Submitted:
09 December 2025
Posted:
11 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Study organism
RNA extraction, library preparation and RNA-sequencing
Alignment and assembly of RNA-seq data and update of the reference gene annotation
DEG analysis
Morphometry
EdU Staining
Microscopy Analysis
Quantification of EdU-positive cells
Brood size measurements
Statistical analysis
Validation of RNAseq datasets using a droplet digital PCR (ddPCR)
3. Results
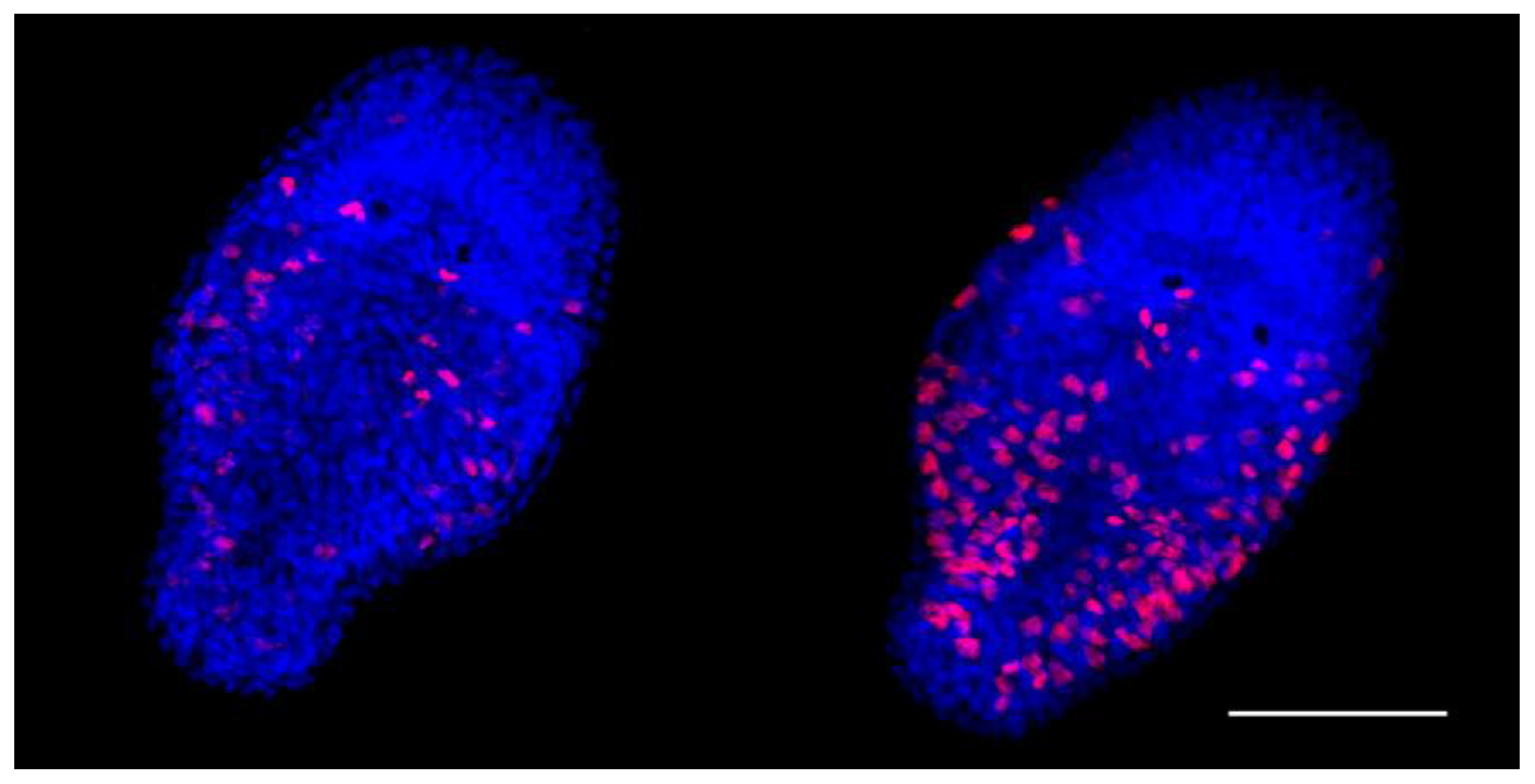
Cell proliferative activity in euploid and aneuploid worms
Brood size measurements of worms from the DV1_8 and DV1_10 sublines
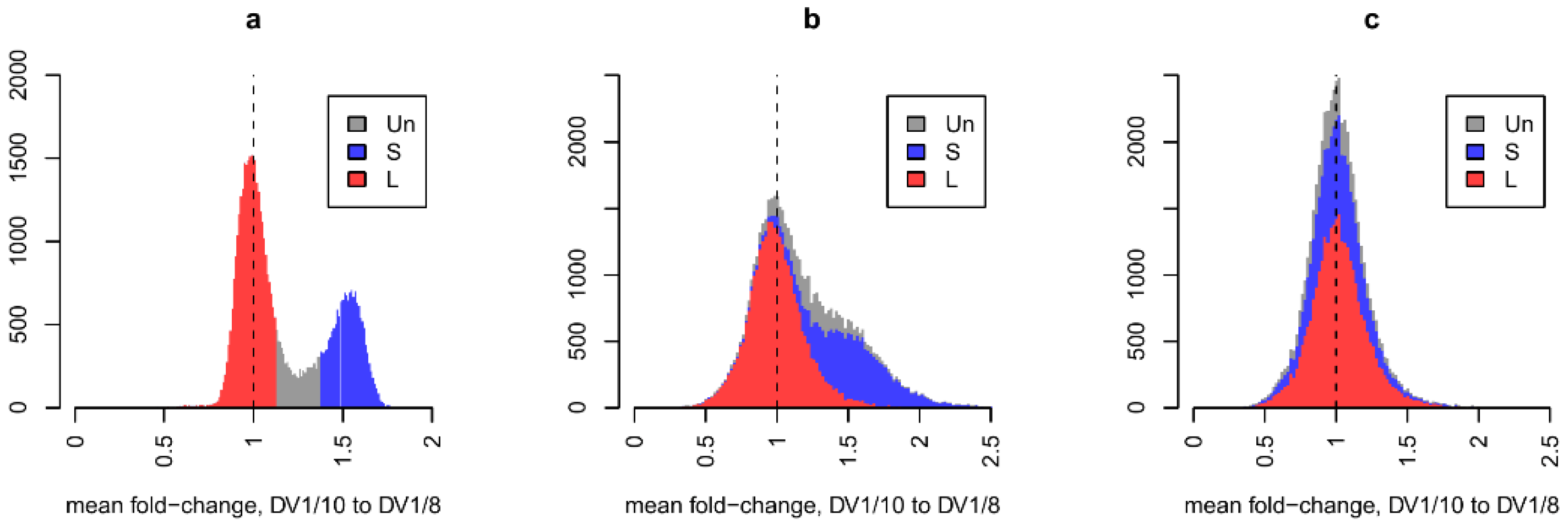
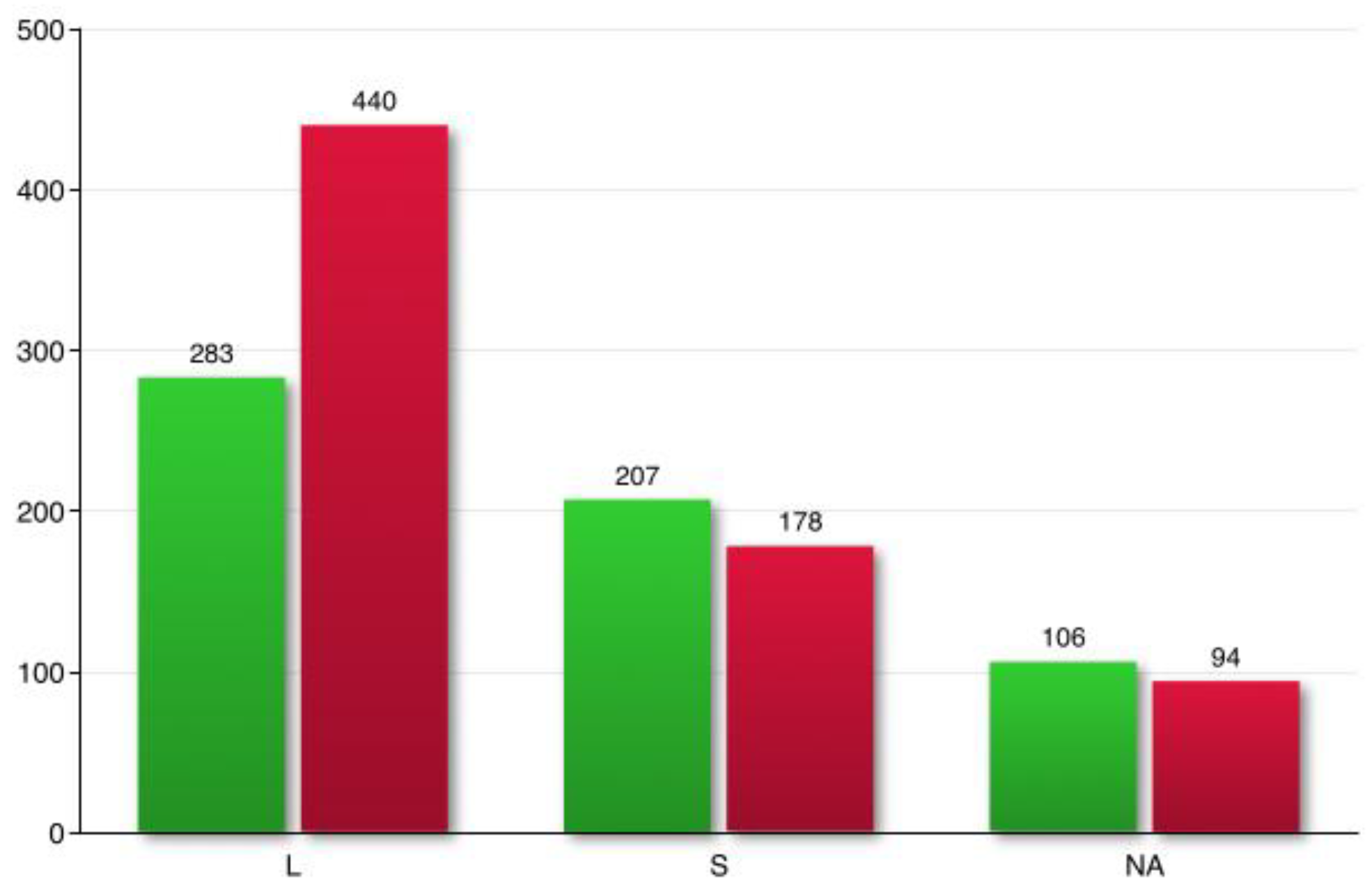
Aneuplody-driven changes in the transcriptome of M. lignano
Functional enrichment analysis for DEGs
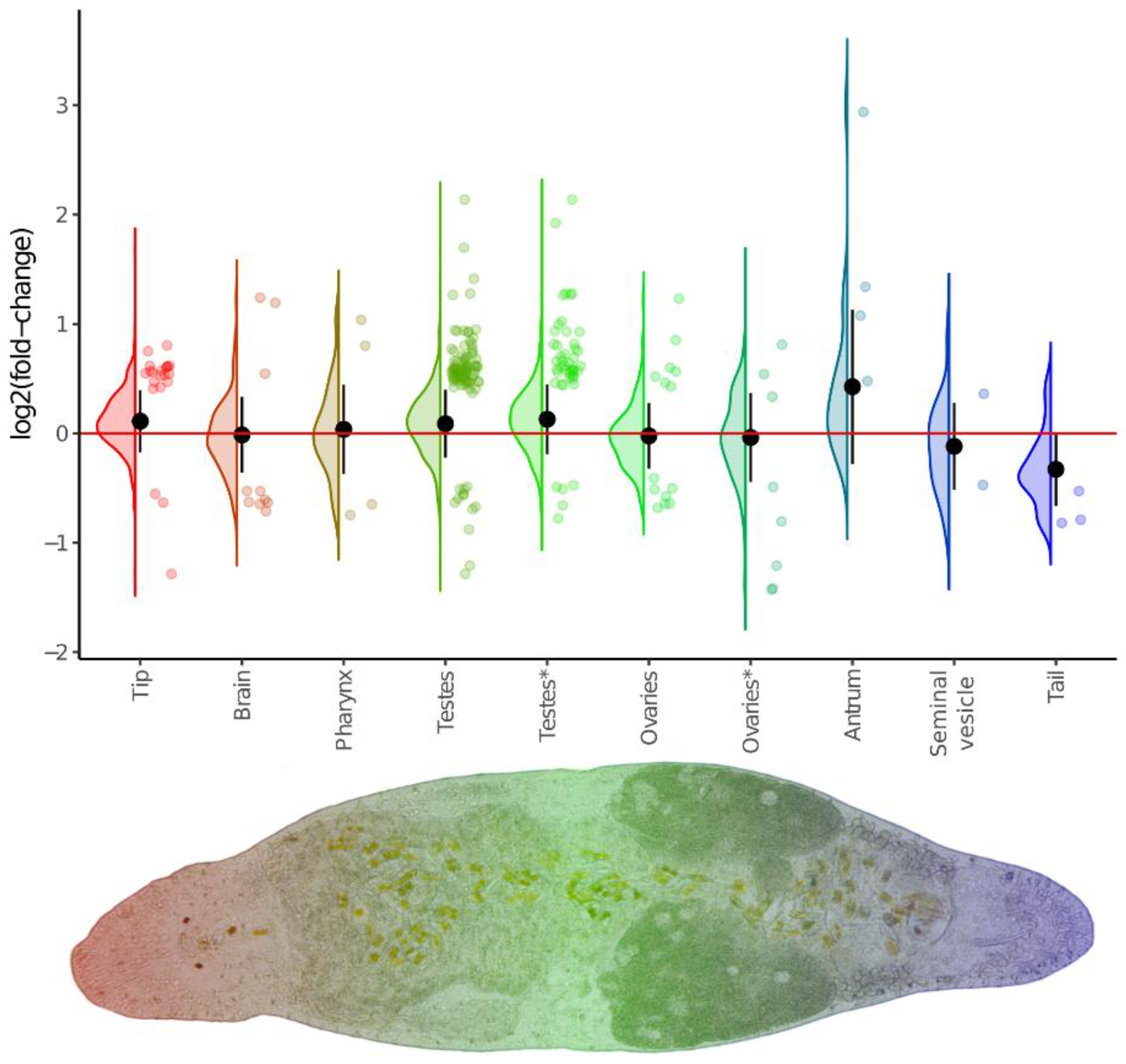
Tissue-specific DEGs linked with changes in ploidy level
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| WGD | Whole Genome Duplication |
| WGS | Whole Genome Sequencing |
| ddPCR | digital droplet PCR |
References
- Lande, R.; Shannon, S. The role of genetic variation in adaptation and population persistence in a changing environment. Evolution 1996, 50, 434–437. [Google Scholar] [CrossRef]
- Merilä, J.; Hendry, A.P. Climate change, adaptation, and phenotypic plasticity: the problem and the evidence. Evol Appl 2014, 7, 1–14. [Google Scholar] [CrossRef]
- Steward, R.A.; de Jong, M.A.; Oostra, V.; Wheat, C.W. Alternative splicing in seasonal plasticity and the potential for adaptation to environmental change. Nat Commun 2022, 13, 755. [Google Scholar] [CrossRef]
- Mantica, F.; Irimia, M. Gene duplication and alternative splicing as evolutionary drivers of proteome specialization. Bioessays 2025, 47, e202400202. [Google Scholar] [CrossRef] [PubMed]
- Fradin, H.; Kiontke, K.; Zegar, C.; Gutwein, M.; Lucas, J.; Kovtun, M.; Corcoran, D.L.; Baugh, L.R.; Fitch, D.H.A.; Piano, F.; Gunsalis, K.S. Genome architecture and evolution of a unichromosomal asexual nematode. Curr Biol 2017, 27, 2928–2939. [Google Scholar] [CrossRef] [PubMed]
- Zadesenets, K.S.; Ershov, N.I.; Bondar, N.P.; Rubtsov, N.B. Unraveling the unusual subgenomic organization in the neopolyploid free-living flatworm Macrostomum lignano. Mol Biol Evol 2023, 40, msad250. [Google Scholar] [CrossRef]
