Submitted:
04 December 2025
Posted:
05 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. NGS sequencing and initial data processing
2.2. Phylogenetics
2.3. Statistics
2.4. Population Genetics
2.5. Phenotypical Characteristics
2.6. Biogeography
3. Results
3.1. RAD Sequencing
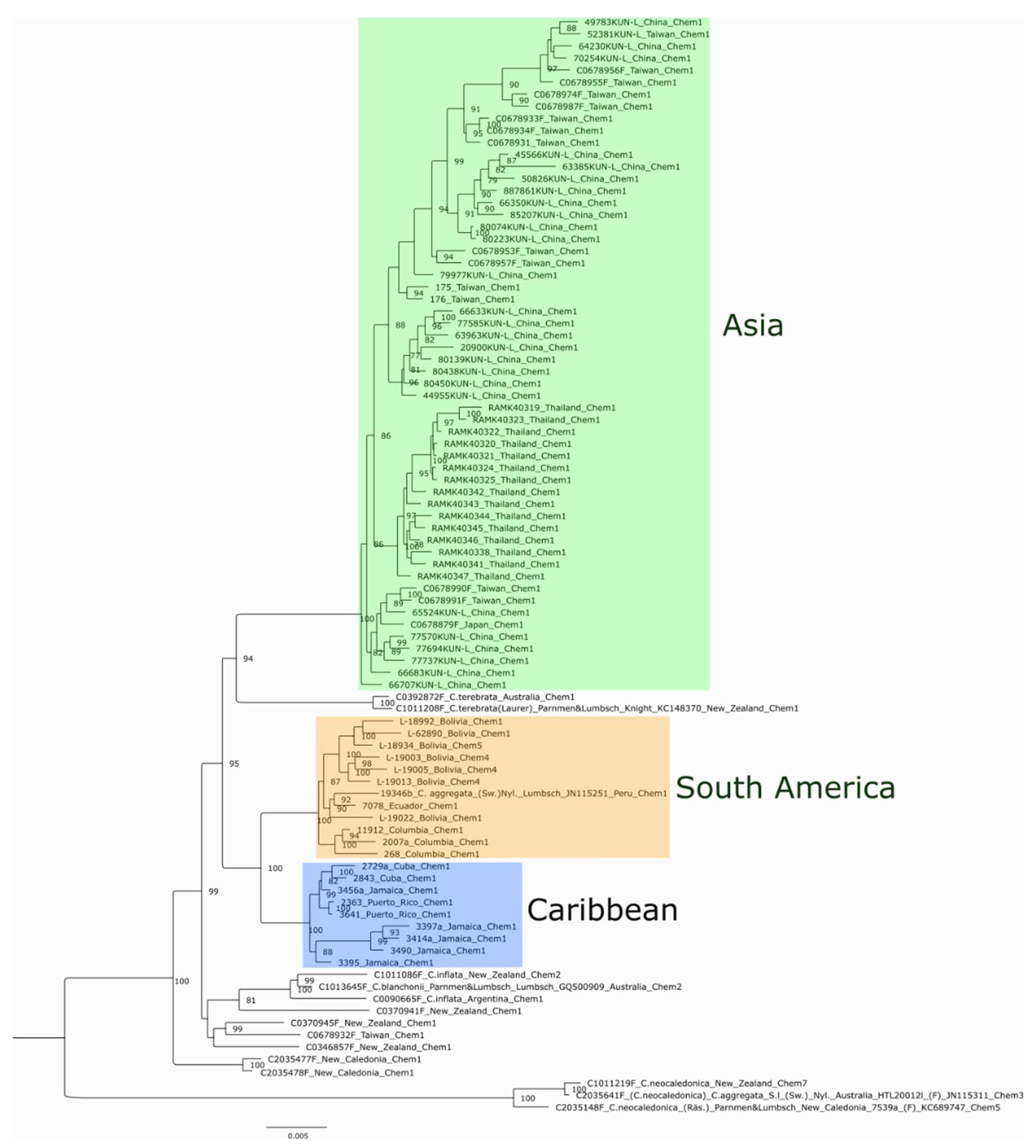
3.2. Phylogenetic Analyses
3.3. Statistics
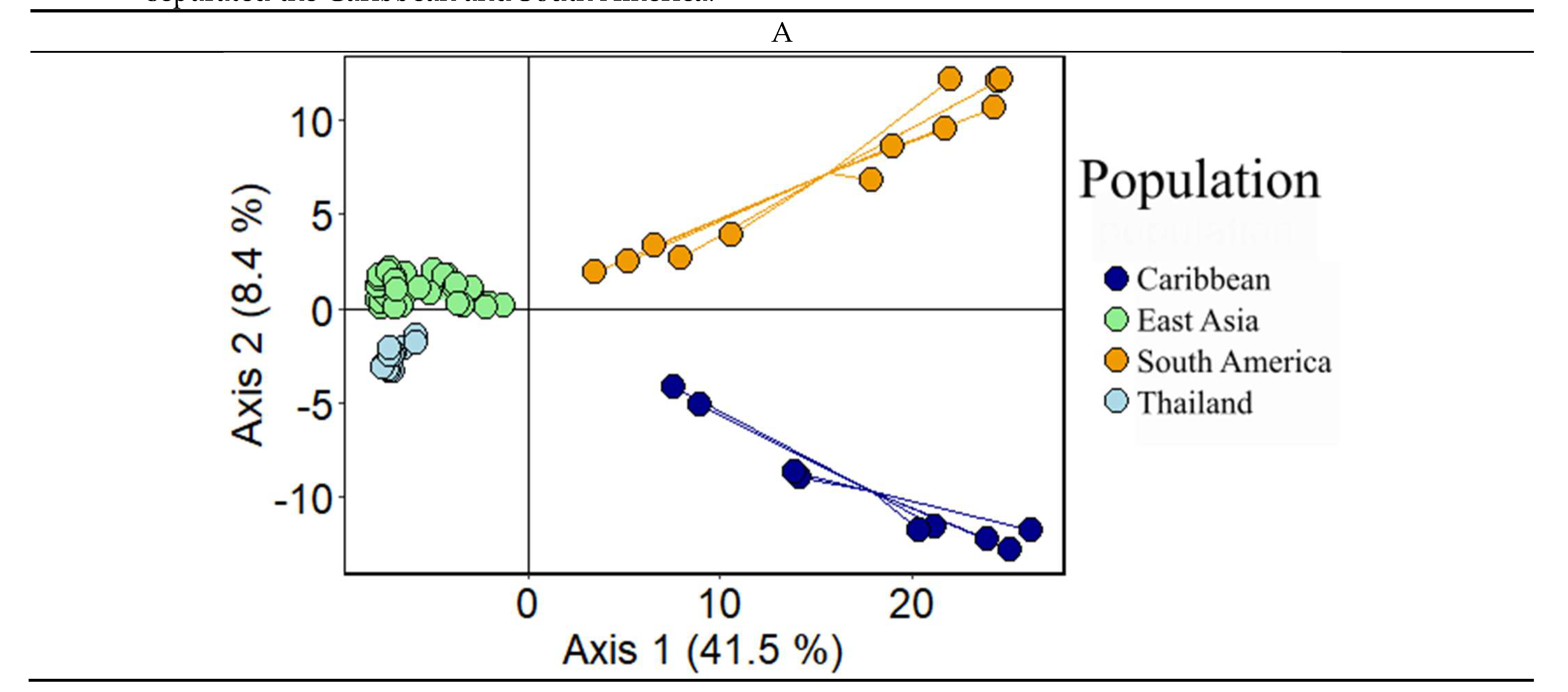
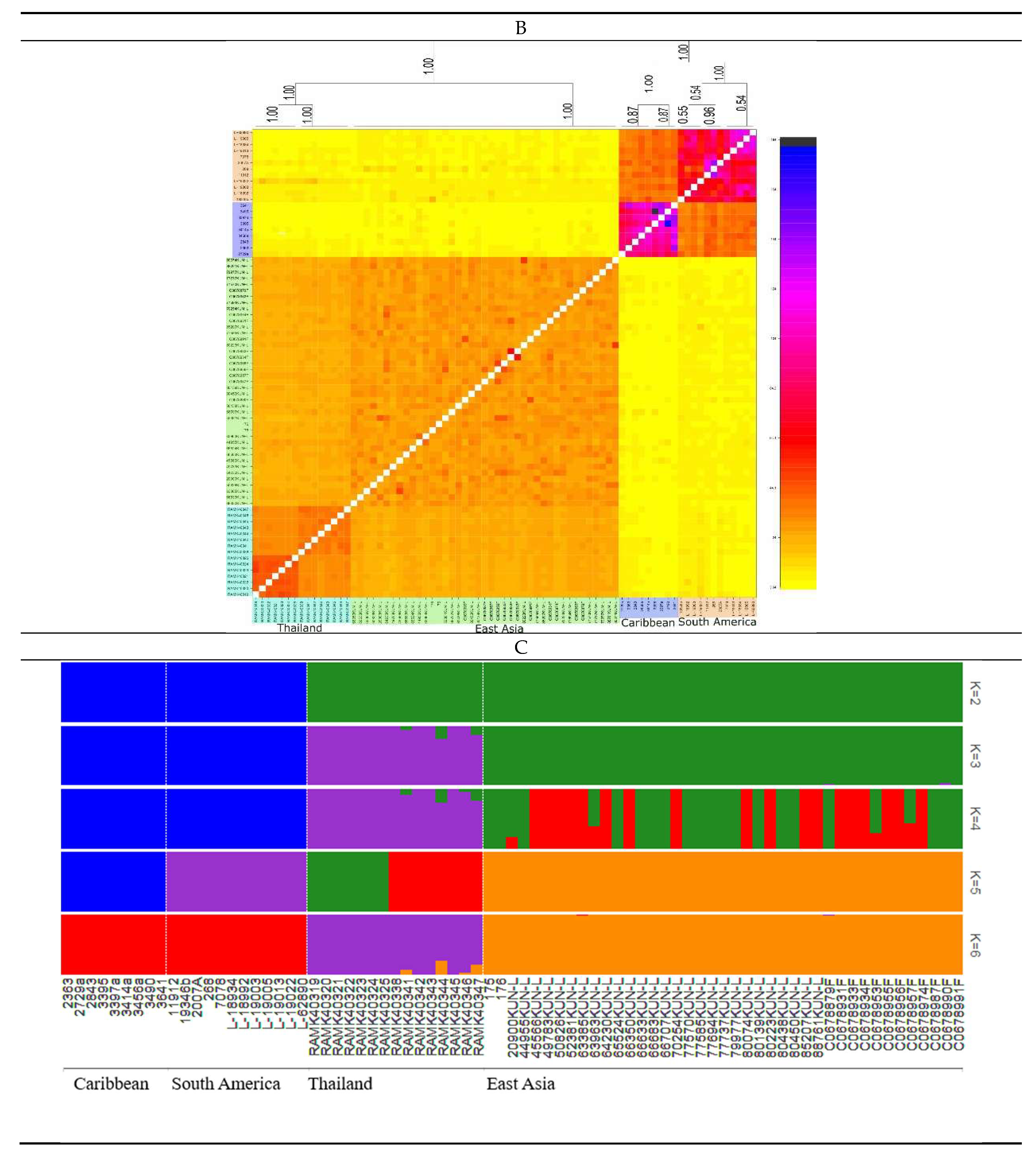
3.4. Population Genetics

3.5. Phenotypic Characteristics
3.6. Biogeography
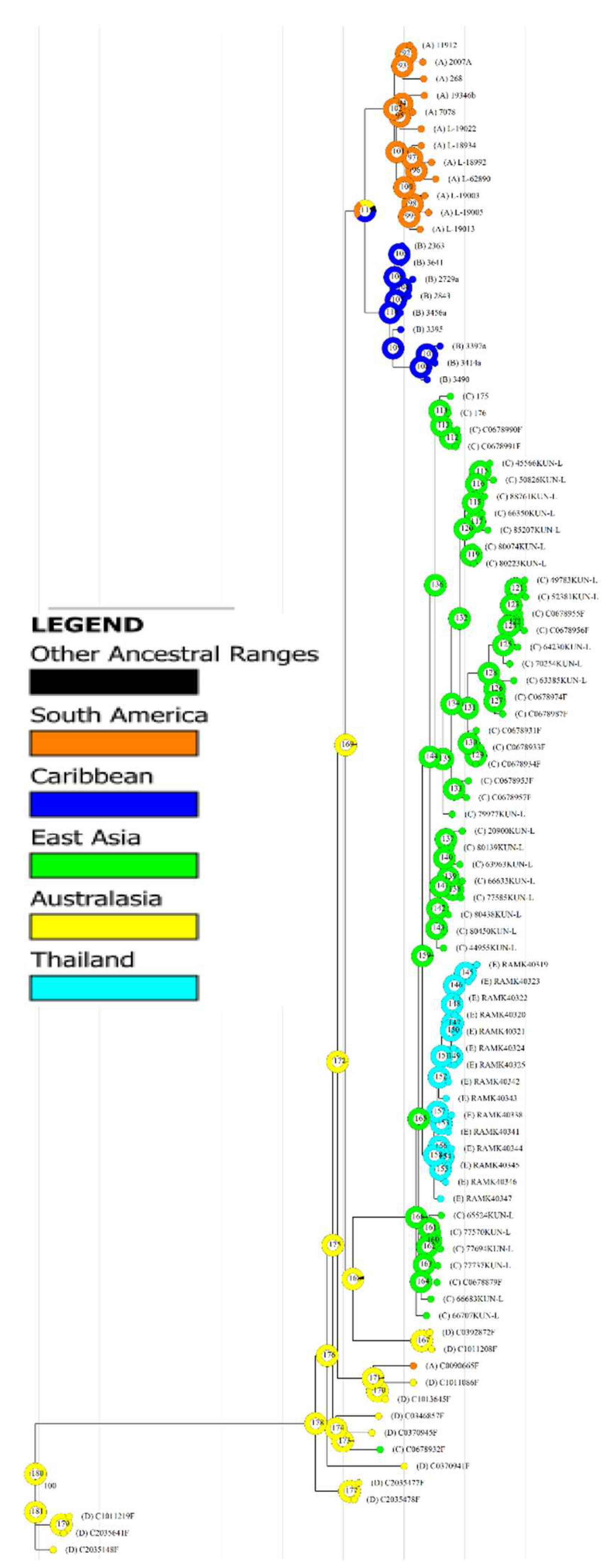

3.7. Taxonomy
4. Discussion
4.1. Species Delimitation
4.2. Biogeography
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rannala, B.; Yang, Z. Species Delimitation. In Phylogenetics in the Genomic Era; No commercial publisher, 2020; p. pp.5.5:1--5.5:18.
- Mora, C.; Tittensor, D.P.; Adl, S.; Simpson, A.G.B.; Worm, B. How Many Species Are There on Earth and in the Ocean? PLOS Biol. 2011, 9, e1001127. [Google Scholar] [CrossRef] [PubMed]
- Honegger, R.; Edwards, D.; Axe, L. The Earliest Records of Internally Stratified Cyanobacterial and Algal Lichens from the Lower Devonian of the Welsh Borderland. New Phytol. 2013, 197, 264–275. [Google Scholar] [CrossRef]
