Submitted:
02 December 2025
Posted:
05 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
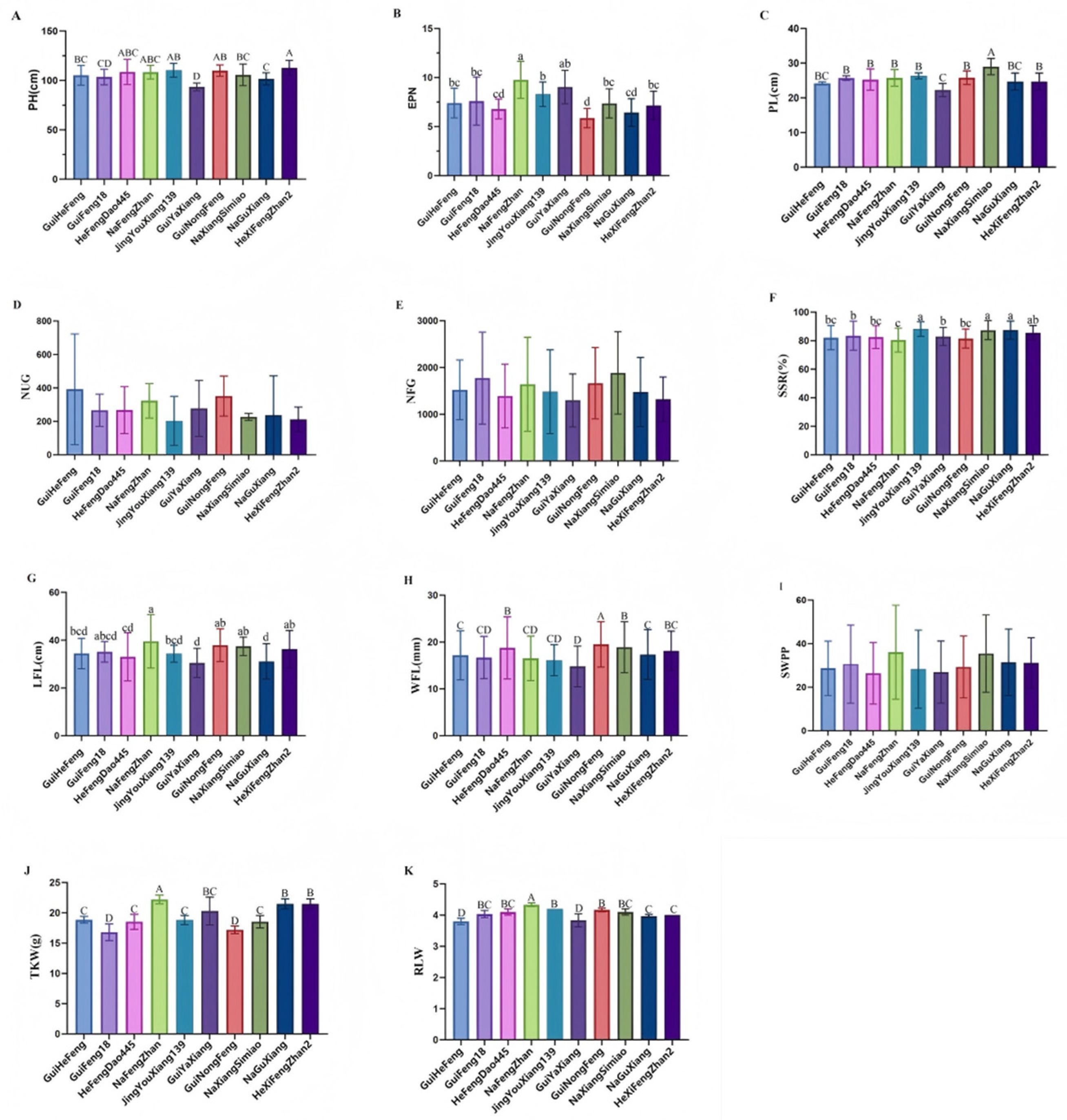
2.1. The Derivatives Exhibited Comparable Agronomic Trait Performance to GuiHeFeng
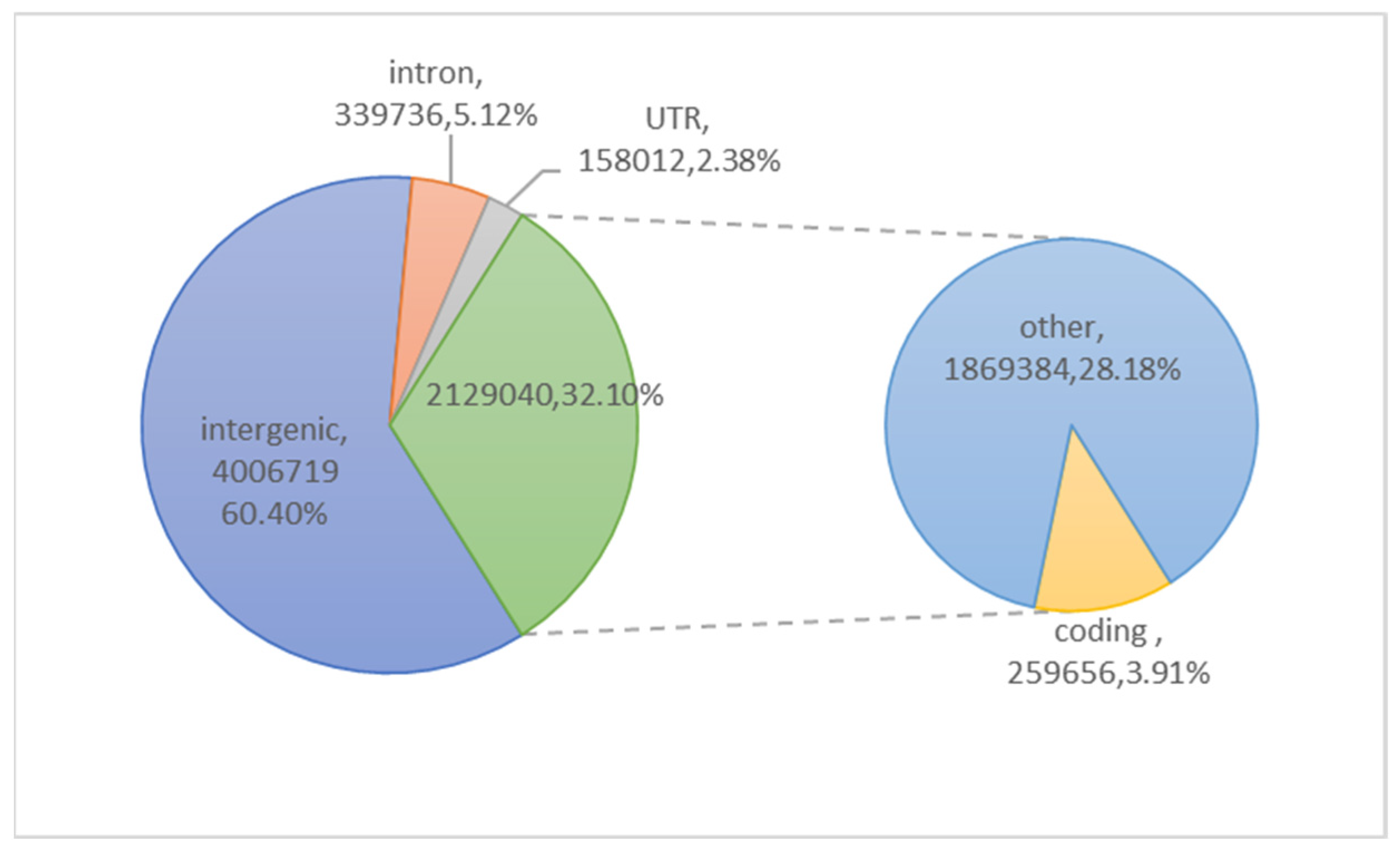
2.2. Sequence and SNPs Information Was Produced by Whole-Genome Resequencing
2.3. Key GuiHeFeng Traceable Blocks Were Found in the Genome of Its Derivatives
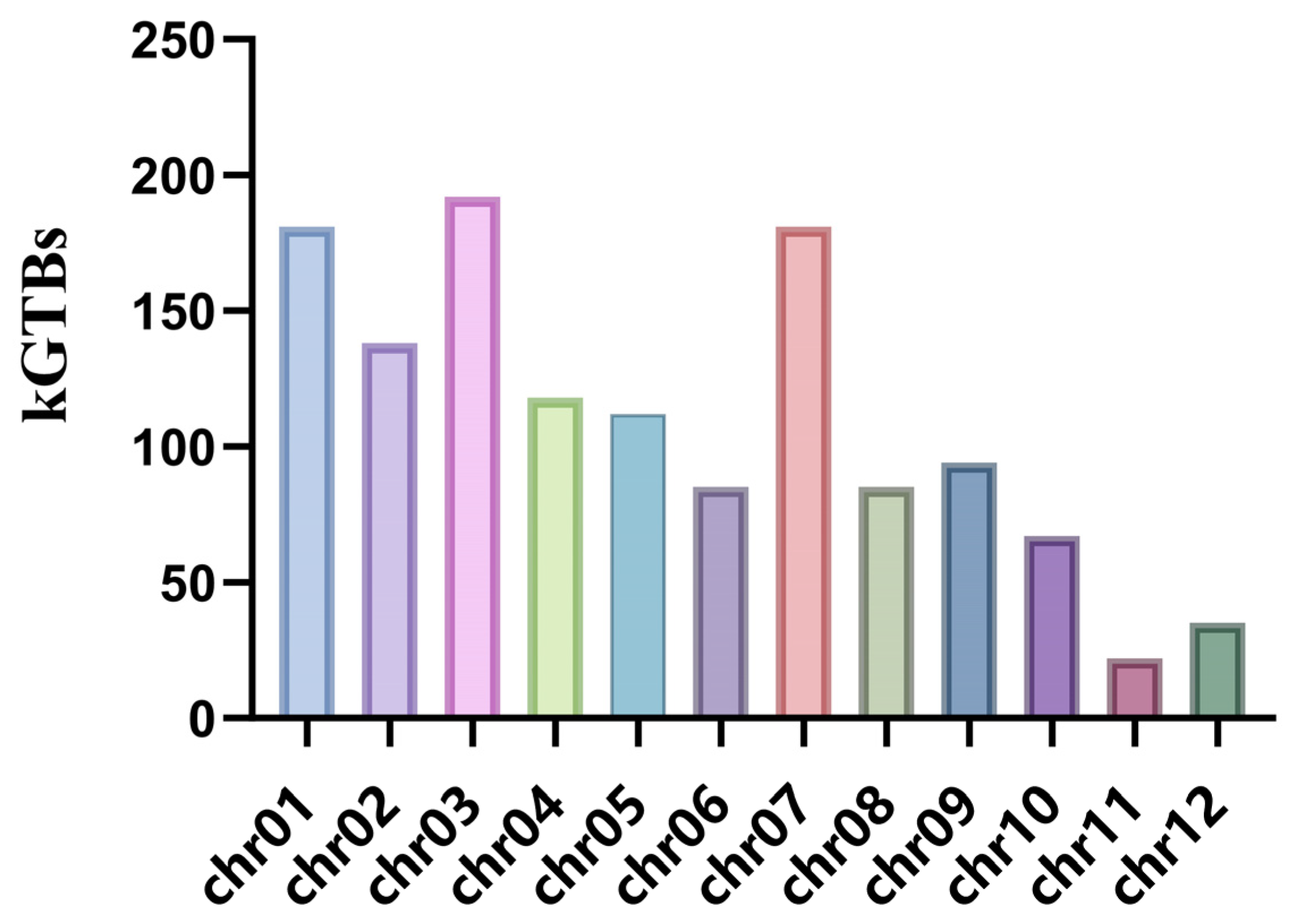
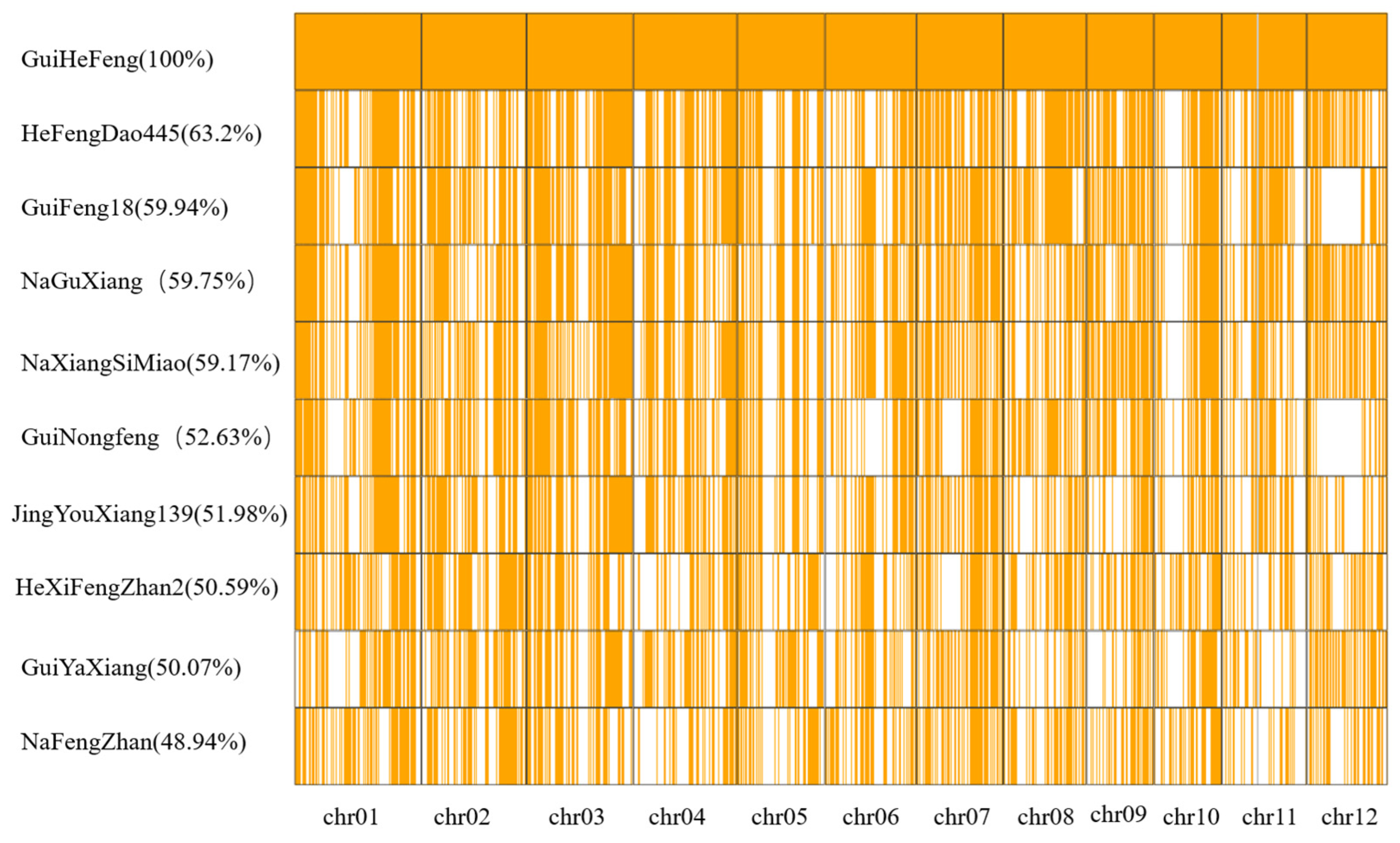
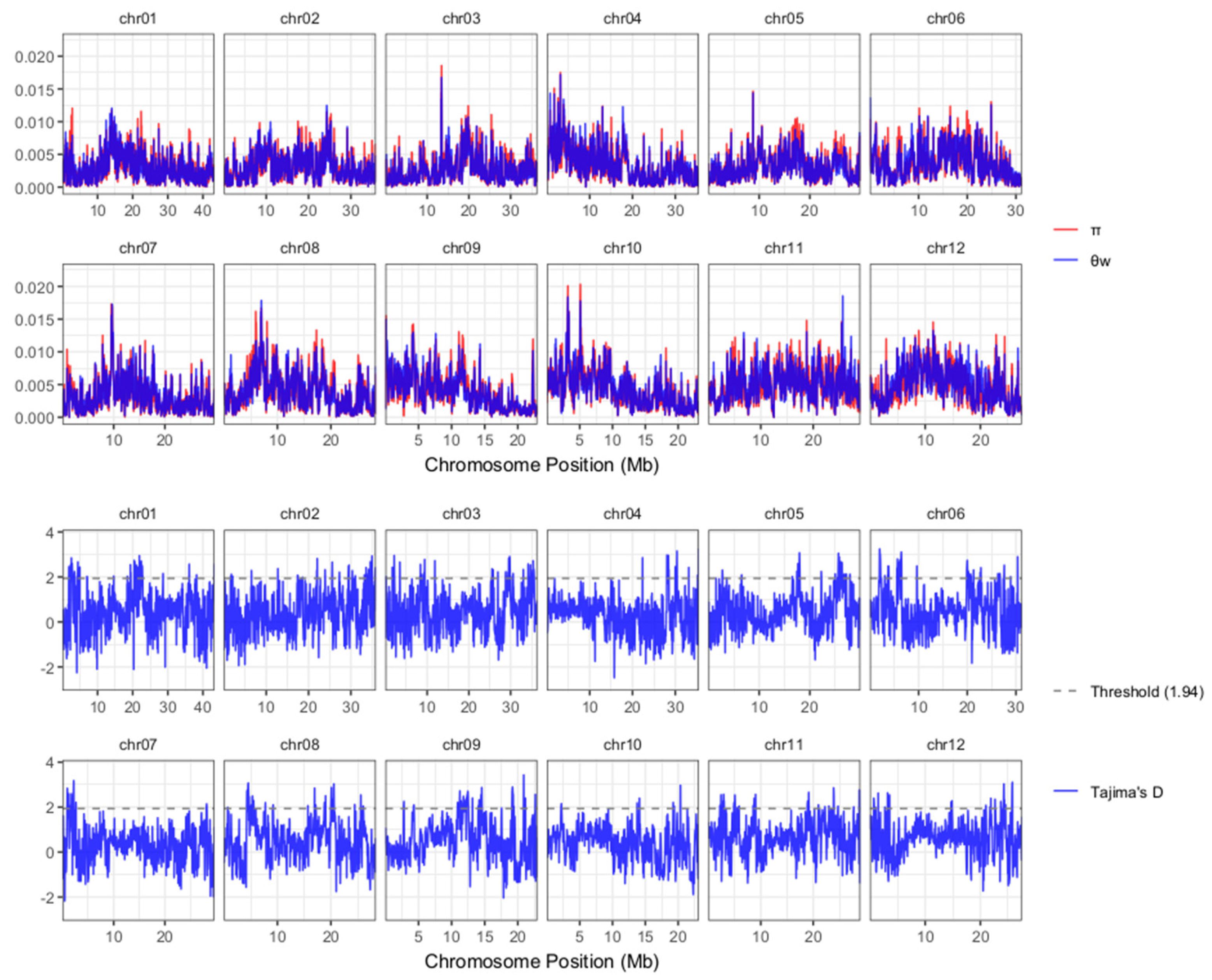
2.4. Key Genomic Regions Were Selected from kGTBs and Selection Sweeps
2.5. Excellent Alleles Were Exploited from kGTBs and Key Genomic Region
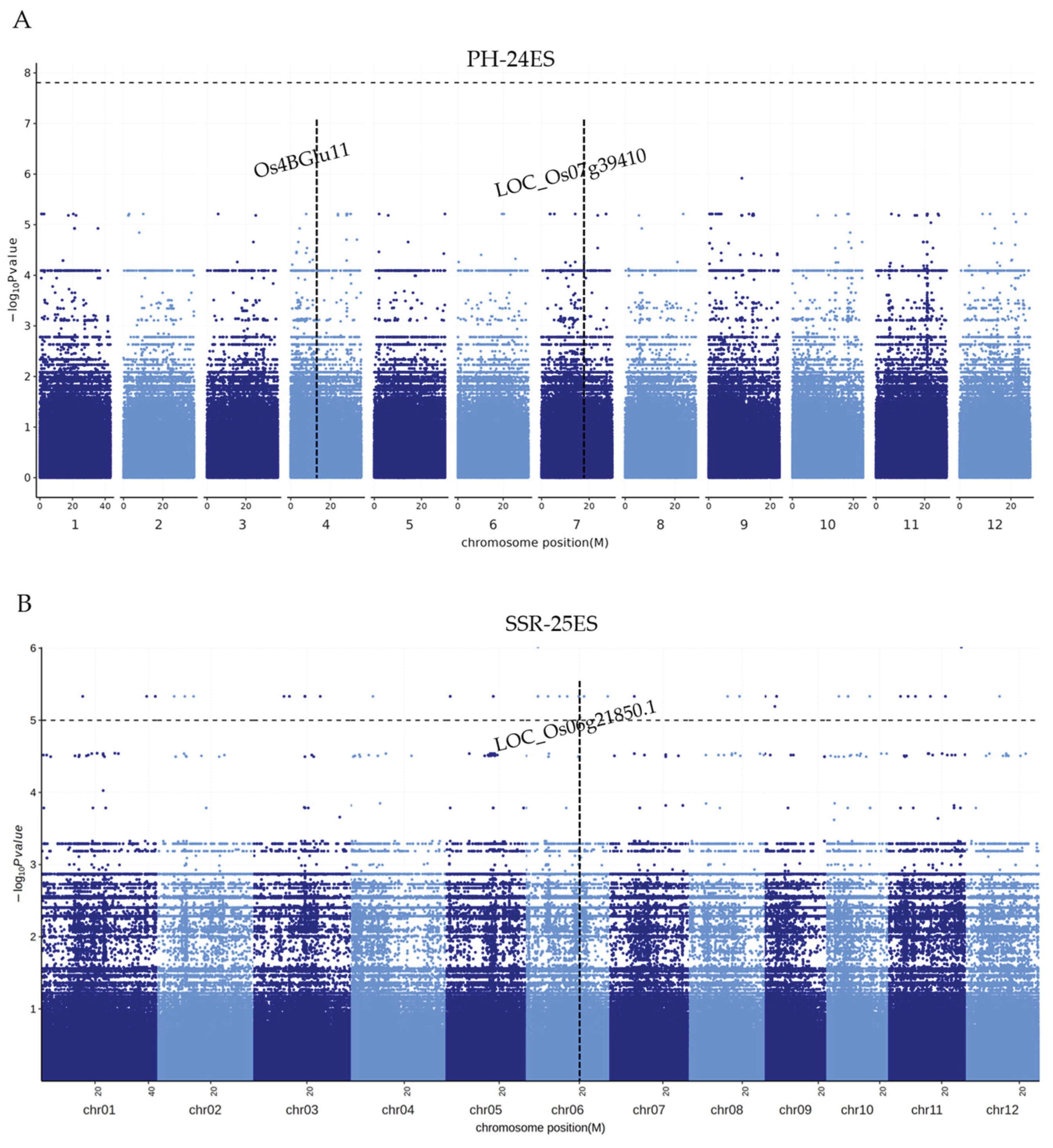
2.6. More SNPs (QTLs) Related to Important Traits Were Detected by GWAS Analysis
3. Discussion
3.1. Important Genes Were Identified from GuiHeFeng and Its Derivatives
3.2. kGTBs and Key Genomic Region Is Useful for Modern Rice Breeding
3.3. GuiHeFeng Is a Backbone Parent for Rice Breeding
4. Materials and Methods
4.1. Plant Materials
4.2. Genome Resequencing and SNP Calling
4.3. Construction of Genome Bins, Identification of Key Genomic Region and Selection Sweep Region
4.4. Association Test and Gene Chip Analysis
4.5. Association Test and Gene Chip Analysis
4.6. Agronomic Trait Investigation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Long-ping, Y. Development of hybrid rice to ensure food security. Rice science. 2014, 21, 1. [Google Scholar]
- Habde, S.V.; Singh, S.K.; Singh, D.K.; Singh, A.K.; Sah, R.P.; Korada, M.; Khaire, A.R.; Majhi, P.K.; Singh, U.M.; Singh, V.K. QTL mapping reveals different set of candidate genes governing stable and location specific QTLs enhancing zinc and iron content in rice. Euphytica. 2024, 220, 179. [Google Scholar] [CrossRef]
- Wang, H.; Deng, X.W. Development of the “third-generation” hybrid rice in China. Genomics, proteomics & bioinformatics. 2018, 16, 393–396. [Google Scholar]
- Peng, S.; Khush, G.S.; Virk, P.; Tang, Q.; Zou, Y. Progress in ideotype breeding to increase rice yield potential. Field Crops Research. 2008, 108, 32–38. [Google Scholar] [CrossRef]
- Zhou, D.; Chen, W.; Lin, Z.; Chen, H.; Wang, C.; Li, H.; Yu, R.; Zhang, F.; Zhen, G.; Yi, J. Pedigree-based analysis of derivation of genome segments of an elite rice reveals key regions during its breeding. Plant Biotechnology Journal. 2016, 14, 638–648. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Lin, Z.; Zhou, D.; Wang, C.; Li, H.; Yu, R.; Deng, H.; Tang, X.; Zhou, S.; Wang Deng, X. Genome-wide study of an elite rice pedigree reveals a complex history of genetic architecture for breeding improvement. Scientific reports. 2017, 7, 45685. [Google Scholar] [CrossRef]
- Abe, A.; Kosugi, S.; Yoshida, K.; Natsume, S.; Takagi, H.; Kanzaki, H.; Matsumura, H.; Yoshida, K.; Mitsuoka, C.; Tamiru, M. Genome sequencing reveals agronomically important loci in rice using MutMap. Nature biotechnology. 2012, 30, 174–178. [Google Scholar] [CrossRef]