- Brand, J.N.; Patil, A.B.; Pandolfini, L.; Zadesenets, K.; Rubtsov, N.; Robledillo, L.; Zhang, M.; Marques, A.; Rink, J.C. Stepwise emergence of recombination suppression precedes fissiparous asexuality in the planarian Schmidtea mediterranea. Biorxiv 2025. [Google Scholar] [CrossRef]
- Ohno, S. Evolution by gene duplication; Springer-Verlag: New York, USA, 1970. [Google Scholar]
- Zhang, J. Evolution by gene duplication: an update. Trends Ecol Evol 2003, 18, 192–198. [Google Scholar] [CrossRef]
- Long, M.; Betran, E.; Thornton, K.; Wang, W. The origin of new genes: glimpses from the young and old. Nat Rev Genet 2003, 4, 865–875. [Google Scholar] [CrossRef]
- Van de Peer, Y.; Mizrachi, E.; Marchal, K. The evolutionary significance of polyploidy. Nat Rev Genet 2017, 18, 411–424. [Google Scholar] [CrossRef]
- Lynch, M.; Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef]
- Zadesenets, K.S.; Rubtsov, N.B. Genome duplication in animal evolution. Russ J Genet 2018, 54, 1125–1136. [Google Scholar] [CrossRef]
- Edger, P.P.; Soltis, D.E.; Yoshioka, S.; Vallejo-Marin, M.; Shimizu-Inatsugi, R.; Shimizu, K.K.; Salmon, A.; Hiscock, S.; Ainouche, M.; Soltis, P.S. Natural neopolyploids: a stimulus for novel research. New Phytol 2025, 246, 78–93. [Google Scholar] [CrossRef]
- Zhao, K.; Dong, J.; Xu, J.; Bai, Y.; Yin, Y.; Long, C.; Wu, L.; Lin, T.; Fan, L.; Wang, Y.; Edger, P.P.; Xiong, Z. Downregulation of the expression of subgenomic chromosome A7 genes promotes plant height in resynthesized allopolyploid Brassica napus. Theor Appl Genet 2023, 137, 11. [Google Scholar] [CrossRef]
- Van de Peer, Y.; Ashman, T.L.; Soltis, P.S.; Soltis, D.E. Polyploidy: an evolutionary and ecological force in stressful times. Plant Cell 2021, 33, 11–26. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.P.; Baslan, T.; Soltis, D.E.; Soltis, P.S.; Fox, D.T. Integrating the study of polyploidy across organisms, tissues, and disease. Annu Rev Genet 2024, 58, 297–318. [Google Scholar] [CrossRef] [PubMed]
- Schärer, L.; Ladurner, P. Phenotypically plastic adjustment of sex allocation in a simultaneous hermaphrodite. P Roy Soc Lond B Bio 2003, 270, 935–941. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.; Nallur, R.; Ladurner, P.; Egger, B.; Rieger, R.; Hartenstein, V. The embryonic development of the flatworm Macrostomum sp. Dev Genes Evol 2004, 214, 220–239. [Google Scholar] [CrossRef]
