Introduction
Human papillomavirus (HPV) is one of the most common sexually transmitted viruses worldwide and responsible for 7.7% of cancers in developing countries, mainly cervical cancer, causing more than 75,000 new cases and 50,000 deaths yearly in sub-Saharan Africa [
1,
2]. Infection with oncogenic types of HPV is the causative factor of nearly all (>99%) cases of cervical cancer [
1,
3]. Fifteen genotypes (including HPV16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, 68, 73, and 82) are classified as oncogenic types of HPV, well known as high-risk HPV (hrHPV), and three genotypes (HPV26, 53, and 66) are considered probably carcinogenic [
4,
5].
Mozambique is among the countries with the highest burden of HPV infection, HIV and cervical cancer. Every year, more than 5300 cervical cancer new cases and more than 3800 deaths are registered [
6], affecting particularly women living with HIV/AIDS [
7,
8]. HPV infection is common among Mozambican women, with data indicating considerable prevalence rates of hrHPV (HPV16, 18, 31, 33, 35, 45, 51, 52, and 58), varying from 26.3% to 75.9%, according to studies [
9,
10,
11,
12,
13].
After the WHO Global strategy for the elimination of cervical cancer was launched [
14], hrHPV-based tests have been increasingly adopted, as primary or co-test for cervical cancer screening, and for monitoring of the effectiveness of HPV vaccination [
15,
16]. The key feature of hrHPV-based tests for cervical cancer screening is their highly sensitivity and accuracy for screening women at risk for cervical lesions, as well as for detecting high-grade cervical intraepithelial neoplasia (CIN) and cervical cancer (CIN2+), particularly in women above 30 years of age [
16,
17,
18].
In Mozambique, the National Cancer Control Program – Ministry of Health has recently introduced (in Maputo) the pilot implementation of hrHPV testing in primary cervical screening to comply with the WHO guidelines. Although there have been many pilot projects and studies using several HPV tests [
12,
13,
19], the assay actually in use within the government health-care system is the recently developed Cobas 5800/6800/8800 HPV-DNA, for use on the Cobas 5800/6800/8800 systems [
20]. The Cobas 4800 HPV-DNA test which has been successfully deployed for primary HPV-based screening [
21,
22], was introduced for hrHPV screening in a cervical screening service, at the DREAM program, Sant’Egídio [
12]. This test is for use on the Cobas 4800 system, an automated system that has been a reference standard for HPV screening programs due to its high sensitivity, reliability and ability to predict CIN2+ or worse [
23,
24,
25].
Guidelines recommend that, in order to enable testing in large centralized laboratories and ensure diagnostic accuracy, clinically validated HPV assays should be used [
14,
17]. The aim of this study was to evaluate the performance concordance of Cobas 4800 and Cobas 5800 tests with Cobas 6800 test for the detection and genotyping of high-risk HPV, using an interlaboratory comparative approach. Yet, there is no information on the performance concordance of these tests; or either on their clinically validation in Mozambican context.
Materials and Methods
Study Design, Population and Ethical Considerations
This is a cross-sectional retrospective study, under the scope of activities of laboratory quality management in 2023 and 2024, of two molecular laboratories (DREAM Sant’Egidio laboratory and laboratory of Mavalane General Hospital), in Maputo, Mozambique. The DREAM Laboratory is accredited by the Portuguese Institute for Accreditation (IPAC) in accordance with ISO 15189 standards. This study used samples form voluntary women aged > 18 years old, who were recruited at DREAM health center in Zimpeto, between July 2021 to May 2023, under the scope of a larger hrHPV screening study (HPV-ISI) that evaluated the feasibility of high-risk HPV testing as primary test compared to visual inspection with acetic acid (VIA) approach [
12]; and further within a routine cervical cancer screening program. The HPV-ISI study was approved by the Mozambican national ethical committee (ref. 688/CNBS/20; amendment nº176/CNBS/22). Furthermore, due to the need to use the collected data for academic purposes, additional approval was obtained from the Institutional Review Board of the Eduardo Mondlane University – Faculty of Medicina & Maputo Central Hospital, Mozambique (ref. CIBSFM&HCM/01/2024). Informed written consents were obtained from all the participants, and all the samples and data were anonymized.
