Submitted:
24 November 2025
Posted:
25 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Results
3.1. Population Structure
3.1.1. Population
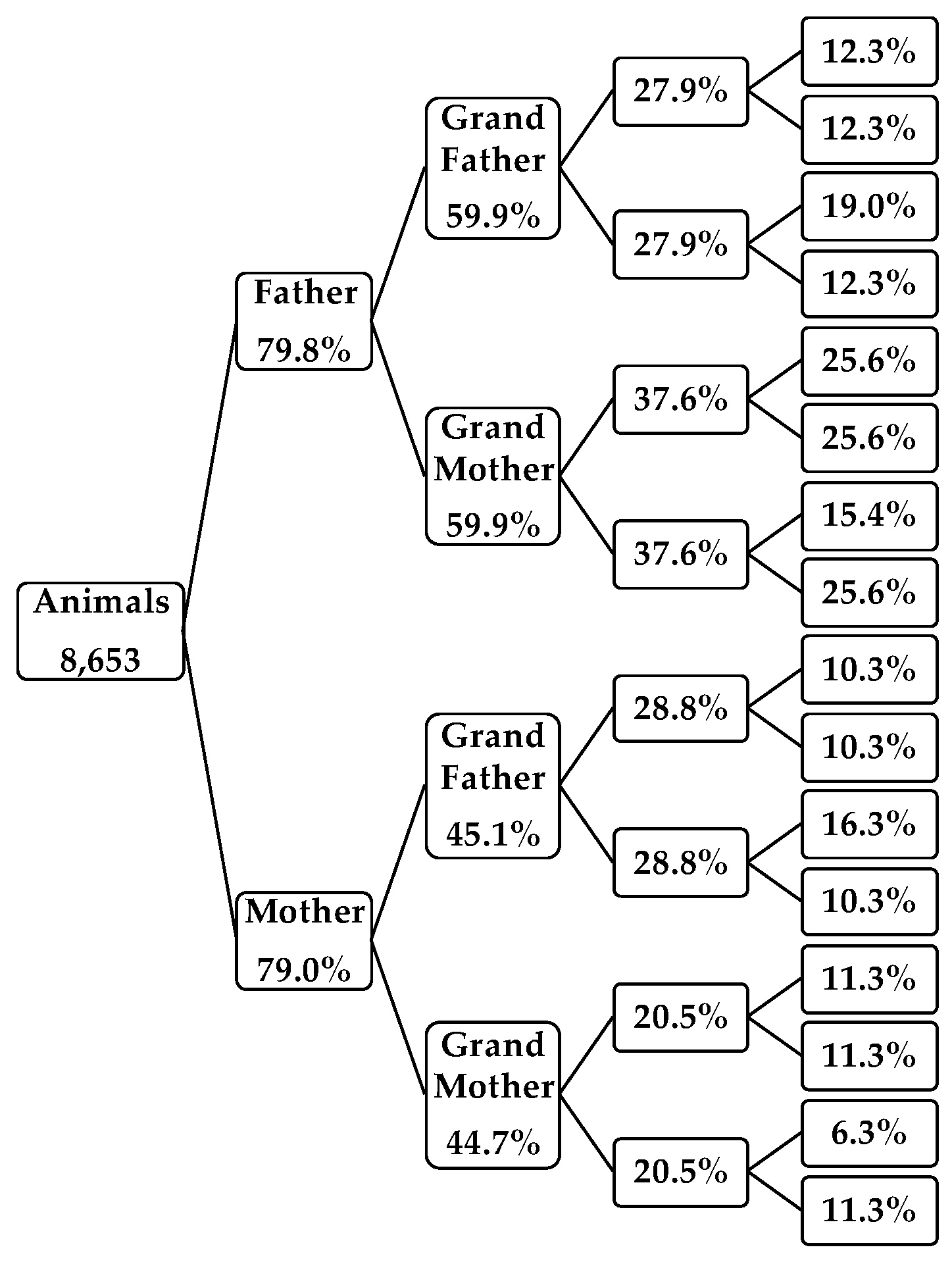
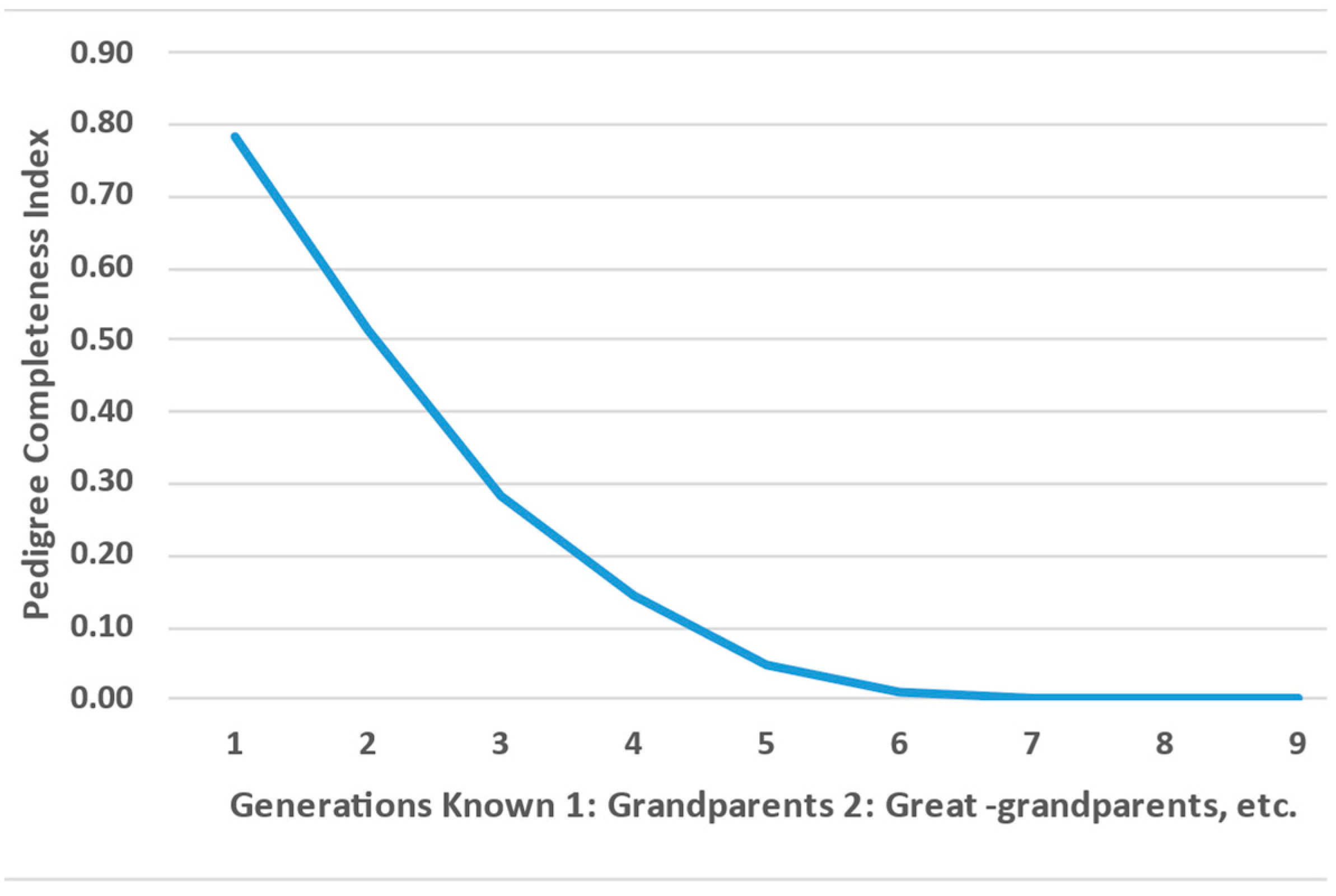
3.1.2. Pedigree Completeness