- Lumbsch, H.T.; Leavitt, S.D. Goodbye Morphology? A Paradigm Shift in the Delimitation of Species in Lichenized Fungi. Fungal Divers. 2011, 50, 59–72. [Google Scholar] [CrossRef]
- Baird, N.A.; Etter, P.D.; Atwood, T.S.; Currey, M.C.; Shiver, A.L.; Lewis, Z.A.; Selker, E.U.; Cresko, W.A.; Johnson, E.A. Rapid SNP Discovery and Genetic Mapping Using Sequenced RAD Markers. PLOS ONE 2008, 3, e3376. [Google Scholar] [CrossRef]
- Barcenas-Peña, A.; Divakar, P.K.; Crespo, A.; Nuñez-Zapata, J.; Lumbsch, H.T.; Grewe, F. Reference-Based Restriction-Site-Associated DNA Sequencing Data Are Useful for Species Delineation in a Recently Diverged Asexually Reproducing Species Complex (Parmeliaceae, Ascomycota). J. Fungi 2023, 9, 1180. [Google Scholar] [CrossRef]
- Grewe, F.; Huang, J.-P.; Leavitt, S.D.; Lumbsch, H.T. Reference-Based RADseq Resolves Robust Relationships among Closely Related Species of Lichen-Forming Fungi Using Metagenomic DNA. Sci. Rep. 2017, 7, 9884. [Google Scholar] [CrossRef] [PubMed]
- Grewe, F.; Lagostina, E.; Wu, H.; Printzen, C.; Lumbsch, H.T. Population Genomic Analyses of RAD Sequences Resolves the Phylogenetic Relationship of the Lichen-Forming Fungal Species Usnea Antarctica and Usnea Aurantiacoatra. MycoKeys 2018, 43, 91–113. [Google Scholar] [CrossRef]