Samples and HPV Testing
A random subset of 124 samples was selected from a larger cohort of 1,323 previously tested for hrHPV-DNA at the DREAM Sant’Egidio laboratory using the Cobas 4800 system (Roche Diagnostics), from 2021-2023 [
12] and Cobas 5800 in 2023Only samples with valid hrHPV-DNA test results on both Cobas 4800 and Cobas 5800 were selected and included in the subset for the interlaboratory evaluation by subsequent analysis on Cobas 6800. Limited laboratory test resources were the primary reason for the choice of the sample size. Samples without age data and those from participants aged under 18 years were excluded.
The samples were collected by a trained health professional using cervical brush into a 20 mL liquid-based cytology media (Roche Cell Collection Medium), then followed by hrHPV DNA test using Cobas 4800/5800 assays. After testing, samples were stored at –20°C, until the until sent to the laboratory of Mavalane General Hospital, for a second/interlaboratory testing, using the Cobas 6800 HPV assay (Roche Diagnostics/Molecular Systems). The Cobas 4800/5800/6800 tests are a real-time PCR based assays targeting HR-HPV L1 gene and human β-globin gene as an internal control. All three tests allow the detection of 14 hrHPV types, genotyping HPV16 and HPV18 and reporting other 12HR-HPVs (including HPV types 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, 66, and 68) as a “pooled” result in the same channel. The testing carried out according to manufacturer’s instructions [
20,
26]. If internal control was not amplified, samples were referred as invalid.
Statistical Analysis
Statistical analysis was performed using SPSS 30 software (IBM Corp., Armonk, New York, USA) and GraphPad QuickCalcs (available on:
https://www.graphpad.com/quickcalcs/). Quantitative variables were presented as median with interquartile range (IQR), and categorical variables were summarized with the number and percentage of subjects in each category. A 2-tailed P value of <.05 was considered significant.
McNemar’s test and Cohen’s kappa (κ) statistic were calculated to assess, respectively, differences in the proportion of positive samples and the level of agreement between the Cobas tests for detecting the 14 hrHPV types. The strength of agreement was classified according to the criteria described by Landis & Koch [
27]. Simple agreement was calculated by adding the agreement of the positive and negative results for both techniques; complete and partial discordant results were evaluated.
Results
Of the 124 samples selected and included in the subset for the interlaboratory evaluation using Cobas 6800 system at the laboratory of Mavalane General Hospital, 94 samples (47 hrHPV-positive and 47 hrHPV-negative) had been previously tested with the Cobas 4800, while 30 samples (15 hrHPV-positive and 15 hrHPV-negative) had been tested with the Cobas 5800.
The median (IQR) age of the sampled participants was 42 (35–48) years, of whom the majority (61.3%) belonged to women aged 30-45 years, followed 31.5% for >45 years, and 4.8% for 18-30 years. Information on age were missing for 3 (2.4%) participants. The median interval between sample collection and testing was 5 days (IQR: 2–6 days) for Cobas 4800 and 10 days (IQR: 8–12 days) for Cobas 5800 at the DREAM Sant’Egidio laboratory, whereas for the testing with Cobas 6800 at the Mavalane General Hospital laboratory, took 76 days (IQR: 59–202 days) for samples tested with Cobas 4800, and 17.5 days (IQR: 16–19 days) for those tested with Cobas 5800.
