies that may be exploited to reverse tolerance or potentiate antibiotic activity (Martínez and Rojo 2011). Collectively, these studies indicate that diverse resistance-associated behaviors are underpinned by distinct but recurring transcriptional programs. To synthesize these findings and clarify how resistance states can be functionally inferred from transcriptomic data, key resistance-associated states, their dominant expression programs, and potential clinical implications are summarized in
Table 2.
3.3. Distinguishing In Vitro and Host-Relevant Transcriptional States
A major translational challenge is that many transcriptomic studies are performed under simplified laboratory conditions that do not fully recapitulate the infection microenvironment, including immune pressures, spatial heterogeneity, and interspecies interactions. Comparative analyses show both overlaps and important discrepancies between in vitro stress responses and transcriptional profiles observed in host-associated or in vivo settings (Avican et al. 2021).
Host-derived stresses such as oxidative and nitrosative stress, iron limitation, and exposure to antimicrobial peptides can induce transcriptional programs that significantly modulate antibiotic susceptibility and are often under-represented in standard culture-based transcriptomes. Dual RNA sequencing approaches, which simultaneously profile host and pathogen transcriptomes, have provided direct evidence that host-derived signals actively shape pathogen gene expression during infection. These studies demonstrate coordinated host-pathogen transcriptional dynamics that influence bacterial stress responses, metabolic adaptation, and survival under antibiotic treatment (Macho Rendón et al. 2021; Hu et al. 2024).
3.4. Classifying Resistance States: Toward Minimal Predictive Signatures
Several recent studies have explored whether compact transcriptomic signatures can classify resistant versus susceptible phenotypes or predict antimicrobial treatment outcomes. Expression-based classifiers using relatively small gene panels have shown promising accuracy in controlled experimental datasets, suggesting that limited transcriptional programs may capture essential features of resistance states (Shahreen et al. 2025). However, the generalizability of these signatures across strains, infection sites, and host backgrounds remains an unresolved challenge, and rigorous validation using diverse clinical cohorts is required before diagnostic application.
Importantly, interpretable and biologically informed models are more likely to be suitable for clinical translation than purely black-box approaches. Mechanistic interpretability enables clinicians to contextualize predictions within known resistance pathways rather than relying solely on algorithmic output (Toprak et al. 2011; Ching et al. 2018).
3.5. Limitations and Technical Considerations
Transcriptomics presents technical challenges that constrain clinical translation. Bacterial RNA is inherently labile, and pathogen-derived transcripts often constitute a small fraction of total nucleic acid in clinical samples dominated by host RNA. Low pathogen biomass, rapid temporal changes in gene expression, and within-host heterogeneity such as biofilm-associated niches further complicate signal detection and interpretation (Joubert et al. 2025).
Additional sources of variability include batch effects, growth-phase differences, and sample handling inconsistencies, all of which can confound biological interpretation if not carefully controlled. Standardized protocols for RNA isolation, stabilization, sequencing, and data normalization, particularly for mixed host-pathogen samples, are therefore essential for reproducible and clinically meaningful transcriptomic measurements. Moreover, because transcriptomic states are transient, diagnostic interpretation must account for sampling time relative to infection progression and antimicrobial exposure (Windels et al. 2019; Andersson and Hughes 2014).
4. Host and Environmental Modulation of Resistance States
Antimicrobial resistance does not emerge or operate in isolation from the host environment. During infection, bacterial pathogens experience a complex and dynamic set of host-derived and environmental pressures that substantially influence gene expression, metabolic activity, and survival strategies. These pressures shape resistance states by modulating transcriptional programs that determine whether resistance mechanisms are activated, suppressed, or bypassed under antimicrobial treatment. Understanding these host- and environment-dependent effects is therefore essential for situating pathogen transcriptomics within a precision medicine framework.
4.1. Host Immune Pressures and Transcriptional Adaptation
Host immune responses impose multiple stresses on invading bacteria, including oxidative and nitrosative stress, nutrient sequestration, pH changes, and exposure to antimicrobial peptides. These stresses trigger well-characterized bacterial transcriptional responses that overlap with pathways involved in antibiotic tolerance and persistence. For example, oxidative stress generated by host phagocytes activates transcriptional programs that enhance DNA repair, redox homeostasis, and metabolic flexibility, which can indirectly reduce antibiotic susceptibility (Fang et al. 2016; Dawan and Ahn 2022).
Iron limitation is another prominent host-imposed constraint that reshapes bacterial transcriptional states. Host iron-sequestration mechanisms induce expression of siderophore systems and global regulatory changes that alter metabolic and respiratory pathways. Such iron-responsive transcriptional programs have been shown to modulate sensitivity to multiple antibiotic classes, linking nutritional immunity directly to resistance-associated phenotypes (McDevitt et al. 2011). These findings highlight that host immunity does not merely restrict bacterial growth but actively shapes the functional state of resistance pathways.
Importantly, immune-mediated stresses vary substantially between patients depending on immune status, inflammation level, and infection site. This variability provides a plausible mechanistic basis for patient-specific resistance states that are not predictable from pathogen genotype alone.
4.2. Infection Site Physiology and Spatial Heterogeneity
The physiological conditions encountered by bacteria differ markedly across infection sites, influencing transcriptional states relevant to antimicrobial response. Oxygen availability, nutrient composition, osmolarity, and pH vary between tissues and even within microenvironments of the same infection. Such spatial heterogeneity is particularly evident in chronic and device-associated infections, where bacteria frequently adopt biofilm-associated lifestyles.
Biofilm growth is associated with transcriptional programs characterized by reduced growth rates, altered metabolism, and increased stress tolerance. These expression states contribute to decreased antibiotic susceptibility without requiring genetic resistance, and they often coexist with planktonic populations exhibiting distinct transcriptional profiles within the same host (Stewart and Franklin 2008). As a result, a single infection may encompass multiple resistance states simultaneously, complicating treatment and contributing to relapse.
Transcriptomic analyses of bacteria grown under oxygen-limited or nutrient-restricted conditions further demonstrate that environmental constraints alone can induce tolerance-associated gene expression programs. These findings emphasize that resistance states are emergent properties of local physiology rather than fixed attributes of the pathogen.
4.3. Microbiome-Mediated Modulation of Resistance States
The surrounding microbial community represents an additional layer of environmental influence on pathogen transcriptional states. Commensal microbes can alter antibiotic efficacy through metabolic interactions, competition for nutrients, and production of signaling molecules. These interactions may indirectly induce or suppress resistance-associated transcriptional programs in pathogens.
Experimental and clinical studies demonstrate that microbiota composition can influence pathogen susceptibility to antibiotics, although the underlying transcriptional mechanisms remain incompletely characterized. For example, microbiome-driven metabolic cross-feeding can sustain pathogen growth under antibiotic stress, while community-level depletion of specific metabolites may sensitize pathogens to treatment (Buffie and Pamer 2013). Such effects suggest that resistance states may be shaped not only by host-pathogen interactions but also by broader ecosystem-level dynamics.
