Submitted:
27 November 2025
Posted:
28 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
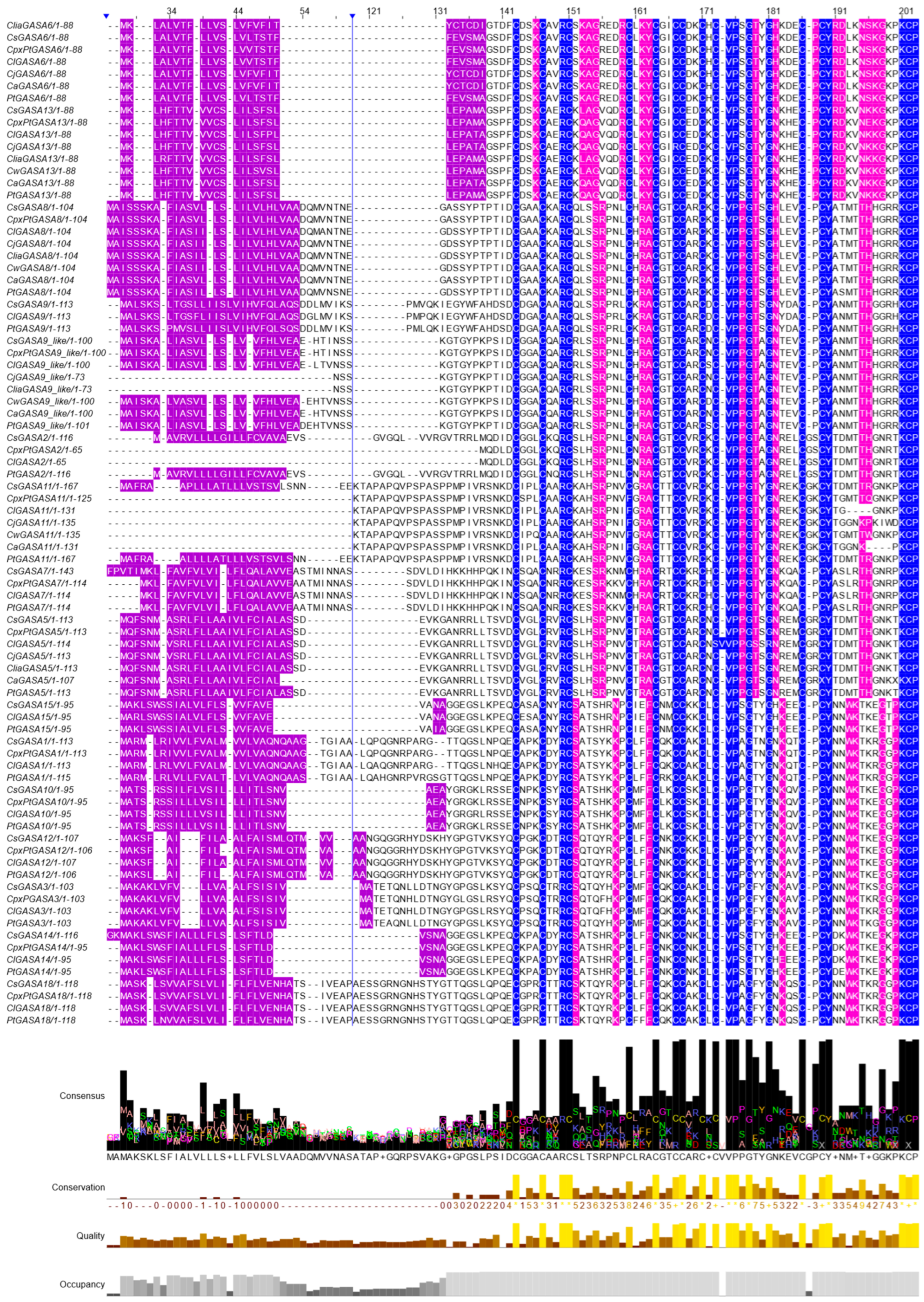
2.1. Identification and Classification of New and Old Genomic Variants of Citrus GASA Genes
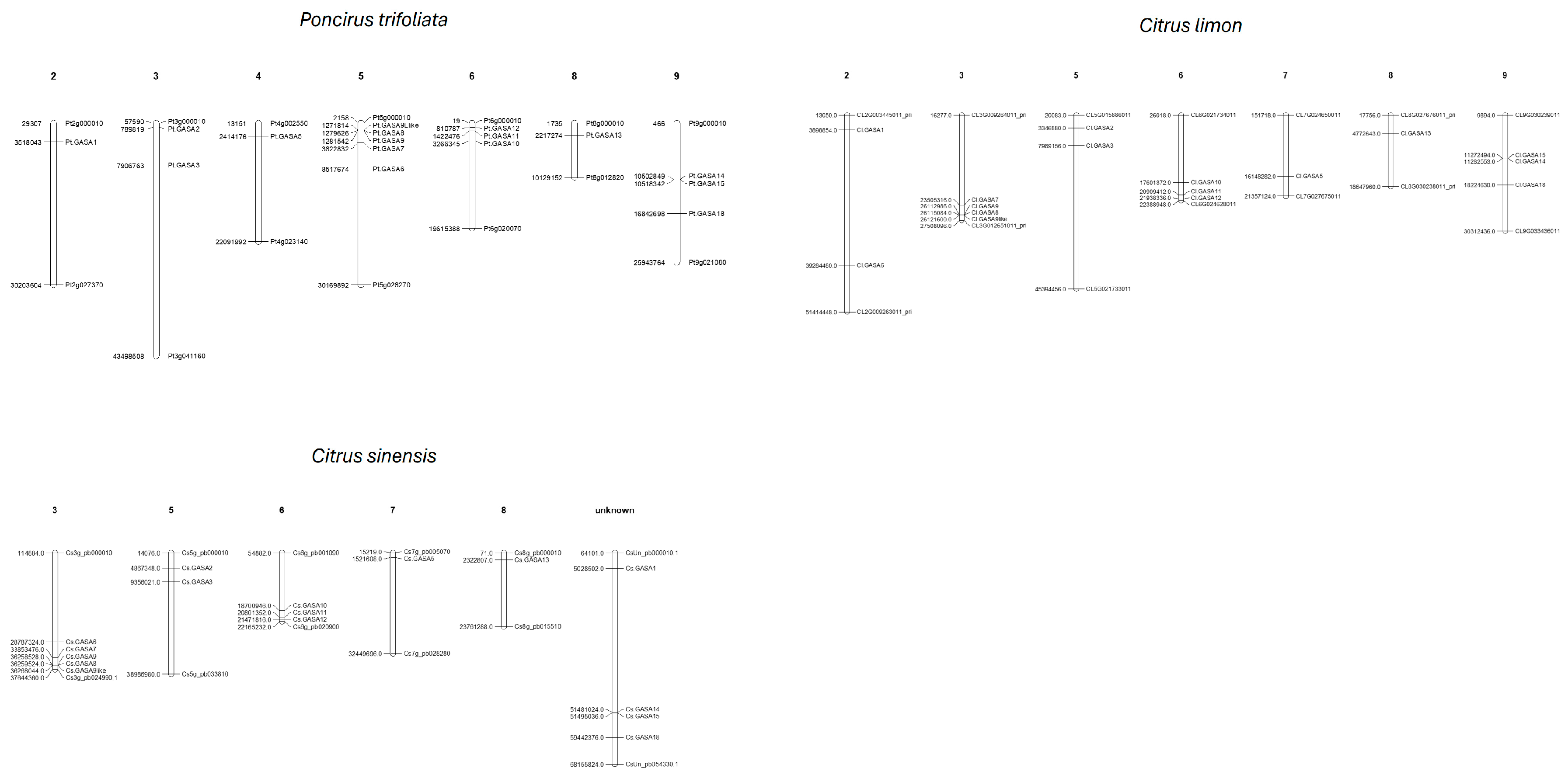
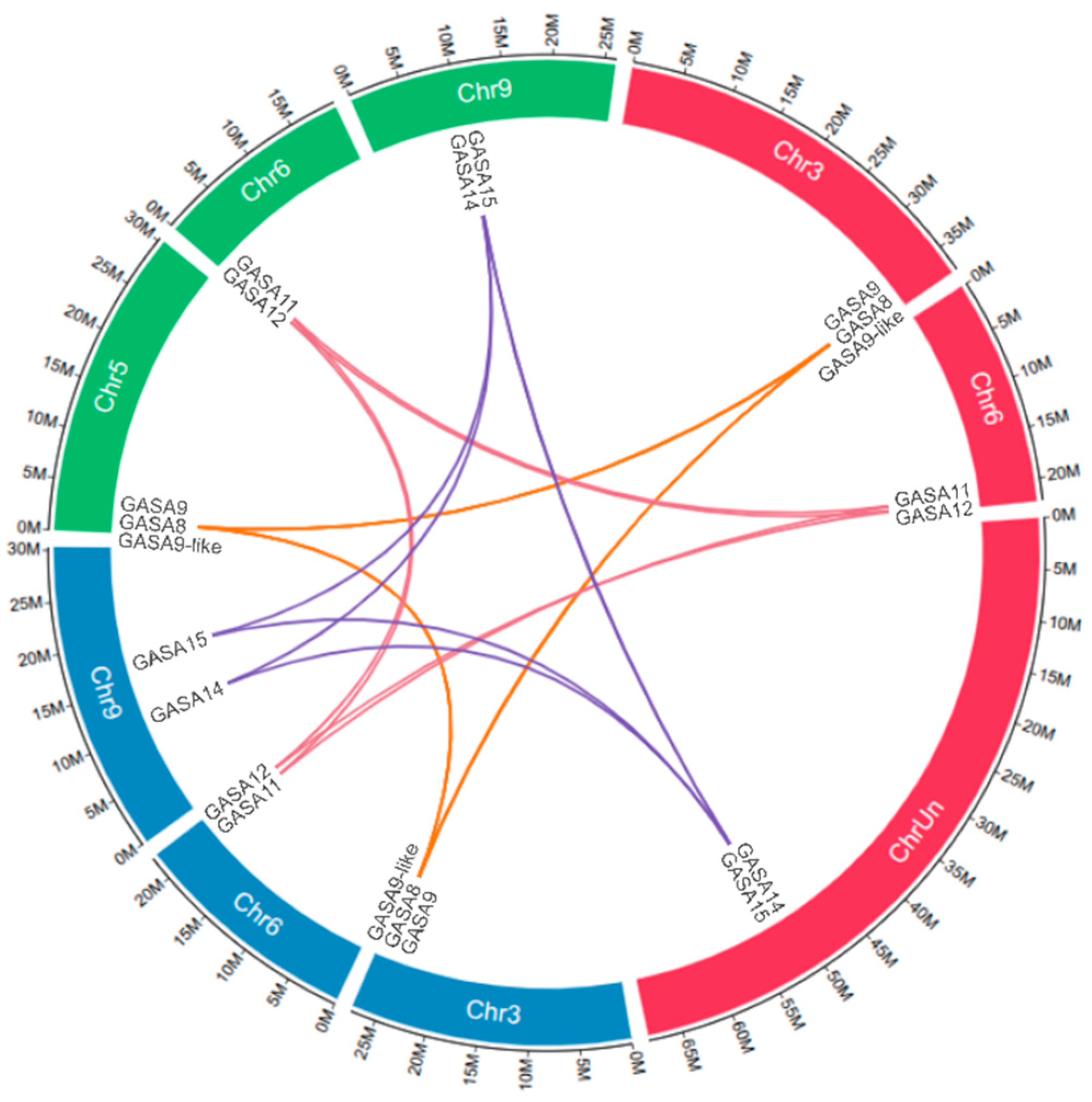
2.2. Location of GASA Genes in Chromosomes and Synteny Blocks
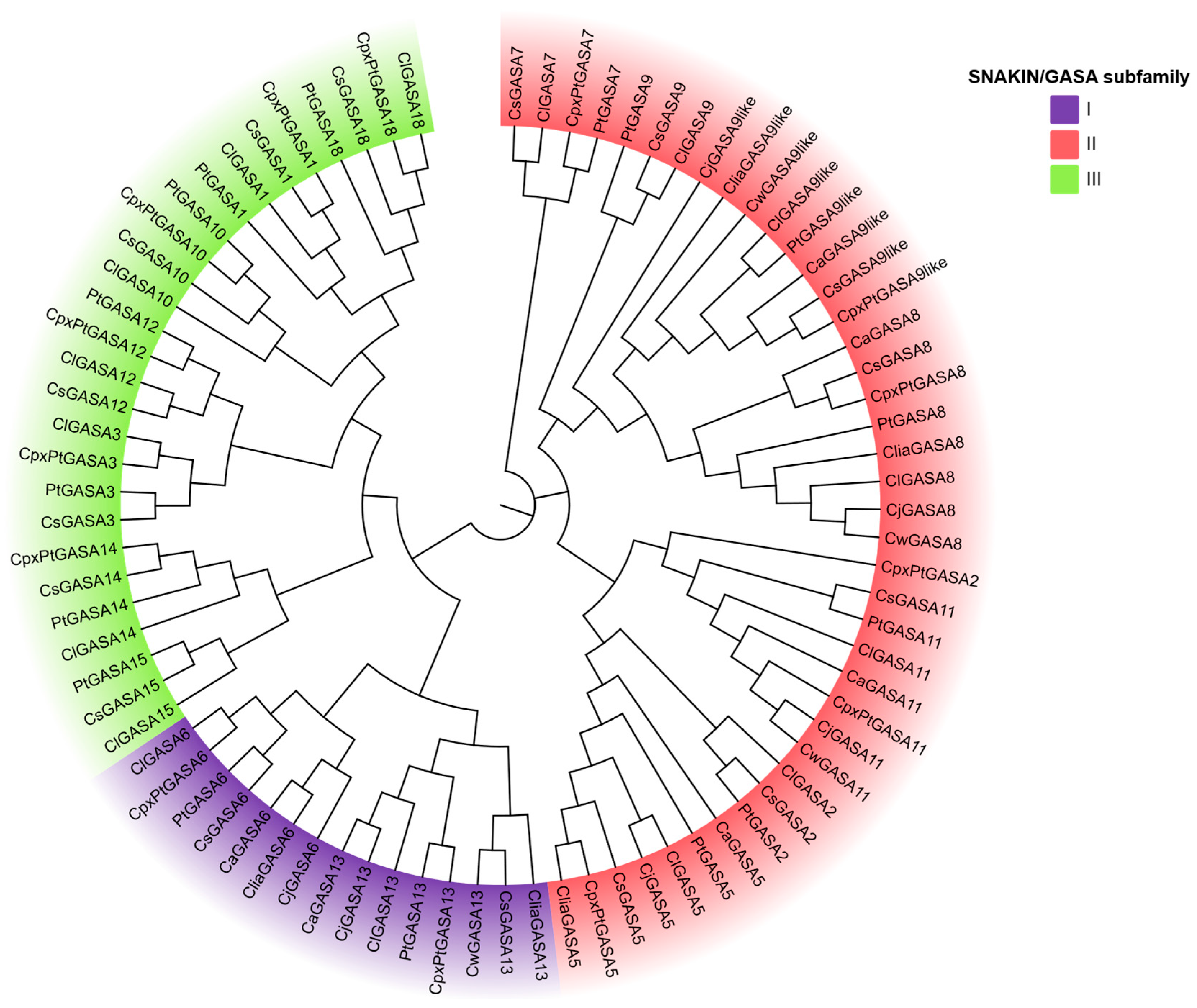
2.3. Phylogenetic Analysis of GASA Genes
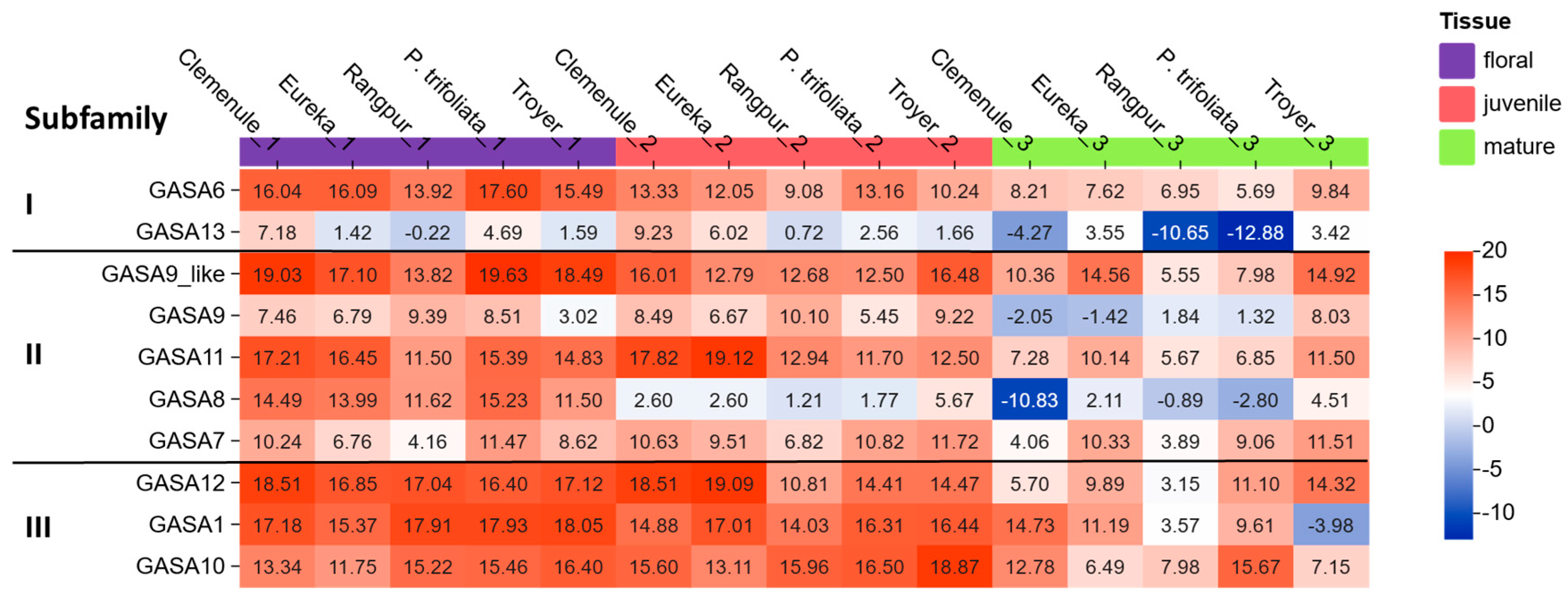