3.1.3. Generational Interval
3.2. Inbreeding and Diversity
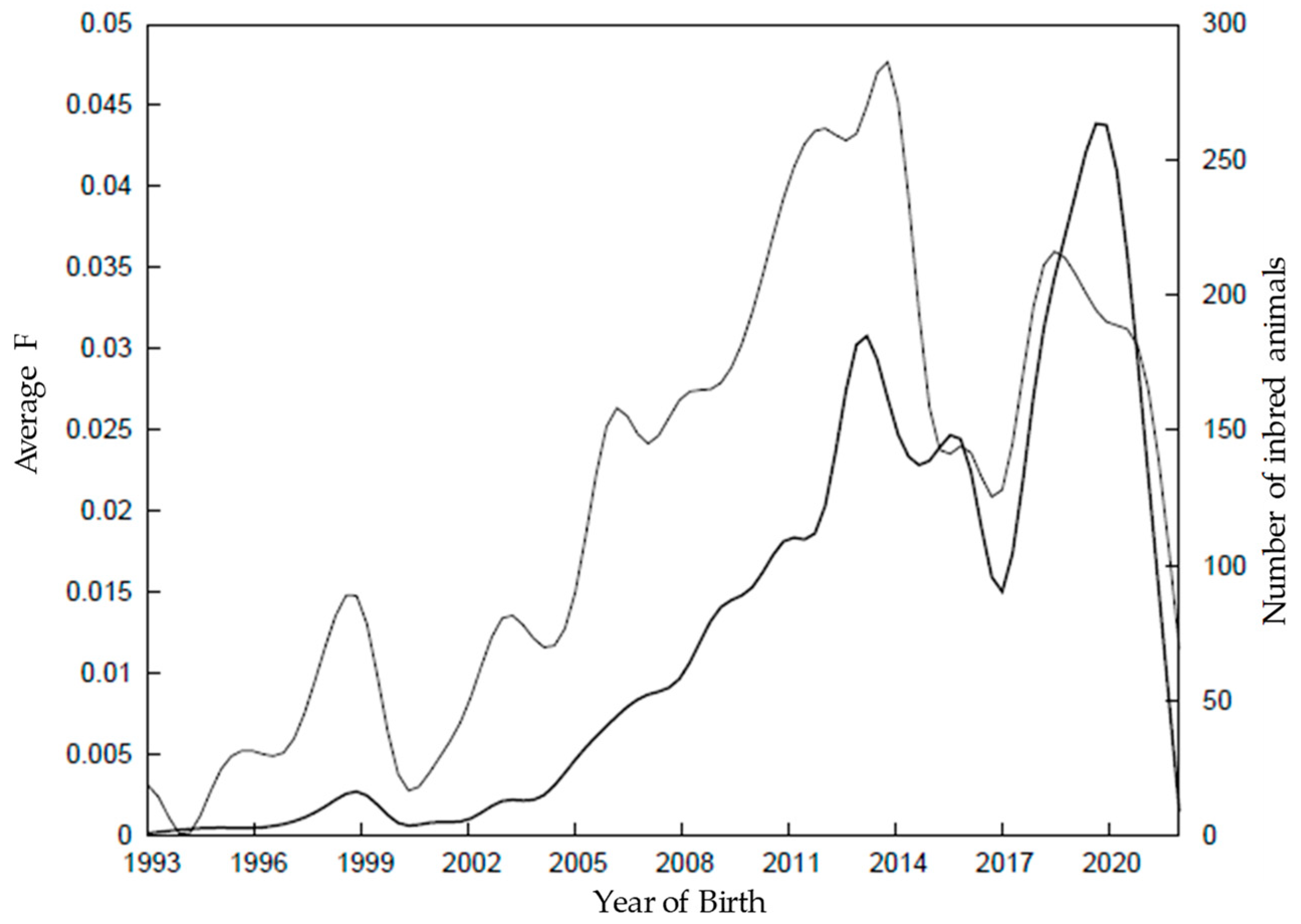
3.2.1. Inbreeding and Relatedness
3.2.2. Effective Population Size
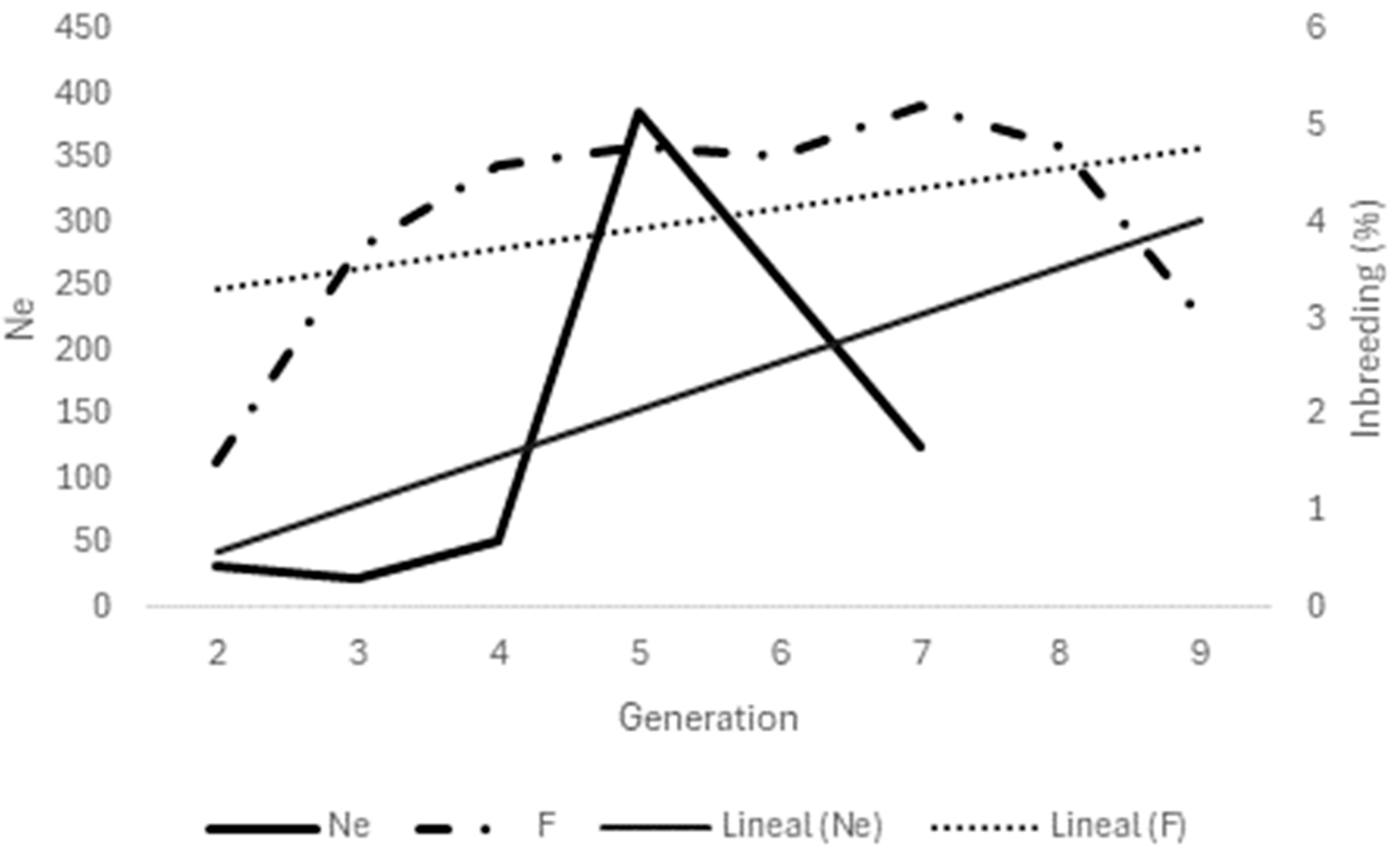
3.2.3. Ancestors