- Nimeth, K.T.; Mahlknecht, M.; Mezzanato, A.; Peter, R.; Rieger, R.; Ladurner, P. Stem cell dynamics during growth, feeding, and starvation in the basal flatworm Macrostomum sp. (Platyhelminthes). Dev Dyn 2004, 230, 91–99. [Google Scholar] [CrossRef]
- Ladurner, P.; Schärer, L.; Salvenmoser, W.; Rieger, R. M. A new model organism among the lower Bilateria and the use of digital microscopy in taxonomy of meiobenthic Platyhelminthes: Macrostomum lignano, n. sp. (Rhabditophora, Macrostomorpha). J Zool Syst Evol Res 2005, 43, 114–126. [Google Scholar] [CrossRef]
- Mouton, S.; Willems, M.; Braeckman, B.P.; Egger, B.; Ladurner, P.; Schärer, L.; Borgonie, G. The free-living flatworm Macrostomum lignano: a new model organism for ageing research. Exp Gerontol 2009, 44, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Lengerer, B.; Pjeta, R.; Wunderer, J.; Rodrigues, M.; Arbore, R.; Schärer, L.; Berezikov, E.; Hess, M.W.; Pfaller, K.; Egger, B.; Obwegeser, S.; Salvenmoser, W.; Ladurner, P. Biological adhesion of the flatworm Macrostomum lignano relies on a duo-gland system and is mediated by a cell type-specific intermediate filament protein. Front Zool 2014, 11, 12. [Google Scholar] [CrossRef]
- Zadesenets, K.S.; Vizoso, D.B.; Schlatter, A.; Konopatskaia, I.D.; Berezikov, E.; Schärer, L.; Rubtsov, N.B. Evidence for karyotype polymorphism in the free-living flatworm, Macrostomum lignano, a model organism for evolutionary and developmental biology. PLoS One 2016, 11, e0164915. [Google Scholar] [CrossRef] [PubMed]
- Ramm, S.A.; Lengerer, B.; Arbore, R.; Pjeta, R.; Wunderer, J.; Giannakara, A.; Berezikov, E.; Ladurner, P.; Schärer, L. Sex allocation plasticity on a transcriptome scale: socially sensitive gene expression in a simultaneous hermaphrodite. Mol Ecol 2019, 28, 2321–2341. [Google Scholar] [CrossRef]
- Wunderer, J.; Lengerer, B.; Pjeta, R.; Bertemes, P.; Kremser, L.; Lindner, H.; Ederth, T.; Hess, M.W.; Stock, D.; Salvenmoser, W.; Ladurner, P. A mechanism for temporary bioadhesion. Proc Natl Acad Sci USA 2019, 116, 4297–306. [Google Scholar] [CrossRef] [PubMed]