2.4. Differential Expression of Selected Citrus GASA Genes in Juvenile, Mature Leaves and Floral Tissues
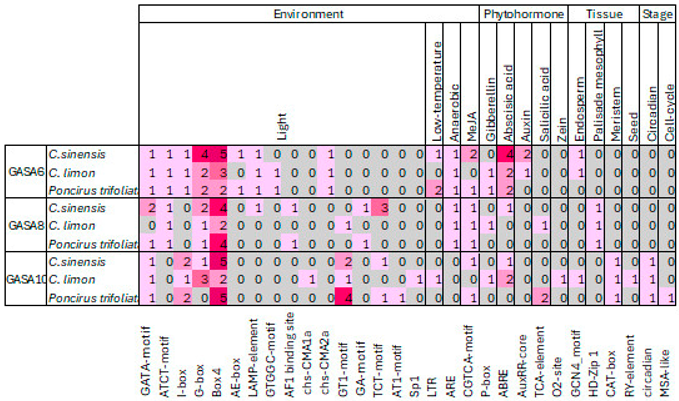
2.5. Promoter Analysis of Citrus GASA 6, 8 and 10 Selected Genes
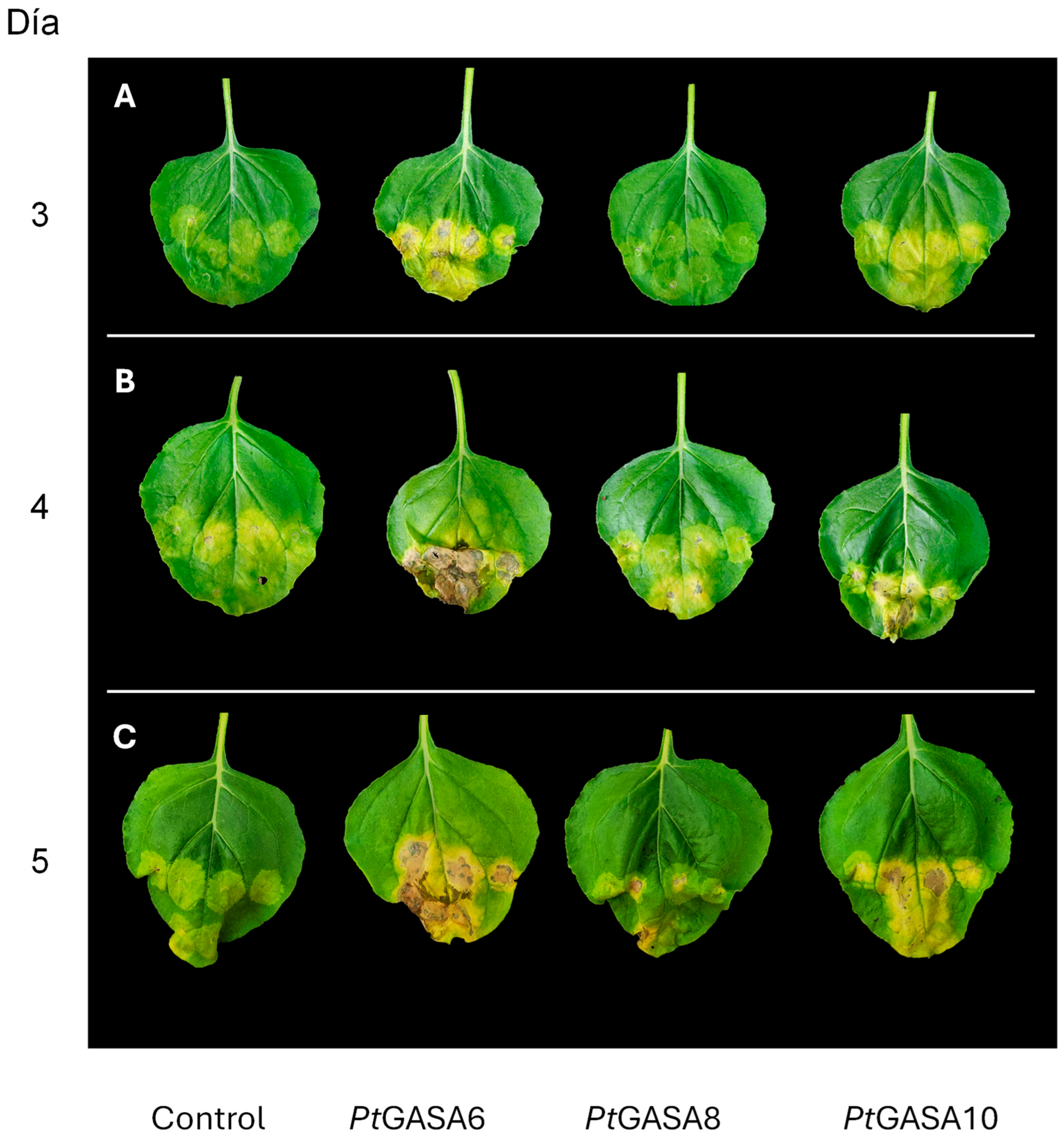
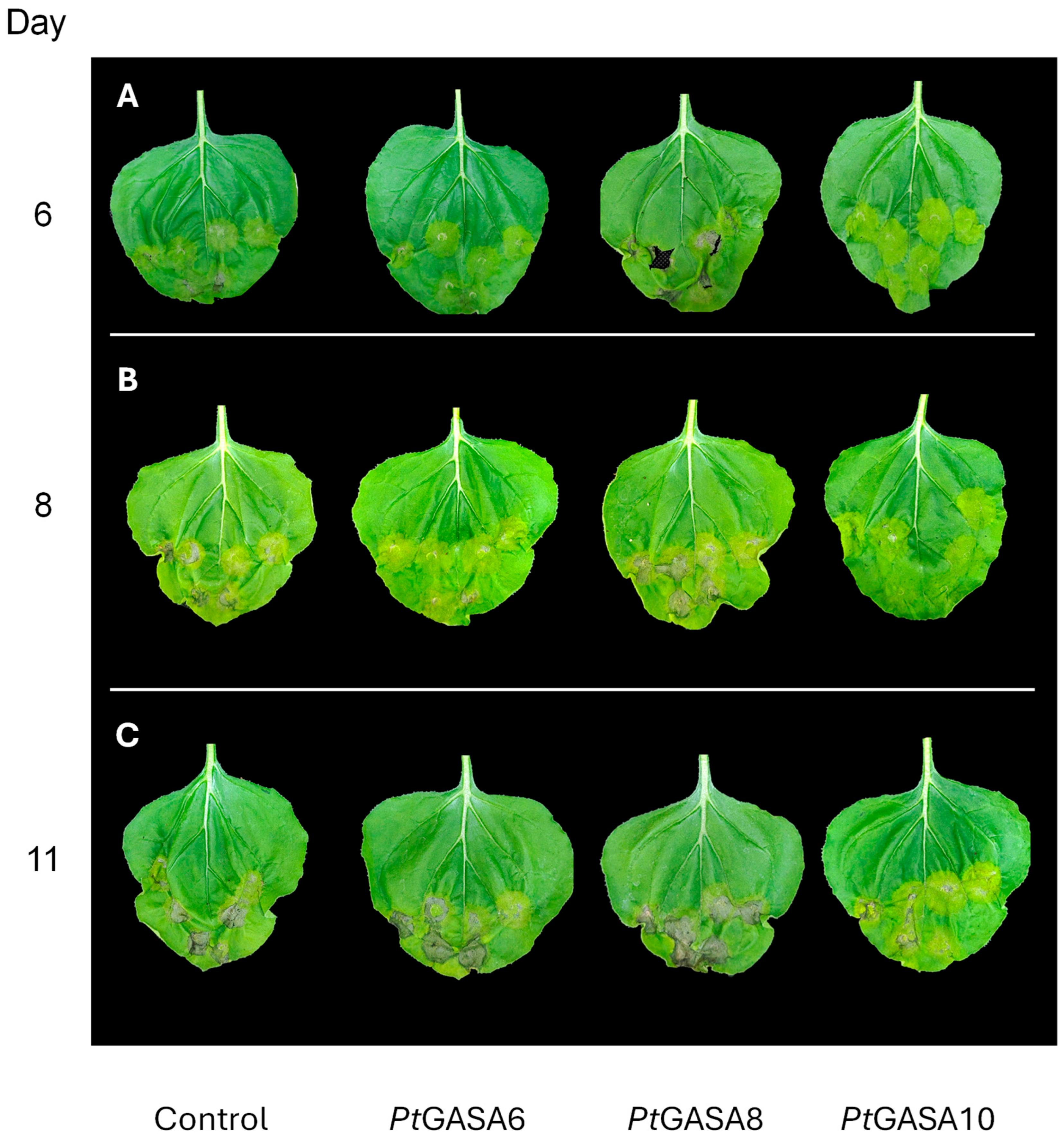
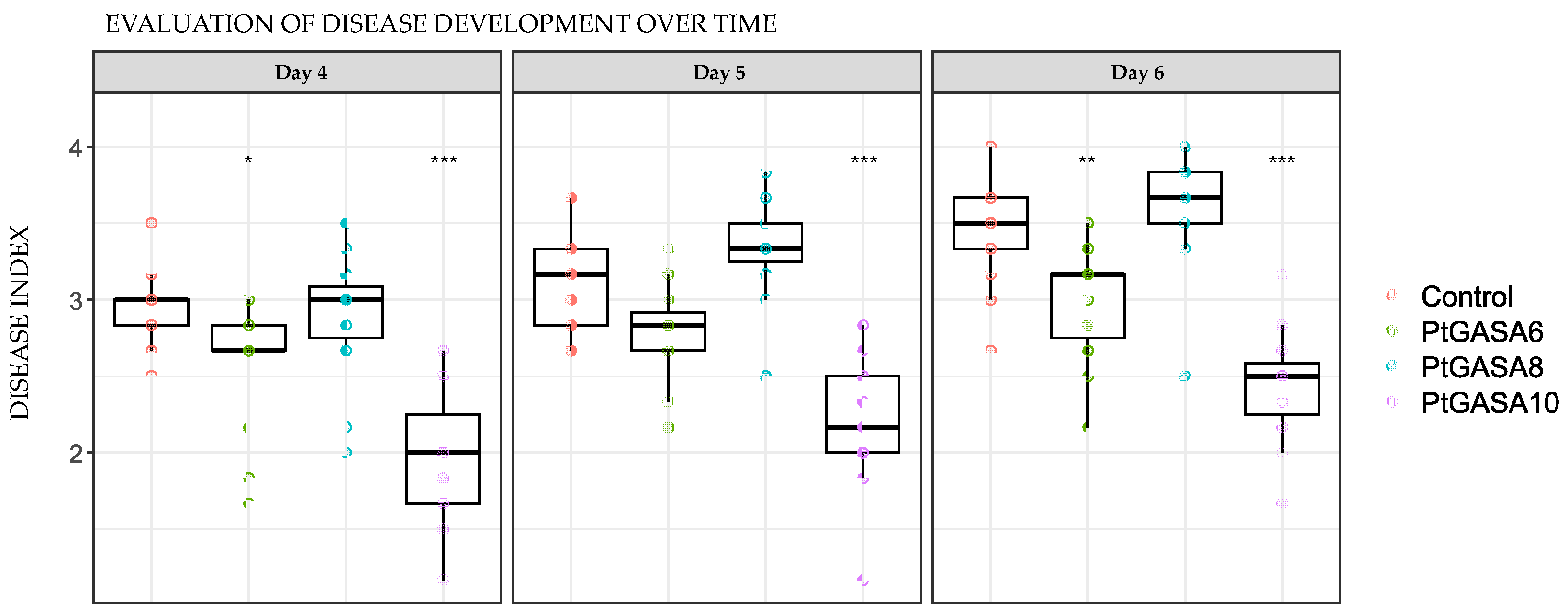
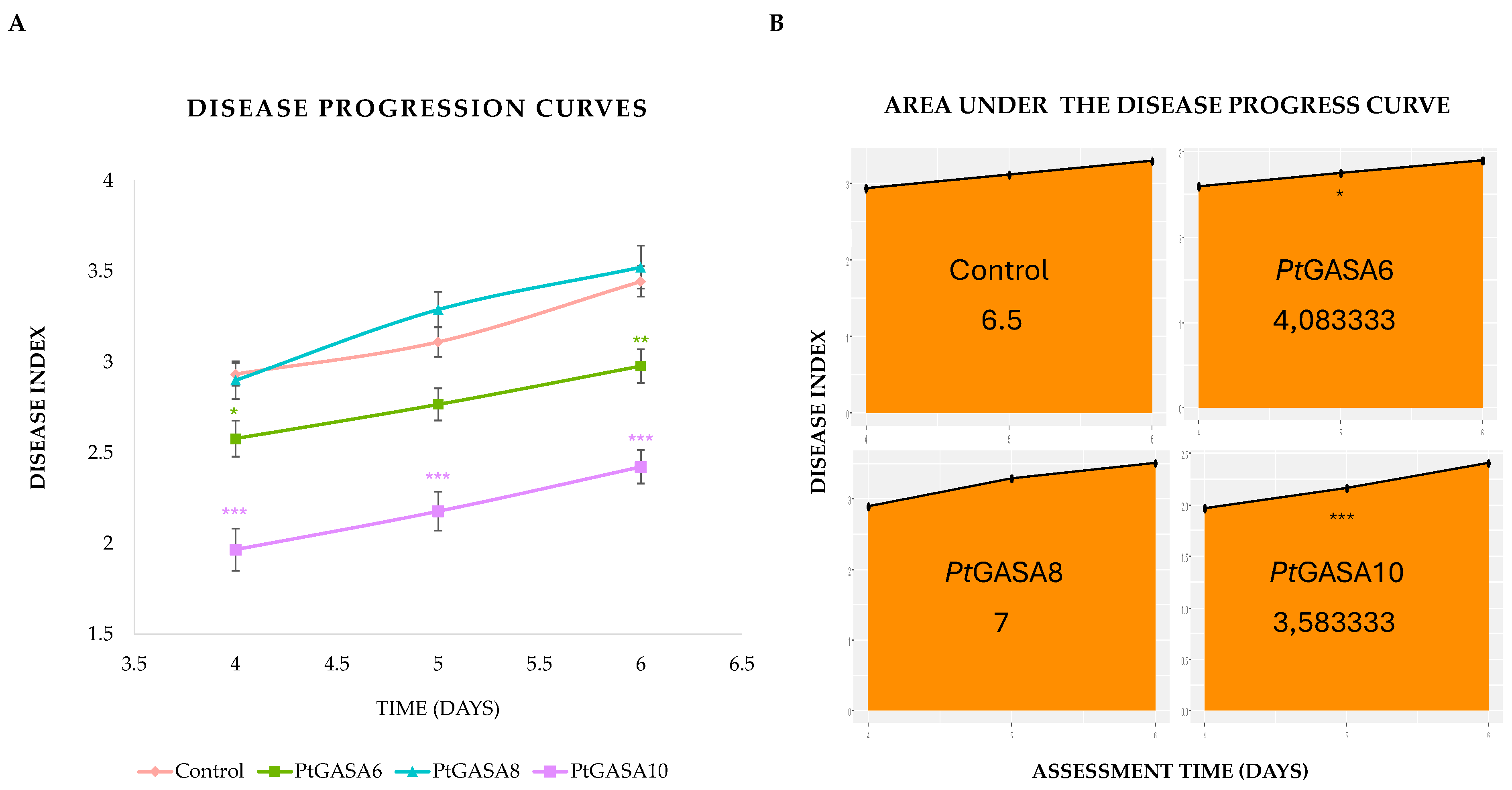
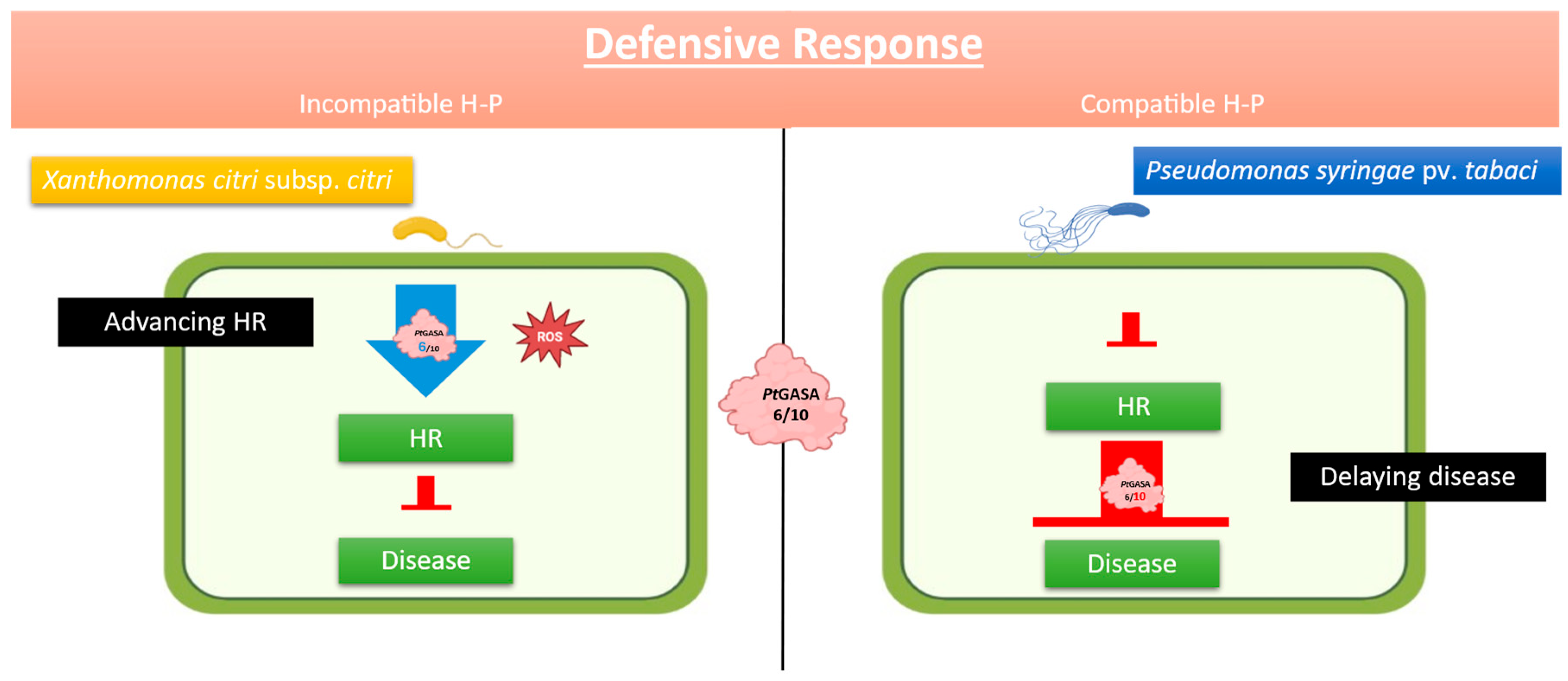
2.6. Overexpression of PtGASA6 and 10 Decreased Disease Symptoms Caused by Pseudomonas syringae pv. Tabaci