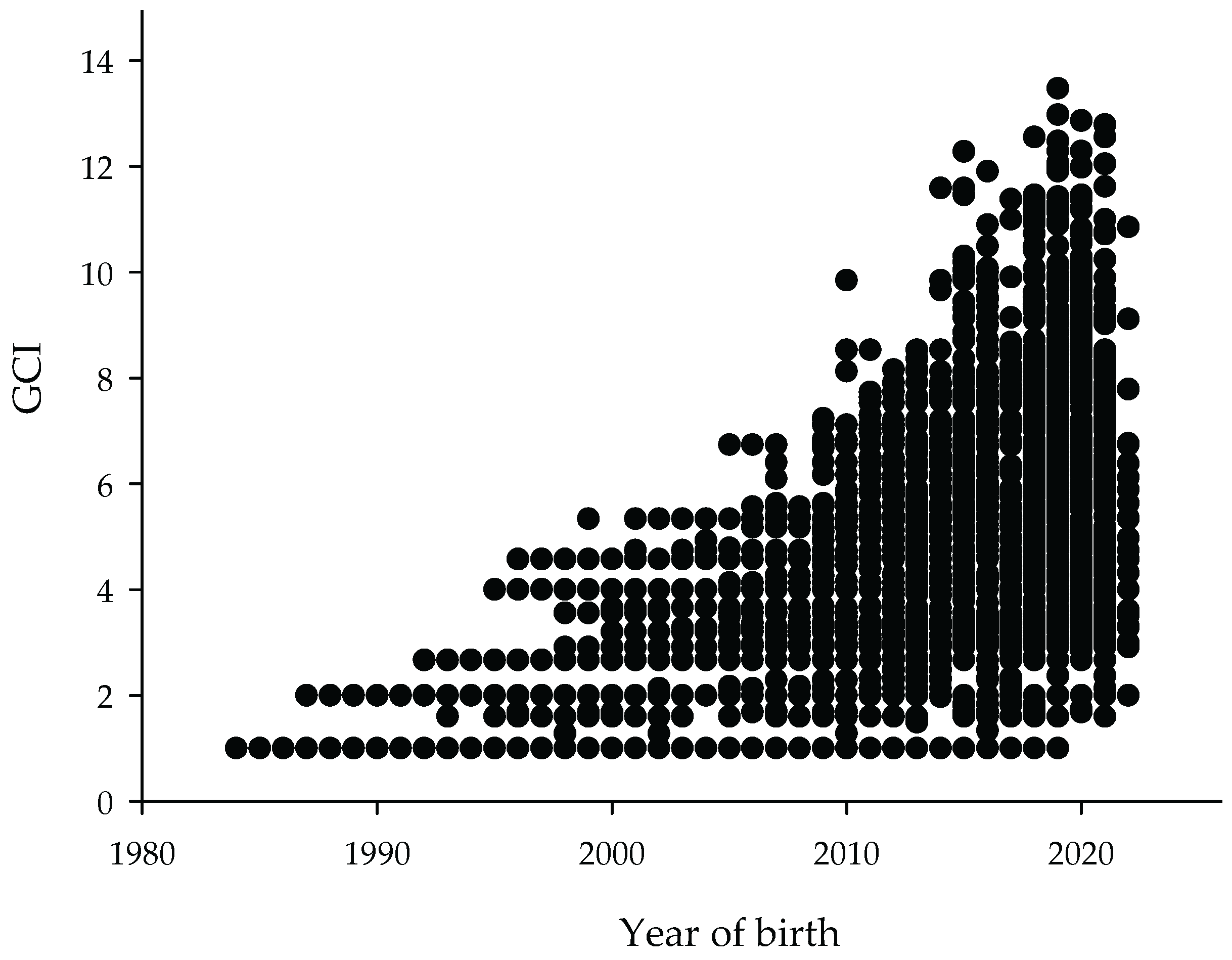
3.2.4. Genetic Conservation Index
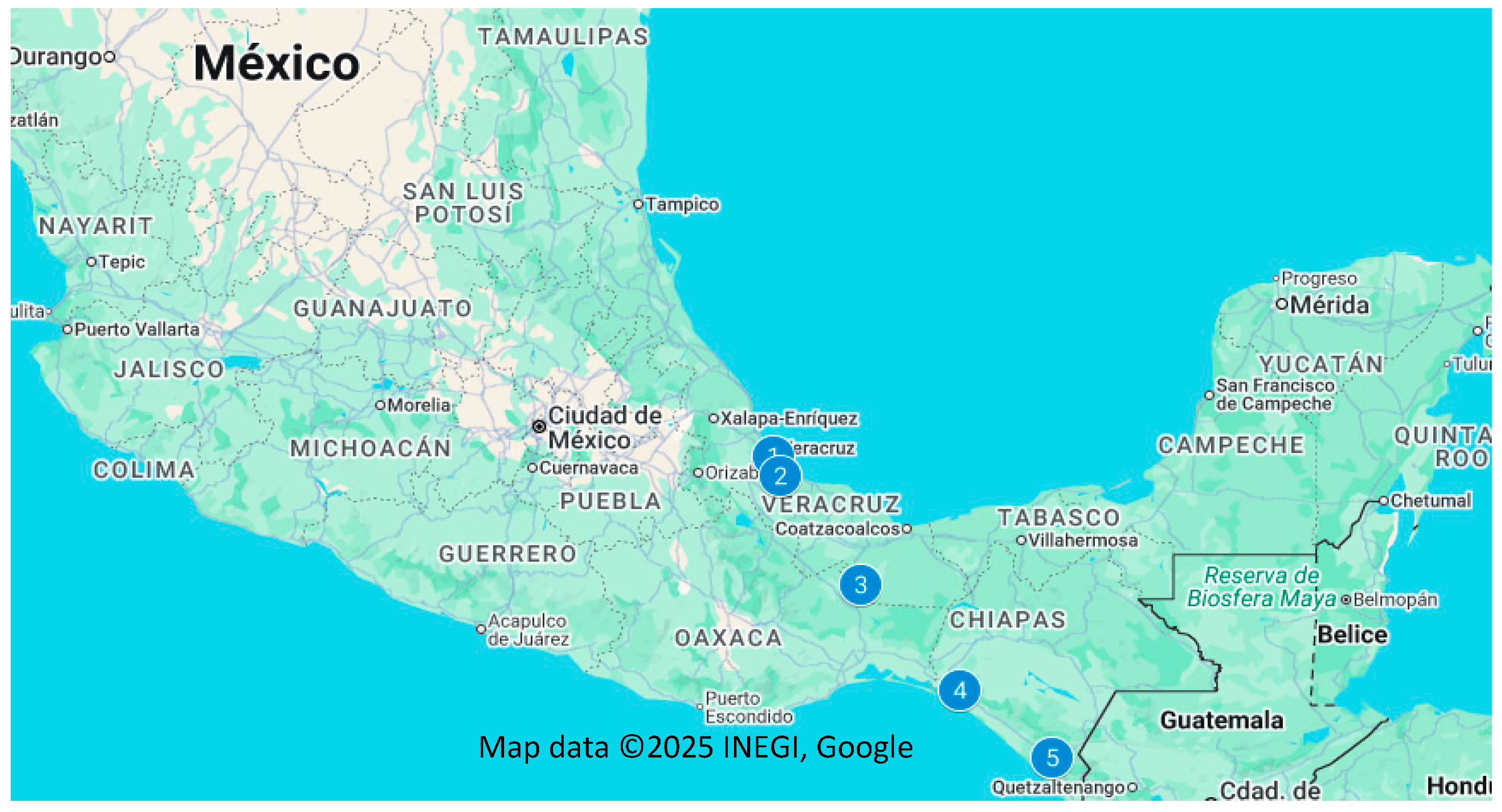
3.2.5. Genetic Differentiation Between Subpopulations
4. Discussion
4.1. Population Structure
4.2. Inbreeding and Diversity
4.3. Implications for Conservation
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Food and Agriculture Organization of the United Nations Sustainable Agriculture for Biodiversity – Biodiversity for Sustainable Agriculture; Roma, Italia, 2018.