All 124 samples were tested with the three Cobas HPV DNA tests (4800, 5800 and 6800), and none of the samples failed for hrHPV testing. Among the 94 samples tested with Cobas 4800
versus Cobas 6800, the prevalence of HPV16 alone, HPV18 alone, HPV16 with Other hrHPV, HPV18 with Other hrHPV, and Other hrHPV types was not significantly different between the 2 tests: 11% versus 12% (
P = 0.82), 9.6% versus 8.5% (
P = 0.80), 8.5% versus 8.5% (
P = 1.00), 9.6% versus 10.6% (
P = 0.81), and 10.6% versus 10.6% (
P = 1.00), respectively. A similar observation was found for the 30 samples tested with Cobas 5800 versus Cobas 6800, with prevalences of HPV16 with Other hrHPV, HPV18 with Other hrHPV, and Other hrHPV, accounting for 10%, 6.7%, and 50%, respectively, in both tests. HPV16 alone and HPV18 alone were not detected (
Table 1).
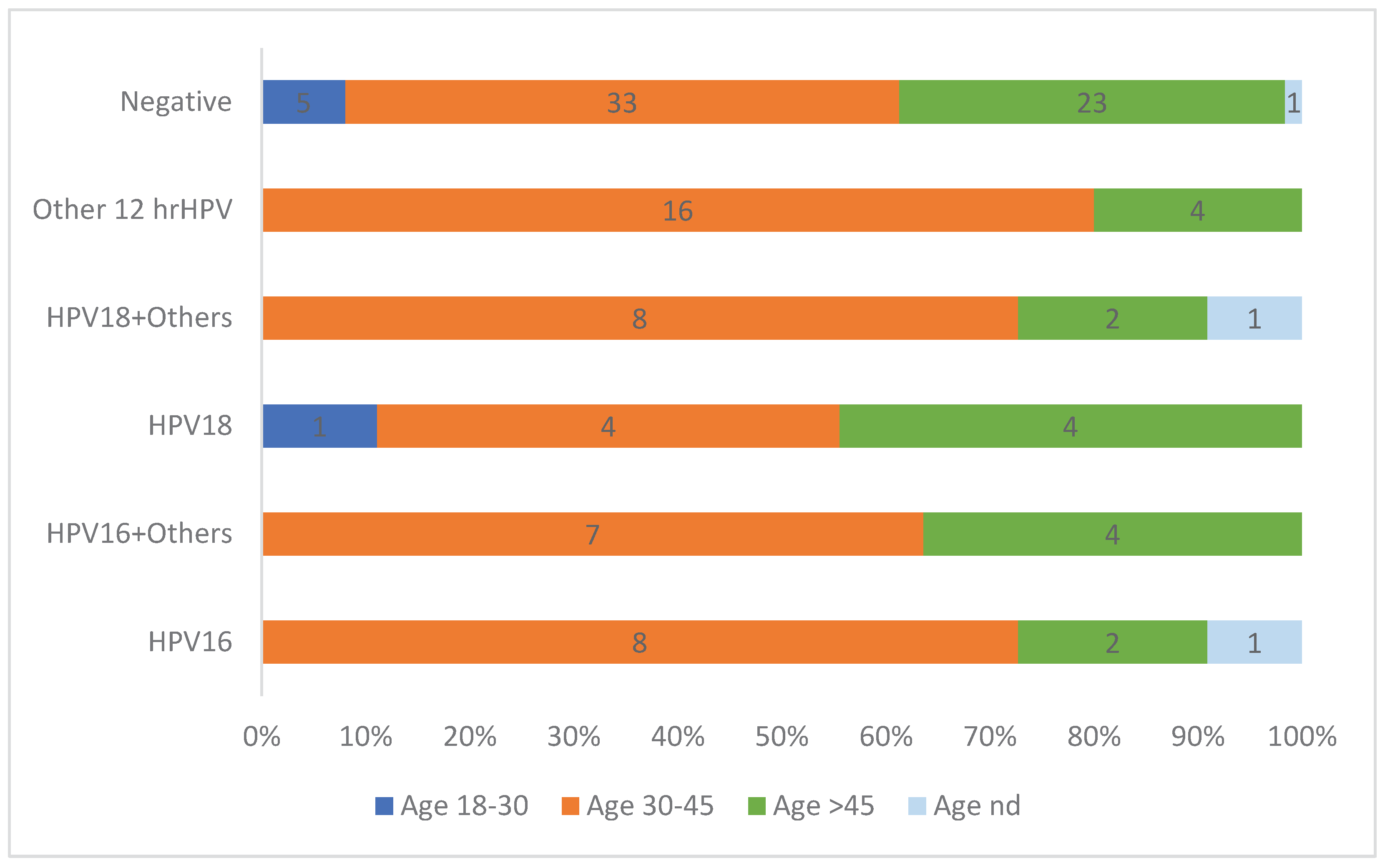
Figure 1 presents the overall distribution of hrHPV by age group, showing a considerable positivity rate among women aged 30-45 years.
