Submitted:
12 December 2025
Posted:
15 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Genetic Engineering
2.2. mRNA Synthesis
2.2.1. In vitro Transcription
2.2.2. Dephosphorylation
2.2.3. Polyadenylation
2.2.4. Purification
2.2.5. Quality Control
2.3. Cell Culture
2.4. Transfection Efficiency and mRNA Expression
2.5. Translation Levels (Western Blotting)
2.5. Cell Response to mRNA Invasion
2.5.1. Multiplex Analysis
2.5.2. RT-qPCR
2.6. Statistical Analysis
3. Results
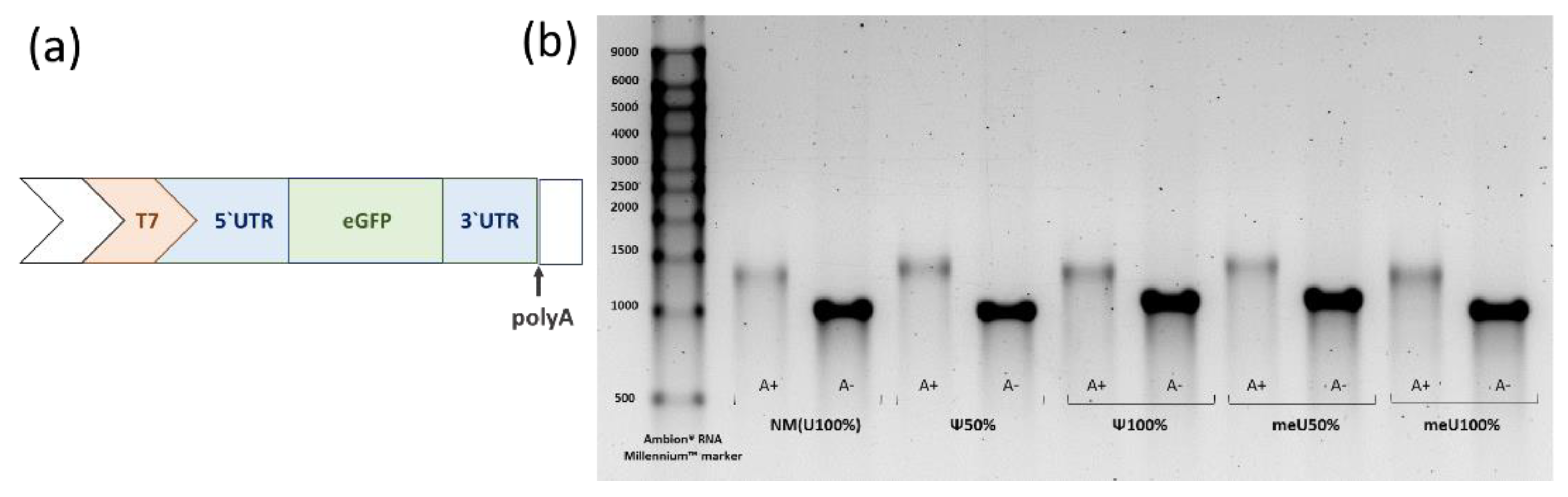
3.1. mRNA Preparations
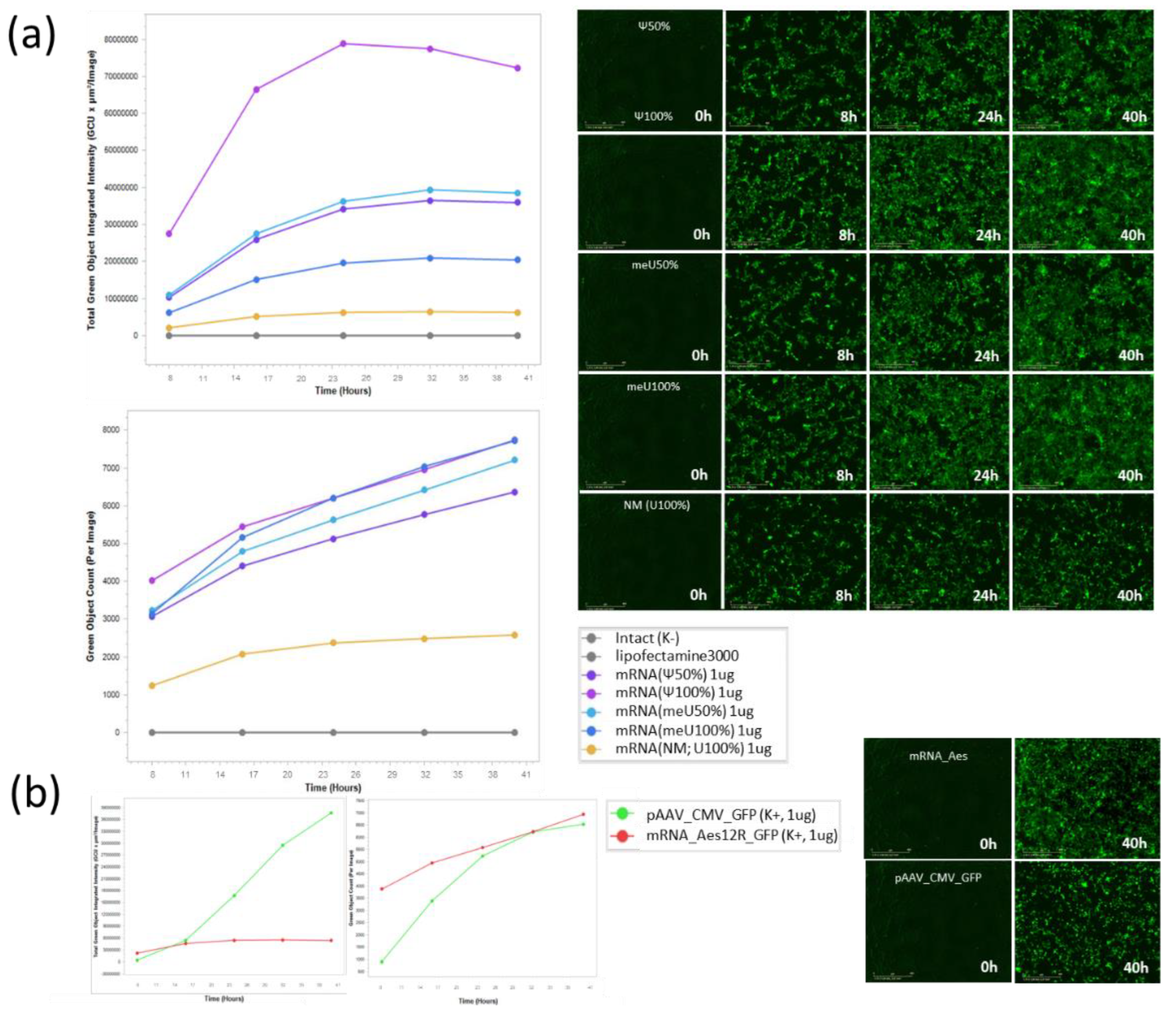