- Robinson, T.P.; William Wint, G.R.; Conchedda, G.; Van Boeckel, T.P.; Ercoli, V.; Palamara, E.; Cinardi, G.; D’Aietti, L.; Hay, S.I.; Gilbert, M. Mapping the Global Distribution of Livestock. PLoS One 2014, 9. [Google Scholar] [CrossRef]
- Leroy, G.; Baumung, R.; Boettcher, P.; Besbes, B.; From, T.; Hoffmann, I. Animal Genetic Resources Diversity and Ecosystem Services. Glob Food Sec 2018, 17. [Google Scholar] [CrossRef]
- Vizmanos Pérez, J.L. Claves de La Genética de Poblaciones; Elsevier España: Barcelona, España, 2014. [Google Scholar]
- Gutiérrez, J.P.; Goyache, F. A Note on ENDOG: A Computer Program for Analysing Pedigree Information. Journal of Animal Breeding and Genetics 2005, 122. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, M.C.; de Rezende, M.P.G.; Dadousis, C.; Biffani, S.; Negrini, R.; Carneiro, P.L.S.; Bozzi, R. Population Structure and Genetic Diversity of Italian Beef Breeds as a Tool for Planning Conservation and Selection Strategies. Animals 2019, 9. [Google Scholar] [CrossRef]
- SIAP (Servicio de información Agroalimentaria y pesquera) Avance de La Producción Mensual Por Estado. Producción Mensual Ganadera. SADER, 2021.
- FAOSTAT Crops and Livestock Production. Statistical database 2021.
- Osorio-Arce, M.M.; Segura-Correa, J.C. Sustentabilidad de Los Sistemas de Producción Bovina En El Trópico: Mejoramiento Genético. Livestock Research for Rural Development 2011, 23. [Google Scholar]
- Mehrabi, Z.; Gill, M.; van Wijk, M.; Herrero, M.; Ramankutty, N. Livestock Policy for Sustainable Development. Nat Food 2020, 1. [Google Scholar] [CrossRef]
- Hansen, P.J. Physiological and Cellular Adaptations of Zebu Cattle to Thermal Stress. In Proceedings of the Animal Reproduction Science; 2004; Vol. 82–83.
- Cooke, R.F.; Cardoso, R.C.; Cerri, R.; Lamb, G.C.; Pohler, K.G.; Riley, D.G.; Vasconcelos, J.L.M. Cattle Adapted to Tropical and Subtropical Environments: Genetic and Reproductive Considerations. J Anim Sci 2020, 98. [Google Scholar] [CrossRef]
- Sponenberg, D.P. Conserving the Genetic Diversity of Domesticated Livestock. Diversity (Basel) 2020, 12. [Google Scholar] [CrossRef]
- Ortega R., E.; Belk, T.; Castañeda, J. El Cebú Origen y Desarrollo En México; Asociación Mexicana de Criadores de Cebú, 2012.
- Castañeda, J.; Ortiz Lanz, C. La Odisea Del Cebú En México. Centenario de La Llegada Del Ganado Brasileño; Asociación Mexicana de Criadores de Cebú, 2024.
- Asociación Mexicana de Criadores de Cebú Patrón de La Raza Sardo Negro Available online: https://cebumexico.com/patrones-raciales/.
- Domínguez V., J.; Rodríguez A., F.A. Resumen de Evaluaciones Genéticas En Bovinos Cebú de México 2018; 2018.
- Asociación Mexicana de Criadores de Cebú Sardo Negro Available online: https://cebumexico.com/sardo-negro/.
- Cooke, R.F. 385 Awardee Talk: Impacts of Temperament on Productive and Reproductive Responses of Bos Taurus and B. Indicus Females. J Anim Sci 2020, 98. [Google Scholar] [CrossRef]
- FAO Sistema de Información Sobre La Diversidad de Los Animales Domésticos (DAD-IS). FAO 2020.