Further statistical analyses for method comparison were also performed on all samples – separately for Cobas 4800 versus Cobas 6800, and Cobas 5800 versus Cobas 6800, with an overall concordance of detecting any hrHPV of 94.68% (agreement on 89/94 samples, Kappa= 0.89; 95% confidence interval: 0.80–0.98), and 100% (agreement on all 30 samples, Kappa= 1.00), respectively.
The concordance of the results between Cobas 4800
versus Cobas 6800 for detection of HPV16 alone, HPV18 alone, HPV16 with Other hrHPV, HPV18 with Other hrHPV, and Other hrHPV types was 98.9%, 98.9%, 100%, 98.9%, and 95.7%, respectively (
Table 2). Five samples were discordant: (
i) 1 for HPV16 positivity and 2 for Other hrHPV positivity were missed by Cobas 4800, and detected by Cobas 6800; and (
ii) 2 for Other hrHPV positivity, detected by Cobas 4800, were missed by Cobas 6800. Additionally, one sample was partially discordant: the Cobas 4800 detected only HPV18 while the Cobas 6800 detected HPV18 with Other hrHPV (
Table 3). The Other hrHPV was the most common type missed by the 2 methods (5/6 cases). In contrast, for Cobas 5800
versus Cobas 6800, all results were 100% concordant (absence of discordance) – same agreement index were observed for genotyping HPV16 + other hrHPV and HPV18 + other hrHPV (
Table 2).
The agreement (Cohen kappa) between Cobas 4800
versus Cobas 6800 for detection of HPV16 alone, HPV18 alone, HPV16 with Other hrHPV, HPV18 with Other hrHPV, and Other hrHPV types was 0.95 (CI: 0.85–1.00), 0.94 (CI: 0.81–1.00), 1.00 (CI: 1.00–1.00), 0.94 (CI: 0.83–1.00) and 0.78 (CI: 0.57–0.99), respectively. The performance to detect HPV-16 (alone/with Other hrHPV), HPV-18 (alone/with Other hrHPV), and Other hrHPV positivity were not significantly different between the 2 tests (McNemar test,
P > 0.05) (
Table 2). In total, the global performance to detect any of the 14 hrHPV types was not significantly different (McNemar test,
P = 1.00).
Discussion
This study aimed to evaluate, within an interlaboratory quality assessment process, the concordance of Cobas 4800 and Cobas 5800 tests with Cobas 6800 test in detecting and genotyping high-risk human papillomavirus (hrHPV) in 124 cervical samples (94 for Cobas 4800 versus Cobas 6800; and 30 for Cobas 5800 versus Cobas 6800), from women aged 18 years or older, screened for hrHPV in a cervical cancer screening service. To our knowledge, this is the first study evaluating the diagnostic performance of the Cobas tests within the recently introduced pilot implementation of hrHPV testing in primary cervical screening in Maputo, Mozambique.
