Submitted:
23 December 2025
Posted:
24 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
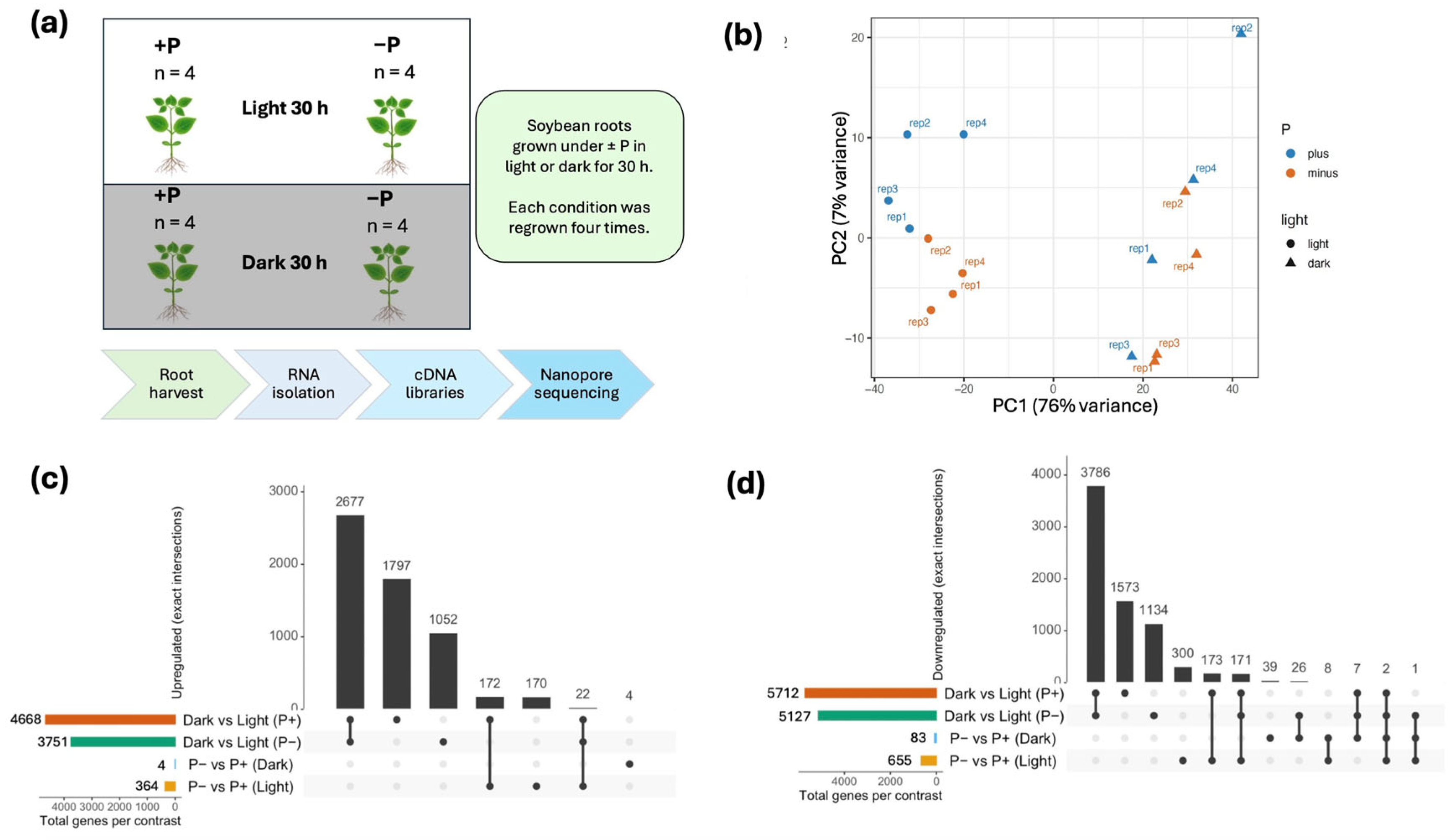
2.1. Overview of Experimental Design and Data Quality
2.2. RNA-Seq Reveals Major Effects of Light and Minor Effects of Pi Deficiency
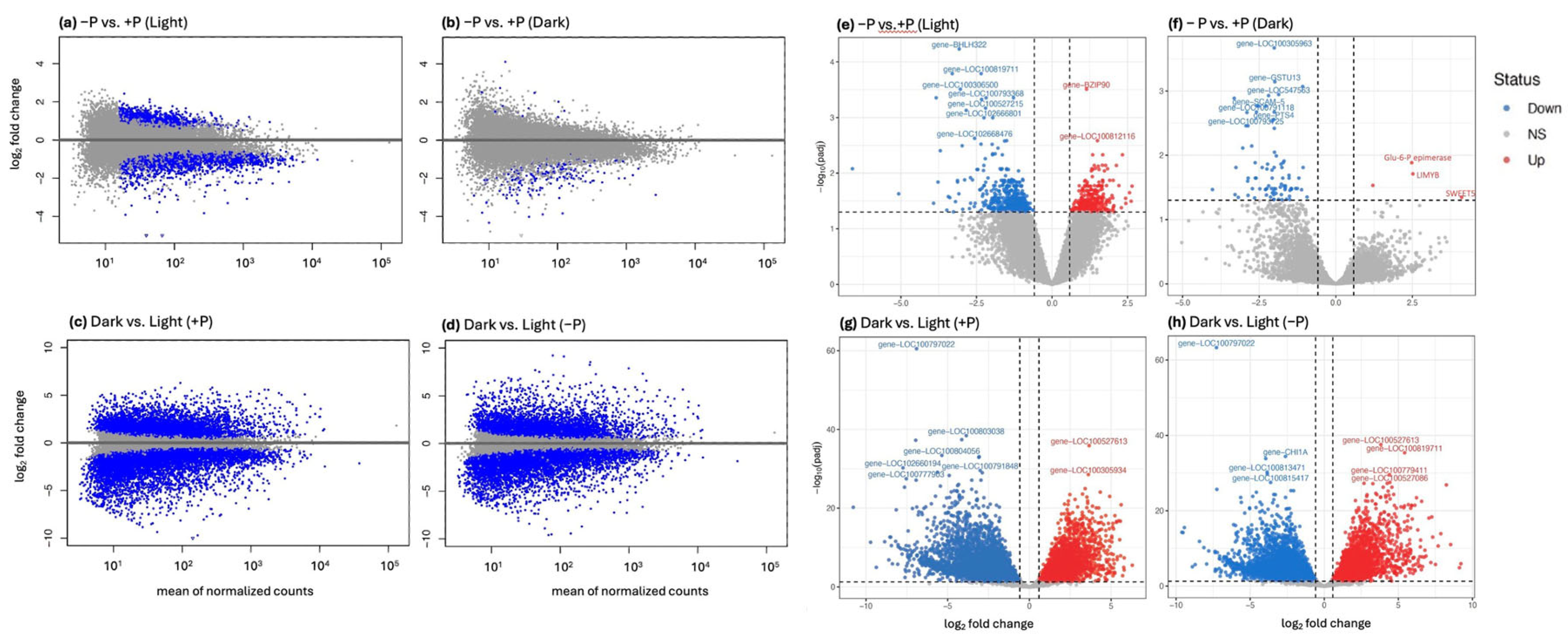
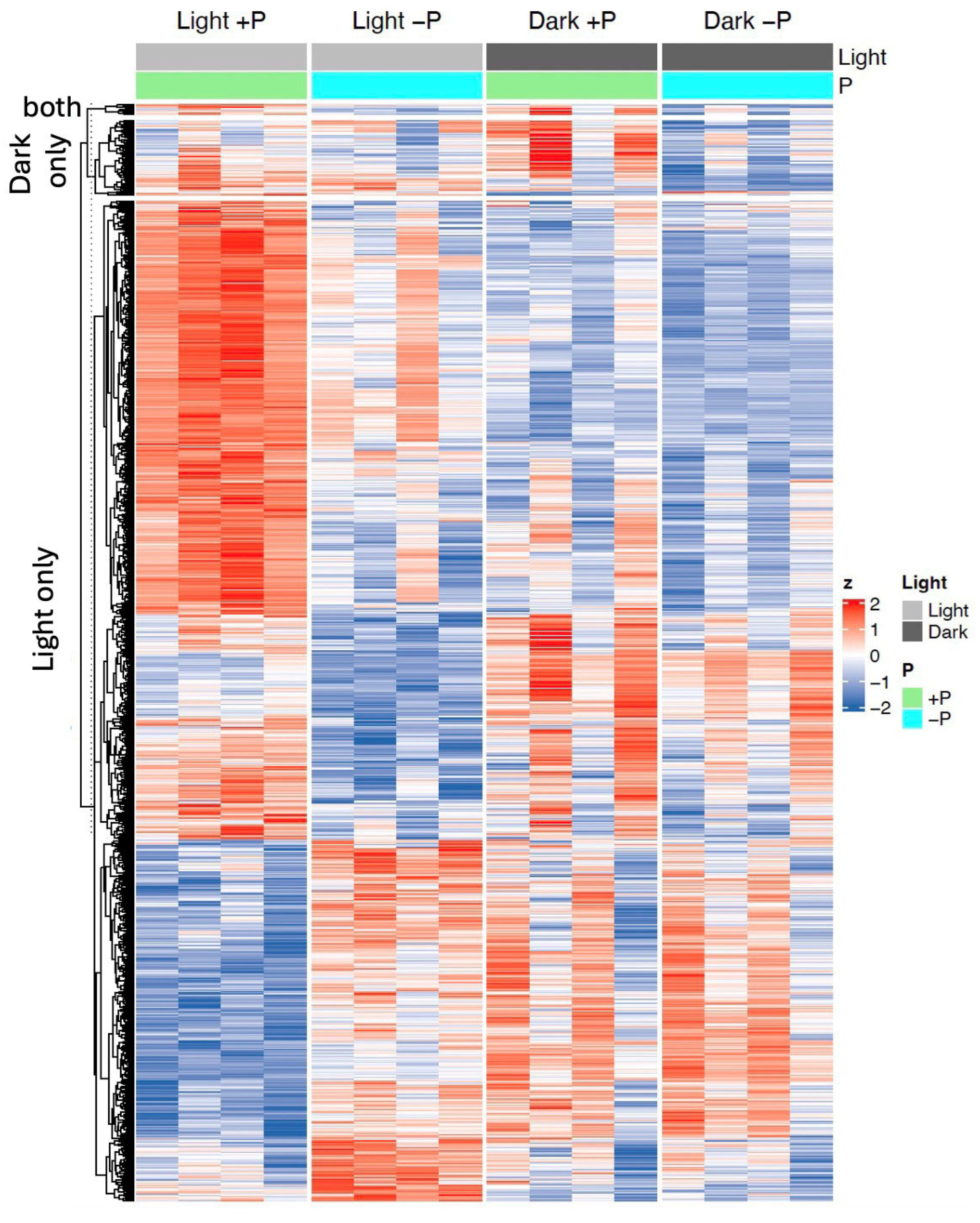
2.3. Light Enables Genome-Wide Activation of the Phosphate Starvation Program
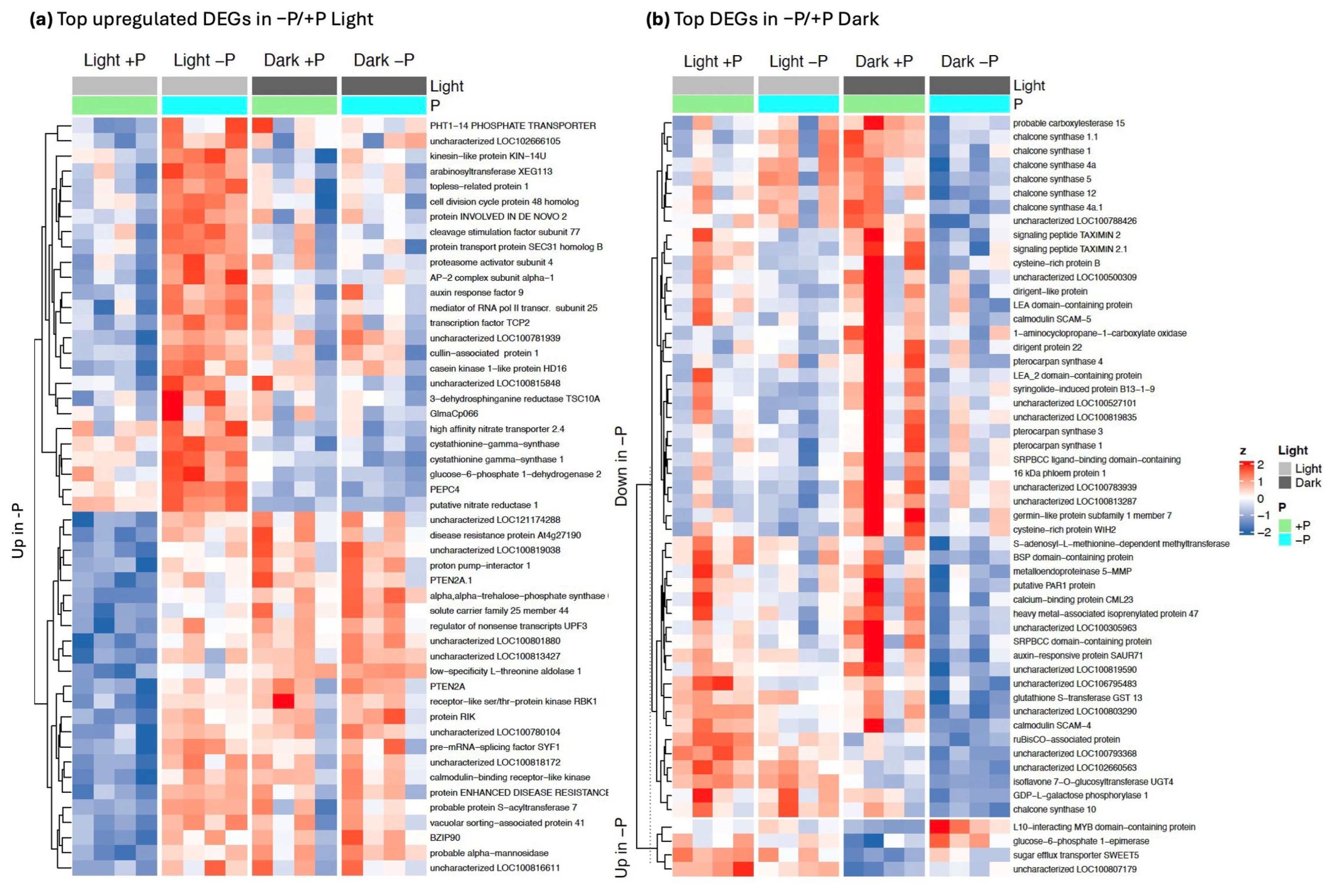
2.4. Canonical Phosphate Starvation Response Genes Are Induced Only in Light
2.5. Sensitivity Analysis Reveals a Small Set of Pi-Responsive Genes in Darkness
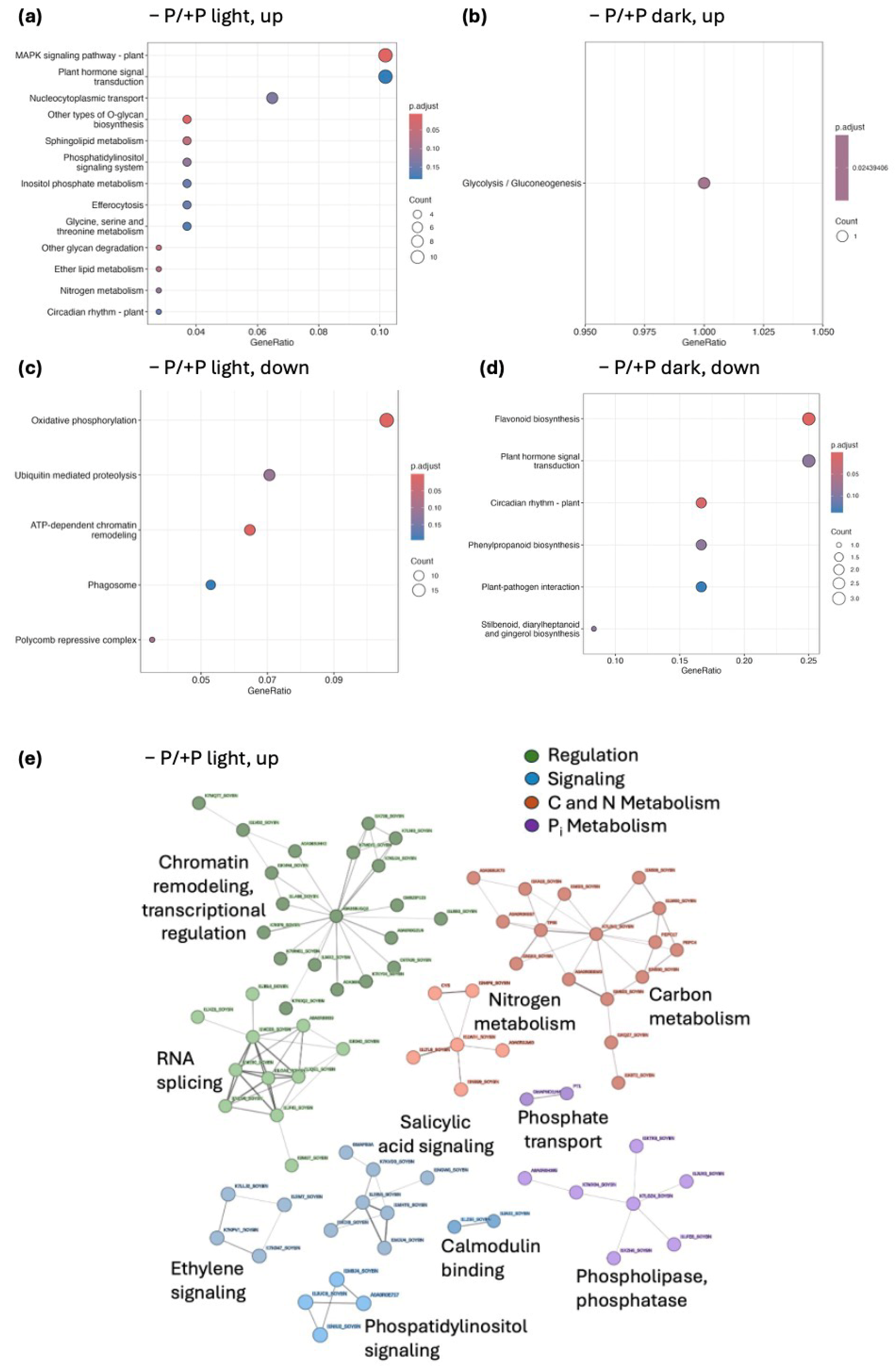
2.6. Functional Enrichment Highlights Light-Dependent Transcriptional Reprogramming During Pi Deficiency
2.7. Protein–Protein Interaction Network Identifies Core Response Modules
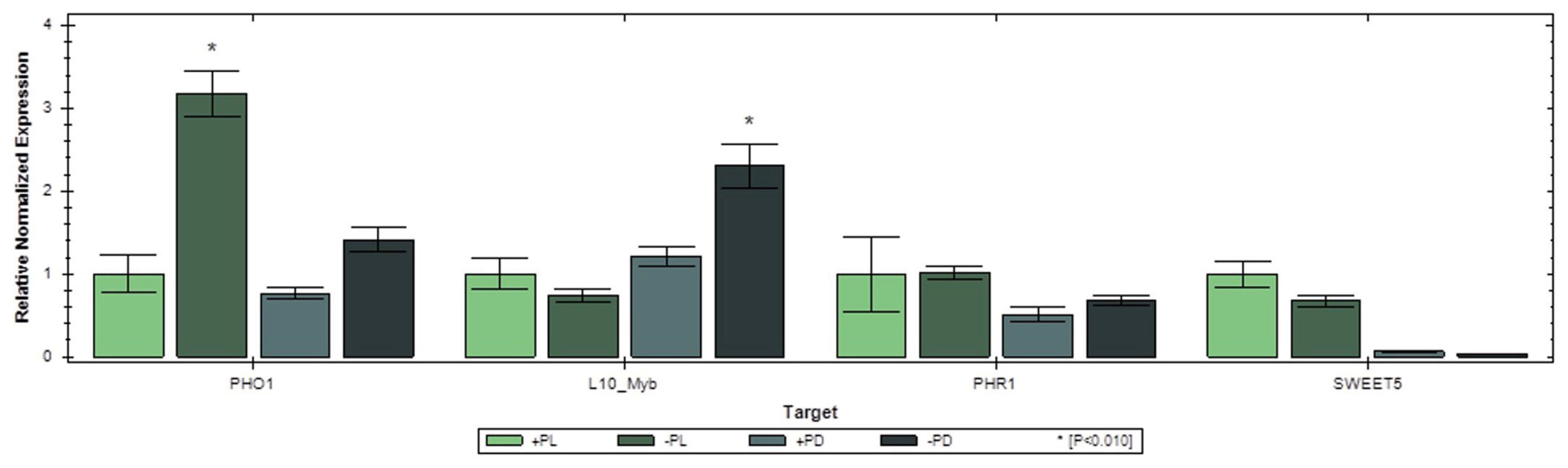
2.8. qRT-PCR Assessment of Selected Pi-Responsive Genes
3. Discussion
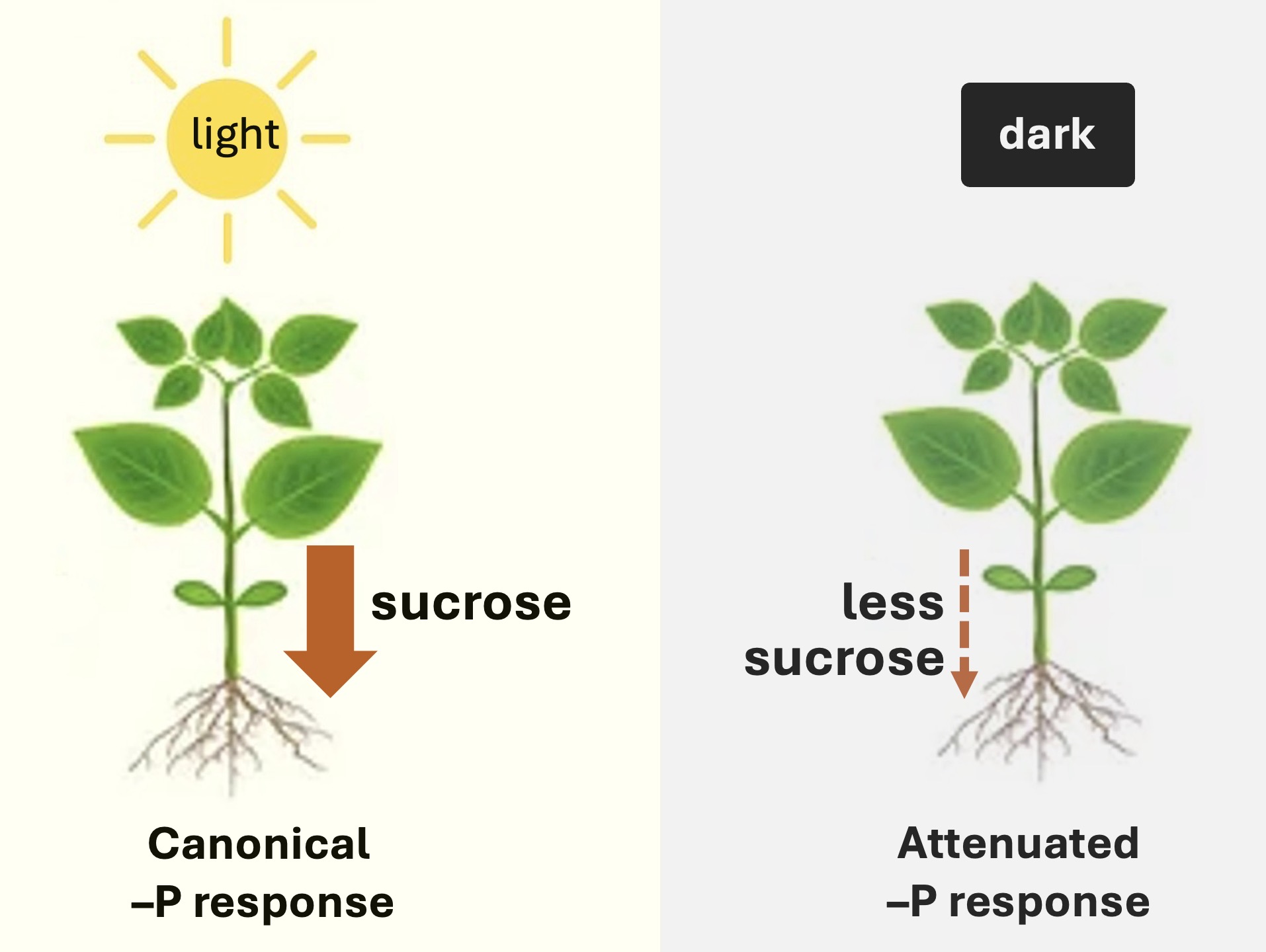
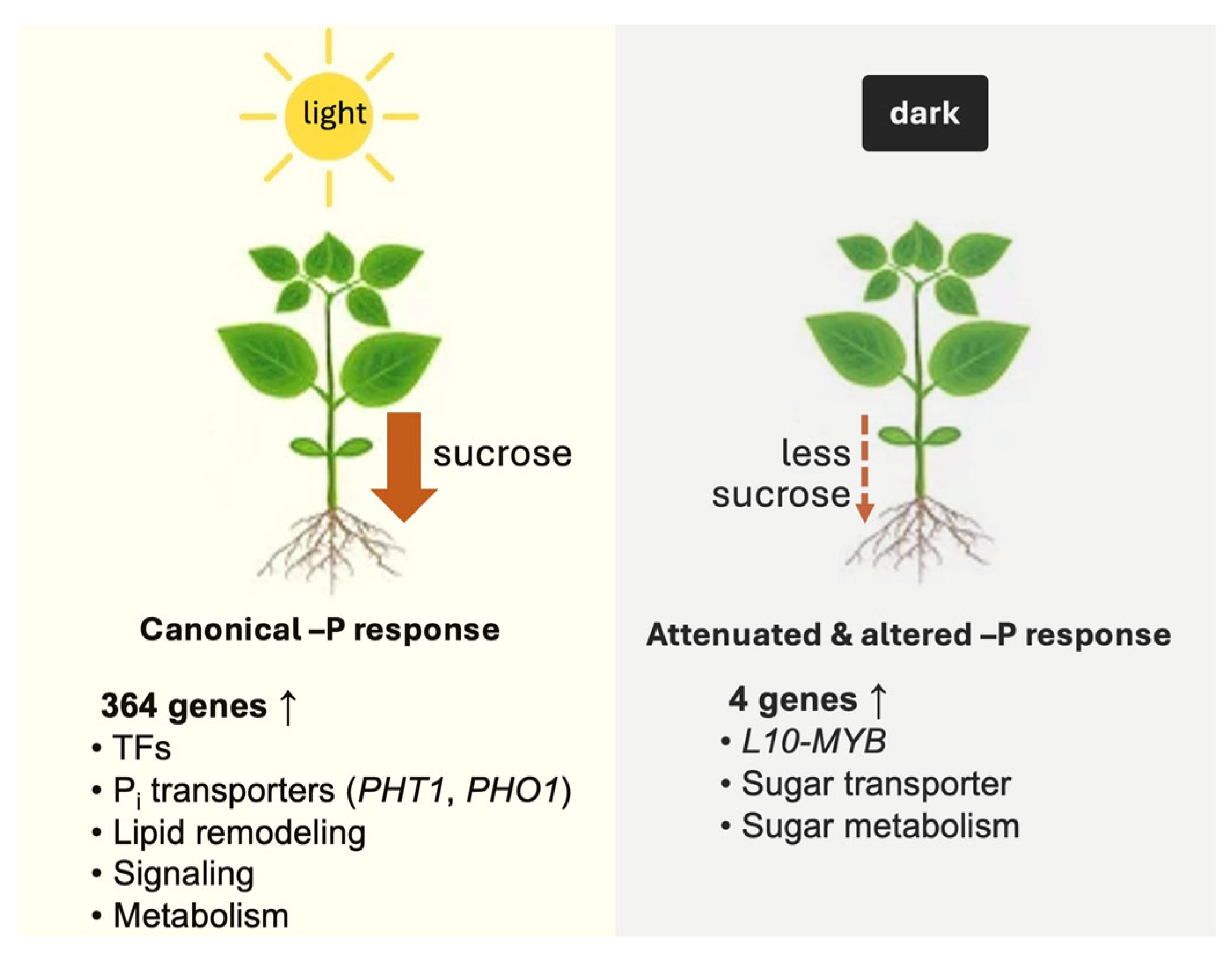
3.1. Systemic Pi Signaling Enabled by Light
3.2. Pi Signaling in Darkness: A Small Sucrose-Independent Response That Includes a MYB Transcription Factor
3.3. Limitations and Future Directions
4. Materials and Methods
4.1. Overview
4.2. Plant Growth and Treatment
4.3. RNA Isolation and Quality Assessment
4.4. cDNA Library Preparation and Quantification
4.5. Nanopore Sequencing
4.6. Nanopore Sequencing Analysis
4.7. Differential Expression Analysis
4.8. Validation by qRT-PCR
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| DEG | Differentially Expressed Genes |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| MA-plot | Mean–average plot (log₂ fold change vs mean expression) |
| padj | Adjusted p-value (Benjamini–Hochberg correction) |
| Pi | inorganic phosphate |
| PCA | Principal Component Analysis |