- Santana, M.L.; Oliveira, P.S.; Pedrosa, V.B.; Eler, J.P.; Groeneveld, E.; Ferraz, J.B.S. Effect of Inbreeding on Growth and Reproductive Traits of Nellore Cattle in Brazil. Livest Sci 2010, 131. [Google Scholar] [CrossRef]
- García-Ruiz, A.; Martínez-Marín, G.J.; Cortes-Hernández, J.; Ruiz-López, F.D.J. Niveles de Consanguinidad y Sus Efectos Sobre La Expresión Fenotípica En Ganado Holstein. Rev Mex Cienc Pecu 2022, 12. [Google Scholar] [CrossRef]
- Mc Parland, S.; Kearney, J.F.; MacHugh, D.E.; Berry, D.P. Inbreeding Effects on Postweaning Production Traits, Conformation, and Calving Performance in Irish Beef Cattle. J Anim Sci 2008, 86. [Google Scholar] [CrossRef] [PubMed]
- Tisdell, C. Socioeconomic Causes of Loss of Animal Genetic Diversity: Analysis and Assessment. Ecological Economics 2003, 45. [Google Scholar] [CrossRef]
- Nardelli, M.; Túnez, J.I. Aportes de La Genética de La Conservación al Estudio de Los Mamíferos Neotropicales: Revisión y Análisis Crítico. Ecología Austral 2017, 27. [Google Scholar] [CrossRef]
- Colli, L.; Milanesi, M.; Talenti, A.; Bertolini, F.; Chen, M.; Crisà, A.; Daly, K.G.; Del Corvo, M.; Guldbrandtsen, B.; Lenstra, J.A.; et al. Genome-Wide SNP Profiling of Worldwide Goat Populations Reveals Strong Partitioning of Diversity and Highlights Post-Domestication Migration Routes. Genetics Selection Evolution 2018, 50. [Google Scholar] [CrossRef] [PubMed]
- Dadar, M.; Mahyari, S.A.; Rokouei, M.; Edriss, M.A. Rates of Inbreeding and Genetic Diversity in Iranian Holstein Cattle. Animal Science Journal 2014, 85. [Google Scholar] [CrossRef]
- Poirier, M.A.; Coltman, D.W.; Pelletier, F.; Jorgenson, J.; Festa-Bianchet, M. Genetic Decline, Restoration and Rescue of an Isolated Ungulate Population. Evol Appl 2019, 12. [Google Scholar] [CrossRef]
- Santana, M.L.; Pereira, R.J.; Bignardi, A.B.; El Faro, L.; Tonhati, H.; Albuquerque, L.G. History, Structure, and Genetic Diversity of Brazilian Gir Cattle. Livest Sci 2014, 163. [Google Scholar] [CrossRef]
- Dash, S.; Singh, A.; Bhatia, A.K.; Jayakumar, S.; Sharma, A.; Singh, S.; Ganguly, I.; Dixit, S.P. Evaluation of Bovine High-Density SNP Genotyping Array in Indigenous Dairy Cattle Breeds. Anim Biotechnol 2018, 29. [Google Scholar] [CrossRef]
- Domínguez-Viveros, J.; Reyes-Cerón, A.; Saiz-Pineda, J.F.; Villegas-Gutiérrez, C.; Aguilar-Palma, G.N.; Rodríguez-Almeida, F.A. Structure and Genetic Variability of the Mexican Sardo Negro Breed. Ciência Rural 2022, 52. [Google Scholar] [CrossRef]
- INEGI (Instituto Nacional de Estadística, G. en I. Compendio de Información Geográfica Municipal 2010 2010.
- Vidal, R. Las Regiones Climáticas de México. Instituto de Geografía, UNAM 2005, 2.
- Groeneveld, E.; Westhuizen, B.D.; Maiwashe, A.; Voordewind, F.; Ferraz, J.B. POPREP: A Generic Report for Population Management. Genet Mol Res 2009, 8. [Google Scholar] [CrossRef] [PubMed]
- Baumung, R.; Farkas, J.; Boichard, D.; Mészáros, G.; Sölkner, J.; Curik, I. Grain: A Computer Program to Calculate Ancestral and Partial Inbreeding Coefficients Using a Gene Dropping Approach. Journal of Animal Breeding and Genetics 2015, 132. [Google Scholar] [CrossRef] [PubMed]
- Maccluer, J.W.; Boyce, A.J.; Dyke, B.; Weitkamp, L.R.; Pfenning, D.W.; Parsons, C.J. Inbreeding and Pedigree Structure in Standardbred Horses. Journal of Heredity 1983, 74. [Google Scholar] [CrossRef]
- Allendorf, F.W.; Funk, W.C.; Aitken, S.N.; Byrne, M.; Luikart, G. Conservation and the Genomics of Populations; 3rd ed.; Oxford University Press, 2022.
- Caballero, R.A. Cuantitative Genetics; 1st ed.; Cambridge University Press, 2020.
- Meuwissen, T.H.E.; Luo, Z. Computing Inbreeding Coefficients in Large Populations. Genetics, Selection, Evolution 1992, 24. [Google Scholar] [CrossRef]
- Alderson, G.L.H. A System to Maximize the Maintenance of Genetic Variability in Small Populations. Genetic Conservation of Domestic Livestock 1992, 2, 18–29. [Google Scholar]
- Vassallo, J.M.; Diaz, C.; Garcia-Medina, J.R. A Note on the Population Structure of the Avileña Breed of Cattle in Spain. Livest Prod Sci 1986, 15. [Google Scholar] [CrossRef]
- Bó, G.; Carballo, D.; Tribulo, A.; Tribulo, H.; Tribulo, R.; Mapletoft, R. Nuevos Tratamientos Hormonales Para La Superovulación De Donantes De Embriones Bovinos. Zona Rural General Paz 2011, 13. [Google Scholar]
- García, D.B. Generalinades de Inseminación Artificial a Tiempo ( IATF ) En Bovinos. Universidad Cooperativa de Colombia 2019. [Google Scholar]
- Florio, J. Consanguinidad En La Ganadería Bovina. In Manual de ganadería doble propósito 2005; González-Stagnaro, C., Soto B., E., Eds.; Ediciones Astro Data: Maracaibo Venezuela.
- FAO In Vivo Conservation of Animal Genetic Resources; 2013.