The results demonstrate a high level of agreement between the Cobas 4800 and Cobas 5800 tests with the Cobas 6800 test across both laboratories, showing an overall concordance (95%, for Cobas 4800
versus Cobas 6800; and 100%, for Cobas 5800
versus Cobas 6800), which indicates an almost perfect agreement [
27,
28]. These findings support the interchangeability of the three platforms in clinical and epidemiological hrHPV screening and surveillance programs. It is widely accepted that, to ensure a robust and highly reliable test performance in clinical settings, the candidate assay must demonstrate inter-laboratory agreement with a lower confidence limit of no less than 87% [
17].
The distribution of hrHPV types—including HPV16, HPV18, and Other hrHPV types—was largely comparable between both methods, with no statistically significant differences observed in their detection frequencies. Notably, the detection concordance for HPV16 and HPV18, the two most oncogenic HPV genotypes, reached 98.9%, underscoring both platforms’ reliability in identifying these high-risk types. The complete agreement observed in cases with co-infection of HPV16 and Other hrHPV further reinforces the robustness of both assays in multi-type detection scenarios.
The discrepancies between the assays were minimal. Only five samples showed discordant results, with the Cobas 6800 slightly outperforming the Cobas 4800 by detecting three additional positive cases (one HPV16 and two Other hrHPV). Two samples were detected exclusively by the Cobas 4800, and one sample was partially discordant with differing co-infection profiles. The marginally higher detection of Other hrHPV by the Cobas 6800 could be attributed to its technological enhancements over the 4800 system, including analytical sensitivity and automation design, which allows for improved nucleic acid extraction and amplification [
17,
20,
29]. However, the difference was not statistically significant, and the kappa agreement for Other hrHPV was slightly lower compared to other genotype categories, suggesting a modest variation in detecting non-HPV16/18 genotypes, which are often present at lower viral loads [
30]. Similar discrepancies were also reported in other studies [
23,
24].
The age distribution of hrHPV positivity aligns with epidemiological patterns, where the highest burden of hrHPV was observed among women aged 30–45 years. This reinforces the importance of targeting this age group for primary HPV screening, especially in low- and middle-income settings where cervical cancer remains a leading cause of mortality among women [
31,
32].
One important procedural difference between the two laboratories involved in this study was the storage time between sample collection and testing. Samples processed at the Mavalane General Hospital laboratory had a significantly longer storage interval (median of 76 days for samples previously tested with Cobas 4800, and 17.5 days for those tested with Cobas 5800) compared to those at DREAM Sant’Egidio (median of 5 days for Cobas 4800 and 10 days for Cobas 5800). Despite this, there was no impact on assay performance, underscoring the stability of cervical specimens in liquid-based cytology collection media under appropriate storage conditions, as previously reported [
33,
34,
35].
Our findings support previous studies that have shown strong agreement between Cobas platforms [
20,
23,
24,
36], emphasizing their validity in clinical practice. Importantly, while the Cobas 4800 system is medium throughput (384 samples/day), the Cobas 5800 and Cobas 6800 offers improved workflow efficiency, higher throughput (528 and
>1500 samples/day, respectively), and reduced hands-on time, making it particularly suitable for large-scale screening programs and molecular testing for other viruses [
20,
26]. Similar conclusions were also reached by studies that clinically evaluated the performance according to the Meijer criteria, and observed high agreement (>98%) for the two assays, and that clinical sensitivity and specificity met the non-inferiority criteria defined to consider an HPV assay as suitable for large-scale laboratory use [
23,
24,
36].
This study has some limitations. The relatively small sample size, due to limited laboratory test resources, archival sample-based testing and the single-country setting may limit generalizability. Additionally, sequencing confirmation of discordant samples was not performed, which could have helped clarify the true-positive status. Nonetheless, the high concordance metrics and low number of discrepancies suggest these limitations did not significantly impact the conclusions.
Conclusion
Based on the results of this study, all three Cobas platforms – 4800, 5800, and 6800 – demonstrated high concordance in detecting and genotyping high-risk HPV in cervical samples. These findings support the suitability of each platform for use in routine hrHPV screening programs and underscore the Cobas 6800 as a reliable, high-throughput option particularly well-suited for scaling up cervical cancer prevention efforts.