| PPi | Pyrophosphate |
| PSR | Phosphate starvation response |
| qRT-PCR | Quantitative reverse transcription PCR |
| RNA-seq | RNA sequencing |
References
- Vance, C.P.; Uhde-Stone, C.; Allan, D.L. Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. New Phytologist 2003, 157, 423–447. [Google Scholar] [CrossRef]
- Cordell, D.; Drangert, J.; White, S. The story of phosphorus: Global food security and food for thought. Global Environmental Change 2009, 19, 292–305. [Google Scholar] [CrossRef]
- Geissler, B.; Mew, M.C.; Matschullat, J.; Steiner, G. Innovation potential along the phosphorus supply chain: A micro and macro perspective on the mining phase. Sci Total Environ 2020, 714, 136701. [Google Scholar] [CrossRef] [PubMed]
- Alewell, C.; Ringeval, B.; Ballabio, C.; Robinson, D.A.; Panagos, P.; Borrelli, P. Global phosphorus shortage will be aggravated by soil erosion. Nature communications 2020, 11, 4546. [Google Scholar] [CrossRef] [PubMed]
- Volkova, E.; Smolyaninova, N. Analysis of world trends in soybean production. In Proceedings of the BIO Web of Conferences, 2024; p. 01026. [Google Scholar]
- Cai, Z.; Cheng, Y.; Xian, P.; Ma, Q.; Wen, K.; Xia, Q.; Zhang, G.; Nian, H. Acid phosphatase gene GmHAD1 linked to low phosphorus tolerance in soybean, through fine mapping. TAG. Theoretical and applied genetics. Theoretische und angewandte Genetik 2018, 131, 1715–1728. [Google Scholar] [CrossRef]
- O'Rourke, J.A.; Graham, M.A. Gene Expression Responses to Sequential Nutrient Deficiency Stresses in Soybean. International journal of molecular sciences 2021, 22. [Google Scholar] [CrossRef]
- Chen, G.; Ran, Q.-X.; Wang, C.; Pang, J.; Ren, M.-J.; Wang, Z.-Y.; He, J.; Lambers, H. Enhancing photosynthetic phosphorus use efficiency through coordination of leaf phosphorus fractions, allocation, and anatomy during soybean domestication. Journal of Experimental Botany 2025, 76, 1446–1457. [Google Scholar] [CrossRef]