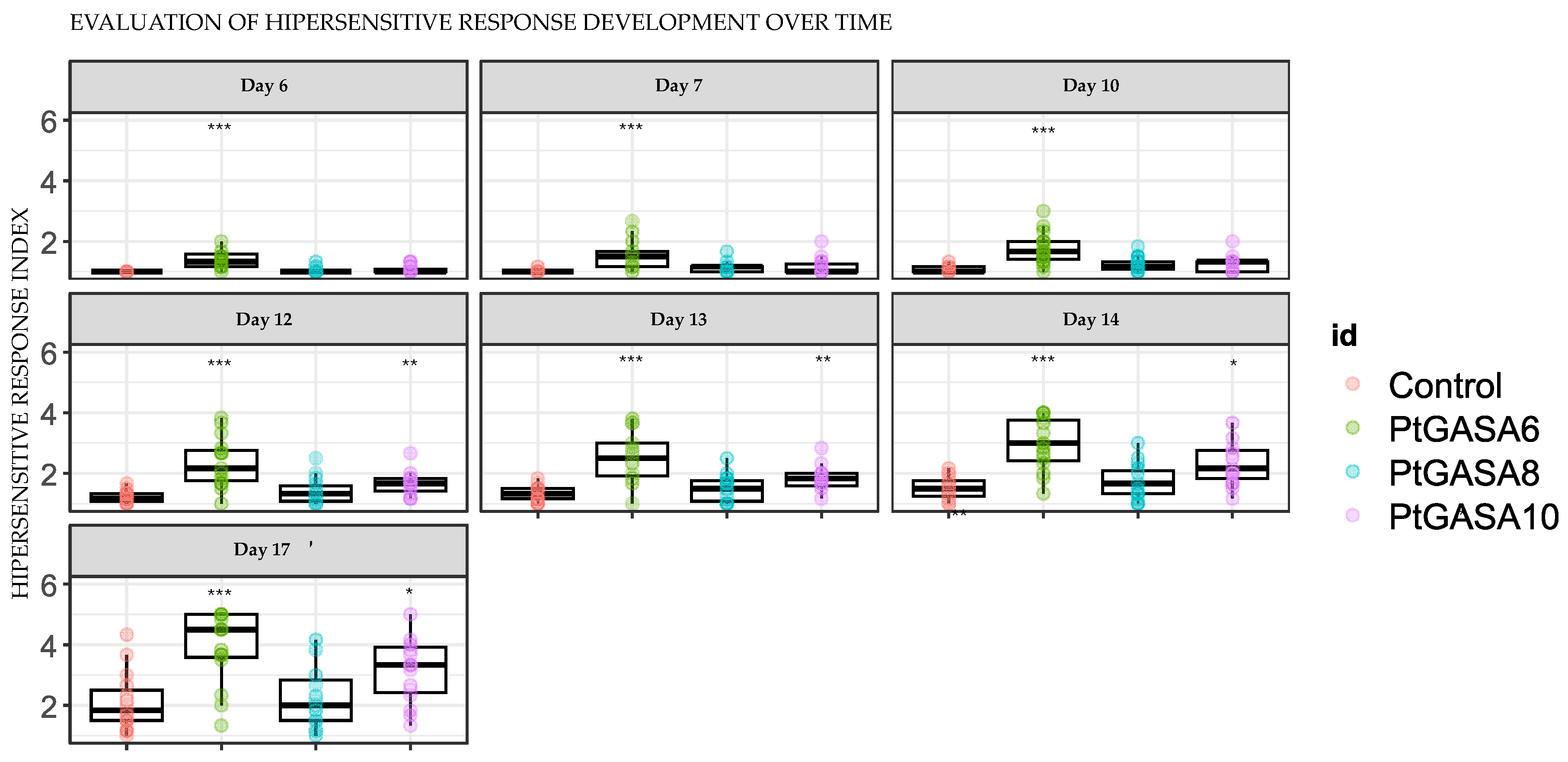
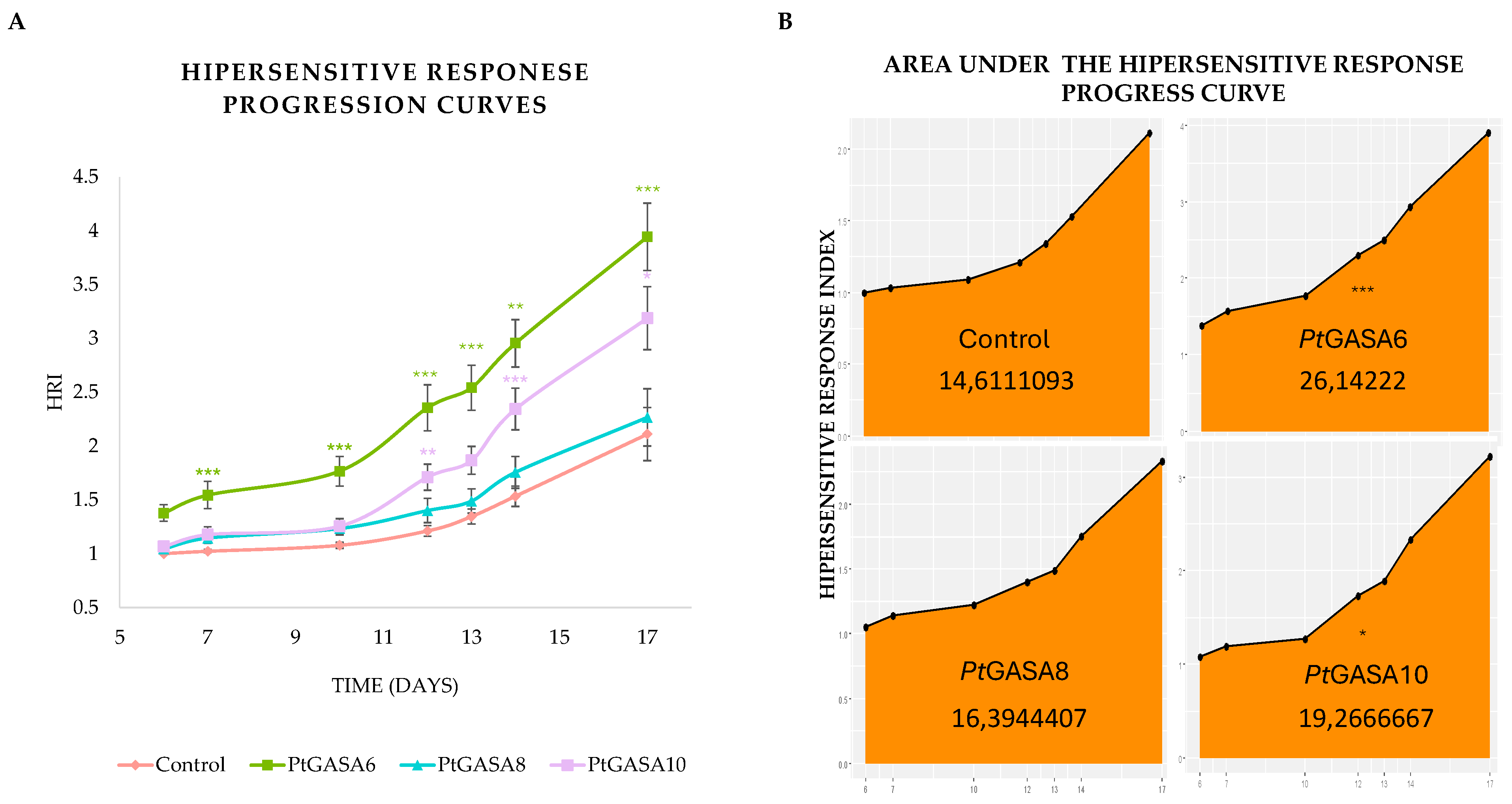
2.7. Overexpression of PtGASA6 and 10 Increased Hypersensitive Response (HR) After Leaf Infiltration with Xanthomonas citri
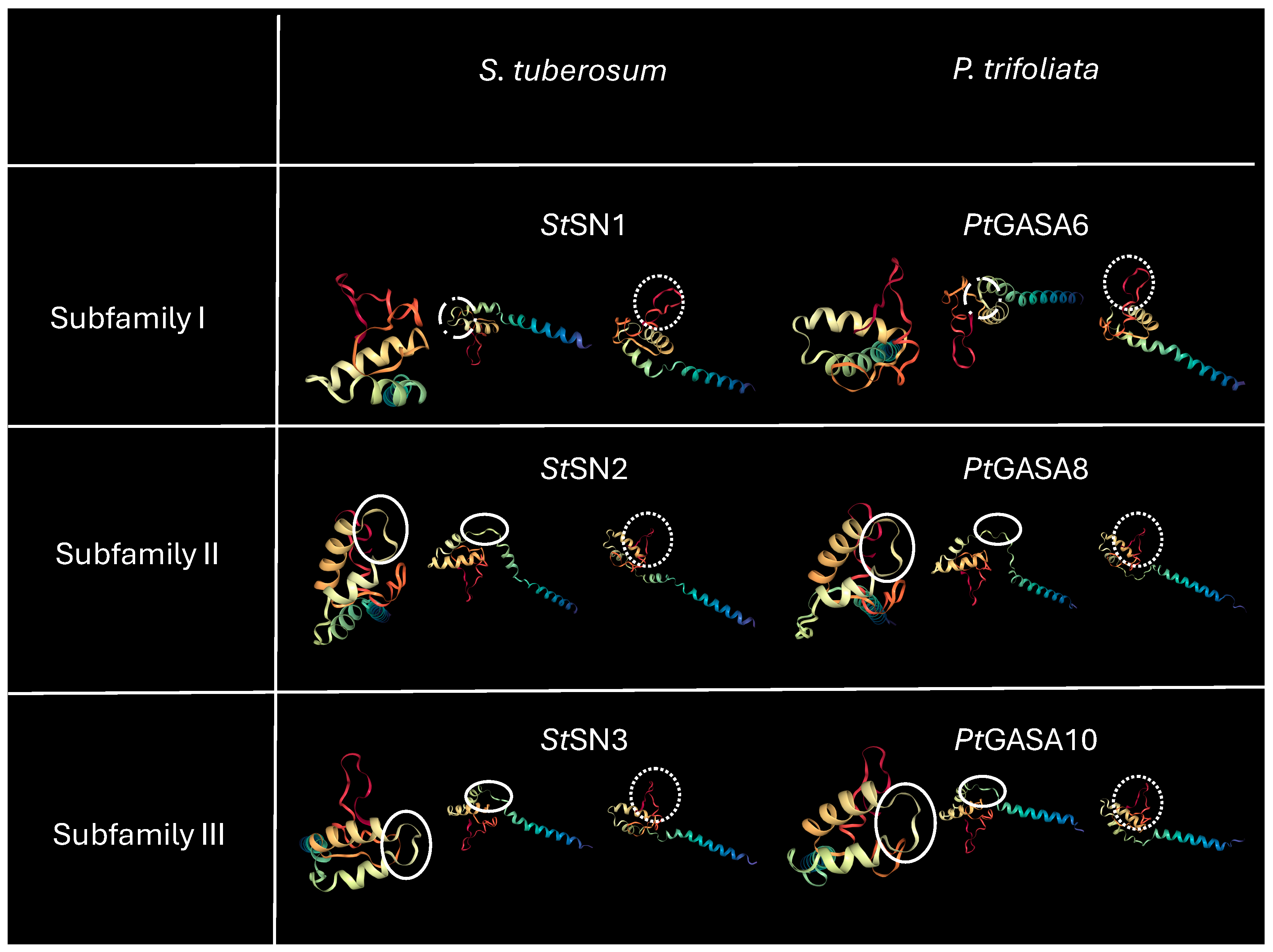
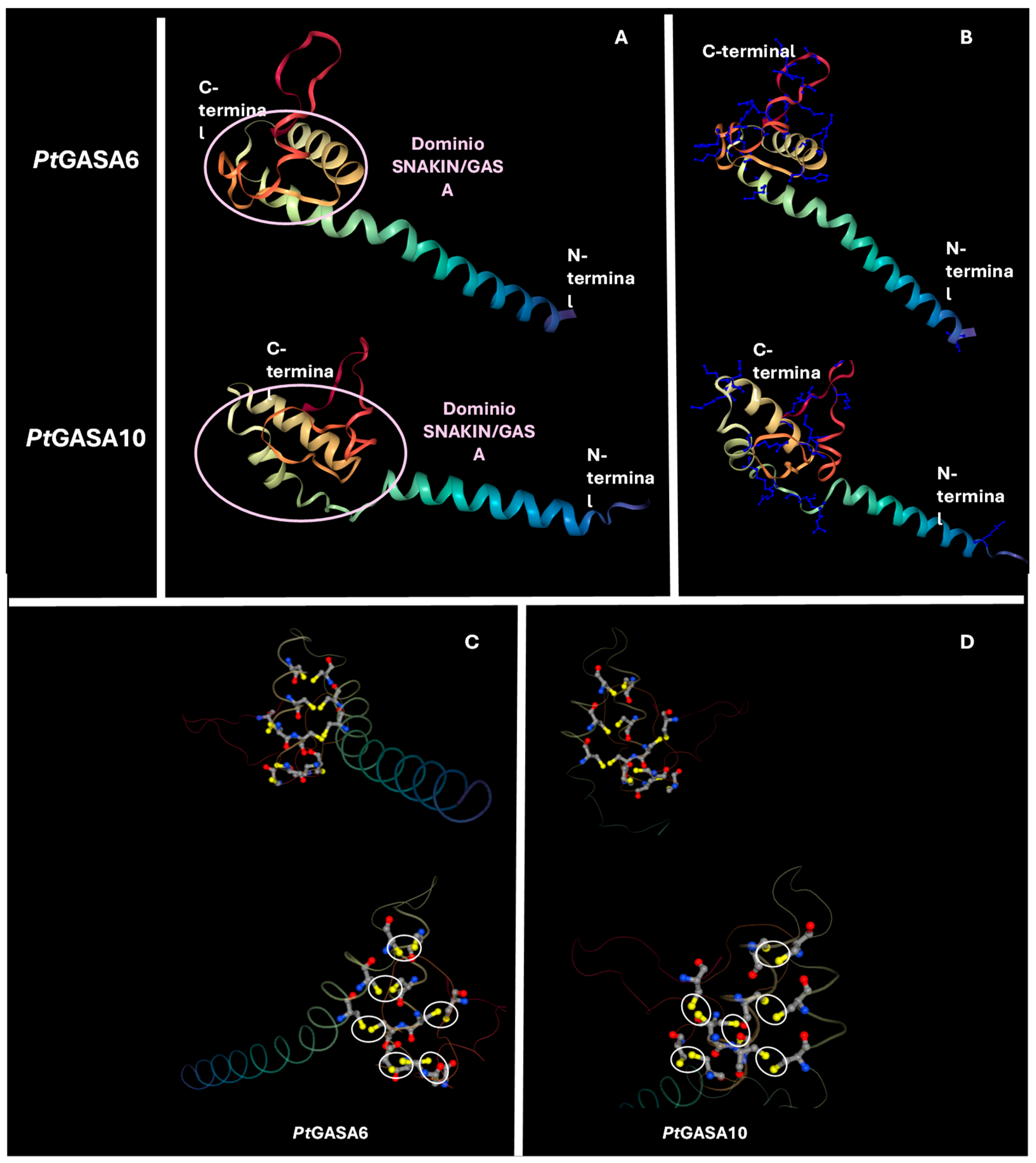
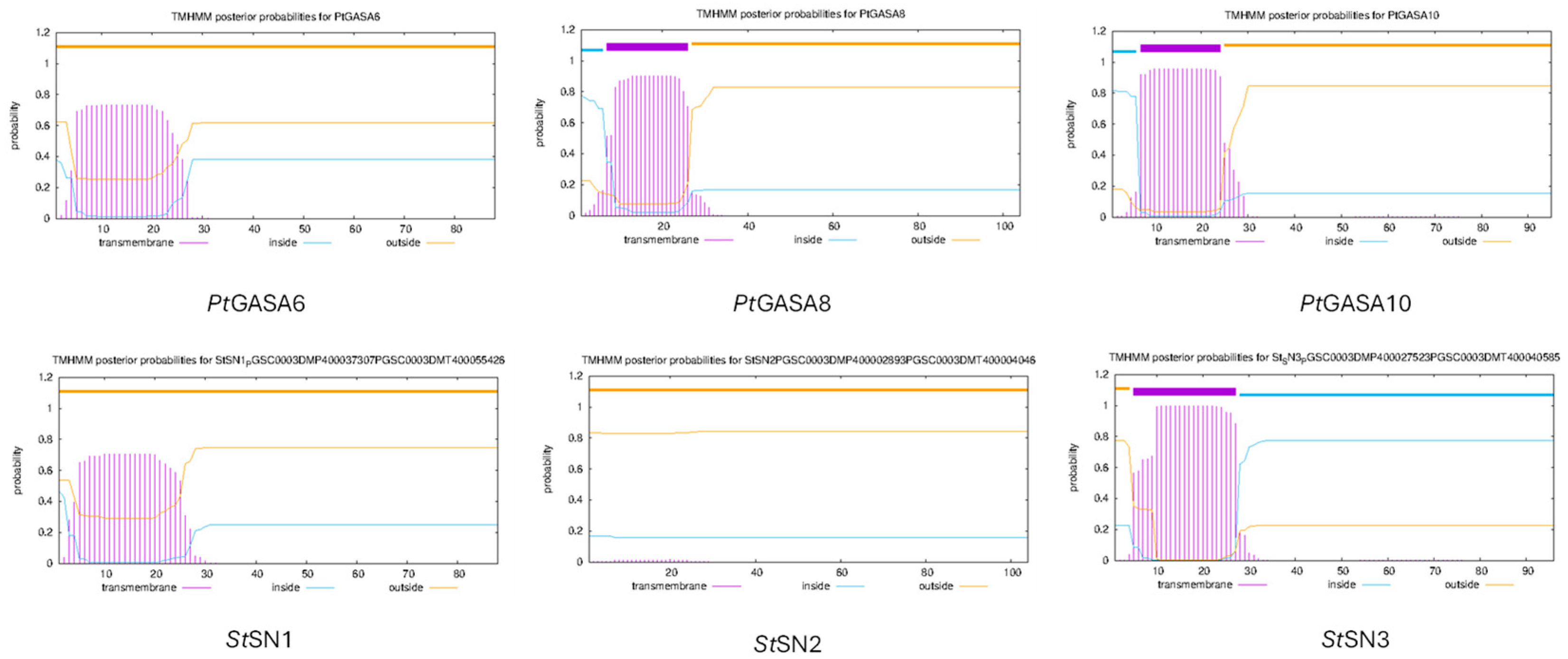
2.8. Structural and Funtional Comparison Between Predicted Tridimensional Structures of Citrus GASA Proteins Versus Reference SNAKIN Proteins from Solanum tuberosum