- Wudarski, J.; Egger, B.; Ramm, S.A.; Schärer, L.; Ladurner, P.; Zadesenets, K.S.; Rubtsov, N.B.; Mouton, S.; Berezikov, E. The free-living flatworm Macrostomum lignano. EvoDevo 2020, 11, 5. [Google Scholar] [CrossRef]
- Zadesenets, K.S.; Schärer, L.; Rubtsov, N.B. New insights into the karyotype evolution of the free-living flatworm Macrostomum lignano (Platyhelminthes, Turbellaria). Sci Rep 2017, 7, 6066. [Google Scholar] [CrossRef]
- Zadesenets, K.S.; Ershov, N.I.; Berezikov, E.; Rubtsov, N.B. Chromosome evolution in the free-living flatworms: first evidence of intrachromosomal rearrangements in karyotype evolution of Macrostomum lignano (Platyhelminthes, Macrostomida). Genes (Basel) 2017, 8, 298. [Google Scholar] [CrossRef]
- Zadesenets, K.S.; Jetybayev, I.Y.; Schärer, L.; Rubtsov, N.B. Genome and karyotype reorganization after whole genome duplication in free-living flatworms of the genus Macrostomum. Int J Mol Sci 2020, 21, 680. [Google Scholar] [CrossRef]
- Egger, B.; Ishida, S. Chromosome fission or duplication in Macrostomum lignano (Macrostomorpha, Plathelminthes) – remarks on chromosome numbers in ‘archoophoran turbellarians’. J Zool Syst Evol Res 2005, 43, 127–132. [Google Scholar] [CrossRef]
- Janicke, T.; Schärer, L. Sperm competition affects sex allocation but not sperm morphology in a flatworm. Behav Ecol Sociobiol 2010, 64, 1367–1375. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Mapleson, D.; Venturini, L.; Kaithakottil, G.; Swarbreck, D. Efficient and accurate detection of splice junctions from RNA-seq with Portcullis. Gigascience 2018, 7, giy131. [Google Scholar] [CrossRef]
- Venturini, L.; Caim, S.; Kaithakottil, G.G.; Mapleson, D.L.; Swarbreck, D. Leveraging multiple transcriptome assembly methods for improved gene structure annotation. Gigascience 2018, 7, giy093. [Google Scholar] [CrossRef]
- Pertea, G.; Pertea, M. GFF Utilities: GffRead and GffCompare. F1000Res. 2020, 9, ISCB Comm J-304. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014, 15, 550. [Google Scholar] [CrossRef]
- Schärer, L.; Ladurner, P.; Rieger, R.M. Bigger testes do work more: experimental evidence that testis size reflects testicular cell proliferation activity in the marine invertebrate, the free-living flatworm Macrostomum sp. Behav Ecol Sociobiol 2004, 56, 420–425. [Google Scholar] [CrossRef]