- Fang, Y.; Lu, L.; Chen, K.; Wang, X. Tradeoffs among root functional traits for phosphorus acquisition in 13 soybean genotypes contrasting in mycorrhizal colonization. Annals of Botany 2024, 134, 179–190. [Google Scholar] [CrossRef]
- Tantriani; Cheng, W.; Oikawa, A.; Tawaraya, K. Phosphorus deficiency alters root length, acid phosphatase activity, organic acids, and metabolites in root exudates of soybean cultivars. Physiol Plant 2023, 175, e14107. [Google Scholar] [CrossRef]
- Péret, B.; Clément, M.; Nussaume, L.; Desnos, T. Root developmental adaptation to phosphate starvation: better safe than sorry. Trends in plant science 2011. [Google Scholar] [CrossRef]
- Xu, H.; Zhang, H.; Fan, Y.; Wang, R.; Cui, R.; Liu, X.; Chu, S.; Jiao, Y.; Zhang, X.; Zhang, D. The purple acid phosphatase GmPAP17 predominantly enhances phosphorus use efficiency in soybean. Plant Sci 2022, 320, 111283. [Google Scholar] [CrossRef]
- Li, F.; Mai, C.; Liu, Y.; Deng, Y.; Wu, L.; Zheng, X.; He, H.; Huang, Y.; Luo, Z.; Wang, J. Soybean PHR1-regulated low phosphorus-responsive GmRALF22 promotes phosphate uptake by stimulating the expression of GmPTs. Plant Sci 2024, 348, 112211. [Google Scholar] [CrossRef]
- Qin, L.; Guo, Y.; Chen, L.; Liang, R.; Gu, M.; Xu, G.; Zhao, J.; Walk, T.; Liao, H. Functional characterization of 14 Pht1 family genes in yeast and their expressions in response to nutrient starvation in soybean. PloS one 2012, 7, e47726. [Google Scholar] [CrossRef]
- Qin, L.; Zhao, J.; Tian, J.; Chen, L.; Sun, Z.; Guo, Y.; Lu, X.; Gu, M.; Xu, G.; Liao, H. The high-affinity phosphate transporter GmPT5 regulates phosphate transport to nodules and nodulation in soybean. Plant physiology 2012, 159, 1634–1643. [Google Scholar] [CrossRef]