- Jorna, J.; Linde, J.B.; Searle, P.C.; Jackson, A.C.; Nielsen, M.-E.; Nate, M.S.; Saxton, N.A.; Grewe, F.; Herrera-Campos, M. de L.A.; Spjut, R.W.; et al. Species Boundaries in the Messy Middle-A Genome-Scale Validation of Species Delimitation in a Recently Diverged Lineage of Coastal Fog Desert Lichen Fungi. Ecol. Evol. 2021, 11, 18615–18632. [Google Scholar] [CrossRef]
- Otero, A.; Barcenas-Peña, A.; Lumbsch, H.T.; Grewe, F. Reference-Based RADseq Unravels the Evolutionary History of Polar Species in ‘the Crux Lichenologorum’ Genus Usnea (Parmeliaceae, Ascomycota). J. Fungi 2023, 9, 99. [Google Scholar] [CrossRef]
- Widhelm, T.J.; Grewe, F.; Huang, J.-P.; Ramanauskas, K.; Mason-Gamer, R.; Lumbsch, H.T. Using RADseq to Understand the Circum-Antarctic Distribution of a Lichenized Fungus, Pseudocyphellaria Glabra. J. Biogeogr. 2021, 48, 78–90. [Google Scholar] [CrossRef]
- Widhelm, T.J.; Rao, A.; Grewe, F.; Lumbsch, H.T. High-Throughput Sequencing Confirms the Boundary between Traditionally Considered Species Pairs in a Group of Lichenized Fungi (Peltigeraceae, Pseudocyphellaria). Bot. J. Linn. Soc. 2023, 201, 471–482. [Google Scholar] [CrossRef]