- Devanna, B.N.; Sucharita, S.; Sunitha, N.; Anilkumar, C.; Singh, P.K.; Pramesh, D.; Samantaray, S.; Behera, L.; Katara, J.L.; Parameswaran, C. Refinement of rice blast disease resistance QTLs and gene networks through meta-QTL analysis. Scientific Reports. 2024, 14, 16458. [Google Scholar] [CrossRef]
- Khahani, B.; Tavakol, E.; Shariati, V.; Rossini, L. Meta-QTL and ortho-MQTL analyses identified genomic regions controlling rice yield, yield-related traits and root architecture under water deficit conditions. Scientific reports. 2021, 11, 6942. [Google Scholar] [CrossRef]
- Tanin, M.J.; Saini, D.K.; Sandhu, K.S.; Pal, N.; Gudi, S.; Chaudhary, J.; Sharma, A. Consensus genomic regions associated with multiple abiotic stress tolerance in wheat and implications for wheat breeding. Scientific reports. 2022, 12, 13680. [Google Scholar] [CrossRef]
- Zhang, N.; Jun, J.; Fa, C. Dissecting the key genomic regions underlying high yield potential in common wheat variety ‘Kenong 9204’. Journal of Integrative Agriculture. 2023, 22, 2603–2616. [Google Scholar]
- Sansaloni, C.; Franco, J.; Santos, B.; Percival-Alwyn, L.; Singh, S.; Petroli, C.; Campos, J.; Dreher, K.; Payne, T.; Marshall, D. Diversity analysis of 80,000 wheat accessions reveals consequences and opportunities of selection footprints. Nature communications. 2020, 11, 4572. [Google Scholar] [CrossRef]
- Stein, J.C.; Yu, Y.; Copetti, D.; Zwickl, D.J.; Zhang, L.; Zhang, C.; Chougule, K.; Gao, D.; Iwata, A.; Goicoechea, J.L. Genomes of 13 domesticated and wild rice relatives highlight genetic conservation, turnover and innovation across the genus Oryza. Nature genetics. 2018, 50, 285–296. [Google Scholar] [CrossRef]
- Tanger, P.; Klassen, S.; Mojica, J.P.; Lovell, J.T.; Moyers, B.T.; Baraoidan, M.; Naredo, M.E.B.; McNally, K.L.; Poland, J.; Bush, D.R. Field-based high throughput phenotyping rapidly identifies genomic regions controlling yield components in rice. Scientific reports. 2017, 7, 42839. [Google Scholar] [CrossRef] [PubMed]
- Breseghello, F.; Coelho, A.S.G. Traditional and modern plant breeding methods with examples in rice (Oryza sativa L.). Journal of agricultural and food chemistry. 2013, 61, 8277–8286. [Google Scholar] [CrossRef] [PubMed]
- Hoque, A.; Anderson, J.V.; Rahman, M. Genomic prediction for agronomic traits in a diverse Flax (Linum usitatissimum L.) germplasm collection. Scientific Reports. 2024, 14, 3196. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Hao, C.; Wang, Z.; Geng, S.; Jia, M.; Wang, F.; Han, X.; Kong, X.; Yin, L.; Tao, S. Wheat breeding history reveals synergistic selection of pleiotropic genomic sites for plant architecture and grain yield. Molecular Plant. 2022, 15, 504–519. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, H.; Deng, S.; Du, H.; Chen, Z.; Zhao, Y.; Dai, D.; Liang, C.; Li, X.; Liang, C. A backbone parent contributes core genomic architecture to pedigree breeding of early-season indica rice. Journal of genetics and genomics= Yi chuan xue bao. 2021, 48, 1040–1043. [Google Scholar] [CrossRef]