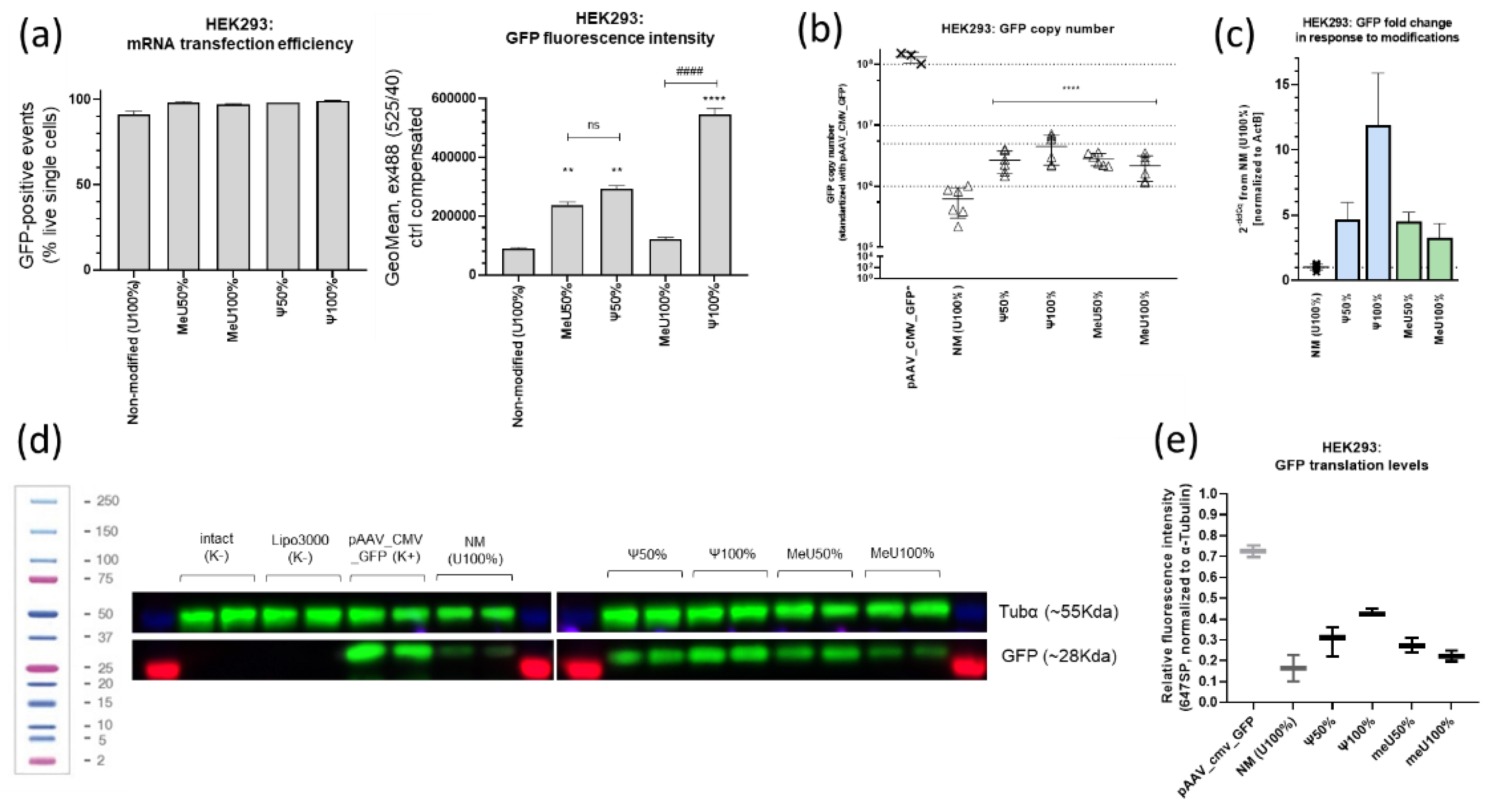
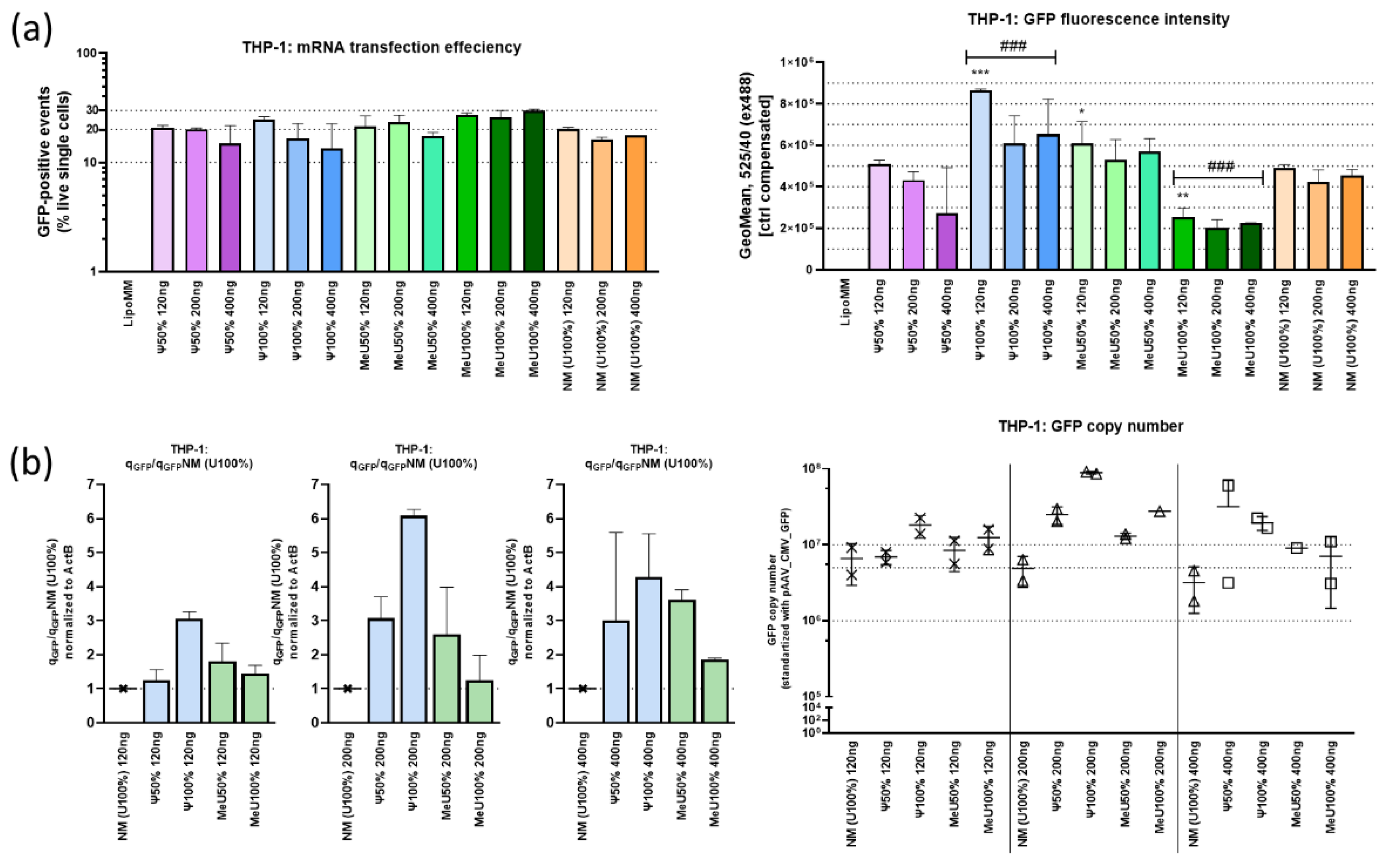
3.2. Comparison of Uridine-Modified mRNA Preparations: Fluorescence Intensity, GFP Expression and Protein Translation
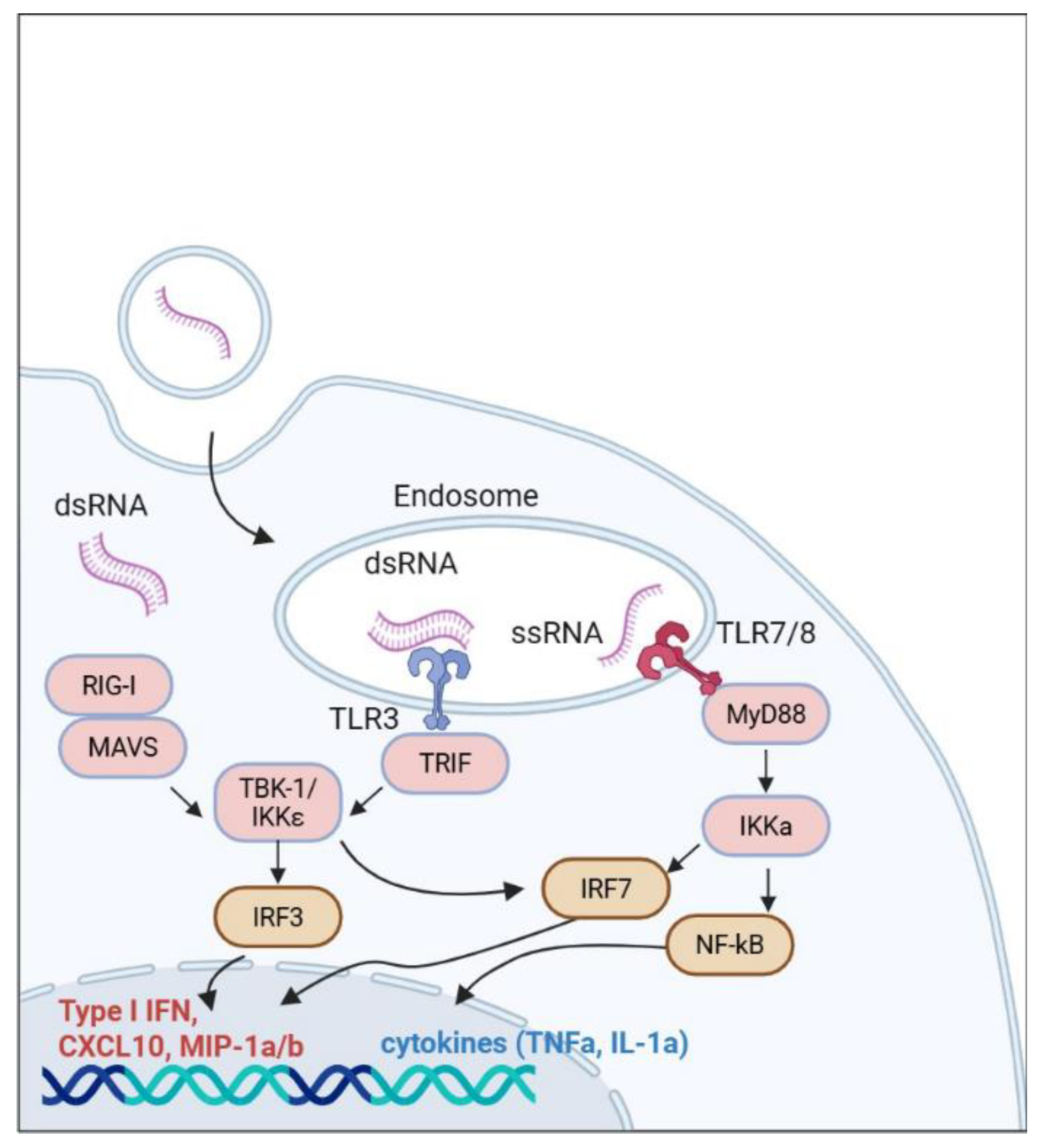
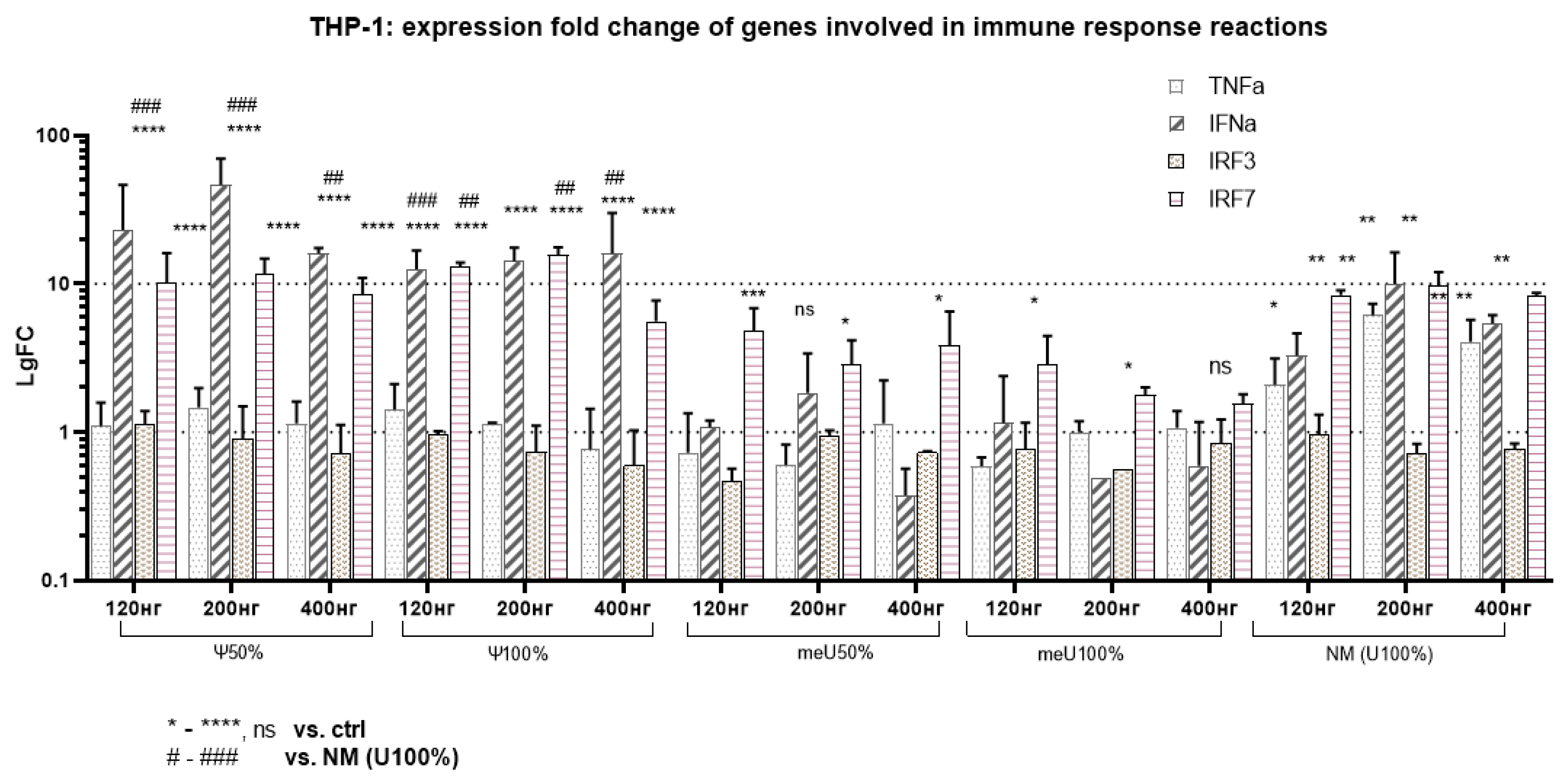
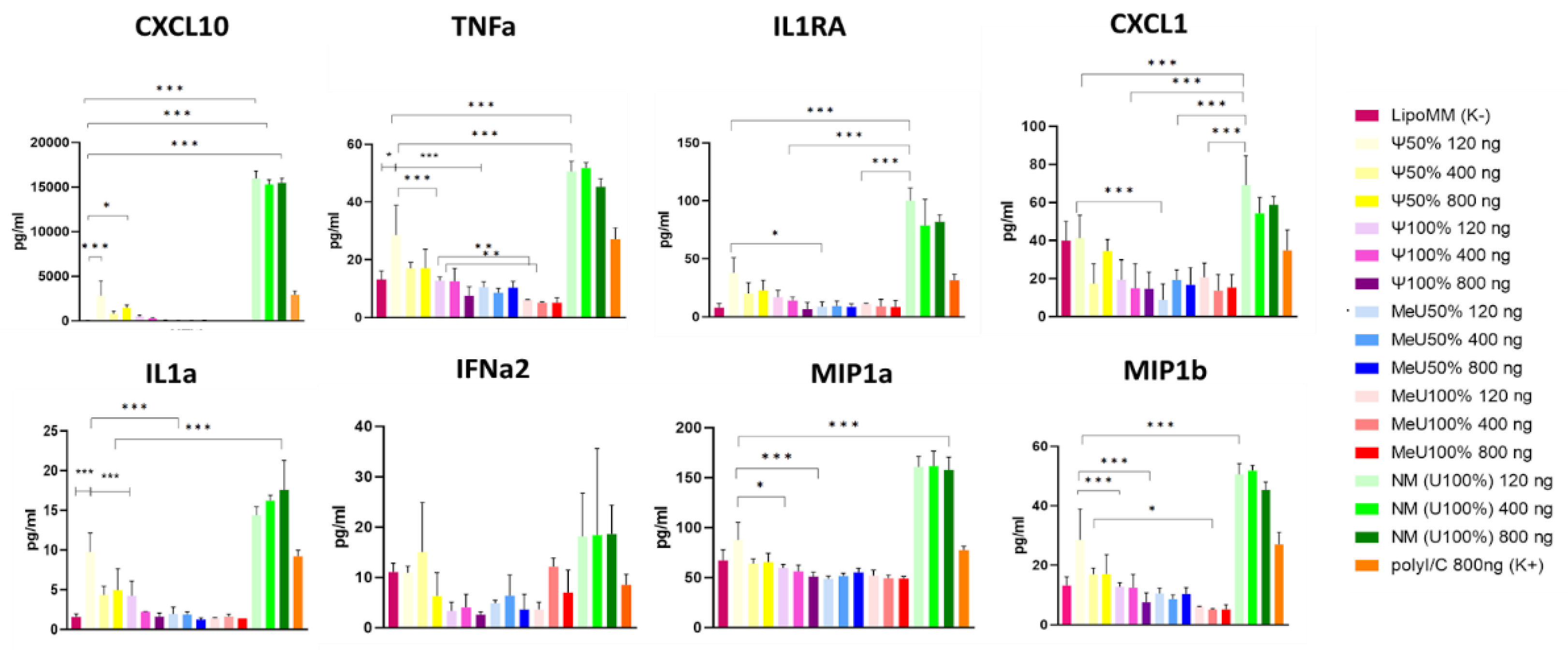
3.3. Altered Expression of Genes Involved in Innate Immune Response
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CCR | CC chemokine receptors (or beta chemokine receptors) |
| CXCL 1 or 10 | C-X-C motif chemokine ligand |
| EGF | Epidermal growth factor |
| eIF-2A | Eukaryotic translation initiation factor 2A |