- Stenroos, S.; Pino-Bodas, R.; Hyvönen, J.; Lumbsch, H.T.; Ahti, T. Phylogeny of the Family Cladoniaceae (Lecanoromycetes, Ascomycota) Based on Sequences of Multiple Loci. Cladistics 2019, 35, 351–384. [Google Scholar] [CrossRef]
- Filson, R.B. A REVISION OF THE LICHEN GENUS CLAD/A NYL. Hattori Bot. Lab. 1981, 49, 1–75. [Google Scholar]
- Kantvilas, G.; Kantvilas, G.; Elix, J.A. Studies on the Lichen Genus Cladia Nyl. in Tasmania: The C. Aggregata Complex. Muelleria Aust. J. Bot. 1999, 12, 135–162. [Google Scholar] [CrossRef]
- Kantvilas, G.; Elix, J. A New Species of Cladia (Lichenized Ascomycotina) from Tasmania. Mycotaxon 1987, 29, 199–205. [Google Scholar] [CrossRef]
- Swartz, O.; Swartz, O.; Nordstrom, A.J.; Swederus, M. Nova Genera & Species Plantarum, Seu, Prodromus Descriptionum Vegetabilium :Maximam Partem Incognitorum Quæ Sub Itinere in Indiam Occidentalem Annis 1783-87; in bibliopoliis Acad. M. Swederi: Holmiae [Stockholm], Upsaliæ & Aboæ, 1788. [Google Scholar]
- Ahti, T. Cladoniaceae; Organization for Flora Neotropica: New York Botanical Garden, 2000; Vol. 78; ISBN 978-0-89327-431-3.
- Parnmen, S.; Rangsiruji, A.; Mongkolsuk, P.; Boonpragob, K.; Nutakki, A.; Lumbsch, H.T. Using Phylogenetic and Coalescent Methods to Understand the Species Diversity in the Cladia Aggregata Complex (Ascomycota, Lecanorales). PLOS ONE 2012, 7, e52245. [Google Scholar] [CrossRef]
- Parnmen, S.; Leavitt, S.; Rangsiruji, A.; Lumbsch, T. Identification of Species in the Cladia Aggregata Group Using DNA Barcoding (Ascomycota: Lecanorales). Phytotaxa 2013, 115, 1. [Google Scholar] [CrossRef]
- Otero, A.; Lumbsch, H.T.; Degtjarenko, P.; Grewe, F. Speciation Continuum in Non-Model Organisms: Revisiting the Species-Pair Concept in Lichens. Mol. Phylogenet. Evol. 2025, 108407. [Google Scholar] [CrossRef]
- Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. A Robust, Simple Genotyping-by-Sequencing (GBS) Approach for High Diversity Species. PLOS ONE 2011, 6, e19379. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML Version 8: A Tool for Phylogenetic Analysis and Post-Analysis of Large Phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Eaton, D.; Overcast, I. Output Formats — Ipyrad Documentation. Available online: https://ipyrad.readthedocs.io/en/latest/output_formats.html (accessed on 29 October 2024).
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef]
- FigTree. Available online: https://tree.bio.ed.ac.uk/software/figtree/ (accessed on 16 May 2025).