- Miao, N.; Zhou, J.; Li, M.; Zhang, J.; Hu, Y.; Guo, J.; Zhang, T.; Shi, L. Remodeling and protecting the membrane system to resist phosphorus deficiency in wild soybean (Glycine soja) seedling leaves. Planta 2022, 255, 53. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Yuan, H.; Liu, D.; Cheng, L. Response of soybean root exudates and related metabolic pathways to low phosphorus stress. PloS one 2024, 19, e0314256. [Google Scholar] [CrossRef]
- Plaxton, W.C.; Tran, H.T. Metabolic adaptations of phosphate-starved plants. Plant physiology 2011, 156, 1006–1015. [Google Scholar] [CrossRef]
- Ried, M.K.; Wild, R.; Zhu, J.; Pipercevic, J.; Sturm, K.; Broger, L.; Harmel, R.K.; Abriata, L.A.; Hothorn, L.A.; Fiedler, D.; et al. Inositol pyrophosphates promote the interaction of SPX domains with the coiled-coil motif of PHR transcription factors to regulate plant phosphate homeostasis. Nature communications 2021, 12, 384. [Google Scholar] [CrossRef] [PubMed]
- Secco, D.; Wang, C.; Arpat, B.A.; Wang, Z.; Poirier, Y.; Tyerman, S.D.; Wu, P.; Shou, H.; Whelan, J. The emerging importance of the SPX domain-containing proteins in phosphate homeostasis. New Phytologist 2012, 193, 842–851. [Google Scholar] [CrossRef]
- Mo, X.; Liu, G.; Zhang, Z.; Lu, X.; Liang, C.; Tian, J. Mechanisms Underlying Soybean Response to Phosphorus Deficiency through Integration of Omics Analysis. International journal of molecular sciences 2022, 23. [Google Scholar] [CrossRef] [PubMed]
- Hamburger, D.; Rezzonico, E.; MacDonald-Comber Petetot, J.; Somerville, C.; Poirier, Y. Identification and characterization of the Arabidopsis PHO1 gene involved in phosphate loading to the xylem. The Plant cell 2002, 14, 889–902. [Google Scholar] [CrossRef] [PubMed]
- Hammond, J.; White, P. Sucrose transport in the phloem: integrating root responses to phosphorus starvation. J Exp Bot 2008, 59, 93–109. [Google Scholar] [CrossRef] [PubMed]