Contribution of Authors
AS, TC and JS conceptualized and designed the study, acquired, analyzed and interpreted the data. All the authors drafted, revised, read, provided critical feedback and approved the manuscript.
Source of Funding
No external funding was received
Data Availability
The data that support the findings made in this study can be made available from the corresponding author, A.S., on request.
Acknowledgements
The co-authors played an invaluable role in completion of this work. Gratitude also goes to the DREAM program administration for allowing the use of the data, and the staff of the two laboratories (DREAM Sant’Egidio laboratory and laboratory of Mavalane General Hospital) who ensured quality data for this study.
Conflicts of Interest
No conflict of interest is declared
Disclaimer
The views expressed in this study are those of the authors and are not an official position of the laboratories administrative: DREAM Program, community of Sant’Egidio and Mavalane General Hospital.
Copyright Notice Agreement Acceptation
The authors agree to the Copyright Notice terms of the African Journal of Clinical and Experimental Microbiology (AJCEM).
References
- de Martel C, Plummer M, Vignat J, Franceschi S. Worldwide burden of cancer attributable to HPV by site, country and HPV type. Int J Cancer [Internet]. 2017 Aug 15;141(4):664–70. Available from: https://onlinelibrary.wiley.com/doi/10.1002/ijc.30716. [CrossRef]
- Mboumba Bouassa RS, Prazuck T, Lethu T, Jenabian MA, Meye JF, Bélec L. Cervical cancer in sub-Saharan Africa: a preventable noncommunicable disease. Expert Rev Anti Infect Ther. 2017 Jun 3;15(6):613–27. [CrossRef]
- Walboomers JMM, Jacobs M V., Manos MM, Bosch FX, Kummer JA, Shah K V., et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. Journal of Pathology. 1999;189(1):12–9.
- Muñoz-Bello JO, Carrillo-García A, Lizano M. Epidemiology and Molecular Biology of HPV Variants in Cervical Cancer: The State of the Art in Mexico. Vol. 23, International Journal of Molecular Sciences. MDPI; 2022. [CrossRef]
- Arbyn M, Tommasino M, Depuydt C, Dillner J. Are 20 human papillomavirus types causing cervical cancer? J Pathol. 2014 Dec 11;234(4):431–5. [CrossRef]
- Bruni L, Albero G, Serrano B, Mena M, Collado JJ, Gómez D, et al. ICO/IARC Information Centre on HPV and Cancer (HPV Information Centre). Human Papillomavirus and Related Diseases in Mozambique. Summary Report [Internet]. 2023 Mar. Available from: www.hpvcentre.net.
- Carrilho C, Fontes F, Tulsidás S, Lorenzoni C, Ferro J, Brandão M, et al. Cancer incidence in Mozambique in 2015–2016: data from the Maputo Central Hospital Cancer Registry. European Journal of Cancer Prevention. 2019 Jul;28(4):373–6. [CrossRef]
- Batman S, Rangeiro R, Monteiro E, Changule D, Daud S, Ribeiro M, et al. Expanding Cervical Cancer Screening in Mozambique: Challenges Associated With Diagnosing and Treating Cervical Cancer. JCO Glob Oncol [Internet]. 2023 Sep;(9). Available from: https://ascopubs.org/doi/10.1200/GO.23.00139. [CrossRef]