3. Discussion
4. Materials and Methods
4.1. Plant Material and Bacterial Strains
4.2. Identification of GASA Genes/Allelic Variants, PCR Cloning and DNA Sequencing
4.3. Structural and Phylogenetic Analyses of GASA Sequences
4.4. Expression Analysis by RT-qPCR
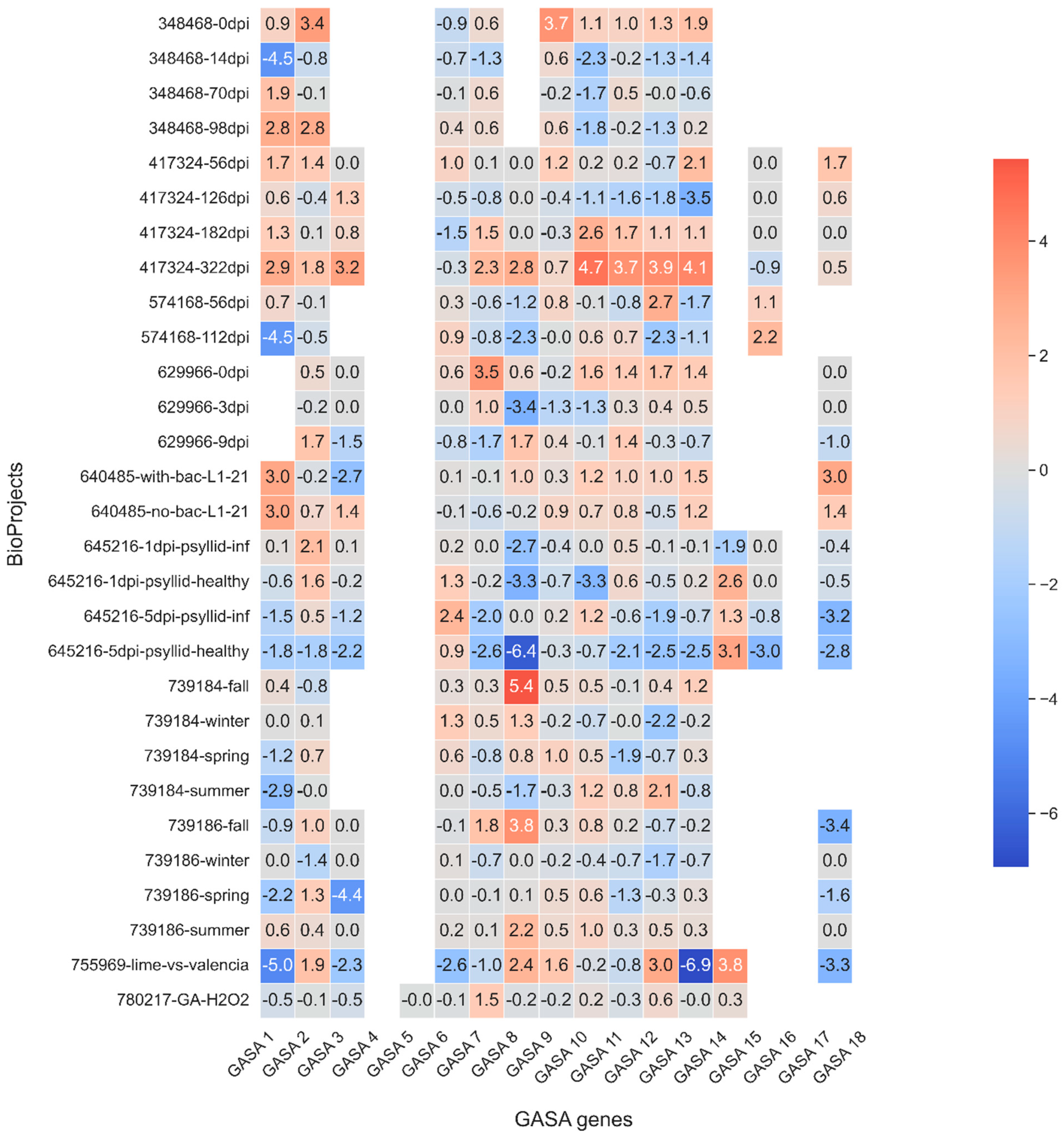
4.5. Transcriptomic Meta-Analysis of Citrus GASA Gene Expression in Public RNA-seq Datasets
4.6. Transient Overexpression and Infection Assays in N. benthamiana
4.7. In Silico Protein Structure Prediction and Subcellular Targeting
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AMP | Anti-Microbial Peptide |
| AUDPC | area under the disease progress curve |
| AUHRPC | area under the hypersensitive response progress curve |
| DI | Disease Index |
| GASA | Gibberellic Acid Stimulated in Arabidopsis |
| HR | Hypersensitive Response |
References
- Nahirñak, V.; Rivarola, M.; de Urreta, M.G.; Paniego, N.; Hopp, H.E.; Almasia, N.I.; Vazquez-Rovere, C. Genome-Wide Analysis of the Snakin/GASA Gene Family in Solanum Tuberosum Cv. Kennebec. American Journal of Potato Research 2016, 93, 172–188. [Google Scholar] [CrossRef]
- Raveh, E.; Goldenberg, L.; Porat, R.; Carmi, N.; Gentile, A.; La Malfa, S. Conventional Breeding of Cultivated Citrus Varieties. The citrus genome 2020, 33–48. [Google Scholar]
- Palacios, J. Citricultura; Tucumán, Argentina, 2005; ISBN 987-43-8326-7.
