Submitted:
26 December 2025
Posted:
29 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
- To confirm the taxonomic status (genus and/or species) of the species collected from cassava leaves in the RC;
- To compare the genetic divergence of these species from others of the same genus that are available in public reference databases.
2. Materials and Methods
2.1. Sampling
2.2. Individual Collection
2.3. DNA Extraction
2.4. Selected Primers and DNA Amplification
| Characteristics of the primers selected to amplify each gene fragment are presented in Table 1. The following primers were used: 18SF and 18SR, used for complete amplification of the internal ITS region, and C1J1718 and 773, used to amplify the COI fragment [24,28]. All amplification reactions were performed in a final volume of 25 µL. The samples were initially denatured at 94 °C for 3 min. PCR was then performed for 35 cycles, with 1 min of denaturation at 94 °C, followed by 1 min of annealing at 51 °C for each sample. The final extension was carried out at 72 °C for 1.5 min, and the PCR products subsequently visualised on a 1% agarose gel saturated with ethidium bromide.Gènes | Taille (pb) | Position | Amorces | Référence | Séquences nucléotidiques |
|
3’de 18S |
18S |
Ben et al.2000 |
5’-AGAGGAAGTAAAAGTCGTAACAAG-3’ |
||
|
ITS |
1200 |
5’de 28S | 18SR | Navajas et al.1998 | 5’-ATATGCTTAAATTCAGCGGG-3’ |
| 5.8S | Inter S | Navia et al.2005 | 5’-GATCACTCGAATTACCAATCG-3’ | ||
| 5.8S | Inter Rev | Navia et al.2005 | 5’-CGATTGGTAATTCGAGTGATC-3’ | ||
|
C1J1718 |
Simon et al.1994 |
5’-GGAGGATTTGGAAATTGATTAGTTCC -3’ |
|||
|
COI |
890 |
COI |
773 Inter COI |
Navia et al.1994 this study |
5’-TACAGCTCCTATAGATAAAAC -3’ 5’- GGNGANCCTATTTTATATCAAC -3’ |