- Chifman, J.; Kubatko, L. Quartet Inference from SNP Data Under the Coalescent Model. Bioinformatics 2014, 30, 3317–3324. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D.L. PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods) 2003.
- Zhang, J.; Kapli, P.; Pavlidis, P.; Stamatakis, A. A General Species Delimitation Method with Applications to Phylogenetic Placements. Bioinformatics 2013, 29, 2869–2876. [Google Scholar] [CrossRef] [PubMed]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R Package for Genetic Analysis of Populations with Clonal, Partially Clonal, and/or Sexual Reproduction. PeerJ 2014, 2, e281. [Google Scholar] [CrossRef]
- Dray, S.; Dufour, A.-B. The Ade4 Package: Implementing the Duality Diagram for Ecologists. J. Stat. Softw. 2007, 22, 1–20. [Google Scholar] [CrossRef]
- Jombart, T.; Ahmed, I. Adegenet 1.3-1: New Tools for the Analysis of Genome-Wide SNP Data. Bioinformatics 2011, 27, 3070–3071. [Google Scholar] [CrossRef]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The Variant Call Format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer-Verlag New York, 2016; ISBN 978-3-319-24277-4.
- Malinsky, M.; Trucchi, E.; Lawson, D.J.; Falush, D. RADpainter and fineRADstructure: Population Inference from RADseq Data. Mol. Biol. Evol. 2018, 35, 1284–1290. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. Ape 5.0: An Environment for Modern Phylogenetics and Evolutionary Analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef]
- Lang, D.T.; Kalibera, T. XML: Tools for Parsing and Generating XML Within R and S-Plus 2025.
- Raj, A.; Stephens, M.; Pritchard, J.K. fastSTRUCTURE: Variational Inference of Population Structure in Large SNP Data Sets. Genetics 2014, 197, 573–589. [Google Scholar] [CrossRef]
- Purcell, S.; Chang, C. PLINK 1. 9 2024.
- Chang, C.C.; Chow, C.C.; Tellier, L.C.; Vattikuti, S.; Purcell, S.M.; Lee, J.J. Second-Generation PLINK: Rising to the Challenge of Larger and Richer Datasets. GigaScience 2015, 4, 7. [Google Scholar] [CrossRef]
- Francis, R.M. Pophelper: An R Package and Web App to Analyse and Visualize Population Structure. Mol. Ecol. Resour. 2017, 17, 27–32. [Google Scholar] [CrossRef]
- Yu, Y.; Blair, C.; He, X. RASP 4: Ancestral State Reconstruction Tool for Multiple Genes and Characters. Mol. Biol. Evol. 2020, 37, 604–606. [Google Scholar] [CrossRef]
- Matzke, N.J. Probabilistic Historical Biogeography: New Models for Founder-Event Speciation, Imperfect Detection, and Fossils Allow Improved Accuracy and Model-Testing. Front. Biogeogr. 2013, 5. [Google Scholar] [CrossRef]
- Ali, S.S.; Yu, Y.; Pfosser, M.; Wetschnig, W. Inferences of Biogeographical Histories within Subfamily Hyacinthoideae Using S-DIVA and Bayesian Binary MCMC Analysis Implemented in RASP (Reconstruct Ancestral State in Phylogenies). Ann. Bot. 2012, 109, 95–107. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian Phylogenetic Inference under Mixed Models. Bioinforma. Oxf. Engl. 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Buschbom, J. Migration between Continents: Geographical Structure and Long-Distance Gene Flow in Porpidia Flavicunda (Lichen-Forming Ascomycota). Mol. Ecol. 2007, 16, 1835–1846. [Google Scholar] [CrossRef] [PubMed]
| Number of Filtered Loci | 5,720 |
| Number of SNPs | 46,731 |
| Percentage of Missing Data | snps matrix size: 81.00% missing sites. sequence matrix size: 87.95% missing sites. |
| Average Number of Loci Per Sample | 1,480.48 |
| Average Number of Raw Reads | 7,068,030.81 |
| Average Number of Mapped Reads | 187,914.98 |
| Average Error Estimate | 0.0017 |
| Variation Between Groups | 544.9762 | 79.19% |
| Variation Within Groups | 143.2486 | 20.81% |
| Total Variation | 688.2249 | 100% |
| Phi-samples-total | 0.7918578 | |
| Monte-Carlo Test P Value | 0.001 | |
| Average Variance of Simulated Groups | 170.36 | |
| Observed Variance Between Actual Groups | 544.98 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