- Edna Omar V, Orvalho A, Nália I, Kaliff M, Lillsunde-Larsson G, Ramqvist T, et al. Human papillomavirus prevalence and genotype distribution among young women and men in Maputo city, Mozambique. BMJ Open. 2017 Jul 1;7(7). [CrossRef]
- Bule YP, Silva J, Carrilho C, Campos C, Sousa H, Tavares A, et al. Human papillomavirus prevalence and distribution in self-collected samples from female university students in Maputo. International Journal of Gynecology & Obstetrics. 2020 May 25;149(2):237–46. [CrossRef]
- Maueia C, Murahwa A, Manjate A, Andersson S, Sacarlal J, Kenga D, et al. Identification of the human papillomavirus genotypes, according to the human immunodeficiency virus status in a cohort of women from maputo, Mozambique. Viruses. 2022 Jan 1;14(1). [CrossRef]
- Sineque A, Catalao C, Ceffa S, Fonseca AM, Parruque F, Guidotti G, et al. Screening approaches for cervical cancer in Mozambique in HIV positive and negative women. European Journal of Cancer Prevention. 2023;32(5). [CrossRef]
- Salcedo MP, Lathrop E, Osman N, Neves A, Rangeiro R, Mariano AAN, et al. The Mulher Study: cervical cancer screening with primary HPV testing in Mozambique. International Journal of Gynecologic Cancer [Internet]. 2023 Dec 1;33(12):1869. Available from: http://ijgc.bmj.com/content/33/12/1869.abstract. [CrossRef]
- World Health Organization. Introducing and scaling up testing for human papillomavirus as part of a comprehensive programme for prevention and control of cervical cancer: A STEP-BY-STEP-GUIDE [Internet]. World Health Organization; 2020 [cited 2025 May 8]. Available from: https://iris.who.int/bitstream/handle/10665/336668/9789240015166-eng.pdf.
- Arbyn M, Snijders PJF, Meijer CJLM, Berkhof J, Cuschieri K, Kocjan BJ, et al. Which high-risk HPV assays fulfil criteria for use in primary cervical cancer screening? Clinical Microbiology and Infection. 2015 Sep;21(9):817–26. [CrossRef]
- Ronco G, Dillner J, Elfström KM, Tunesi S, Snijders PJF, Arbyn M, et al. Efficacy of HPV-based screening for prevention of invasive cervical cancer: follow-up of four European randomised controlled trials. The Lancet. 2014 Feb;383(9916):524–32. [CrossRef]
- Meijer CJLM, Berkhof J, Castle PE, Hesselink AT, Franco EL, Ronco G, et al. Guidelines for human papillomavirus DNA test requirements for primary cervical cancer screening in women 30 years and older. Int J Cancer. 2009 Feb 1;124(3):516–20. [CrossRef]
- Stanczuk G, Baxter G, Currie H, Lawrence J, Cuschieri K, Wilson A, et al. Clinical validation of hrHPV testing on vaginal and urine self-samples in primary cervical screening (cross-sectional results from the Papillomavirus Dumfries and Galloway-PaVDaG study). BMJ Open. 2016 Apr 25;6(4):e010660. [CrossRef]
- de Oliveira CM, Salcedo MMP, Mariano A, Mariano A, Verdinho MJC, Julião RDAM, et al. HPV testing for cervical cancer screening in Mozambique: Challenges and recommendations. J Glob Health Rep. 2022;6.
- Roche Molecular Systems Inc. Cobas ® HPV Qualitative nucleic acid test for use on the cobas ® 5800/6800/8800 Systems. Doc Rev. 09652736001-01. [Internet]. 2023 Nov [cited 2025 May 8]. Available from: https://elabdoc-prod.roche.com/eLD/api/downloads/f5944ece-fcae-ee11-2191-005056a772fd?countryIsoCode=XG.