| Inter Rev | this study | 5’- GTTGATATAAAATAGGNTCNCC -3’ |
2.5. Sequencing, Correction and Processing of Chromatograms: Obtaining Consensus Sequences
2.6. NCBI BLAST
2.7. Methodology for Data Collection from GenBank
2.8. Multiple Sequence Alignment
2.9. Nucleotide Evolution Models, Genetic Divergence, and Phylogenetic Tree
3. Results
3.1. Amplification Rate
3.2. BOLD Blast
3.3. GenBank Blast
3.4. Submission of Sequences to GenBank
3.5. Total Data
3.6. General Characteristics of the COI Alignment
3.7. Selection of the Best-Fitting Nucleotide Substitution Model
3.8. Intra- and Inter-group Genetic Divergence
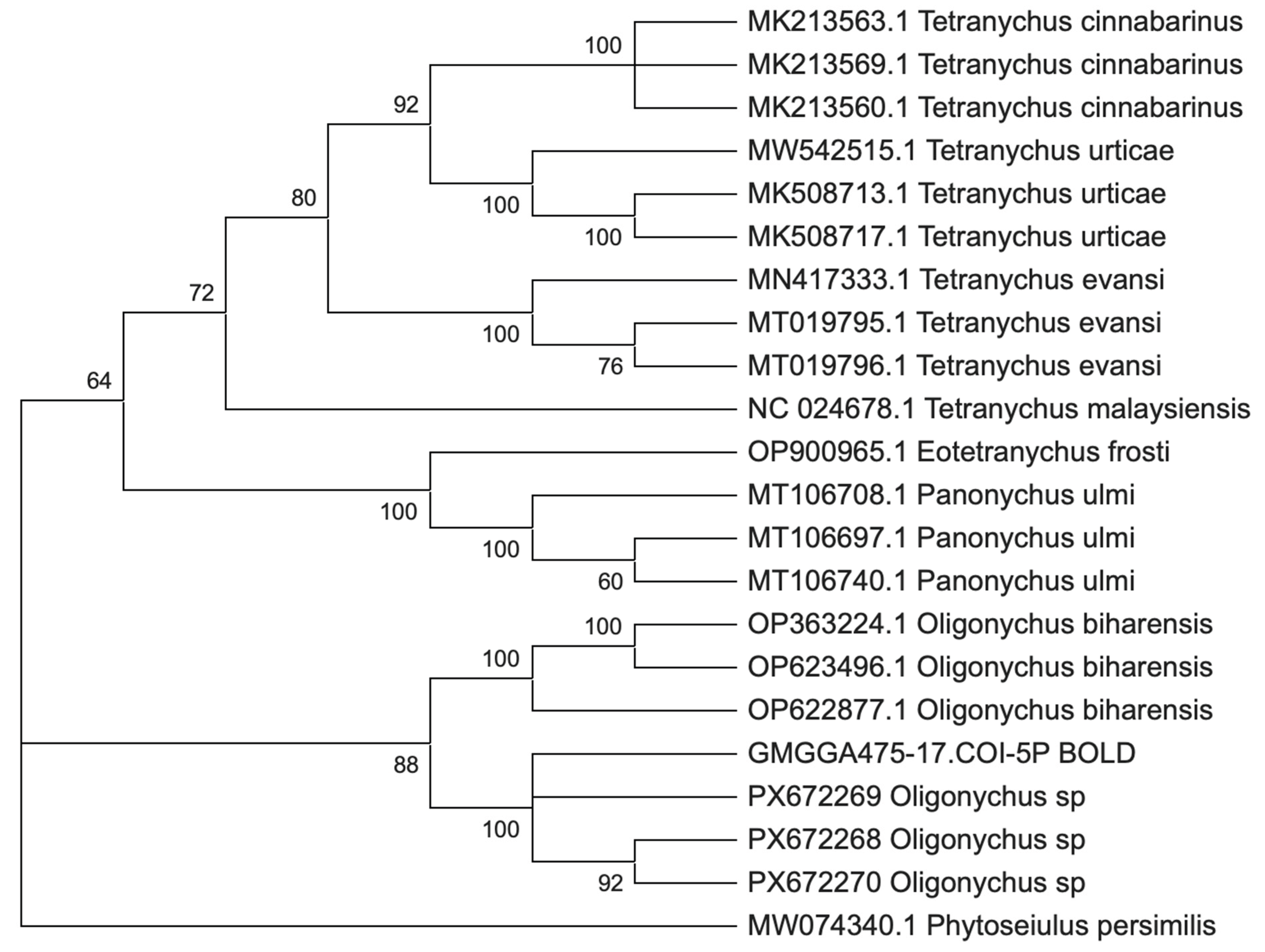
3.9. Phylogenetic Analysis
4. Discussion
4.1. Failure to Amplify ITS: Implications and Hypotheses
4.2. Study Limitations
5. Conclusions
Author Contributions
Funding
Use of Generative AI
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| COI | Cytochrome c oxidase I |
| RC | Republic of Congo |
References
- Bolland, H.; Gutierrez, J.; Flechtmann, C. World Catalogue of the Spider Mite Family (Acari: Tetranychidae). 1998.
- Migeon, A.; Nouguier, E.; Dorkeld, F. Spider Mites Web: A Comprehensive Database for the Tetranychidae. Trends Acarol. 2011, 557–560.
- Montagnac, J.A.; Davis, C.R.; Tanumihardjo, S.A. Nutritional Value of Cassava for Use as a Staple Food and Recent Advances for Improvement. Compr. Rev. Food Sci. Food Saf. 2009, 8, 181–194. [CrossRef]
- In FAO Save and Grow: Cassava: A Guide to Sustainable Production Intensification; FAO: Rome, Italy, 2013; ISBN 978-92-5-107641-5. 4. FAO Save and Grow: Cassava: A Guide to Sustainable Production Intensification; FAO: Rome, Italy, 2013; ISBN 978-92-5-107641-5.