- Huang, J.; Li, J.; Zhou, J.; Wang, L.; Yang, S.; Hurst, L.D.; Li, W.-H.; Tian, D. Identifying a large number of high-yield genes in rice by pedigree analysis, whole-genome sequencing, and CRISPR-Cas9 gene knockout. Proceedings of the National Academy of Sciences. 2018, 115, E7559–E7567. [Google Scholar] [CrossRef]
- Yun, R.; Dan, C.; Wen-jie, L.; Luo, T.; Guo-qiang, Y.; Ye, C.; Xue-mei, L.; Qiming, D.; Shi-quan, W.; Jun, Z. Genome-wide pedigree analysis of elite rice Shuhui 527 reveals key regions for breeding. Journal of Integrative Agriculture. 2021, 20, 35–45. [Google Scholar]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome research. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [PubMed]
- Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics. 1989, 123, 585–595. [Google Scholar] [CrossRef]
- Hutter, S.; Vilella, A.J.; Rozas, J. Genome-wide DNA polymorphism analyses using VariScan. BMC bioinformatics. 2006, 7, 409. [Google Scholar] [CrossRef]
- Kongdin, M.; Mahong, B.; Lee, S.-K.; Shim, S.-H.; Jeon, J.-S.; Ketudat Cairns, J.R. Action of multiple rice β-glucosidases on abscisic acid glucose ester. International Journal of Molecular Sciences. 2021, 22, 7593. [Google Scholar] [CrossRef]
- Terao, T.; Hirose, T. Control of grain protein contents through SEMIDWARF1 mutant alleles: sd1 increases the grain protein content in Dee-geo-woo-gen but not in Reimei. Molecular Genetics and Genomics. 2015, 290, 939–954. [Google Scholar] [CrossRef]
- Song, W.-Y.; Wang, G.-L.; Chen, L.-L.; Kim, H.-S.; Pi, L.-Y.; Holsten, T.; Gardner, J.; Wang, B.; Zhai, W.-X.; Zhu, L.-H. A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science. 1995, 270, 1804–1806. [Google Scholar] [CrossRef]
- Ashikari, M.; Sakakibara, H.; Lin, S.; Yamamoto, T.; Takashi, T.; Nishimura, A.; Angeles, E.R.; Qian, Q.; Kitano, H.; Matsuoka, M. Cytokinin oxidase regulates rice grain production. Science. 2005, 309, 741–745. [Google Scholar] [CrossRef]
- Tian, Z.; Qian, Q.; Liu, Q.; Yan, M.; Liu, X.; Yan, C.; Liu, G.; Gao, Z.; Tang, S.; Zeng, D. Allelic diversities in rice starch biosynthesis lead to a diverse array of rice eating and cooking qualities. Proceedings of the National Academy of Sciences. 2009, 106, 21760–21765. [Google Scholar] [CrossRef]
- Li, Y.; Fan, C.; Xing, Y.; Jiang, Y.; Luo, L.; Sun, L.; Shao, D.; Xu, C.; Li, X.; Xiao, J. Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nature genetics. 2011, 43, 1266–1269. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Wang, Y.; Xue, D.; Wang, J.; Yan, M.; Liu, G.; Dong, G.; Zeng, D.; Lu, Z.; Zhu, X. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nature genetics. 2010, 42, 541–544. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Lin, Z.; Li, H.; Li, X.; Li, J.; Wang, Y.; Zhang, X.; Zhu, Z.; Zhai, W.; Wang, X. TAC1, a major quantitative trait locus controlling tiller angle in rice. The Plant Journal. 2007, 52, 891–898. [Google Scholar] [CrossRef]
- Monna, L.; Kitazawa, N.; Yoshino, R.; Suzuki, J.; Masuda, H.; Maehara, Y.; Tanji, M.; Sato, M.; Nasu, S.; Minobe, Y. Positional cloning of rice semi dwarfing gene, sd-1: rice “green revolution gene” encodes a mutant enzyme involved in gibberellin synthesis. DNA research. 2002, 9, 11–17. [Google Scholar] [CrossRef]