- Orr-Weaver, T.L. When bigger is better: the role of polyploidy in organogenesis. Trends Genet 2015, 31, 307–315. [Google Scholar] [CrossRef]
- Vizoso, D.B.; Rieger, G.; Schärer, L. Goings-on inside a worm: functional hypotheses derived from sexual conflict thinking. Biol J Linn Soc 2010, 99, 370–383. [Google Scholar] [CrossRef]
- Schärer, L.; Joss, G.; Sandner, P. Mating behaviour of the marine turbellarian Macrostomum sp.: These worms suck. Mar Biol 2004, 145, 373–380. [Google Scholar] [CrossRef]
- Santhosh, S.; Ebert, D.; Janicke, T. Sperm competition favours intermediate sperm size in a hermaphrodite. J Evol Biol 2024, 37, 829–838. [Google Scholar] [CrossRef]
- Egger, B.; Ladurner, P.; Nimeth, K.; Gschwentner, R.; Rieger, R. The regeneration capacity of the flatworm Macrostomum lignano - on repeated regeneration, rejuvenation, and the minimal size needed for regeneration. Dev Genes Evol 2006, 216, 565–577. [Google Scholar] [CrossRef]
- Pfister, D.; De Mulder, K.; Philipp, I.; Kuales, G.; Hrouda, M.; Eichberger, P.; Borgonie, G.; Hartenstein, V.; Ladurner, P. The exceptional stem cell system of Macrostomum lignano: screening for gene expression and studying cell proliferation by hydroxyurea treatment and irradiation. Front Zool 2007, 4, 9. [Google Scholar] [CrossRef] [PubMed]
- Sekii, K.; Salvenmoser, W.; De Mulder, K.; Schärer, L.; Ladurner, P. Melav2, an elav-like gene, is essential for spermatid differentiation in the flatworm Macrostomum lignano. BMC Dev Biol 2009, 9, 62. [Google Scholar] [CrossRef] [PubMed]
- Grudniewska, M.; Mouton, S.; Grelling, M.; Wolters, A.H.G.; Kuipers, J.; Giepmans, B.N.G.; Berezikov, E. A novel flatworm-specific gene implicated in reproduction in Macrostomum lignano. Sci Rep 2018, 8, 3192. [Google Scholar] [CrossRef] [PubMed]
- Arbore, R.; Sekii, K.; Beisel, C.; Ladurner, P.; Berezikov, E.; Schärer, L. Positional RNA-Seq identifies candidate genes for phenotypic engineering of sexual traits. Front Zool 2015, 12, 14. [Google Scholar] [CrossRef]
- Sheltzer, J.M.; Torres, E.M.; Dunham, M.J.; Amon, A. Transcriptional consequences of aneuploidy. Proc Natl Acad Sci U S A 2012, 109, 12644–12649. [Google Scholar] [CrossRef]