- Karthikeyan, A.S.; Varadarajan, D.K.; Jain, A.; Held, M.A.; Carpita, N.C.; Raghothama, K.G. Phosphate starvation responses are mediated by sugar signaling in Arabidopsis. Planta 2007, 225, 907–918. [Google Scholar] [CrossRef] [PubMed]
- Ruffel, S. Nutrient-Related Long-Distance Signals: Common Players and Possible Cross-Talk. Plant Cell Physiol 2018, 59, 1723–1732. [Google Scholar] [CrossRef]
- Hammond, J.P.; White, P.J. Sugar signaling in root responses to low phosphorus availability. Plant physiology 2011, 156, 1033–1040. [Google Scholar] [CrossRef]
- Lei, M.; Liu, Y.; Zhang, B.; Zhao, Y.; Wang, X.; Zhou, Y.; Raghothama, K.G.; Liu, D. Genetic and genomic evidence that sucrose is a global regulator of plant responses to phosphate starvation in Arabidopsis. Plant physiology 2011, 156, 1116–1130. [Google Scholar] [CrossRef] [PubMed]
- Nidumolu, L.C.M.; Lorilla, K.M.; Chakravarty, I.; Uhde-Stone, C. Soybean Root Transcriptomics: Insights into Sucrose Signaling at the Crossroads of Nutrient Deficiency and Biotic Stress Responses. Plants (Basel) 2023, 12. [Google Scholar] [CrossRef]
- Franco-Zorrilla, J.; González, E.; Bustos, R.; Linhares, F.; Leyva, A.; Paz-Ares, J. The transcriptional control of plant responses to phosphate limitation. J Exp Bot 2004, 55, 285–293. [Google Scholar] [CrossRef]
- Chien, P.S.; Chiang, C.P.; Leong, S.J.; Chiou, T.J. Sensing and Signaling of Phosphate Starvation - from Local to Long Distance. Plant Cell Physiol 2018. [Google Scholar] [CrossRef]
- Lin, W.-Y.; Huang, T.-K.; Leong, S.J.; Chiou, T.-J. Long-distance call from phosphate: systemic regulation of phosphate starvation responses. Journal of experimental botany 2014, 65, 1817–1827. [Google Scholar] [CrossRef]