- Li, M.; Tang, D.; Wang, K.; Wu, X.; Lu, L.; Yu, H.; Gu, M.; Yan, C.; Cheng, Z. Mutations in the F-box gene LARGER PANICLE improve the panicle architecture and enhance the grain yield in rice. Plant biotechnology journal. 2011, 9, 1002–1013. [Google Scholar] [CrossRef]
- Wan, X.; Weng, J.; Zhai, H.; Wang, J.; Lei, C.; Liu, X.; Guo, T.; Jiang, L.; Su, N.; Wan, J. Quantitative trait loci (QTL) analysis for rice grain width and fine mapping of an identified QTL allele gw-5 in a recombination hotspot region on chromosome 5. Genetics. 2008, 179, 2239–2252. [Google Scholar] [CrossRef]
- Zhou, Y.; Li, S.; Qian, Q.; Zeng, D.; Zhang, M.; Guo, L.; Liu, X.; Zhang, B.; Deng, L.; Liu, X. BC10, a DUF266-containing and Golgi-located type II membrane protein, is required for cell-wall biosynthesis in rice (Oryza sativa L.). The Plant Journal. 2009, 57, 446–462. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Zhao, F.; Cong, Y.; Sang, X.; Du, Q.; Wang, D.; Li, Y.; Ling, Y.; Yang, Z.; He, G. Rolling-leaf14 is a 2OG-Fe (II) oxygenase family protein that modulates rice leaf rolling by affecting secondary cell wall formation in leaves. Plant biotechnology journal. 2012, 10, 524–532. [Google Scholar] [CrossRef]
- Nakashima, K.; Tran, L.S.P.; Van Nguyen, D.; Fujita, M.; Maruyama, K.; Todaka, D.; Ito, Y.; Hayashi, N.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. The Plant Journal. 2007, 51, 617–630. [Google Scholar] [CrossRef]
- Dong, H.; Zhao, H.; Xie, W.; Han, Z.; Li, G.; Yao, W.; Bai, X.; Hu, Y.; Guo, Z.; Lu, K. A novel tiller angle gene, TAC3, together with TAC1 and D2 largely determine the natural variation of tiller angle in rice cultivars. PLoS genetics. 2016, 12, e1006412. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, T.; Maekawa, M.; Oki, T.; Suda, I.; Iida, S.; Shimada, H.; Takamure, I.; Kadowaki, K.i. The Rc and Rd genes are involved in proanthocyanidin synthesis in rice pericarp. The Plant Journal. 2007, 49, 91–102. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Wu, T.; Ye, J.; Sun, J.; Jiang, Y.; Yu, J.; Tang, J.; Chen, G.; Wang, C.; Wan, J. SNP-based analysis of genetic diversity reveals important alleles associated with seed size in rice. BMC plant biology. 2016, 16, 93. [Google Scholar]
- Leng, Y.; Gao, Y.; Chen, L.; Yang, Y.; Huang, L.; Dai, L.; Ren, D.; Xu, Q.; Zhang, Y.; Ponce, K. Using Heading date 1 preponderant alleles from indica cultivars to breed high-yield, high-quality japonica rice varieties for cultivation in south China. Plant Biotechnology Journal. 2020, 18, 119–128. [Google Scholar] [CrossRef]
- Wu, W.; Zheng, X.-M.; Lu, G.; Zhong, Z.; Gao, H.; Chen, L.; Wu, C.; Wang, H.-J.; Wang, Q.; Zhou, K. Association of functional nucleotide polymorphisms at DTH2 with the northward expansion of rice cultivation in Asia. Proceedings of the National Academy of Sciences. 2013, 110, 2775–2780. [Google Scholar] [CrossRef] [PubMed]