- Santana Jr., M. L.; Pereira, R.J.; Bignardi, A.B.; Ayres, D.R.; Menezes, G.R.O.; Silva, L.O.C.; Leroy, G.; Machado, C.H.C.; Josahkian, L.A.; Albuquerque, L.G. Structure and Genetic Diversity of Brazilian Zebu Cattle Breeds Assessed by Pedigree Analysis. Livest Sci 2016, 187, 6–15. [Google Scholar] [CrossRef]
- Roelofs, J.B.; Bouwman, E.B.; Pedersen, H.G.; Rasmussen, Z.R.; Soede, N.M.; Thomsen, P.D.; Kemp, B. Effect of Time of Artificial Insemination on Embryo Sex Ratio in Dairy Cattle. Anim Reprod Sci 2006, 93. [Google Scholar] [CrossRef] [PubMed]
- Delesa, E.K.; Yohannes, A.; Alemayehu, M.; Samuel, T.; Yehualaeshet, T. Calves’ Sex Ratio in Naturally and Artificially Bred Cattle in Central Ethiopia. Theriogenology 2014, 82. [Google Scholar] [CrossRef]
- Ramírez-Valverde, R.; Delgadillo-Zapata, A.R.; Domínguez-Viveros, J.; Hidalgo-Moreno, J.Á.; Núñez-Domínguez, R.; Rodríguez-Almeida, F.A.; Reyes-Quiroz, C.; García-Muñiz, J.G. Análisis de Pedigrí En La Determinación de La Diversidad Genética de Poblaciones Bovinas Para Carne Mexicanas. Rev Mex Cienc Pecu 2018, 9. [Google Scholar] [CrossRef]
- Filipini, V.T.; Isola, J.V.V.; Neves, A.P.; Barbosa, M.R.; Wienke, B.C.D.S.; Scherer, N.P.; da Fontoura Júnior, J.A.S. Simulation Model for Bull: Cow Ratio in Beef Cattle. Braz J Vet Res Anim Sci 2020, 57. [Google Scholar] [CrossRef]
- Román, P.H.; Aguilera, S.R.; Patraca, F.A. Producción y Comercialización de Ganado y Carne de Bovino En El Estado de Veracruz. Comté Nacional del Sistema Producto Bovinos Carne 2012, Volume 1.
- Kie, J.G.; Johnson, B.K.; Noyes, J.H.; Williams, C.L.; Dick, B.L.; Rhodes, O.E.; Stussy, R.J.; Bowyer, R.T. Reproduction in North American Elk Cervus Elaphus: Paternity of Calves Sired by Males of Mixed Age Classes. Wildlife Biol 2013, 19. [Google Scholar] [CrossRef] [PubMed]
- González-Cano, R.; González-Martínez, A.; Muñoz-Mejías, M.E.; Valera, P.; Rodero, E. Analyses of Genetic Diversity in the Endangered"Berrenda" Spanish Cattle Breeds Using Pedigree Data. Animals 2022, 12. [Google Scholar] [CrossRef]
- Elayadeth-Meethal, M.; Thazhathu Veettil, A.; Maloney, S.K.; Hawkins, N.; Misselbrook, T.H.; Sejian, V.; Rivero, M.J.; Lee, M.R.F. Size Does Matter: Parallel Evolution of Adaptive Thermal Tolerance and Body Size Facilitates Adaptation to Climate Change in Domestic Cattle. Ecol Evol 2018, 8. [Google Scholar] [CrossRef]
- Murray-Tortarolo, G.N.; Jaramillo, V.J. Precipitation Extremes in Recent Decades Impact Cattle Populations at the Global and National Scales. Science of the Total Environment 2020, 736. [Google Scholar] [CrossRef]
- Chawala, A.R.; Banos, G.; Peters, A.; Chagunda, M.G.G. Farmer-Preferred Traits in Smallholder Dairy Farming Systems in Tanzania. Trop Anim Health Prod 2019, 51. [Google Scholar] [CrossRef]
- Blackburn, H.; Plante, Y. North American Animal Breeding and Production: Meeting the Needs of a Changing Landscape. Journal of Animal Breeding and Genetics 2014, 131. [Google Scholar] [CrossRef] [PubMed]
- Cartuche Macas, L.F.; Camacho Vallejo, M.E.; González Ariza, A.; León Jurado, J.M.; Delgado Bermejo, J.V.; Marín Navas, C.; Navas González, F.J. Analysis of Endangered Andalusian Black Cattle (Negra Andaluza) Reveals Genetic Reservoir for Bovine Black Trunk. Animals 2024, 14. [Google Scholar] [CrossRef]
- Mc Parland, S.; Kearney, J.F.; Rath, M.; Berry, D.P. Inbreeding Trends and Pedigree Analysis of Irish Dairy and Beef Cattle Populations. J Anim Sci 2007, 85. [Google Scholar] [CrossRef]
- Ablondi, M.; Sabbioni, A.; Stocco, G.; Cipolat-Gotet, C.; Dadousis, C.; Kaam, J.T. van; Finocchiaro, R.; Summer, A. Genetic Diversity in the Italian Holstein Dairy Cattle Based on Pedigree and SNP Data Prior and After Genomic Selection. Front Vet Sci 2022, 8. [Google Scholar] [CrossRef]
- François, L.; Wijnrocx, K.; Colinet, F.G.; Gengler, N.; Hulsegge, B.; Windig, J.J.; Buys, N.; Janssens, S. Genomics of a Revived Breed: Case Study of the Belgian Campine Cattle. PLoS One 2017, 12. [Google Scholar] [CrossRef]
- Cassell, B.G.; Adamec, V.; Pearson, R.E. Effect of Incomplete Pedigrees on Estimates of Inbreeding and Inbreeding Depression for Days to First Service and Summit Milk Yield in Holsteins and Jerseys. J Dairy Sci 2003, 86. [Google Scholar] [CrossRef] [PubMed]
- Núñez-Domínguez, R.; Martínez-Rocha, R.E.; Hidalgo-Moreno, J.A.; Ramírez-Valverde, R.; García-Muñiz, J.G. Evaluation of the Romosinuano Cattle Population Structure in Mexico Using Pedigree Analysis. Revista Colombiana de Ciencias Pecuarias 2020, 33. [Google Scholar] [CrossRef]