- Marro, N.; Lidoy, J.; Chico, M.A.; Rial, C.; Garcia, J.; Varela, R.M.; Macias, F.A.; Pozo, M.J.; Janouskova, M.; Lopez-Raez, J.A. Strigolactones: New players in the nitrogen-phosphorus signalling interplay. Plant Cell Environ 2022, 45, 512–527. [Google Scholar] [CrossRef] [PubMed]
- Poirier, Y.; Jaskolowski, A.; Clua, J. Phosphate acquisition and metabolism in plants. Current biology: CB 2022, 32, R623–R629. [Google Scholar] [CrossRef]
- Rouached, H.; Arpat, A.B.; Poirier, Y. Regulation of phosphate starvation responses in plants: signaling players and cross-talks. Molecular plant 2010, 3, 288–299. [Google Scholar] [CrossRef]
- Lin, X.Y.; Ye, Y.Q.; Fan, S.K.; Jin, C.W.; Zheng, S.J. Increased Sucrose Accumulation Regulates Iron-Deficiency Responses by Promoting Auxin Signaling in Arabidopsis Plants. Plant physiology 2016, 170, 907–920. [Google Scholar] [CrossRef]
- Madison, I.; Gillan, L.; Peace, J.; Gabrieli, F.; Van den Broeck, L.; Jones, J.L.; Sozzani, R. Phosphate starvation: response mechanisms and solutions. J Exp Bot 2023, 74, 6417–6430. [Google Scholar] [CrossRef]
- Pribat, A.; Sormani, R.; Rousseau-Gueutin, M.; Julkowska, M.M.; Testerink, C.; Joubes, J.; Castroviejo, M.; Laguerre, M.; Meyer, C.; Germain, V.; et al. A novel class of PTEN protein in Arabidopsis displays unusual phosphoinositide phosphatase activity and efficiently binds phosphatidic acid. Biochem J 2012, 441, 161–171. [Google Scholar] [CrossRef]
- Price, J.; Laxmi, A.; St Martin, S.; Jang, J. Global transcription profiling reveals multiple sugar signal transduction mechanisms in Arabidopsis. The Plant cell 2004, 16, 2128–2150. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Samac, D.; Bucciarelli, B.; Allan, D.; Vance, C. Signaling of phosphorus deficiency-induced gene expression in white lupin requires sugar and phloem transport. The Plant journal: for cell and molecular biology 2005, 41, 257–268. [Google Scholar] [CrossRef] [PubMed]