| FGF-2 | Fibroblast growth factor |
| Flt-3L | Fms-related tyrosine kinase 3 ligand |
| G-CSF, GM-CSF | Granulocyte and granulocyte-macrophage colony-stimulating factor |
| GFP | Green Fluorescent Protein |
| GRO | Growth-Regulated Oncogene |
| IFN | Interferon |
| IL | Interleukin |
| IP-10 | Interferon-gamma-inducible protein 10 |
| IRF | Interferon Regulatory Factor |
| MCP | Monocyte chemoattractant protein |
| MDA | Melanoma differentiation-associated protein |
| MDC | Macrophage-Derived Chemokine (CCL22) |
| MIP | Macrophage Inflammatory Protein |
| MyD88 | Myeloid differentiation primary response 88 |
| NF-κB | Nuclear factor kappa-light-chain-enhancer of activated B cells |
| PKR | Protein Kinase R |
| poly(I:C) | Polyinosinic:polycytidylic acid |
| RIG-1 | Retinoic acid-inducible gene 1 |
| sCD40L | soluble CD40 ligand, |
| TGF | Transforming growth factor |
| TRIF | TIR-domain containing adaptor protein |
| TLR | Toll-like receptor |
| TNF | Tumor necrosis factor |
| VEGF | Vascular endothelial growth factor |
References
- Shi, Y.; et al. Progress and prospects of mRNA-based drugs in pre-clinical and clinical applications. Signal Transduct. Target. Ther. 2024; 9(1): pp.1–20. [CrossRef]