- Si, L.; Chen, J.; Huang, X.; Gong, H.; Luo, J.; Hou, Q.; Zhou, T.; Lu, T.; Zhu, J.; Shangguan, Y. OsSPL13 controls grain size in cultivated rice. Nature genetics. 2016, 48, 447–456. [Google Scholar] [CrossRef]
- Cai, J.; Liao, Q.; Dai, Z.; Zhu, H.; Zeng, R.; Zhang, Z.; Zhang, G.-Q. Allelic differentiations and effects of the Rf3 and Rf4 genes on fertility restoration in rice with wild abortive cytoplasmic male sterility. Biologia plantarum. 2013, 57, 274–280. [Google Scholar] [CrossRef]
- Mizuta, Y.; Harushima, Y.; Kurata, N. Rice pollen hybrid incompatibility caused by reciprocal gene loss of duplicated genes. Proceedings of the National Academy of Sciences. 2010, 107, 20417–20422. [Google Scholar] [CrossRef]
- Kubo, T.; Takashi, T.; Ashikari, M.; Yoshimura, A.; Kurata, N. Two tightly linked genes at the hsa1 locus cause both F1 and F2 hybrid sterility in rice. Molecular plant. 2016, 9, 221–232. [Google Scholar] [CrossRef] [PubMed]
- Xiong, H.; Yu, J.; Miao, J.; Li, J.; Zhang, H.; Wang, X.; Liu, P.; Zhao, Y.; Jiang, C.; Yin, Z. Natural variation in OsLG3 increases drought tolerance in rice by inducing ROS scavenging. Plant Physiology. 2018, 178, 451–467. [Google Scholar] [CrossRef]
- Ochiai, K.; Shimizu, A.; Okumoto, Y.; Fujiwara, T.; Matoh, T. Suppression of a NAC-like transcription factor gene improves boron-toxicity tolerance in rice. Plant physiology. 2011, 156, 1457–1463. [Google Scholar] [CrossRef]
- Fu, J.; Wu, H.; Ma, S.; Xiang, D.; Liu, R.; Xiong, L. OsJAZ1 attenuates drought resistance by regulating JA and ABA signaling in rice. Frontiers in plant science. 2017, 8, 2108. [Google Scholar] [CrossRef]
- Liu, C.; Schläppi, M.R.; Mao, B.; Wang, W.; Wang, A.; Chu, C. The bZIP 73 transcription factor controls rice cold tolerance at the reproductive stage. Plant Biotechnology Journal. 2019, 17, 1834–1849. [Google Scholar] [CrossRef]
- Liu, M.; Zhang, S.; Hu, J.; Sun, W.; Padilla, J.; He, Y.; Li, Y.; Yin, Z.; Liu, X.; Wang, W. Phosphorylation-guarded light-harvesting complex II contributes to broad-spectrum blast resistance in rice. Proceedings of the National Academy of Sciences. 2019, 116, 17572–17577. [Google Scholar] [CrossRef]
- Liu, M.; Fan, F.; He, S.; Guo, Y.; Chen, G.; Li, N.; Li, N.; Yuan, H.; Si, F.; Yang, F. Creation of elite rice with high-yield, superior-quality and high resistance to brown planthopper based on molecular design. Rice. 2022, 15, 17. [Google Scholar] [CrossRef]
- Reinke, R.; Kim, S.-M.; Kim, B.-K. Developing Japonica rice introgression lines with multiple resistance genes for brown planthopper, bacterial blight, rice blast, and rice stripe virus using molecular breeding. Molecular Genetics and Genomics. 2018, 293, 1565–1575. [Google Scholar]
- Lei, L.; Zheng, H.; Wang, J.; Liu, H.; Sun, J.; Zhao, H.; Yang, L.; Zou, D. Genetic dissection of rice (Oryza sativa L.) tiller, plant height, and grain yield based on QTL mapping and meta-analysis. Euphotic. 2018, 214, 109. [Google Scholar] [CrossRef]