- Yang, A.; Kong, L.; Wang, H.; Yao, X.; Xie, F.; Wang, H.; Ao, X. Response of Soybean Root to Phosphorus Deficiency under Sucrose Feeding: Insight from Morphological and Metabolome Characterizations. BioMed research international 2020, 2020, 2148032. [Google Scholar] [CrossRef]
- Rubio, V.; Linhares, F.; Solano, R.; Martín, A.; Iglesias, J.; Leyva, A.; Paz-Ares, J. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes & development 2001, 15, 2122–2133. [Google Scholar] [CrossRef]
- Ferreira, M.A.; Teixeira, R.M.; Brustolini, O.J.B.; Saia, T.F.F.; Jean-Baptiste, J.; Ribeiro, N.G.A.; Breves, S.S.; Sampaio, F.R.; Santos, E.G.D.; Leon, B.A.; et al. The immune NIK1/RPL10/LIMYB signaling module regulates photosynthesis and translation under biotic and abiotic stresses. Nature communications 2025, 16, 4433. [Google Scholar] [CrossRef]
- Puga, M.I.; Poza-Carrion, C.; Martinez-Hevia, I.; Perez-Liens, L.; Paz-Ares, J. Recent advances in research on phosphate starvation signaling in plants. J Plant Res 2024, 137, 315–330. [Google Scholar] [CrossRef] [PubMed]
- Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Genome Project Data Processing, S. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014, 15, 550. [Google Scholar] [CrossRef]
- Conway, J.R.; Lex, A.; Gehlenborg, N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 2017, 33, 2938–2940. [Google Scholar] [CrossRef]
- Gu, Z.; Hubschmann, D. Make Interactive Complex Heatmaps in R. Bioinformatics 2022, 38, 1460–1462. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation (Camb) 2021, 2, 100141. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Nastou, K.; Koutrouli, M.; Kirsch, R.; Mehryary, F.; Hachilif, R.; Hu, D.; Peluso, M.E.; Huang, Q.; Fang, T.; et al. The STRING database in 2025: protein networks with directionality of regulation. Nucleic acids research 2025, 53, D730–D737. [Google Scholar] [CrossRef]