Importantly, microbiome composition varies widely across individuals due to diet, prior antibiotic exposure, geography, and health status. This variability reinforces the need for precision approaches that consider microbiome context when interpreting pathogen transcriptomic data (Shahid 2025).
4.4. Evidence from Host-Relevant and Dual RNA-Seq Studies
Dual RNA sequencing has provided direct evidence that host-derived signals actively shape pathogen transcriptional states during infection. By simultaneously profiling host and bacterial transcriptomes, these studies reveal coordinated responses in which host immune activation correlates with pathogen stress-response and metabolic gene expression (Westermann et al. 2012; Hu et al. 2024). Such datasets demonstrate that resistance-associated transcriptional programs cannot be fully understood without reference to the host environment in which they arise.
Host-relevant transcriptomic studies further show that bacterial gene expression profiles measured in vivo or in host-mimicking models differ substantially from those observed under standard laboratory conditions. These discrepancies underscore the limitations of extrapolating resistance mechanisms from simplified in vitro systems and highlight the importance of physiologically informed experimental designs for identifying clinically relevant resistance states (Fang et al. 2016; Westermann et al. 2012).
4.5. Implications for Precision Antimicrobial Therapy
Collectively, these observations indicate that resistance states are shaped by an interplay of host immunity, infection site physiology, and microbial community context. From a precision medicine perspective, this implies that effective antimicrobial therapy must account for patient-specific physiological and environmental factors in addition to pathogen genotype.
Integrating pathogen transcriptomics with host and environmental context offers a route toward more individualized treatment strategies. For example, expression signatures reflecting immune-induced stress or biofilm-associated states could inform antibiotic selection, dosing, or combination therapy. However, translating such insights into clinical practice will require systematic validation across diverse patient populations and infection types.
5. Transcriptomics-Informed Precision AMR Diagnostics
Precision medicine requires diagnostic tools that capture functional biological states rather than static markers alone. In the context of antimicrobial resistance, current diagnostics largely rely on phenotypic susceptibility testing or genomic detection of resistance determinants. While these approaches are indispensable, they provide limited insight into whether resistance mechanisms are actively engaged during infection or how bacterial responses evolve over the course of treatment. Transcriptomics offers an opportunity to complement existing diagnostics by enabling state-aware assessment of resistance pathways, potentially supporting more individualized antimicrobial decision-making.
5.1. Moving Beyond Binary Resistance Classification
Conventional antimicrobial susceptibility testing classifies pathogens as “susceptible,” “intermediate,” or “resistant” based on standardized breakpoints, implicitly assuming stable resistance phenotypes. However, transcriptionally mediated resistance states such as tolerance, persistence, or inducible resistance can enable bacterial survival despite apparent susceptibility in vitro. Transcriptomic profiling provides a means to detect activation of resistance-associated pathways even when classical resistance genes are absent or phenotypic resistance thresholds are not exceeded.
Several studies have demonstrated that expression levels of efflux systems, stress-response regulators, and metabolic pathways correlate with survival under antibiotic exposure, suggesting that resistance states may be inferred from gene expression patterns rather than genotype alone (Martínez and Rojo 2011; El Meouche et al. 2024). Importantly, transcriptomic readouts can capture reversible and temporal adaptations that are invisible to genomic diagnostics, offering a functional layer of information relevant to treatment outcome.
5.2. Expression-Based Classifiers and Resistance State Signatures
Recent work has explored whether reduced transcriptomic signatures can classify antimicrobial resistance phenotypes or predict treatment response. Using machine-learning and statistical approaches, several studies have shown that relatively small gene sets, often on the order of tens of transcripts, can distinguish resistant from susceptible states under controlled experimental conditions (Shahreen et al. 2025). These findings indicate that resistance states may be encoded in compact transcriptional programs rather than requiring whole-transcriptome profiling.
However, the clinical applicability of expression-based classifiers remains limited by issues of generalizability. Many reported models are trained on specific species, strains, or experimental conditions and have not been validated across diverse clinical isolates or infection contexts. Moreover, purely data-driven classifiers may lack mechanistic interpretability, therefore reduce clinician confidence and complicate regulatory approval. Approaches that integrate pathway-level information or constrain models using known resistance mechanisms may improve robustness and translational potential (Shahreen et al. 2025).
5.3. Validation Requirements for Clinical Translation
For transcriptomics-informed diagnostics to be clinically useful, rigorous validation is essential. First, candidate expression signatures must be reproducible across independent datasets and robust to technical variability in RNA extraction, sequencing depth, and normalization. Second, validation should prioritize patient-derived isolates and clinically relevant conditions, including antibiotic exposure levels and host-associated stresses.
Longitudinal sampling represents an additional requirement for validating resistance state diagnostics. Because transcriptional responses can change rapidly during treatment, static single-time-point measurements may be insufficient to guide therapy. Studies incorporating time-series transcriptomic data have demonstrated dynamic shifts in resistance-associated expression programs, underscoring the importance of temporal context when interpreting diagnostic readouts (Kirchner et al. 2024).
Finally, diagnostic performance must be evaluated against clinically meaningful endpoints, such as treatment failure, relapse, or time to clearance, rather than laboratory susceptibility alone. Without such outcome-based validation, transcriptomic diagnostics risk remaining exploratory rather than actionable. To consolidate the translational opportunities and constraints associated with transcriptomics-informed antimicrobial resistance diagnostics, key diagnostic dimensions, added value, and implementation challenges are summarized in
Table 3.
5.4. Practical Considerations for Implementation
Despite advances in sequencing technologies, several practical challenges constrain the implementation of transcriptomic diagnostics in routine clinical settings. RNA instability, low pathogen biomass, and host RNA contamination complicate sample processing, particularly for blood and tissue specimens. Targeted expression panels focusing on predefined resistance-associated transcripts may offer a pragmatic alternative to whole-transcriptome sequencing, reducing cost and turnaround time while retaining functional insight (Joubert et al. 2025; Shahid 2025b).
Integration of transcriptomic diagnostics into existing clinical workflows will also require careful consideration of reporting formats and decision thresholds. Rather than replacing established susceptibility testing, transcriptomic data are more likely to serve as an adjunct, informing decisions such as antibiotic escalation, combination therapy, or treatment duration. Clear interpretive frameworks will be necessary to translate expression-based readouts into actionable clinical recommendations.
5.5. Ethical and Regulatory Considerations
The introduction of transcriptomics into AMR diagnostics raises ethical and regulatory questions related to data interpretation, responsibility, and equity of access. Expression-based diagnostics may generate probabilistic or state-dependent outputs rather than binary classifications, requiring transparent communication of uncertainty to clinicians and patients. Additionally, regulatory pathways for RNA-based diagnostics remain less established than those for genomic assays, emphasizing the need for standardized validation criteria and reporting guidelines.