- Heideman DAM, Hesselink AT, Berkhof J, van Kemenade F, Melchers WJG, Daalmeijer NF, et al. Clinical Validation of the cobas 4800 HPV Test for Cervical Screening Purposes. J Clin Microbiol. 2011 Nov;49(11):3983–5. [CrossRef]
- Lamin H, Eklund C, Elfström KM, Carlsten-Thor A, Hortlund M, Elfgren K, et al. Randomised healthcare policy evaluation of organised primary human papillomavirus screening of women aged 56–60. BMJ Open. 2017 May 30;7(5):e014788. [CrossRef]
- Frayle H, Gori S, Rizzi M, Graziani BN, Vian E, Giorgi Rossi P, et al. HPV testing for cervical cancer screening: Technical improvement of laboratory logistics and good clinical performance of the cobas 6800 in comparison to the 4800 system. BMC Womens Health. 2019 Mar 25;19(1). [CrossRef]
- Sundström K, Lamin H, Dillner J. Validation of the cobas 6800 human papillomavirus test in primary cervical screening. PLoS One. 2021 Feb 1;16(2 February). [CrossRef]
- Liu SS, Chan KKL, Wei TN, Tse KY, Ngu SF, Chu MMY, et al. Clinical performance of the Roche Cobas 4800 HPV test for primary cervical cancer screening in a Chinese population. PLoS One. 2022 Aug 5;17(8):e0272721. [CrossRef]
- Roche Molecular Systems Inc. Cobas ® HPV Test for use on the cobas ® 4800 System. Doc Rev. 05641268001-12 [Internet]. 2016 Nov [cited 2025 May 8]. Available from: https://www.accessdata.fda.gov/cdrh_docs/pdf10/p100020s017c.pdf.
- Landis JR, Koch GG. The Measurement of Observer Agreement for Categorical Data. Biometrics [Internet]. 1977 Mar [cited 2025 May 7];33(1):159. Available from: https://www.jstor.org/stable/2529310?origin=crossref.
- Miot HA. Análise de concordância em estudos clínicos e experimentais. Vol. 15, Jornal Vascular Brasileiro. Sociedade Brasileira de Angiologia e Cirurgia Vascular; 2016. p. 89–92.
- Ogilvie GS, Krajden M, van Niekerk DJ, Martin RE, Ehlen TG, Ceballos K, et al. Primary cervical cancer screening with HPV testing compared with liquid-based cytology: results of round 1 of a randomised controlled trial – the HPV FOCAL Study. Br J Cancer. 2012 Dec 20;107(12):1917–24. [CrossRef]
- Castle PE, Solomon D, Wheeler CM, Gravitt PE, Wacholder S, Schiffman M. Human Papillomavirus Genotype Specificity of Hybrid Capture 2. J Clin Microbiol. 2008 Aug;46(8):2595–604. [CrossRef]
- Arbyn M, Weiderpass E, Bruni L, de Sanjosé S, Saraiya M, Ferlay J, et al. Estimates of incidence and mortality of cervical cancer in 2018: a worldwide analysis. Lancet Glob Health. 2020 Feb;8(2):e191–203. [CrossRef]
- Bruni L, Albero G, Serrano B, Mena M, Collado JJ, Gómez D, et al. ICO/IARC Information Centre on HPV and Cancer (HPV Information Centre). Human Papillomavirus and Related Diseases in the World. Summary Report [Internet]. 2023 Mar. Available from: www.hpvcentre.net.
- Castle PE, Solomon D, Hildesheim A, Herrero R, Concepcion Bratti M, Sherman ME, et al. Stability of archived liquid-based cervical cytologic specimens. Cancer. 2003 Jan 30;99(2):89–96.
- Agreda PM, Beitman GH, Gutierrez EC, Harris JM, Koch KR, LaViers WD, et al. Long-Term Stability of Human Genomic and Human Papillomavirus DNA Stored in BD SurePath and Hologic PreservCyt Liquid-Based Cytology Media. J Clin Microbiol. 2013 Aug;51(8):2702–6. [CrossRef]
- Gilbert L, Oates E, Ratnam S. Stability of Cervical Specimens in SurePath Medium for Human Papillomavirus Testing with the Roche cobas 4800 Assay. J Clin Microbiol. 2013 Oct;51(10):3412–4. [CrossRef]
- Saville M, Sultana F, Malloy MJ, Velentzis LS, Caruana M, Ip ELO, et al. Clinical Validation of the cobas HPV Test on the cobas 6800 System for the Purpose of Cervical Screening. J Clin Microbiol. 2019 Feb 1;57(2). [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).