- Zadesenets, K.S.; Rubtsov, N.B. From cytogenetics to proteogenomics: new horizons in the study of aneuploidies. Vavilovskii Zhurnal Genet Selektsii 2025, 29, 335–348. [Google Scholar] [CrossRef]
- Sullivan, K.D.; Lewis, H.C.; Hill, A.A.; Pandey, A.; Jackson, L.P.; Cabral, J.M.; Smith, K.P.; Liggett, L.A.; Gomez, E.B.; Galbraith, M.D.; DeGregori, J.; Espinosa, J.M. Trisomy 21 consistently activates the interferon response. Elife 2016, 5, e16220. [Google Scholar] [CrossRef]
- Raznahan, A.; Parikshak, N.N.; Chandran, V.; Blumenthal, J.D.; Clasen, L.S.; Alexander-Bloch, A.F.; Zinn, A.R.; Wangsa, D.; Wise, J.; Murphy, D.G.M.; Bolton, P.F.; Ried, T.; Ross, J.; Giedd, J.N.; Geschwind, D.H. Sex-chromosome dosage effects on gene expression in humans. Proc. Natl. Acad. Sci. U S A 2018, 115, 7398–7403. [Google Scholar] [CrossRef]
- Li, R.; Zhu, J. Effects of aneuploidy on cell behaviour and function. Nat Rev Mol Cell Biol 2022, 23, 250–265. [Google Scholar] [CrossRef]
- Pavelka, N.; Rancati, G.; Zhu, J.; Bradford, W.D.; Saraf, A.; Florens, L.; Sanderson, B.W.; Hattem, G.L.; Li, R. Aneuploidy confers quantitative proteome changes and phenotypic variation in budding yeast. Nature 2010, 468, 321–325. [Google Scholar] [CrossRef]
- Veitia, R.A.; Bottani, S.; Birchler, J.A. Gene dosage effects: nonlinearities, genetic interactions, and dosage compensation. Trends Genet 2013, 29, 385–393. [Google Scholar] [CrossRef]
- Pires, J.C.; Conant, G.C. Robust yet fragile: expression noise, protein misfolding, and gene dosage in the evolution of genomes. Annu Rev Genet 2016, 50, 113–131. [Google Scholar] [PubMed]
- Birchler, J.A.; Veitia, R.A. Gene balance hypothesis: connecting issues of dosage sensitivity across biological disciplines. Proc Natl Acad Sci U S A 2012, 109, 14746–14753. [Google Scholar] [CrossRef]
- Veitia, R.A.; Potier, M.C. Gene dosage imbalances: action, reaction, and models. Trends Biochem Sci 2015, 40, 309–317. [Google Scholar] [CrossRef]
- Lindemans, M.; Liu, F.; Janssen, T.; Husson, S.J.; Mertens, I.; Gäde, G.; Schoofs, L. Adipokinetic hormone signaling through the gonadotropin-releasing hormone receptor modulates egg-laying in Caenorhabditis elegans. Proc Natl Acad Sci U S A 2009, 106, 1642–1647. [Google Scholar] [CrossRef]