From a precision medicine perspective, these considerations reinforce the importance of aligning technological innovation with clinical feasibility and governance frameworks. Transcriptomics-informed AMR diagnostics should be developed not as standalone tools, but as components of integrated decision-support systems grounded in validated biological mechanisms and real-world evidence (Joubert et al. 2025).
6. Implications for Personalized Antimicrobial Therapy
The recognition that antimicrobial resistance manifests as a dynamic, context-dependent state rather than a fixed genetic trait has important implications for how antimicrobial therapy is selected, optimized, and monitored. Integrating transcriptomic information into therapeutic decision-making offers a means to align treatment strategies with the functional state of the pathogen within an individual patient, while remaining grounded in existing clinical frameworks.
6.1. Antibiotic Selection and Therapeutic Stratification
Current antimicrobial selection is largely guided by phenotypic susceptibility testing and, increasingly, by genomic prediction of resistance determinants. However, these approaches provide limited insight into whether resistance pathways are actively expressed during infection. Transcriptomic data can complement these methods by indicating the activation state of efflux systems, stress-response regulators, or metabolic adaptations that may reduce antibiotic efficacy despite apparent susceptibility (Martínez and Rojo 2011; El Meouche et al. 2024).
From a precision medicine perspective, such information could support stratified therapeutic decisions. For example, detection of transcriptional signatures associated with tolerance or stress adaptation may justify the use of antibiotics with alternative mechanisms of action, combination therapy, or adjunctive agents targeting metabolic or stress-response pathways. Importantly, transcriptomic insights should be interpreted alongside conventional diagnostics rather than in isolation, serving as an additional layer of functional evidence rather than a replacement for established standards of care.
6.2. Optimizing Treatment Duration and Monitoring Response
Treatment duration remains a major source of variability in antimicrobial therapy, often relying on standardized protocols rather than individualized assessment of bacterial clearance. Transcriptomic profiling offers a potential means to monitor changes in bacterial physiological state during therapy. Downregulation of stress-response and survival-associated transcriptional programs may indicate effective bacterial killing, whereas persistence of such signatures could signal incomplete eradication or elevated risk of relapse (Shahreen et al. 2025).
Time-resolved transcriptomic studies demonstrate that resistance-associated expression programs can shift rapidly in response to antibiotic exposure, emphasizing the importance of temporal context when interpreting these data (Kirchner et al. 2024). While routine longitudinal transcriptomic monitoring is not currently feasible in most clinical settings, these observations highlight a conceptual framework in which treatment duration could eventually be informed by functional response rather than fixed time points alone.
6.3. Preventing Resistance Emergence Through State-Aware Therapy
Antimicrobial exposure is a key driver of resistance evolution, particularly when bacteria survive treatment through transcriptionally mediated tolerance or persistence. By identifying resistance states associated with survival rather than outright resistance, transcriptomic data may help inform strategies to minimize the selective pressures that promote stable resistance mutations.
For example, recognition of metabolic or stress-response states linked to tolerance could guide the use of metabolic adjuvants or treatment regimens designed to limit the survival of non-growing or slow-growing subpopulations (Martínez and Rojo 2011; Lobritz et al. 2015). Such state-aware therapeutic approaches align with broader goals of antimicrobial stewardship by seeking not only to treat infection effectively but also to reduce the long-term risk of resistance emergence.
Figure 3 outlines a pragmatic workflow for integrating transcriptomic resistance-state profiling into existing antimicrobial treatment pipelines.
6.4. Clinical Feasibility and Translational Boundaries
Despite its conceptual promise, the incorporation of transcriptomics into personalized antimicrobial therapy faces practical constraints. Turnaround time, cost, and sample quality remain significant barriers, and most transcriptomic insights currently derive from research settings rather than routine clinical practice. As such, near-term applications are likely to focus on high-risk or refractory infections, where additional functional information may justify increased diagnostic complexity.
Crucially, the translation of transcriptomic insights into therapy must be supported by evidence demonstrating improved clinical outcomes. Without such validation, transcriptomics risks remaining an informative but non-actionable layer. Precision antimicrobial therapy informed by transcriptomics should therefore be pursued through carefully designed clinical studies that integrate molecular data with treatment decisions and patient outcomes (Shahreen et al. 2025).
7. Knowledge Gaps and Future Directions
Despite growing evidence that transcriptionally mediated resistance states play an important role in antimicrobial treatment outcomes, significant conceptual, technical, and translational gaps remain. Addressing these limitations is essential before transcriptomics can be meaningfully integrated into precision antimicrobial resistance (AMR) frameworks.
7.1. Defining and Standardizing Resistance States
A central unresolved challenge is the lack of standardized definitions for resistance states at the transcriptional level. While concepts such as tolerance, persistence, and heteroresistance are widely discussed, their molecular boundaries often overlap, and transcriptional signatures reported across studies are not always consistent. Variability in experimental design, antibiotic exposure conditions, growth phase, and analytical pipelines complicates cross-study comparison and limits reproducibility(Brauner et al. 2016; El Meouche et al. 2024).
Future work will need to establish consensus frameworks for defining resistance states that integrate transcriptional data with phenotypic and clinical endpoints. Such frameworks should distinguish between adaptive expression programs that transiently promote survival and those that meaningfully alter treatment outcomes. Without standardized reference states, transcriptomic diagnostics risk producing context-specific results that are difficult to generalize across pathogens, infection types, or clinical settings.
7.2. Improving Physiological Relevance of Transcriptomic Models
Many transcriptomic studies of antimicrobial response are conducted under simplified laboratory conditions that only partially reflect the host environment. Although host-mimicking models and dual RNA sequencing have improved physiological relevance, these approaches remain underutilized and technically demanding (Westermann et al. 2012; Avican et al. 2021). As a result, transcriptional signatures identified in vitro may not accurately represent resistance states operating during infection.
Future research should prioritize experimental designs that incorporate host-derived stresses, spatial heterogeneity, and microbial community context. This includes greater use of infection-relevant growth conditions, animal models, and ex vivo systems that capture immune pressure and nutrient limitation. Importantly, transcriptomic findings from such models should be systematically compared with patient-derived data to assess translational validity.
7.3. Validation Across Clinical Diversity
A major barrier to clinical translation is the limited validation of transcriptomic resistance signatures across diverse patient populations. Most studies to date focus on a narrow range of strains, species, or experimental conditions, raising concerns about generalizability. Differences in host immunity, microbiome composition, antibiotic exposure history, and infection site may all influence transcriptional states in ways that are not captured by existing datasets (Fang et al. 2016).