- Yang, J.; Hu, C.; Hu, H.; Yu, R.; Xia, Z.; Ye, X.; Zhu, J. QTL Network: mapping and visualizing genetic architecture of complex traits in experimental populations. Bioinformatics. 2008, 24, 721–723. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Zhong, X.; Pan, J.; Liang, K.; Liu, Y.; Peng, B.; Hu, X.; Huang, N. QTLs identification for nitrogen and phosphorus uptake-related traits using ultra-high density SNP linkage. Plant Science. 2019, 288, 110209. [Google Scholar] [CrossRef] [PubMed]
- Gu, R.; Chen, F.; Long, L.; Cai, H.; Liu, Z.; Yang, J.; Wang, L.; Li, H.; Li, J.; Liu, W. Enhancing phosphorus uptake efficiency through QTL-based selection for root system architecture in maize. Journal of Genetics and Genomics. 2016, 43, 663–672. [Google Scholar] [CrossRef]
- Han, B.; Cui, D.; Ma, X.; Cao, G.; Zhang, H.; Koh, H.J.; Han, L. Evidence for evolution and selection of drought-resistant genes based on high-throughput resequencing in weedy rice. Journal of Experimental Botany. 2022, 73, 1949–1962. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics. 2010, 26, 589–595. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Subgroup, G.P.D.P. The sequence alignment/map format and SAMtools. bioinformatics. 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly. 2012, 6, 80–92. [Google Scholar] [PubMed]
- Crossa, J.; Campos, G.d.l.; Pérez, P.; Gianola, D.; Burgueno, J.; Araus, J.L.; Makumbi, D.; Singh, R.P.; Dreisigacker, S.; Yan, J. Prediction of genetic values of quantitative traits in plant breeding using pedigree and molecular markers. Genetics. 2010, 186, 713–724. [Google Scholar] [PubMed]
| No. | Variety | Pedigree |
| 1 | GuiHeFeng | HeFengZhan/YueTaiZhan |
| 2 | GuiFeng18 | GuiHeFeng/MeiXiangZhan |
| 3 | HeFengDao445 | GuiHeFeng/GuiYu9Hao |
| 4 | NaFengZhan | GuiHeFeng//ZaoHui3Hao/GuiHui1561 |
| 5 | JingYouXiang139 | BaiXiang139/GuiHeFeng |
| 6 | GuiYaXiang | GuiHeFeng/XiangChangMang |
| 7 | GuiNongFeng | GuiHeFeng/YeXiangZhan |
| 8 | NaXiangSiMiao | GuiHeFeng//BaiXiang139/GuiHui110 |
| 9 | NaGuXiang | GuiHeFeng//BaiXiang139/HuangHuaZhan |
| 10 | HeXiFengZhan2Hao | HeXiXiang/GuiHeFeng |
| Variety |
Reads (M) |
Bases (G) |
Map Reads (%) |
Map Reads |
Depth X |
Cov_ratio (%) |
| GuiHeFeng | 124.74 | 18.59 | 98.59 | 122978517 | 51.78 | 89.69 |
| GuiFeng18 | 58.54 | 8.72 | 98.73 | 57794774 | 24.51 | 85.53 |
| HeFengDao445 | 66.84 | 9.93 | 98.77 | 66016096 | 27.99 | 86.67 |
| NaFengZhan | 70.66 | 10.53 | 98.62 | 69682802 | 29.65 | 86.98 |
| JingYouXiang139 | 84.91 | 12.60 | 98.70 | 83805328 | 35.4 | 87.92 |
| GuiYaXiang | 76.20 | 11.34 | 98.58 | 75121141 | 31.73 | 87.22 |
| GuiNongFeng | 71.49 | 10.66 | 98.61 | 70495489 | 29.83 | 87.66 |
| NaXiangSiMiao | 81.89 | 12.17 | 98.62 | 80756694 | 34.09 | 87.74 |
| NaGuXiang | 68.9 | 10.23 | 98.62 | 67954951 | 28.74 | 86.36 |
| HeXiFengZhan2 | 47.15 | 7.04 | 98.46 | 46426567 | 19.98 | 82.42 |
| Sum | 751.32 | 111.18 |
| No. | Chr | Star | End | Gene | Function | Category | Genechip result |
| 1 | chr01 | 25383093 | 25383093 | Rd/DFR/OsDfr | red seed coat | Seed Morphology | T |
| 2 | chr01 | 5244076 | 5244076 | D2/CYP90D2/SMG11 | larger tiller angle | Plant Architecture | T |