- Wu, T.; Cheng, C.; Zhong, Y.; Lv, Y.; Ma, Y.; Zhong, G. Molecular Characterization of the Gibberellin-Stimulated Transcript of GASA4 in Citrus. Plant Growth Regul 2020, 91, 89–99. [Google Scholar] [CrossRef]
- Xu, Q.; Chen, L.L.; Ruan, X.; Chen, D.; Zhu, A.; Chen, C.; Bertrand, D.; Jiao, W.B.; Hao, B.H.; Lyon, M.P.; et al. The Draft Genome of Sweet Orange (Citrus Sinensis). Nat Genet 2013, 45, 59–66. [Google Scholar] [CrossRef]
- Xu, Q.; Roose, M.L. Citrus Genomes: From Sequence Variations to Epigenetic Modifications. The Citrus genome 2020, 141–165. [Google Scholar]
- Nakandala, U.; Furtado, A.; Henry, R.J.; Jester B, U.T.J.D. Citrus Genomes: Past, Present and Future. Hortic Res 2025, 186, uhaf033. [Google Scholar] [CrossRef] [PubMed]
- Voorrips, R.E. MapChart: Software for the Graphical Presentation of Linkage Maps and QTLs. Journal of Heredity 2002, 93, 77–78. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The Neighbor-Joining Method: A New Method for Reconstructing Phylogenetic Trees. Mol Biol Evol 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Nahirñak, V.; Almasia, N.I.; Lia, V.V.; Hopp, H.E.; Vazquez Rovere, C. Unveiling the Defensive Role of Snakin-3, a Member of the Subfamily III of Snakin/GASA Peptides in Potatoes. Plant Cell Rep 2024, 43, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Zhong, Y.; Chen, M.; Wu, B.; Wang, T.; Jiang, B.; Zhong, G. Analysis of CcGASA Family Members in Citrus Clementina (Hort. Ex Tan.) by a Genome-Wide Approach. BMC Plant Biol 2021, 21, 1–18. [Google Scholar] [CrossRef]
- Berrocal-Lobo, M.; Segura, A.; Moreno, M.; López, G.; García-Olmedo, F.; Molina, A. Snakin-2, an Antimicrobial Peptide from Potato Whose Gene Is Locally Induced by Wounding and Responds to Pathogen Infection. Plant Physiol 2002, 128, 951–961. [Google Scholar] [CrossRef]
- Gao, C.; Li, Z.; Zhang, H.; Li, C.; Sun, H.; Li, S.; Ma, N.; Qi, X.; Cui, Y.; Yang, P.; et al. Genome-Wide Identification and Characterization of the GASA Gene Family in Medicago Truncatula, and Expression Patterns under Abiotic Stress and Hormone Treatments. Plants 2024, 13. [Google Scholar] [CrossRef]
- Alves, M.N.; Lopes, S.A.; Raiol-Junior, L.L.; Wulff, N.A.; Girardi, E.A.; Ollitrault, P.; Peña, L. Resistance to ‘Candidatus Liberibacter Asiaticus,’ the Huanglongbing Associated Bacterium, in Sexually and/or Graft-Compatible Citrus Relatives. Front Plant Sci 2021, 11, 1–16. [Google Scholar] [CrossRef]
- Ramadugu, C.; Keremane, M.L.; Halbert, S.E.; Duan, Y.P.; Roose, M.L.; Stover, E.; Lee, R.F. Long-Term Field Evaluation Reveals Huanglongbing Resistance in Citrus Relatives. Plant Dis 2016, 100, 1858–1869. [Google Scholar] [CrossRef]
- Munir, S.; He, P.; Wu, Y.; He, P.; Khan, S.; Huang, M.; Cui, W.; He, P.; He, Y. Huanglongbing Control: Perhaps the End of the Beginning. Microb Ecol 2017, 76, 192–204. [Google Scholar] [CrossRef]
- Bouteraa, M.T.; Romdhane, W. Ben; Baazaoui, N.; Alfaifi, M.Y.; Chouaibi, Y.; Akacha, B. Ben; Hsouna, A. Ben; Kaˇ, M.; Garzoli, S.; Saad, R. Ben; et al. GASA Proteins: Review of Their Functions in Plant Environmental Stress Tolerance. 2023, 1–16. [CrossRef]
- Almasia, N.I.; Bazzini, A.A.; Hopp, H.E.; Vazquez-Rovere, C. Overexpression of Snakin-1 Gene Enhances Resistance to Rhizoctonia Solani and Erwinia Carotovora in Transgenic Potato Plants. Mol Plant Pathol 2008, 9, 329–338. [Google Scholar] [CrossRef]
- Conti, G.; Gardella, V.; Vandecaveye, M.A.; Gomez, C.A.; Joris, G.; Hauteville, C.; Burdyn, L.; Almasia, N.I.; Nahirñak, V.; Vazquez-Rovere, C.; et al. Transgenic Citrange Troyer Rootstocks Overexpressing Antimicrobial Potato Snakin-1 Show Reduced Citrus Canker Disease Symptoms. J Biotechnol 2020, 324, 99–102. [Google Scholar] [CrossRef] [PubMed]
- Darqui, F.S.; Radonic, L.M.; Trotz, P.M.; López, N.; Vázquez Rovere, C.; Hopp, H.E.; López Bilbao, M. Potato Snakin-1 Gene Enhances Tolerance to Rhizoctonia Solani and Sclerotinia Sclerotiorum in Transgenic Lettuce Plants. J Biotechnol 2018, 283, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Das, K.; Datta, K.; Sarkar, S.N.; Datta, S.K. Expression of Antimicrobial Peptide Snakin-1 Confers Effective Protection in Rice against Sheath Blight Pathogen, Rhizoctonia Solani. Plant Biotechnol Rep 2021, 15, 39–54. [Google Scholar] [CrossRef]
- Segura, A.; Moreno, M.; Madueño, F.; Molina, A.; García-Olmedo, F. Snakin-1, a Peptide from Potato That Is Active against Plant Pathogens. Molecular Plant-Microbe Interactions 1999, 12, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Weber, K.C.; Mahmoud, L.M.; Stanton, D.; Welker, S.; Qiu, W.; Grosser, J.W.; Levy, A.; Dutt, M. Insights into the Mechanism of Huanglongbing Tolerance in the Australian Finger Lime (Citrus Australasica). Front Plant Sci 2022, 13, 1–23. [Google Scholar] [CrossRef]
- Younas, M.; Wang, C.; Hassan, M.F.; Li, W.; Zheng, Z.; Bin, Y.; Wei, T.; Li, Y. Comprehensive Transcriptomic Profiling of Citrus Australasica Unveils Antimicrobial Peptides and Immune Pathways for Huanglongbing Tolerance. J Agric Food Chem 2025. [Google Scholar] [CrossRef] [PubMed]
- Chin, E.L.; Ramsey, J.; Saha, S.; Mishchuk, D.; Chavez, J.; Howe, K.; Zhong, X.; Flores-Gonzalez, M.; Mitrovic, E.; Polek, M.; et al. Multi-Omics Comparison Reveals Landscape of Citrus Limon and Citrus Sinensis Response to ‘ Candidatus Liberibacter Asiaticus’. PhytoFrontiersTM 2021, 1, 76–84. [Google Scholar] [CrossRef]
- Wei, X.; Mira, A.; Yu, Q.; Gmitter, F.G. The Mechanism of Citrus Host Defense Response Repression at Early Stages of Infection by Feeding of Diaphorina Citri Transmitting Candidatus Liberibacter Asiaticus. Front Plant Sci 2021, 12, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.Y.; Niu, D.D.; Kund, G.; Jones, M.; Albrecht, U.; Nguyen, L.; Bui, C.; Ramadugu, C.; Bowman, K.D.; Trumble, J.; et al. Identification of Citrus Immune Regulators Involved in Defence against Huanglongbing Using a New Functional Screening System. Plant Biotechnol J 2021, 19, 757–766. [Google Scholar] [CrossRef]
- Almasia, N.I.; Nahirñak, V.; Hopp, H.E.; Vazquez-Rovere, C. Potato Snakin-1: An Antimicrobial Player of the Trade-off between Host Defense and Development. Plant Cell Rep 2020, 39, 839–849. [Google Scholar] [CrossRef] [PubMed]
- Nahirñak, V.; Almasia, N.I.; Hopp, H.E.; Vazquez-Rovere, C. Snakin/GASA Proteins. Plant Signal Behav 2012, 7, 1004–1008. [Google Scholar] [CrossRef]
- Steinegger, M.; Söding, J. MMseqs2 Enables Sensitive Protein Sequence Searching for the Analysis of Massive Data Sets. Nat Biotechnol 2017, 35, 1026–1028. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: Making Protein Folding Accessible to All. Nat Methods 2022, 19, 679–682. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.; Lee, S.; Levy Karin, E.; Kim, H.; Moriwaki, Y.; Ovchinnikov, S.; Steinegger, M.; Mirdita, M. Easy and Accurate Protein Structure Prediction Using ColabFold. Nat Protoc 2024, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Rose, A.; Hildebrand, P. NGL Viewer: A Web Application for Molecular Visualization. Nucleic Acids Res 2015, 43. [Google Scholar] [CrossRef]
- Yeung, H.; Squire, C.J.; Yosaatmadja, Y.; Panjikar, S.; López, G.; Molina, A.; Baker, E.N.; Harris, P.W.R.; Brimble, M.A. Radiation Damage and Racemic Protein Crystallography Reveal the Unique Structure of the GASA/Snakin Protein Superfamily. Angewandte Chemie 2016, 128, 8062–8065. [Google Scholar] [CrossRef]
- Krogh, A.; Larsson, B.; Von Heijne, G.; Sonnhammer, E.L.L. Predicting Transmembrane Protein Topology with a Hidden Markov Model: Application to Complete Genomes. J Mol Biol 2001, 305, 567–580. [Google Scholar] [CrossRef]
- Liu, H.; Wang, X.; Liu, S.; Huang, Y.; Guo, Y.X.; Xie, W.Z.; Liu, H.; Tahir ul Qamar, M.; Xu, Q.; Chen, L.L. Citrus Pan-Genome to Breeding Database (CPBD): A Comprehensive Genome Database for Citrus Breeding. Mol Plant 2022, 15, 1503–1505. [Google Scholar] [CrossRef] [PubMed]
- Geer, L.Y.; Marchler-Bauer, A.; Geer, R.C.; Han, L.; He, J.; He, S.; Liu, C.; Shi, W.; Bryant, S.H. The NCBI Biosystems Database. Nucleic Acids Res 2010, 38, D492–D496. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J.; Doyle, J.L. A Rapid DNA Isolation Procedure for Small Quantities of Fresh Leaf Tissue. Phytochemical bulletin 1987. [Google Scholar]
- Sievers, F.; Higgins, D.G. Clustal Omega for Making Accurate Alignments of Many Protein Sequences. Protein Science 2018, 27, 135–145. [Google Scholar] [CrossRef]
- Drozdetskiy, A.; Cole, C.; Procter, J.; Barton, G.J. JPred4: A Protein Secondary Structure Prediction Server. Nucleic Acids Res 2015, 43, W389–W394. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Zuckerkandl, E.; Pauling, L. Evolutionary Divergence and Convergence, in Proteins.