- Kobayashi, A.; Hamada, M.; Yoshida, M.A.; Kobayashi, Y.; Tsutsui, N.; Sekiguchi, T.; Matsukawa, Y.; Maejima, S.; Gingell, J.J.; Sekiguchi, S.; Hamamoto, A.; Hay, D.L.; Morris, J.F.; Sakamoto, T.; Sakamoto, H. Vasopressin-oxytocin-type signaling is ancient and has a conserved water homeostasis role in euryhaline marine planarians. Sci Adv 2022, 8, eabk0331. [Google Scholar] [CrossRef] [PubMed]
- Mäkelä, J.A.; Saario, V.; Bourguiba-Hachemi, S.; Nurmio, M.; Jahnukainen, K.; Parvinen, M.; Toppari, J. Hedgehog signalling promotes germ cell survival in the rat testis. Reproduction 2011, 142, 711–721. [Google Scholar] [CrossRef]
- Kibet, M.K.; Hilchenbach, J.; Neumann, L.; Mayer, R.; Aigner, G.P.; Höckner, M.; Hobmayer, B.; Egger, B. Silencing of β-catenin1 blocks tail regeneration, but does not induce head regeneration in the flatworm Macrostomum lignano. Discov Dev Biol 2025, 235, 1. [Google Scholar] [CrossRef]
- Demircan, T.; Berezikov, E. The Hippo pathway regulates stem cells during homeostasis and regeneration of the flatworm Macrostomum lignano. Stem Cells Dev 2013, 22, 2174–2185. [Google Scholar] [CrossRef]
- Choudhury, D.; Ghosh, D.; Mondal, M.; Singha, D.; Pothuraju, R.; Malakar, P. Polyploidy and mTOR signaling: a possible molecular link. Cell Commun Signal 2024, 22, 196. [Google Scholar] [CrossRef] [PubMed]
- Muenzner, J.; Trébulle, P.; Agostini, F.; Zauber, H.; Messner, C.B.; Steger, M.; Kilian, C.; Lau, K.; Barthel, N.; Lehmann, A.; Textoris-Taube, K.; Caudal, E.; Egger, A.S.; Amari, F.; De Chiara, M.; Demichev, V.; Gossmann, T.I.; Mülleder, M.; Liti, G.; Schacherer, J.; Selbach, M.; Berman, J.; Ralser, M. Natural proteome diversity links aneuploidy tolerance to protein turnover. Nature 2024, 630, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Storchová, Z. Cells cope with altered chromosome numbers by enhancing protein breakdown. Nature 2024, 630, 41–43. [Google Scholar] [CrossRef] [PubMed]
- Shyh-Chang, N.; Ng, H. H. The metabolic programming of stem cells. Genes Dev 2017, 31, 336–346. [Google Scholar] [CrossRef]

| Subline | Body | Eye | Testis | Ovary | Stylet | Segmented L | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| L | W | A | d | L | A | L | A | Distal opening | Proximal opening | Straight L | ||
| DV1_8 | 1255,16 ±197,92 |