Robust validation will therefore require multicentre studies encompassing diverse geographic regions, healthcare settings, and patient demographics. Importantly, validation should extend beyond laboratory correlates of resistance to include clinically meaningful outcomes such as treatment failure, relapse, and duration of infection. Without such evidence, transcriptomics-informed approaches are unlikely to gain acceptance in routine clinical practice.
7.4. Balancing Model Complexity and Interpretability
As transcriptomic datasets grow in size and complexity, there is increasing interest in computational approaches for resistance state classification. However, highly complex or black-box models may undermine clinical trust and limit interpretability. Precision medicine frameworks emphasize not only predictive accuracy but also mechanistic understanding and transparency.
Future methodological development should therefore focus on models that balance complexity with interpretability, integrating prior biological knowledge where possible. Pathway-informed classifiers and constrained learning approaches may offer a compromise between data-driven discovery and mechanistic insight, improving both robustness and clinical acceptability (Toprak et al. 2011).
7.5. Ethical, Logistical, and Implementation Challenges
Finally, the broader implementation of transcriptomics in AMR diagnostics raises ethical and logistical considerations. Access to advanced sequencing technologies remains uneven across healthcare systems, raising concerns about equity in precision medicine. In addition, transcriptomic diagnostics may generate probabilistic or state-dependent outputs that complicate clinical decision-making and communication with patients.
Clear guidelines for data interpretation, reporting standards, and regulatory evaluation will be required to ensure responsible use of transcriptomic information. From a translational perspective, future efforts should focus on defining use cases where transcriptomics provides clear added value over existing diagnostics, rather than pursuing broad adoption without demonstrated benefit (Byron et al. 2016).
8. Conclusions
Antimicrobial resistance (AMR) remains one of the most persistent challenges in infectious disease management, in part because resistance is commonly conceptualized as a static genetic property rather than a dynamic biological phenomenon. While genomic approaches have substantially advanced resistance detection and surveillance, they provide limited insight into whether resistance mechanisms are functionally engaged during infection or how bacterial responses evolve over the course of treatment. As highlighted throughout this review, transcriptomics offers a complementary and mechanistically informative layer by capturing the regulated resistance states and context-dependent expression programs that shape antimicrobial response in ways not predictable from genotype alone (Brauner et al. 2016; El Meouche et al. 2024).
Evidence from pathogen transcriptomics demonstrates that bacterial survival under antibiotic exposure is frequently mediated by inducible and reversible transcriptional programs involving stress responses, metabolic adaptation, and regulatory network activation. These programs underpin clinically relevant phenomena such as tolerance and persistence, which contribute to treatment failure and relapse despite apparent susceptibility in standard diagnostics (Brauner et al. 2016; Lewis 2010). Importantly, transcriptomic studies also reveal that resistance-associated expression states are strongly modulated by host immune pressures, infection site physiology, and microbiome context, reinforcing the notion that antimicrobial response is inherently patient- and environment-specific (Fang et al. 2016; Avican et al. 2021).
From a precision medicine perspective, these findings challenge the sufficiency of binary resistance classifications and motivate a shift toward state-aware frameworks for antimicrobial decision-making. Transcriptomics-informed diagnostics have the potential to complement existing phenotypic and genomic tools by identifying active resistance pathways, monitoring adaptive responses during therapy, and informing stratified treatment strategies. However, as emphasized in this review, such applications remain constrained by technical, logistical, and validation-related barriers. Transcriptomic resistance signatures are often context-dependent, insufficiently standardized, and rarely validated against clinically meaningful outcomes across diverse patient populations (Chappell et al. 2015).
The path toward clinical integration therefore requires cautious and methodical progress. Priority should be given to defining reproducible resistance state signatures, improving physiological relevance of experimental models, and conducting outcome-linked validation studies using patient-derived samples. Equally important is the development of interpretable analytical frameworks that align transcriptomic data with known resistance mechanisms, supporting clinical trust and regulatory acceptance (Toprak et al. 2011). In the near term, transcriptomics is most likely to add value in complex or refractory infections, where conventional diagnostics fail to explain treatment response and additional functional insight may justify increased diagnostic complexity.
In conclusion, reframing antimicrobial resistance as a dynamic, transcriptionally regulated state provides a more nuanced and biologically grounded foundation for precision infection medicine. Transcriptomics does not replace genomics or phenotypic testing but rather extends them by revealing how resistance potential is realized within specific physiological contexts. By integrating transcriptomic insights with host, environmental, and clinical data, future precision medicine approaches may move closer to individualized antimicrobial therapy that is both mechanistically informed and clinically actionable. Achieving this goal will depend not on technological acceleration alone, but on rigorous validation, thoughtful implementation, and sustained integration of molecular insight with patient-centered care.
Conflicts of Interest
The author declares no conflict of interest and received no external funding for this work. No new data were created or analysed in this study. The author is solely responsible for the conceptualization, literature review, writing, and editing of the manuscript. The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Andersson, D.I.; Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 2014, 12, 465–478. [Google Scholar] [CrossRef]
- Avican, K.; Aldahdooh, J.; Togninalli, M.; Mahmud, A.K.M.F.; Tang, J.; Borgwardt, K.M.; Rhen, M.; Fällman, M. RNA atlas of human bacterial pathogens uncovers stress dynamics linked to infection. Nat. Commun. 2021, 12, 3282. [Google Scholar] [CrossRef] [PubMed]
- Balaban, N.Q.; Helaine, S.; Lewis, K.; Ackermann, M.; Aldridge, B.; Andersson, D.I.; Brynildsen, M.P.; Bumann, D.; Camilli, A.; Collins, J.J.; et al. Definitions and guidelines for research on antibiotic persistence. Nat. Rev. Microbiol. 2019, 17, 441–448. [Google Scholar] [CrossRef] [PubMed]
- Band, V.I.; Weiss, D.S. Heteroresistance: A cause of unexplained antibiotic treatment failure? PLOS Pathog. 2019, 15, e1007726. [Google Scholar] [CrossRef]