- Xie, C.; Yao, R.; Xia, X. The advances of adjuvants in mRNA vaccines. NPJ Vaccines 2023; 8(1): 162. [CrossRef]
- Verbeke, R.; Lentacker, I.; De Smedt, S.C.; Dewitte, H. Three decades of messenger RNA vaccine development. Nano Today 2019; 28 (100766). [CrossRef]
- Onomoto, K.; Onoguchi, K.; Yoneyama, M. Regulation of RIG-I-like receptor-mediated signaling: interaction between host and viral factors. Cell Mol Immunol 2021; 18: pp. 539–555. [CrossRef]
- Markiewicz, L.; Drazkowska, K.; Sikorski, P.J. Tricks and threats of RNA viruses - towards understanding the fate of viral RNA. RNA Biol 2021; 18(5): pp. 669–687. [CrossRef]
- Hühn, M.H.; McCartney, S.A.; Lind, K.; Svedin, E.; Colonna, M.; Flodström-Tullberg, M. et al. Melanoma differentiation-associated protein-5 (MDA-5) limits early viral replication but is not essential for the induction of type 1 interferons after Coxsackievirus infection. Virology 2010; 401(1): pp. 42–48. [CrossRef]
- Gitlin, L.; et al. Essential role of mda-5 in type I IFN responses to polyriboinosinic: polyribocytidylic acid and encephalomyo-carditis picornavirus. Proc. Natl. Acad. Sci. U. S. A. 2006; 103(22): pp. 8459–8464.