166,89 ±15,87 |
508642,71 ±37772,35 |
8.12 ±0.56 |
109.73 ±16.5 |
4147.32 ±1112.47 |
54.74 ±13.24 |
1510.19 ±485.92 |
4.82 ±0,66 |
14.58 ±2,41 |
69.88 ±3.91 |
70.39 ±4.28 |
| DV1_10 | 1480,83 ±156,27* |
183,44 ±26,59* |
520780,83 ±36558,49 |
8.39 ±0.62 |
144.89 ±16.61* |
7272.15 ±1331.13* |
72.76 ±23.89* |
2933.12 ±1741.63* |
4.76 ±0.6 |
15.09 ±2.46 |
75.68 ±4.69* |
74.39 ±12.21 |
| Subline | Sperm | ||||
|---|---|---|---|---|---|
| total L | body, L | feeler, L | brush, L | bristle, L | |
| DV1_8 | 53,85 ±6.06 |
11,86 ±1,50 |
17,01 ±1,63 |
5.15 ±0,86 |
11,79 ±0,86 |
| DV1_10 | 65,40 ±3,06* |
13,21 ±1,06* |
20,31 ±1,48* |
6,01 ±0,51* |
12,74 ±0,76* |
| Subline | Hatchlings (h) | Adult worms (a) | ||||
|---|---|---|---|---|---|---|
| N | p-value | N | p-value | N | p-value | |
| testes | ovaries | |||||
| DV1_8 (2n=8) |
71.03 ±19.79 |
52.47 ±16.48 |
5.22 ±3.8 |
|||
| DV1_10 (2n=10) |
102.33 ±22.82* |
81.97 ±31.24* |
8.11 ±3.5* |
|||
| <0.00001 | 0.000015 | 4*10^-5 | ||||
| Karyotype | N | Laid eggs (N) | Hatchlings (N) | Juveniles (N) | Abnormal/Dead worms (N) | Adults (N) |
|---|---|---|---|---|---|---|
| 2n=8 | 80 | 304 (209)* | 129 | 103 | 59 | 44 (34.11%)** |
| 2n=10 | 78 | 249 (161)* | 105 | 99 | 2 | 97 (92.38%)** |
| Transcript ID | Log2FC | Adjusted p-value | Subgenome |
|---|---|---|---|
| mikado.scaf1795G14 | 5.67 | 0.004391 | L |
| mikado.scaf3150G9 | 5.49 | 0.003809 | L |
| mikado.scaf656G9 | 4.26 | 1.92E-70 | S |
| mikado.scaf1804G1 | 3.59 | 9.03E-09 | L |
| mikado.scaf1592G1 | 3.16 | 0.022669 | L |
| mikado.scaf1151G13 | 3.16 | 0.002523 | L |
| mikado.scaf3332G6 | 2.99 | 1.8E-05 | S |
| mikado.scaf3157G7 | 2.89 | 0.001061 | NA |
| mikado.scaf3771G4 | 2.88 | 0.024537 | NA |
| mikado.scaf794G14 | 2.83 | 0.0418 | L |
| mikado.scaf514G9 | -2.29 | 0.00114 | L |
| mikado.scaf1021G14 | -2.32 | 0.007259 | NA |
| mikado.scaf4546G2 | -2.51 | 0.00025 | L |
| mikado.scaf572G34 | -2.55 | 0.009102 | L |
| mikado.scaf528G22 | -2.63 | 0.011372 | L |
| mikado.scaf77G16 | -3.16 | 1.31E-05 | S |
| mikado.scaf756G13 | -3.31 | 0.002773 | L |
| mikado.scaf1160G12 | -3.33 | 5.58E-05 | L |
| mikado.scaf1116G34 | -3.67 | 0.00151 | L |
| mikado.scaf249G20 | -4.93 | 0.01119 | S |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