- Ezenwaka, L.; Del Carpio, D.P.; Jannink, J.-L.; Rabbi, I.; Danquah, E.; Asante, I.; Danquah, A.; Blay, E.; Egesi, C. Genome-Wide Association Study of Resistance to Cassava Green Mite Pest and Related Traits in Cassava. Crop Sci. 2018, 58, 1907–1918. [CrossRef]
- Ezenwaka, L.; Rabbi, I.; Onyeka, J.; Kulakow, P.; Egesi, C. Identification of Additional /Novel QTL Associated with Resistance to Cassava Green Mite in a Biparental Mapping Population. PLOS ONE 2020, 15, e0231008. [CrossRef]
- Gutierrez, J.; Bonato, O. Les Acariens Tetranychidae Attaquant Le Manioc Au Congo et Quelques-Uns de Leurs Prédateurs. Rev. Zool. Afr. 1994, 108, 191–200.
- Belle Mbou Okassa, M.; Louzeymo, J.B.; Nianga Bikouta, G.; Lenga, A. Diversité Des Acariens Ravageurs de La Famille Des Tetranychidae Présents Sur Manihot Esculenta Dans La Commune de Brazzaville : État Des Lieux et Perspectives. Rev. Afr. D’Environnement D’Agriculture 2024, 7, 136–142. [CrossRef]
- Dayrat, B. Towards Integrative Taxonomy. Biol. J. Linn. Soc. 2005, 85, 407–415. [CrossRef]
- Padial, J.M.; Miralles, A.; De la Riva, I.; Vences, M. The Integrative Future of Taxonomy. Front. Zool. 2010, 7, 16. [CrossRef]
- Schlick-Steiner, B.C.; Steiner, F.M.; Seifert, B.; Stauffer, C.; Christian, E.; Crozier, R.H. Integrative Taxonomy: A Multisource Approach to Exploring Biodiversity. Annu. Rev. Entomol. 2010, 55, 421–438. [CrossRef]
- Okassa, M.; Tixier, M.-S.; Kreiter, S. Morphological and molecular diagnostics of Phytoseiulus persimilis and Phytoseiulus macropilis (Acari: Phytoseiidae. Exp Appl Acarol 2010, 52, 291–303.
- Okassa, M.B.M.; Ntabi, D.M.; Lenga, A. Morphological and Molecular Identification of Specimens in the Genus Euseius (Acari: Phytoseiidae) from the Republic of Congo. Zootaxa 2020, 4768, 479–498. [CrossRef]
- Tixier, M.-S.; Kreiter, S.; Barbar, Z.; Ragusa, S.; Cheval, B. Status of two cryptic species, Typhlodromus exhilaratus Ragusa and Typhlodromus phialatus Athias-Henriot (Acari: Phytoseiidae): consequences for taxonomy. Zool Scr 2006, 35, 115–122.
- Tixier, M.-S.; Auger, P.; Migeon, A.; Douin, M.; Fossoud, A.; Navajas, M.; Arabuli, T. Integrated Taxonomy Supports the Identification of Some Species of Phytoseiidae (Acari: Mesostigmata) from Georgia. Acarologia 2021, 61, 824–844. [CrossRef]
- Hebert, P.D.N.; Ratnasingham, S.; deWaard, J.R. Barcoding Animal Life: Cytochrome c Oxidase Subunit 1 Divergences among Closely Related Species. Proc. R. Soc. B Biol. Sci. 2003, 270, S96–S99. [CrossRef]
- Hebert, P.D.N.; Penton, E.H.; Burns, J.M.; Janzen, D.H.; Hallwachs, W. Ten Species in One: DNA Barcoding Reveals Cryptic Species in the Neotropical Skipper Butterfly Astraptes Fulgerator. Proc. Natl. Acad. Sci. 2004, 101, 14812–14817. [CrossRef]
- Matsuda, T.; Fukumoto, C.; Hinomoto, N.; Gotoh, T. DNA-Based Identification of Spider Mites: Molecular Evidence for Cryptic Species of the Genus Tetranychus (Acari: Tetranychidae). J. Econ. Entomol. 2013, 106, 463–472. [CrossRef]