| 3 | chr01 | 5270928 | 5270928 | Gn1a/OsCKX2 | increasing grain number | Yield components | T |
| 4 | chr01 | 5275530 | 5275530 | Gn1a/OsCKX2 | increasing grain number | Yield components | T |
| 5 | chr01 | 5275544 | 5275544 | Gn1a/OsCKX2 | increasing grain number | Yield components | T |
| 6 | chr01 | 5568692 | 5568692 | Rf3/OsMADS3 | fertility restoration | Yield components | T |
| 7 | chr02 | 30096330 | 30096330 | DTH2/Hd7 | delaying heading date under LD | Heading date | T |
| 8 | chr03 | 4353347 | 4353347 | OsLG3 | increasing drought tolerance | Yield components | T |
| 9 | chr03 | 4353103 | 4353103 | OsLG3 | increasing drought tolerance | Yield components | T |
| 10 | chr04 | 23886659 | 23886659 | BET1 | Increasing boron-toxicity tolerance | Abiotic Stress | T |
| 11 | chr04 | 28894753 | 28894753 | OsCYP704A3 | Longer seed size | Seed Morphology | T |
| 12 | chr04 | 33304910 | 33304910 | OsJAZ1 | decreasing root length and weight | Abiotic Stress | T |
| 13 | chr06 | 4201227 | 4201227 | DPL2 | hybrid incompatibility | Yield components | T |
| 14 | chr06 | 9338220 | 9338220 | Hd1 | Promoting heading date under LD | Heading date | T |
| 15 | chr07 | 19060398 | 19060398 | OsUGT707A2 | more 5-O-glucoside | Secondary metabolism | T |
| 16 | chr07 | 19103249 | 19103249 | OsSPL13/GLW7 | increasing grain size | Yield components | T |
| 17 | chr09 | 18122850 | 18122850 | bZIP73 | decreasing chilling tolerance | Abiotic Stress | T |
| 18 | chr09 | 20731844 | 20731844 | TAC1 | Spread-out plant architecture | Plant Architecture | T |
| 19 | chr11 | 7659694 | 7659694 | LHCB5 | increasing blast resistance | Biotic Stress | T |
| 20 | chr12 | 24669797 | 24669797 | HSA1b | hybrid incompatibility | Yield components | T |
| No. | Chr | Star | End | Gene | Function | Category |
| 1 | chr01 | 2053583 | 2057638 | LRK10L-2.1 | resistance gene analogs (RGAs) | Biotic stress |
| 2 | chr01 | 28666309 | 28668106 | Xa21 | bacterial blight resistance | Biotic stress |
| 3 | chr01 | 28669479 | 28673568 | OsLRR-RLK | Regulate defence reaction | Biotic stress |
| 4 | chr02 | 12798344 | 12804729 | Retrovirus-related Pol polyprotein from transposon RE1 | Increase the Resistance for Broad bean wilt virus 2 | Biotic stress |
| 5 | chr03 | 26952048 | 26959200 | OsTHIC | positively REGULATE vitamin B 1 synthesis | Other |
| 6 | chr03 | 3489869 | 3500130 | TOP3α | regulates meiotic recombination | Other |
| 7 | chr04 | 22369632 | 22376812 | OsABA1 | Positively regulate plant development and adaptation to abiotic and biotic stresses | Biotic/Abiotic Stress |
| 8 | chr04 | 22353707 | 22355207 | OsAP37 | Mediate the tolerance to drought | Abiotic Stress |
| 9 | chr04 | 22362239 | 22367204 | OsPT17 | Involved in Chilling Response and salt stress | Abiotic Stress |
| 10 | chr04 | 22389303 | 22393831 | OsPP65 | Decrease rice resistance to chilling | Abiotic Stress |
| 11 | chr04 | 33185813 | 33186889 | OsWAK54 | plays important roles in cell expansion, pathogen resistance | Biotic stress |