- Van Belkum, A.; Burnham, C.-A.D.; Rossen, J.W.A.; Mallard, F.; Rochas, O.; Dunne, W.M., Jr. Innovative and rapid antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 2020, 18, 299–311. [Google Scholar] [CrossRef]
- Bie, L.; Zhang, M.; Wang, J.; Fang, M.; Li, L.; Xu, H.; Wang, M. Comparative Analysis of Transcriptomic Response of Escherichia coli K-12 MG1655 to Nine Representative Classes of Antibiotics. Microbiol. Spectr. 2023, 11, e0031723. [Google Scholar] [CrossRef]
- Brauner, A.; Fridman, O.; Gefen, O.; Balaban, N.Q. Distinguishing between resistance, tolerance and persistence to antibiotic treatment. Nat. Rev. Microbiol. 2016, 14, 320–330. [Google Scholar] [CrossRef]
- Buffie, C.G.; Pamer, E.G. Microbiota-mediated colonization resistance against intestinal pathogens. Nat. Rev. Immunol. 2013, 13, 790–801. [Google Scholar] [CrossRef]
- Byron, S.A.; Van Keuren-Jensen, K.R.; Engelthaler, D.M.; Carpten, J.D.; Craig, D.W. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat. Rev. Genet. 2016, 17, 257–271. [Google Scholar] [CrossRef]
- Chappell, J.; Takahashi, M.K.; Lucks, J.B. Creating small transcription activating RNAs. Nat. Chem. Biol. 2015, 11, 214–220. [Google Scholar] [CrossRef] [PubMed]
- Ching, T.; Himmelstein, D.S.; Beaulieu-Jones, B.K.; Kalinin, A.A.; Do, B.T.; Way, G.P.; Ferrero, E.; Agapow, P.-M.; Zietz, M.; Hoffman, M.M.; et al. Opportunities and obstacles for deep learning in biology and medicine. J. R. Soc. Interface 2018, 15. [Google Scholar] [CrossRef]
- Collins, F.S.; Varmus, H. A New Initiative on Precision Medicine. New Engl. J. Med. 2015, 372, 793–795. [Google Scholar] [CrossRef]
- Croucher, N.J.; Thomson, N.R. Studying bacterial transcriptomes using RNA-seq. Curr. Opin. Microbiol. 2010, 13, 619–624. [Google Scholar] [CrossRef]
- Dawan, J.; Ahn, J. Bacterial Stress Responses as Potential Targets in Overcoming Antibiotic Resistance. Microorganisms 2022, 10, 1385. [Google Scholar] [CrossRef] [PubMed]
- Didelot, X.; Maiden, M.C. Impact of recombination on bacterial evolution. Trends Microbiol. 2010, 18, 315–322. [Google Scholar] [CrossRef]
- El Meouche, I.; Jain, P.; Jolly, M.K.; Capp, J.-P. Drug tolerance and persistence in bacteria, fungi and cancer cells: Role of non-genetic heterogeneity. Transl. Oncol. 2024, 49, 102069. [Google Scholar] [CrossRef]
- Ellington, M.; Ekelund, O.; Aarestrup, F.; Canton, R.; Doumith, M.; Giske, C.; Grundman, H.; Hasman, H.; Holden, M.; Hopkins, K.; et al. The role of whole genome sequencing in antimicrobial susceptibility testing of bacteria: report from the EUCAST Subcommittee. Clin. Microbiol. Infect. 2017, 23, 2–22. [Google Scholar] [CrossRef]
- Fang, F.C.; Frawley, E.R.; Tapscott, T.; Vázquez-Torres, A. Bacterial Stress Responses during Host Infection. Cell Host Microbe 2016, 20, 133–143. [Google Scholar] [CrossRef] [PubMed]
- Hassall, J.; Coxon, C.; Patel, V.C.; Goldenberg, S.D.; Sergaki, C. Limitations of current techniques in clinical antimicrobial resistance diagnosis: examples and future prospects. npj Antimicrob. Resist. 2024, 2, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Zhai, Z.Y.; Huang, Z.Y.; Chen, Z.M.; Zhou, P.; Li, X.X.; Yang, G.H.; Bao, C.J.; You, L.J.; Cui, X.B.; et al. Dual RNA sequencing of Helicobacter pylori and host cell transcriptomes reveals ontologically distinct host-pathogen interaction. mSystems 2024, 9, e0020624. [Google Scholar] [CrossRef]
- Joubert, I.A.; Mullally, C.; Litton, E.; Raby, E.; Chopra, A.; Strunk, T.; Wang, P.; Currie, A. Dual RNA isolation from blood: an optimized protocol for host and bacterial RNA purification for dual RNA-sequencing analysis in whole blood sepsis samples. Microb. Genom. 2025, 11, 001501. [Google Scholar] [CrossRef]
- Kirchner, M.; Nunez-Garcia, J.; Duggett, N.; Gosling, R.J.; Anjum, M.F. Use of transcriptomics and genomics to assess the effect of disinfectant exposure on the survival and resistance of Escherichia coli O157:H7, a human pathogen. Front. Microbiol. 2024, 15, 1477683. [Google Scholar] [CrossRef] [PubMed]
- Levin-Reisman, I.; Brauner, A.; Ronin, I.; Balaban, N.Q. Epistasis between antibiotic tolerance, persistence, and resistance mutations. Proc. Natl. Acad. Sci. 2019, 116, 14734–14739. [Google Scholar] [CrossRef]
- Lewis, K. Persister Cells. Annu. Rev. Microbiol. 2010, 64, 357–372. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.; Tan, J.; Li, X.; Huang, D.; Wang, L.; Zhu, L.; Chen, Z.; Gan, L.; Liu, W.; Jin, X. Combined transcriptome and metabolome analysis revealed the antimicrobial mechanism of Griseorhodin C against Methicillin-resistant Staphylococcus aureus. Sci. Rep. 2024, 14, 30242. [Google Scholar] [CrossRef]
- Lobritz, M.A.; Belenky, P.; Porter, C.B.M.; Gutierrez, A.; Yang, J.H.; Schwarz, E.G.; Dwyer, D.J.; Khalil, A.S.; Collins, J.J. Antibiotic efficacy is linked to bacterial cellular respiration. Proc. Natl. Acad. Sci. 2015, 112, 8173–8180. [Google Scholar] [CrossRef] [PubMed]
- Rendón, J.M.; Lang, B.; Llorens, M.R.; Tartaglia, G.G.; Burgas, M.T. DualSeqDB: the host–pathogen dual RNA sequencing database for infection processes. Nucleic Acids Res. 49, D687–D693. [CrossRef]
- Martínez, J.L.; Rojo, F. Metabolic regulation of antibiotic resistance. FEMS Microbiol. Rev. 2011, 35, 768–789. [Google Scholar] [CrossRef]
- A McDevitt, C.; Ogunniyi, A.D.; Valkov, E.; Lawrence, M.C.; Kobe, B.; McEwan, A.G.; Paton, J.C. A Molecular Mechanism for Bacterial Susceptibility to Zinc. PLOS Pathog. 2011, 7, e1002357. [Google Scholar] [CrossRef]
- Ransom, E.M.; Potter, R.F.; Dantas, G.; Burnham, C.-A.D. Genomic Prediction of Antimicrobial Resistance: Ready or Not, Here It Comes! Clin. Chem. 2020, 66, 1278–1289. [Google Scholar] [CrossRef]
- Shahid, U. Microbiome-guided precision medicine: Mechanistic insights, multi-omics integration, and translational horizons. J. Precis. Med. Heal. Dis. 2025, 3. [Google Scholar] [CrossRef]
- Shahid, U. Advances in RNA therapeutics: Classes, innovations and clinical applications. J. Precis. Med. Heal. Dis. 2025, 3, 100016. [Google Scholar] [CrossRef]
- Shahid, U.; Hooi, S.L.; Lim, S.Y.; Aris, A.M.; Khor, B.C.; Ayub, Q.; Tan, H.S. Metagenomic surveillance of microbial community and antibiotic resistant genes associated with Malaysian wastewater during the COVID-19 pandemic. Curr. Genet. 2024, 71, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Shahreen, N.; Shahid, S.A.; Subhani, M.; Al-Siyabi, A.; Saha, R. Minimal gene signatures enable high-accuracy prediction of antibiotic resistance in Pseudomonas aeruginosa. npj Syst. Biol. Appl. 2025, 11, 108. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S.; Franklin, M.J. Physiological heterogeneity in biofilms. Nat. Rev. Microbiol. 2008, 6, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Su, M.; Satola, S.W.; Read, T.D. Genome-Based Prediction of Bacterial Antibiotic Resistance. J. Clin. Microbiol. 2019, 57, 1–15. [Google Scholar] [CrossRef]
- Toprak, E.; Veres, A.; Michel, J.-B.; Chait, R.; Hartl, D.L.; Kishony, R. Evolutionary paths to antibiotic resistance under dynamically sustained drug selection. Nat. Genet. 2011, 44, 101–105. [Google Scholar] [CrossRef]
- Westermann, A.J.; Gorski, S.A.; Vogel, J. Dual RNA-seq of pathogen and host. Nat. Rev. Microbiol. 2012, 10, 618–630. [Google Scholar] [CrossRef]
- Windels, E.M.; Michiels, J.E.; Fauvart, M.; Wenseleers, T.; Van den Bergh, B.; Michiels, J. Bacterial persistence promotes the evolution of antibiotic resistance by increasing survival and mutation rates. ISME J. 2019, 13, 1239–1251. [Google Scholar] [CrossRef]
Figure 1.