- Chan-Chable, R.J.; Martínez-Arce, A.; Mis-Avila, P.C.; Ortega-Morales, A.I. DNA Barcodes and Evidence of Cryptic Diversity of Anthropophagous Mosquitoes in Quintana Roo, Mexico. Ecol. Evol. 2019, 9, 4692–4705. [CrossRef]
- Ballard, J.W.O.; Whitlock, M.C. The Incomplete Natural History of Mitochondria. Mol. Ecol. 2004, 13, 729–744. [CrossRef]
- Okassa, M.; Tixier, M.-S.; Cheval, B.; Kreiter, S. Molecular and Morphological Evidence for a New Species Status within the Genus Euseius (Acari: Phytoseiidae). Can. J. Zool. 2009, 87, 689–698. [CrossRef]
- Tixier, M.-S.; Okassa, M.; Kreiter, S. An Integrative Morphological and Molecular Diagnostic for Typhlodromus Pyri (Acari: Phytoseiidae). Zool. Scr. 2012, 41, 68–78. [CrossRef]
- Tixier, M.-S.; Hernandes, F.A.; Guichou, S.; Kreiter, S. The Puzzle of DNA Sequences of Phytoseiidae (Acari : Mesostigmata) in the Public GenBank Database. Invertebr. Syst. 2012, 25, 389–406. [CrossRef]
- Navajas, M.; Gutierrez, J.; Williams, M.; Gotoh, T. Synonymy between Two Spider Mite Species, Tetranychus Kanzawai and T. Hydrangeae (Acari: Tetranychidae), Shown by Ribosomal ITS2 Sequences and Cross-Breeding Experiments. Bull. Entomol. Res. 2001, 91, 117–123.
- Mushtaq, H.M.S.; Alatawi, F.J.; Kamran, M.; Flechtmann, C.H.W. The Genus Oligonychus Berlese (Acari, Prostigmata, Tetranychidae): Taxonomic Assessment and a Key to Subgenera, Species Groups, and Subgroups. ZooKeys 2021, 1079, 89–127. [CrossRef]
- Mushtaq, H.M.S.; Kamran, M.; Saleh, A.A.; Alatawi, F.J.; Mushtaq, H.M.S.; Kamran, M.; Saleh, A.A.; Alatawi, F.J. Evidence for Reconsidering the Taxonomic Status of Closely Related Oligonychus Species in Punicae Complex (Acari: Prostigmata: Tetranychidae). Insects 2022, 14. [CrossRef]
- Mushtaq, H.M.S.; Saleh, A.A.; Kamran, M.; Alatawi, F.J. Molecular-Based Taxonomic Inferences of Some Spider Mite Species of the Genus Oligonychus Berlese (Acari, Prostigmata, Tetranychidae). Insects 2023, 14, 192. [CrossRef]
- de Mendonça, R.S.; Navia, D.; Diniz, I.R.; Auger, P.; Navajas, M. A Critical Review on Some Closely Related Species of Tetranychus Sensu Stricto (Acari: Tetranychidae) in the Public DNA Sequences Databases. Exp. Appl. Acarol. 2011, 55, 1–23. [CrossRef]
- Edgar, R.C. MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [CrossRef]
- Kumar, S.; Stecher, G.; Suleski, M.; Sanderford, M.; Sharma, S.; Tamura, K. MEGA12: Molecular Evolutionary Genetic Analysis Version 12 for Adaptive and Green Computing. Mol. Biol. Evol. 2024, 41, msae263. [CrossRef]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; deWaard, J.R. Biological Identifications through DNA Barcodes. Proc. Biol. Sci. 2003, 270, 313–321. [CrossRef]
- Navajas, M.; Lagnel, J.; Gutierrez, J.; Boursot, P. Species-Wide Homogeneity of Nuclear Ribosomal ITS2 Sequences in the Spider Mite Tetranychus Urticae Contrasts with Extensive Mitochondrial COI Polymorphism. Heredity 1998, 80, 742–752. [CrossRef]