| Condition | Passed reads (×10⁶) | Avg. read Quality (Q) |
N50 (bp) |
Mapped reads (%) |
| +P Light (n = 4) | 6.9 ± 2.5 | 11 ± 0 | 1,034 ± 75 | 96.9 ± 0.5 |
| −P Light (n = 4) | 4.8 ± 1.9 | 10 ± 0 | 814 ± 96 | 93.4 ± 2.6 |
| +P Dark (n = 4) | 6.7 ± 2.2 | 10 ± 0 | 879 ± 113 | 95.6 ± 1.5 |
| −P Dark (n = 4) | 4.9 ± 1.1 | 11 ± 0 | 972 ± 103 | 96.8 ± 0.4 |
|
Canonical PSR markers* |
Light (+P) | Light (-P) | Dark (+P) | Dark (-P) |
| PHO1 | 28.1 | 68.7 | 23.6 | 31.7 |
| PHT1 | 537.5 | 1111.0 | 943.6 | 806.5 |
| PTEN2α-1 | 38.6 | 84.0 | 111.8 | 97.5 |
| PTEN2α-2 | 76.9 | 155.0 | 172.2 | 184.9 |
| SAP | 8.9 | 24.3 | 53.9 | 71.4 |
| Gene_ID | Description |
−P/+P dark log2FC |
−P/+P dark padj |
−P/+P light log2FC |
−P/+P light padj |
| SWEET5 | sugar efflux transporter SWEET5 | 4.1 | 0.045 | -0.9 | 0.215 |
| LOC100787217 | glucose-6-phosphate 1-epimerase | 2.5 | 0.013 | -0.4 | 0.612 |
| LOC100820117 | L10-interacting MYB domain-containing protein | 2.5 | 0.019 | 0.1 | NA |
| LOC100807179 | uncharacterized LOC100807179 | 1.2 | 0.029 | -0.7 | 0.072 |
| LOC100818214 | WEB family protein | 2.7 | 0.061 | 0.6 | NA |
| LOC100776105 | arabinogalactan protein 9 |
2.0 | 0.091 | -0.7 | 0.351 |
| PHR11 | PSR 11 | 1.8 | 0.061 | -0.2 | 0.738 |
| LOC100775446 | aspartyl protease AED3 | 1.7 | 0.055 | -0.1 | 0.878 |
| LOC113000488 | L10-interacting MYB domain-containing protein | 1.5 | 0.077 | 0.6 | NA |
| LOC100808302 | cyclin-dependent kinase B2-2 |
1.5 | 0.078 | -0.6 | 0.352 |
| LOC100785581 | kinesin-like protein KIN-14U | 1.5 | 0.089 | 1.7 | 0.006 |
| LOC100804267 | uncharacterized | 1.5 | 0.091 | 0.2 | 0.766 |
| LOC100802287 | glucan endo-1,3-beta-glucosidase 7 | 1.3 | 0.077 | -0.3 | 0.588 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