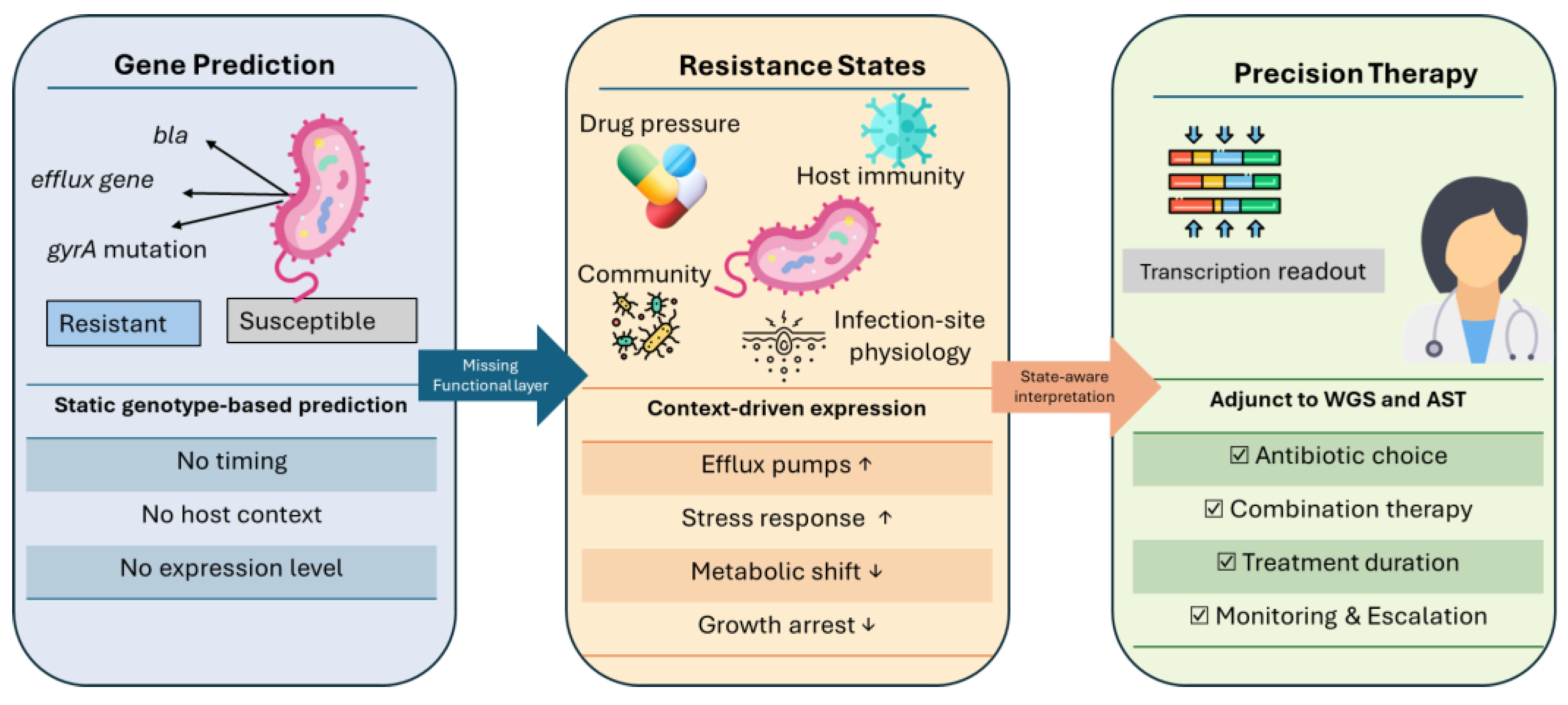
From resistance genes to resistance states in precision antimicrobial therapy. The schematic illustrates the conceptual transition from static, gene-centric antimicrobial resistance prediction (left panel), based on the presence of resistance determinants identified by whole-genome sequencing, to dynamic resistance states shaped by context-dependent transcriptional responses (middle panel). These resistance states emerge from the interaction of drug pressure, host immunity, infection-site physiology, and microbial community context, resulting in regulated changes in efflux activity, stress responses, metabolism, and growth. Incorporation of transcriptomic readouts enables state-aware interpretation that complements conventional WGS and phenotypic susceptibility testing, informing personalized therapeutic decisions such as antibiotic selection, combination therapy, treatment duration, and monitoring strategies (right panel).
Figure 1.
From resistance genes to resistance states in precision antimicrobial therapy. The schematic illustrates the conceptual transition from static, gene-centric antimicrobial resistance prediction (left panel), based on the presence of resistance determinants identified by whole-genome sequencing, to dynamic resistance states shaped by context-dependent transcriptional responses (middle panel). These resistance states emerge from the interaction of drug pressure, host immunity, infection-site physiology, and microbial community context, resulting in regulated changes in efflux activity, stress responses, metabolism, and growth. Incorporation of transcriptomic readouts enables state-aware interpretation that complements conventional WGS and phenotypic susceptibility testing, informing personalized therapeutic decisions such as antibiotic selection, combination therapy, treatment duration, and monitoring strategies (right panel).