- Matsuda, T.; Fukumoto, C.; Hinomoto, N.; Gotoh, T. DNA-Based Identification of Spider Mites: Molecular Evidence for Cryptic Species of the Genus Tetranychus (Acari: Tetranychidae). J. Econ. Entomol. 2013, 106, 463–472. [CrossRef]
- Knihinicki, D.K.; Flechtmann, C.H. A New Species of Spider Mite, Oligonychus Calicicola (Acari: Tetranychidae), Damaging Date Fruit, Phoenix Dactylifera L. (Arecaceae), in Australia. Aust. J. Entomol. 1999, 38, 176–178. [CrossRef]
- Krehenwinkel, H.; Fong, M.; Kennedy, S.; Huang, E.G.; Noriyuki, S.; Cayetano, L.; Gillespie, R. The Effect of DNA Degradation Bias in Passive Sampling Devices on Metabarcoding Studies of Arthropod Communities and Their Associated Microbiota. PLoS ONE 2018, 13, e0189188. [CrossRef]
- Pérez-Sayas, C.; Pina, T.; Sabater-Muñoz, B.; Gómez-Martínez, M.A.; Jaques, J.A.; Hurtado-Ruiz, M.A. DNA Barcoding and Phylogeny of Acari Species Based on ITS and COI Markers. J. Zool. Syst. Evol. Res. 2022, 2022, 5317995. [CrossRef]
- Ben-David, T.; Melamed, S.; Gerson, U.; Morin, S. ITS2 Sequences as Barcodes for Identifying and Analyzing Spider Mites (Acari: Tetranychidae). Exp. Appl. Acarol. 2007, 41, 169–181. [CrossRef]
- Matsuda, T.; Hinomoto, N.; Singh, R.N.; Gotoh, T. Molecular-Based Identification and Phylogeny of Oligonychus Species (Acari: Tetranychidae). J. Econ. Entomol. 2012, 105, 1043–1050. [CrossRef]
- Shim, J.; Khaing, T.M.; Seo, H.; Ahn, J.; Jung, D.; Lee, J.; Lee, K. Development of Species-specific Primers for Rapid Diagnosis of T Etranychus Urticae , T. Kanzawai , T. Phaselus and T. Truncatus ( A Cari: T Etranychidae). Entomol. Res. 2016, 46, 162–169. [CrossRef]
- Matsuda, T.; Kozaki, T.; Ishii, K.; Gotoh, T. Phylogeny of the Spider Mite Sub-Family Tetranychinae (Acari: Tetranychidae) Inferred from RNA-Seq Data. PLOS ONE 2018, 13, e0203136. [CrossRef]
- Razuvaeva, A.V.; Ulyanova, E.G.; Skolotneva, E.S.; Andreeva, I.V. Species Identification of Spider Mites (Tetranychidae: Tetranychinae): A Review of Methods. Vavilov J. Genet. Breed. 2023, 27, 240–249. [CrossRef]
| Species | Intraspecific divergence (%) |
| Oligonychus sp. (RC) | 0.59 (0–0.88) |
| Oligonychus biharensis | 3.14 (0–4.75) |
| Tetranychus cinnabarinus | 0.00 (0–0) |
| Tetranychus urticae | 2.10 (0–3.10) |
| Tetranychus evansi | 0.00 (0–0) |
| Panonychus ulmi | 0.00 (0–0) |
| Eotetranychus frosti | 0.00 (0–0) |
| Phytoseiulus persimilis | 0.00 (0–0) |
| Oligonychus sp. (Congo) | O. biharensis | T. cinnabarinus | T. urticae | T. evansi | P. ulmi | E. frosti | Phytoseiulus persimilis (outgroup) | |
|---|---|---|---|---|---|---|---|---|
| Oligonychus sp. (RC) | 0.00 | |||||||
| O. biharensis | 0.13 (0.1193–0.1410) | 0.00 | ||||||
| T. cinnabarinus | 0.16 (0.1552–0.1558) | 0.00 | ||||||
| T. urticae | 0.16 (0.1548–0.1618) | 0.00 | ||||||
| T. evansi | 0.16 (0.1491–0.1626) | 0.00 | ||||||
| P. ulmi | 0.15 (0.1521–0.1547) | 0.00 | ||||||
| E. frosti | 0.16 (0.1565–0.1571) | 0.00 | ||||||
| Phytoseiulus persimilis (outgroup) | 0.39 (0.3866–0.3881) | 0.00 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