- Uehata, T.; Takeuchi, O. RNA Recognition and Immunity-Innate Immune Sensing and Its Posttranscriptional Regulation Mechanisms. Cells 2020; 9(7): pp. 1701. [CrossRef]
- Chai, Z.; Zhou, Y.; Yang, L.; et al. MyD88 Contributes to TLR3-Mediated NF-κB Activation and Cytokine Production in Macrophages. Cells 2025; 14(19): pp. 1507. [CrossRef]
- Kato, H.; Takeuchi, O.; Sato, S.. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 2006; 441(1): pp. 101–105. [CrossRef]
- McNab, F.; Mayer-Barber, K.; Sher, A.; Wack, A.; O’Garra, A. Type I interferons in infectious disease. Nat Rev Immunol 2015; 15(2): pp. 87–103. [CrossRef]
- Dias Junior, A.G.; Sampaio, N.G.; Rehwinkel, J. A Balancing Act: MDA5 in Antiviral Immunity and Autoinflammation. Trends Microbiol 2019; 27(1): pp. 75–85. [CrossRef]
- Stumpo, D.J.; Lai, W.S.; Blackshear, P.J. Inflammation: cytokines and RNA-based regulation. Wiley Interdiscip Rev RNA 2010; 1(1): pp. 60–80. [CrossRef]
- Singh, S.; Anshita, D.; Ravichandiran, V. MCP-1: Function, regulation, and involvement in disease. Int Immunopharmacol 2021; 101(Pt B): 107598. [CrossRef]
- Jafarzadeh, A.; Chauhan, P.; Saha, B.; Jafarzadeh, S.; Nemati, M. Contribution of monocytes and macrophages to the local tissue inflammation and cytokine storm in COVID-19: Lessons from SARS and MERS, and potential therapeutic interventions. Life Sci 2020; 257: pp. 118102. [CrossRef]
- Freund, I.; Eigenbrod, T.; Helm, M.; Dalpke, A.H. RNA Modifications Modulate Activation of Innate Toll-Like Receptors. Genes (Basel) 2019; 10(2): 92. [CrossRef]
- Verbeke, R.; Hogan, M.J.; Loré, K.; Pardi, N. Innate immune mechanisms of mRNA vaccines. Immunity 2022; 55(11): pp. 1993–2005. [CrossRef]
- Thompson, M.G.; Sacco, M.T.; Horner, S.M. How RNA modifications regulate the antiviral response. Immunol Rev 2021; 304(1): pp. 169–180. [CrossRef]
- Svitkin, Y.V.; Cheng, Y.M.; Chakraborty, T.; Presnyak, V.; John, M.; Sonenberg, N. N1-methyl-pseudouridine in mRNA enhances translation through eIF2α-dependent and independent mechanisms by increasing ribosome density. Nucleic Acids Res 2017; 45(10): pp. 6023–6036. [CrossRef]
- Borchardt, E.K.; Martinez, N.M.; Gilbert, W.V. Regulation and Function of RNA Pseudouridylation in Human Cells. Annu Rev Genet. 2020; 54: pp. 309–336. [CrossRef]
- Andries, O.; Mc Cafferty, S.; De Smedt, S.C.; Weiss, R.; Sanders, N.N.; Kitada, T. N(1)-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice. J Control Release 2015; 217: pp. 337–344. [CrossRef]
- Mulroney, T.E.; Pöyry, T.; Yam-Puc, J.C.; et al. N1-methylpseudouridylation of mRNA causes +1 ribosomal frameshifting. Nature 2024; 625(7993): pp. 189–194. [CrossRef]
- Liu, A.; Wang, X. The Pivotal Role of Chemical Modifications in mRNA Therapeutics. Front. Cell Dev. Biol. 2023; 11: pp. 1–15. [CrossRef]
- Monroe, J.; Eyler, D.E.; Mitchell, L.; et al. N1-Methylpseudouridine and pseudouridine modifications modulate mRNA decoding during translation. Nat Commun 2024; 15: 8119. [CrossRef]
- Acevedo-Whitehouse, K.; Bruno, R. Potential health risks of mRNA-based vaccine therapy: A hypothesis. Med Hypotheses 2023; 171: 111015. [CrossRef]