Figure 2.
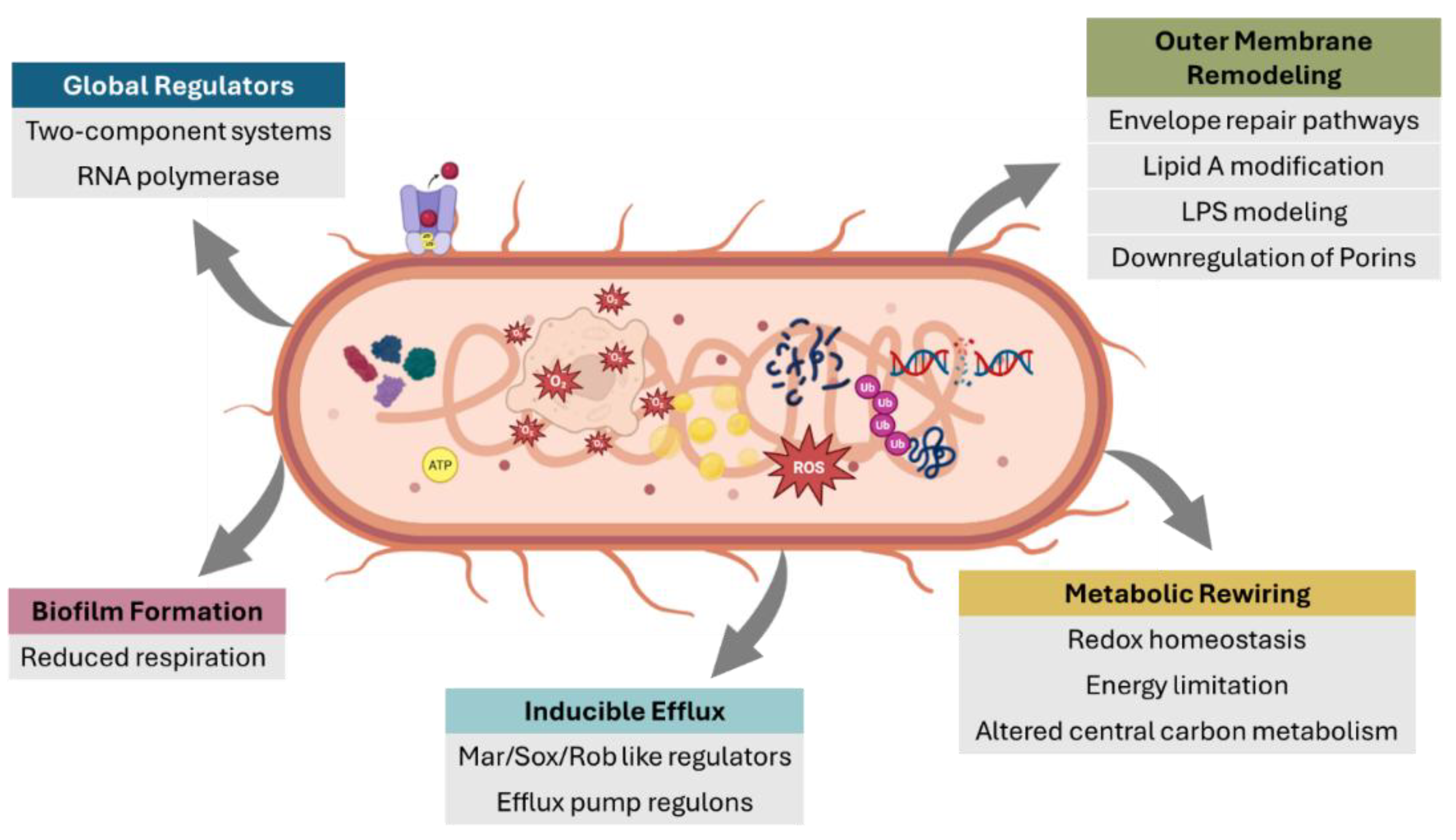
Molecular architecture of transcriptionally mediated resistance states in bacterial pathogens. The schematic illustrates major transcriptional programs that contribute to non-genetic resistance phenotypes, including inducible efflux, metabolic rewiring, stress-response activation, membrane remodeling, and biofilm-associated states. These programs are regulated by global transcriptional regulators and two-component systems and can operate simultaneously within the same bacterial population. Their coordinated activation gives rise to resistance-associated behaviors such as antibiotic tolerance, persistence, and survival without stable shifts in minimum inhibitory concentration (MIC), highlighting resistance as a dynamic, state-dependent phenotype rather than a fixed genetic trait.
Figure 2.
Molecular architecture of transcriptionally mediated resistance states in bacterial pathogens. The schematic illustrates major transcriptional programs that contribute to non-genetic resistance phenotypes, including inducible efflux, metabolic rewiring, stress-response activation, membrane remodeling, and biofilm-associated states. These programs are regulated by global transcriptional regulators and two-component systems and can operate simultaneously within the same bacterial population. Their coordinated activation gives rise to resistance-associated behaviors such as antibiotic tolerance, persistence, and survival without stable shifts in minimum inhibitory concentration (MIC), highlighting resistance as a dynamic, state-dependent phenotype rather than a fixed genetic trait.
Figure 3.