| 12 | chr04 | 33192623 | 33196131 | OsWAK55 | plays important roles in cell expansion, pathogen resistance | Biotic stress |
| 13 | chr04 | 35287781 | 35289156 | OsPR5 | increase pathogen resistance | Biotic stress |
| 14 | chr04 | 35270952 | 35276805 | OsSPARK2 | negatively regulation the tolerance | Biotic/Abiotic Stress |
| 15 | chr06 | 28941271 | 28943704 | OsRRK1 | Positively regulate brown planthopper resistance | Biotic stress |
| 16 | chr06 | 28905577 | 28909089 | OsLRR-RLK1 | initiates striped stem borer resistance | Biotic stress |
| 17 | chr06 | 30357699 | 30361201 | OsNPSN11 | Positively regulate the blast resistance | Biotic stress |
| 18 | chr08 | 15695534 | 15703960 | Protein PHR1-LIKE 3 | enhances tolerance to Pi deficiency and salt stress in rice | Abiotic Stress |
| 19 | chr09 | 13154943 | 13155832 | OsSAP17 | Enhancing plant resistance to drought and salt | Abiotic Stress |
| 20 | chr09 | 13181330 | 13184741 | OsPHD38 | Mediate the tolerance to drought and salt stress | Abiotic Stress |
| 21 | chr09 | 17556929 | 17558591 | OsDjC69 | Mediate flowering and the tolerance to drought and salt stress | Abiotic Stress |
| 22 | chr09 | 17565471 | 17566197 | OsbHLH043 | Mediate the tolerance to drought and arsenic stress | Abiotic Stress |
| 23 | chr09 | 20915301 | 20919808 | OsMYB85 | cell wall regulators | Other |
| 24 | chr09 | 21151736 | 21154358 | OsCYP-24 | Mediate the tolerance to drought and salt stress | Abiotic Stress |
| 25 | chr09 | 21155956 | 21157945 | OsRNS4 | enhanced tolerance to high salinity | Abiotic Stress |
| 26 | chr09 | 21171653 | 21174067 | OsPAD1 | regulate pollen aperture formation | Fertility |
| 27 | chr09 | 21189381 | 21190738 | OsMYB31 | Increase yield | Yield components |
| 28 | chr09 | 21197503 | 21199723 | MS5 | regulate pollen formation | Fertility |
| 29 | chr09 | 21199731 | 21202763 | OsAPX9 | Increase the tolerance to drought, plant height and heading date | Abiotic Stress/Heading date/Plant Architecture |
| 30 | chr09 | 22653849 | 22657046 | Ohp2 | Positively Mediate the tolerance to salt stress | Abiotic Stress |
| 31 | chr09 | 22666306 | 22671392 | OsWD40-174 | take important role in rice-Xoo interactions | Biotic stress |
| 32 | chr10 | 20355076 | 20355657 | OsERF18 | enhances tolerance to Pi deficiency | Abiotic Stress |
| 33 | chr10 | 20374252 | 20375455 | OsEMSA1 | Involved in Embryo sac Development | Fertility |
| 34 | chr10 | 20377195 | 20380235 | OsNP1 | required for another cuticle formation and pollen exine patterning | Fertility |
| 35 | chr10 | 20377031 | 20386096 | OsPLDbeta1 | activates defense responses and increases disease resistance in rice | Biotic stress |
| 36 | chr11 | 28804248 | 28808550 | OsHSP70 | Induces the tolerance to high temperature stress | Abiotic Stress |
| 37 | chr11 | 28827676 | 28828513 | OsMT1a | positively regulated rice resistance to blast | Biotic stress |
| 38 | chr11 | 28845905 | 28852938 | OsSCL57 | Regulate the phosphorus homeostasis of rice | Other |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