- Lin, T.Y.; Mehta, R.; Glatt, S. Pseudouridines in RNAs: switching atoms means shifting paradigms. FEBS Lett. 2021; 595(18): pp.2310–2322. [CrossRef]
- Johnson, R. Medicine from modified mRNA. Nat Chem Biol 2023; 19: 1443. [CrossRef]
- Xu, S.; Yang, K.; Li, R.; Zhang, L. mRNA Vaccine Era-Mechanisms, Drug Platform and Clinical Prospection. Int J Mol Sci 2020; 21(18): 6582. [CrossRef]
- Vivier, E.; Malissen, B. Innate and adaptive immunity: specificities and signaling hierarchies revisited. Nat Immunol 2005; 6: pp. 17–21. [CrossRef]
- Moradian, H.; Roch, T.; Anthofer, L.; Lendlein, A.; Gossen, M. Chemical modification of uridine modulates mRNA-mediated pro-inflammatory and antiviral response in primary human macrophages. Mol Ther Nucleic Acids 2022; 27: pp. 854–869. [CrossRef]
- Drzeniek, N.M.; Kahwaji, N.; Picht, S.; Dimitriou, I.M.; Schlickeiser, S.; Moradian, H.; Geissler, S.; Schmueck-Henneresse, M.; Gossen, M.; Volk, H.D. In Vitro Transcribed mRNA Immunogenicity Induces Chemokine-Mediated Lymphocyte Recruitment and Can Be Gradually Tailored by Uridine Modification. Adv Sci (Weinh) 2024; 11(21): e2308447. [CrossRef]
- Vaidyanathan, S.; Azizian, K.T.; Haque, A.K.M.A.; Henderson, J.M.; Hendel, A.; Shore, S.; Antony, J.S.; Hogrefe, R.I.; Kormann, M.S.D.; Porteus, M.H.; McCaffrey, A.P. Uridine Depletion and Chemical Modification Increase Cas9 mRNA Activity and Reduce Immunogenicity without HPLC Purification. Mol Ther Nucleic Acids 2018; 12: pp. 530–542. [CrossRef]
- Schnoor, M.; Buers, I.; Sietmann, A.; Brodde, M.F.; Hofnagel, O.; Robenek, H.; Lorkowski, S. Efficient non-viral transfection of THP-1 cells. J Immunol Methods 2009; 344(2): pp. 109–115. [CrossRef]
- Tang, X.; Aljahdali, B.; Alasiri, M.; Bamashmous, A.; Cao, F.; Dibart, S.; Salih, E. A method for high transfection efficiency in THP-1 suspension cells without PMA treatment. Anal Biochem 2018; 544: pp. 93–97. [CrossRef]
- Uehata, T.; Takeuchi, O. RNA Recognition and Immunity-Innate Immune Sensing and Its Posttranscriptional Regulation Mechanisms. Cells 2020; 9(7): 1701. [CrossRef]
- Fleming, S.B. Viral Inhibition of the IFN-Induced JAK/STAT Signalling Pathway: Development of Live Attenuated Vaccines by Mutation of Viral-Encoded IFN-Antagonists. Vaccines (Basel) 2016; 4(3): 23. [CrossRef]
- Sioud, M.; Juzeniene, A.; Sæbøe-Larssen, S. Exploring the Impact of mRNA Modifications on Translation Efficiency and Immune Tolerance to Self-Antigens. Vaccines (Basel) 2024; 12(6): 624. [CrossRef]
| Gene of interest | FW primer sequence (5′- 3′) | RV primer sequence (5′- 3′) |
|---|---|---|
| ActB | CACCATTGGCAATGAGCGGTTC | AGGTCTTTGCGGATGTCCACGT |
| IRF7 | CCACGCTATACCATCTACCTGG | GCTGCTATCCAGGGAAGACACA |
| IRF3 | TCTGCCCTCAACCGCAAAGAAG | TACTGCCTCCACCATTGGTGTC |
| IFNa | TGGGCTGTGATCTGCCTCAAAC | CAGCCTTTTGGAACTGGTTGCC |
| TNFa | CTCTTCTGCCTGCTGCACTTTG | ATGGGCTACAGGCTTGTCACTC |
| GFP | TCAAGATCCGCCACAACATC | GTGCTCAGGTAGTGGTTGTC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