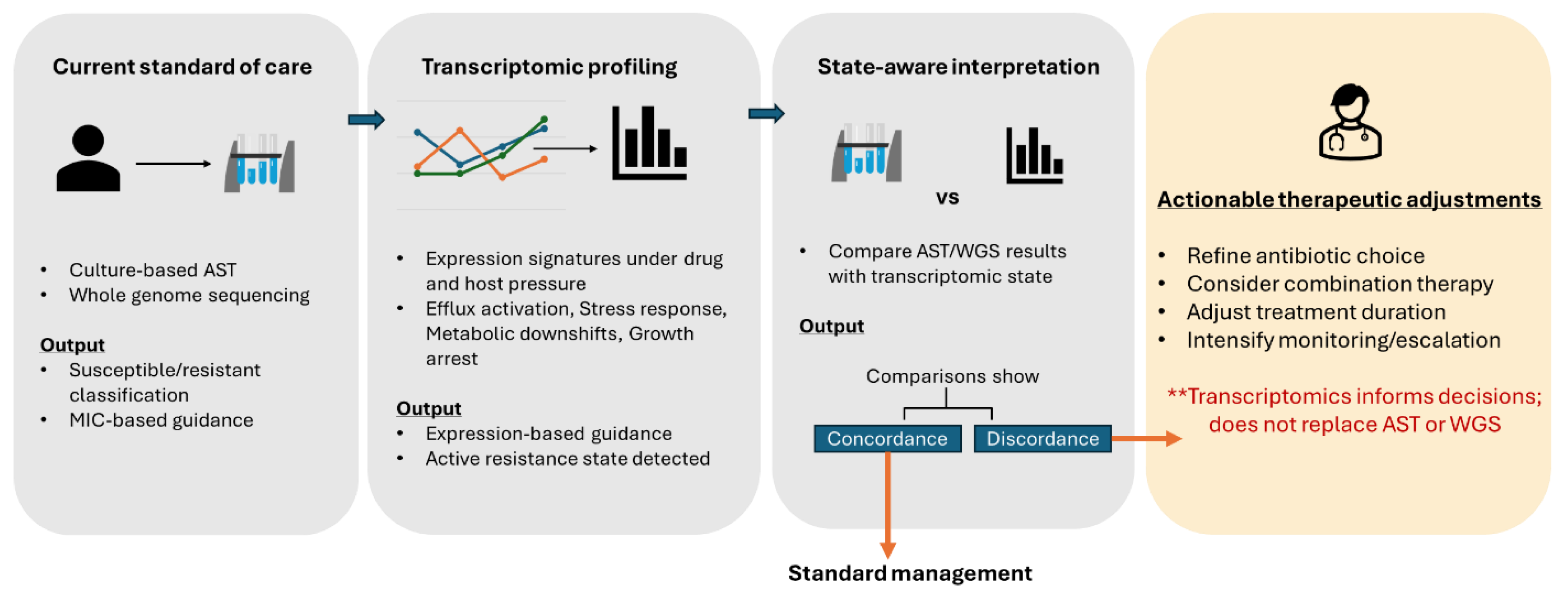
Integrating pathogen transcriptomics into contemporary antimicrobial treatment workflows. Schematic illustrating how transcriptomic resistance-state profiling can function as an adjunct to standard-of-care diagnostics. Conventional culture-based antimicrobial susceptibility testing (AST) and genomic prediction provide baseline classifications of susceptibility or resistance. Optional transcriptomic profiling captures context-dependent resistance states driven by gene expression under host and drug pressure. State-aware interpretation enables identification of discordance between predicted susceptibility and active resistance states, informing refined antibiotic selection, treatment duration, and monitoring strategies. Transcriptomics is positioned as a complementary tool rather than a replacement for existing diagnostic frameworks.
Figure 3.
Integrating pathogen transcriptomics into contemporary antimicrobial treatment workflows. Schematic illustrating how transcriptomic resistance-state profiling can function as an adjunct to standard-of-care diagnostics. Conventional culture-based antimicrobial susceptibility testing (AST) and genomic prediction provide baseline classifications of susceptibility or resistance. Optional transcriptomic profiling captures context-dependent resistance states driven by gene expression under host and drug pressure. State-aware interpretation enables identification of discordance between predicted susceptibility and active resistance states, informing refined antibiotic selection, treatment duration, and monitoring strategies. Transcriptomics is positioned as a complementary tool rather than a replacement for existing diagnostic frameworks.
Table 1.
Conceptual and diagnostic differences between gene-centric and state-based frameworks of antimicrobial resistance.
Table 1.
Conceptual and diagnostic differences between gene-centric and state-based frameworks of antimicrobial resistance.
| Dimension |
Gene-Centric AMR Framework |
Resistance-State Framework |
| Primary unit of analysis |
Presence/absence of resistance genes or mutations |
Transcriptionally regulated functional states |
| Temporal resolution |
Static |
Dynamic and time-dependent |
| Sensitivity to environment |
Limited |
High (host, antibiotic exposure, microbiome, site physiology) |
| Captures inducible resistance |
Poorly |
Explicitly |
| Explains tolerance and persistence |
Largely no |
Yes |
| Predictive power for treatment outcome |
Moderate and population-averaged |
Potentially patient-specific |
| Diagnostic modality |
WGS, PCR-based assays |
Transcriptomics (whole or targeted panels) |
| Clinical interpretation |
Binary (resistant/susceptible) |
State-aware and probabilistic |
| Precision medicine compatibility |
Limited |
High |
Table 2.
Transcriptionally defined resistance-associated states and their functional and clinical implications.
Table 2.
Transcriptionally defined resistance-associated states and their functional and clinical implications.
| Resistance state |
Dominant transcriptomic features |
Biological interpretation |
Potential clinical relevance |
| Antibiotic tolerance |
Upregulation of global stress-response regulons; reduced expression of growth-associated genes |
Transient survival strategy without genetic resistance |
Explains treatment failure despite in vitro susceptibility |
| Persistence |
Suppression of replication, transcription, and translation programs |
Entry into dormant or low-activity subpopulation |
Predicts relapse and prolonged infection |
| Inducible resistance |
Conditional expression of resistance determinants (e.g., efflux systems, modifying enzymes) |
Context-dependent activation of resistance pathways |
Missed by static genotypic diagnostics |
| Metabolic adaptation |
Reprogramming of central carbon metabolism and redox balance |
Reduced antibiotic lethality through altered physiology |
Identifies metabolic vulnerabilities for adjuvant therapy |
| Biofilm-associated resistance |
Expression of matrix synthesis genes; altered metabolic and stress-response profiles |
Spatial and physiological protection from antibiotics |
Relevant to chronic and device-associated infections |
| Host-induced resistance states |
Activation of oxidative, nitrosative, or nutrient-limitation stress responses |
Resistance behavior shaped by immune and tissue context |
Explains discordance between in vitro and in vivo response |
Table 3.
Translational potential and limitations of transcriptomics-informed AMR diagnostics.
Table 3.
Translational potential and limitations of transcriptomics-informed AMR diagnostics.
| Diagnostic dimension |
What transcriptomics can reveal |
Added value beyond genomics |
Key limitations / considerations |
| Resistance pathway activation |
Active expression of efflux systems, stress-response regulons, metabolic adaptations |
Distinguishes expressed vs silent resistance potential |
Expression is context- and time-dependent |
| Early adaptive responses |
Transcriptional changes preceding phenotypic resistance |
Enables detection of pre-resistance or tolerance states |
Requires precise timing of sampling |
| Host-influenced resistance states |
Immune- and environment-induced expression programs |
Captures in vivo-relevant resistance behavior |
Host RNA dominance in clinical samples |
| Treatment monitoring |
Dynamic shifts in resistance-associated expression during therapy |
Potential marker of treatment response or persistence |
Longitudinal sampling often impractical |
| Diagnostic stratification |
Compact expression signatures associated with survival outcomes |
Supports state-aware risk stratification |
Limited cross-cohort validation |
| Clinical feasibility |
Targeted expression panels (vs whole transcriptome) |
Reduced cost and turnaround time |
Requires standardization and regulatory approval |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).