- Rombauts, S.; Déhais, P.; Van Montagu, M.; Rouzé, P. PlantCARE, a Plant Cis-Acting Regulatory Element Database. Nucleic Acids Res 1999, 27, 295–296. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y. TBtools-II: A “One for All, All for One” Bioinformatics Platform for Biological Big-Data Mining. Mol Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef]
- Mafra, V.; Kubo, K.S.; Alves-Ferreira, M.; Ribeiro-Alves, M.; Stuart, R.M.; Boava, L.P.; Rodrigues, C.M.; Machado, M.A. Reference Genes for Accurate Transcript Normalization in Citrus Genotypes under Different Experimental Conditions. PLoS One 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A New Mathematical Model for Relative Quantification in Real-Time RT-PCR. Nucleic Acids Res 2001, 29. [Google Scholar] [CrossRef]
- Di Rienzo, J.A. FgStatistics, Statistical Software for the Analysis of Experiments of Functional Genomics. 2009.
- Machado, R.; Moschen, S.; Conti, G.; González, S.; Rivarola, M.; Gómez, C.; Hopp, E.; Fernández, P. Exploring the Genetic Networks of Hlb Tolerance in Citrus : Insights Across Species and Tissues Exploring the Genetic Networks of HLB Tolerance in Citrus : Insights Across Species and Tissues. 2025, 0–24. [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. FeatureCounts: An Efficient General Purpose Program for Assigning Sequence Reads to Genomic Features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol 2014, 15, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Schandry, N. A Practical Guide to Visualization and Statistical Analysis of R. Solanacearum Infection Data Using R. Front Plant Sci 2017, 8. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Hallgren, J.; Tsirigos, K.D.; Pedersen, M.D.; Almagro Armenteros, J.J.; Marcatili, P.; Nielsen, H.; Krogh, A.; Winther, O. DeepTMHMM Predicts Alpha and Beta Transmembrane Proteins Using Deep Neural Networks. biorxiv 2022, 2022–2024. [Google Scholar]
- Bekier, F.N. Análisis de La Inmunidad Específica y de Amplio Espectro: Caracterización y Evaluación Funcional de Genes Candidatos Para La Obtención de Cultivos Resistentes a Bacterias Fitopatógenas. Doctoral thesis, Universidad de Buenos Aires, Facultad de Ciencias Exactas y Naturales: Buenos Aires, Argentina, 2025.
- Arce-Leal, Á.P.; Bautista, R.; Rodríguez-Negrete, E.A.; Manzanilla-Ramírez, M.Á.; Velázquez-Monreal, J.J.; Santos-Cervantes, M.E.; Méndez-Lozano, J.; Beuzón, C.R.; Bejarano, E.R.; Castillo, A.G.; et al. Gene Expression Profile of Mexican Lime (Citrus Aurantifolia) Trees in Response to Huanglongbing Disease Caused by Candidatus Liberibacter Asiaticus. Microorganisms 2020, 8. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Li, T.; Cui, L.; Wang, N.; Huang, G.; Li, R. OrangeExpDB: An Integrative Gene Expression Database for Citrus Spp. BMC Genomics 2024, 25, 1–7. [Google Scholar] [CrossRef]

| Sequence Name | Gene ID | Exons | CDS (bp) |
Protein (aa) | |
|---|---|---|---|---|---|
| Subfamily I | CsGASA6 | LOC102614811 | 2 | 267 | 88 |
| CpxPtGASA6 | OP728331 | 2 | 267 | 88 | |
| ClGASA6 | OP728332 | 2 | 267 | 88 | |
| CjGASA6 | OP728333 | 2 | 267 | 88 | |
| CliaGASA6 | OP728334 | 2 | 267 | 88 | |
| CaGASA6 | OP728335 | 2 | 267 | 88 | |
| PtGASA6 | OP728336 | 2 | 267 | 88 | |
| CsGASA13 | LOC102619143 | 2 | 267 | 88 | |
| CpxPtGASA13 | OP791894 | 2 | 267 | 88 | |
| ClGASA13 | OP791895 | 2 | 267 | 88 | |
| CjGASA13 | OP791896 | 2 | 267 | 88 | |
| CliaGASA13 | OP791897 | 2 | 267 | 88 | |
| CwGASA13 | OP791898 | 2 | 267 | 88 | |
| CaGASA13 | OP791899 | 2 | 267 | 88 | |
| PtGASA13 | OP791900 | 2 | 267 | 88 | |
| Subfamily II | CsGASA8 | LOC102628220 | 3 | 315 | 104 |
| CpxPtGASA8 | OP947003 | 3 | 315 | 104 | |
| ClGASA8 | OP947004 | 3 | 315 | 104 | |
| CjGASA8 | OP947005 | 3 | 315 | 104 | |
| CliaGASA8 | OP947006 | 3 | 315 | 104 | |
| CwGASA8 | OP947007 | 3 | 315 | 104 | |
| CaGASA8 | OP947008 | 3 | 315 | 104 | |
| PtGASA8 | OP947009 | 3 | 315 | 104 | |
| CsGASA9 | LOC102627925 | 3 | 342 | 113 | |
| ClGASA9 | OP946990 | 3 | 342 | 113 | |
| PtGASA9 | OP946991 | 3 | 342 | 113 | |
| CsGASA9-like | LOC102628717 | 3 | 303 | 100 | |
| CpxPtGASA9-like | OQ053283 | 3 | 303 | 100 | |
| ClGASA9-like | OQ053284 | 3 | 303 | 100 | |
| CjGASA9-like | OQ053285 | 3 | 226 | 73 (partial CDS) | |
| CliaGASA9-like | OQ053286 | 3 | 226 | 73 (partial CDS) | |
| CwGASA9-like | OQ053287 | 3 | 303 | 100 | |
| CaGASA9-like | OQ053288 | 3 | 303 | 100 | |
| PtGASA9-like | OQ053289 | 3 | 306 | 101 | |
| CsGASA2 | LOC102628910 | 4 | 351 | 116 | |
| CpxPtGASA2 | OP946981 | 2 | 213 | 65 (partial CDS) | |
| ClGASA2 | OP946982 | 4 | 213 | 65 (partial CDS) | |
| PtGASA2 | OP946983 | 351 | 116 | ||
| CsGASA11 | LOC102631482 | 4 | 504 | 167 | |
| CpxPtGASA11 | OP946975 | 2 | 408 | 125 (partial CDS) | |
| ClGASA11 | OP946976 | 2 | 396 | 130 (partial CDS) | |
| CliaGASA11 | OP946977 | 2 | 396 | 135 (partial CDS) | |
| CwGASA11 | OP946978 | 2 | 408 | 135 (partial CDS) | |
| CaGASA11 | OP946979 | 2 | 396 | 130 (partial CDS) | |
| PtGASA11 | OP946980 | 4 | 504 | 167 | |
| CsGASA7 | LOC102625142 | 4 | 432 | 143 | |
| CpxPtGASA7 | OP947000 | 4 | 345 | 114 | |
| ClGASA7 | OP947001 | 4 | 345 | 114 | |
| PtGASA7 | OP947002 | 4 | 345 | 114 | |
| CsGASA5 | LOC102625142 | 3 | 324 | 113 | |
| CpxPtGASA5 | OP946984 | 3 | 324 | 113 | |
| ClGASA5 | OP946985 | 3 | 327 | 114 | |
| CjGASA5 | OP946986 | 3 | 324 | 113 | |
| CliaGASA5 | OP946987 | 3 | 324 | 113 | |
| CaGASA5 | OP946988 | 3 | 324 | 113 | |
| PtGASA5 | OP946989 | 3 | 324 | 113 | |
| Subfamily III | CsGASA15 | LOC102625588 | 3 | 286 | 95 |
| ClGASA15 | OP946995 | 3 | 285 | 95 | |
| PtGASA15 | OP946996 | 3 | 285 | 95 | |
| CsGASA1 | LOC102629636 | 4 | 342 | 113 | |
| CpxPtGASA1 | OQ053290 | 4 | 342 | 113 | |
| ClGASA1 | OQ053291 | 4 | 342 | 113 | |
| PtGASA1 | OQ053292 | 4 | 348 | 115 | |
| CsGASA10 | LOC102618255 | 3 | 288 | 95 | |
| CpxPtGASA10 | OP830506 | 3 | 288 | 95 | |
| ClGASA10 | OP830507 | 3 | 288 | 95 | |
| PtGASA10 | OP830508 | 3 | 288 | 95 | |
| CsGASA12 | LOC102611788 | 4 | 324 | 107 | |
| CpxPtGASA12 | OQ053280 | 4 | 321 | 106 | |
| ClGASA12 | OQ053281 | 4 | 324 | 107 | |
| PtGASA12 | OQ053282 | 4 | 321 | 106 | |
| CsGASA3 | LOC102626694 | 3 | 312 | 103 | |
| CpxPtGASA3 | OP946997 | 3 | 312 | 103 | |
| ClGASA3 | OP946998 | 3 | 312 | 103 | |
| PtGASA3 | OP946999 | 2 | 312 | 103 | |
| CsGASA14 | LOC102625321 | 3 | 351 | 116 | |
| CpxPtGASA14 | OP946992 | 3 | 286 | 95 | |
| ClGASA14 | OP946993 | 3 | 285 | 95 | |
| PtGASA14 | OP946994 | 3 | 285 | 95 | |
| CsGASA18 | LOC102630614 | 4 | 357 | 118 | |
| CpxPtGASA18 | OP830509 | 4 | 357 | 118 | |
| ClGASA18 | OP830510 | 4 | 357 | 118 | |
| PtGASA18 | OP830511 | 4 | 357 | 118 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