- Stachowicz, K.; Sargolzaei, M.; Miglior, F.; Schenkel, F.S. Rates of Inbreeding and Genetic Diversity in Canadian Holstein and Jersey Cattle. J Dairy Sci 2011, 94. [Google Scholar] [CrossRef]
- Sartori, R.; Bastos, M.R.; Baruselli, P.S.; Gimenes, L.U.; Ereno, R.L.; Barros, C.M. Physiological Differences and Implications to Reproductive Management of Bos Taurus and Bos Indicus Cattle in a Tropical Environment. Soc Reprod Fertil Suppl 2010, 67. [Google Scholar] [CrossRef]
- González-Cano, R.; González-Martínez, A.; Muñoz-Mejías, M.E.; Valera, P.; Rodero, E. Analyses of Genetic Diversity in the Endangered"Berrenda" Spanish Cattle Breeds Using Pedigree Data. Animals 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- de Rezende, M.P.G.; Malhado, C.H.M.; Biffani, S.; Souza Carneiro, P.L.; Bozzi, R. Genetic Diversity Derived from Pedigree Information and Estimation of Genetic Parameters for Reproductive Traits of Limousine and Charolais Cattle Raised in Italy. Ital J Anim Sci 2020, 19. [Google Scholar] [CrossRef]
- García-Atance, M.A.; Carleos, C.; Andrino, S.; Justo, J.R.; Rivero, C.J.; Fernández, M.; Cañon, J.; Cortes, O. Genetic Diversity of Five Galician (Northwestern Spain) Local Primitive Bovine Breeds Using Pedigree Records. Diversity (Basel) 2023, 15. [Google Scholar] [CrossRef]
- Domínguez Viveros, J.; Rodríguez Almeida, F.A.; Núñez Domínguez, R.; Ramírez Valverde, R.; Ortega Gutierrez, J.A.; Ruíz Flores, A. Análisis Del Pedigrí y Efectos de La Consanguinidad En El Comportamiento Del Ganado de Lidia Mexicano. Archivos de Zootecnia 2010, 59. [Google Scholar] [CrossRef]
- Lozada-Soto, E.A.; Maltecca, C.; Lu, D.; Miller, S.; Cole, J.B.; Tiezzi, F. Trends in Genetic Diversity and the Effect of Inbreeding in American Angus Cattle under Genomic Selection. Genetics Selection Evolution 2021, 53. [Google Scholar] [CrossRef]
- Hinrichs, D.; Bennewitz, J.; Wellmann, R.; Thaller, G. Estimation of Ancestral Inbreeding Effects on Stillbirth, Calving Ease and Birthweight in German Holstein Dairy Cattle. Journal of Animal Breeding and Genetics 2015, 132. [Google Scholar] [CrossRef]
- Wirth, A.; Duda, J.; Distl, O. Impact of Inbreeding and Ancestral Inbreeding on Longevity Traits in German Brown Cows. Animals 2023, 13. [Google Scholar] [CrossRef] [PubMed]
- Frankham, R.; Gilligan, D.M.; Morris, D.; Briscoe, D.A. Inbreeding and Extinction: Effects of Purging. Conservation Genetics 2001, 2. [Google Scholar] [CrossRef]
- J. , A.; Crow, J.F.; Kimura, M. An Introduction to Population Genetics Theory. Population (French Edition) 1971, 26. [Google Scholar] [CrossRef]
- Mwangi, S.I.; Muasya, T.K.; Ilatsia, E.D.; Kahi, A.K. Effect of Controlling Future Rate of Inbreeding on Expected Genetic Gain and Genetic Variability in Small Livestock Populations. Anim Prod Sci 2020, 60. [Google Scholar] [CrossRef]
- Ríos-Utrera, Á.; Montaño-Bermúdez, M.; Vega-Murillo, V.E.; Martínez-Velázquez, G.; Baeza-Rodríguez, J.J.; Román-Ponce, S.I. Genetic Diversity Evolution in the Mexican Charolais Cattle Population. Anim Biosci 2021, 34. [Google Scholar] [CrossRef]
- Figueredo, J.S.; Cruz, J.F.; Sousa, L.S.; Teixeira Neto, M.R.; Carneiro, P.L.S.; Brito, N.D.; Pinheiro, R.G.S.; Lacerda, K.S.O.; Mottin, V.D. Genetic Diversity and Population Structure Estimation of Brazilian Somali Sheep from Pedigree Data. Small Ruminant Research 2019, 179. [Google Scholar] [CrossRef]
- Oliveira, R.R.; Brasil, L.H.A.; Delgado, J. V.; Peguezuelos, J.; León, J.M.; Guedes, D.G.P.; Arandas, J.K.G.; Ribeiro, M.N. Genetic Diversity and Population Structure of the Spanish Murciano–Granadina Goat Breed According to Pedigree Data. Small Ruminant Research 2016, 144. [Google Scholar] [CrossRef]
- Domínguez-Viveros, J.; Ortega-Gutiérrez, J.Á.; Rodríguez-Almeida, F.A.; Cárdenas-Rivera, J.Á. Variabilidad Genética En Ganaderías de Lidia Mexicanas a Partir de La Información Del Registro Genealógico. ACTA ZOOLÓGICA MEXICANA (N.S.) 2014, 30. [Google Scholar] [CrossRef]
- Martínez-Rocha, R.; Reyes-Ceron, A.; Domínguez-Viveros, J.; Hidalgo, J.; Núñez-Domínguez, R.; Ramírez-Valverde, R.; Larios-Sarabia, N.; Villegas-Gutiérrez, C. Genome-Wide Assessment of Genetic Diversity in Mexican Sardo Negro Breed. Livest Sci 2023, 274. [Google Scholar] [CrossRef]
- Honda, T.; Nomura, T.; Mukai, F. Conservation of Genetic Diversity in the Japanese Black Cattle Population by the Construction of Partially Isolated Lines. Journal of Animal Breeding and Genetics 2005, 122. [Google Scholar] [CrossRef] [PubMed]
- Taberlet, P.; Coissac, E.; Pansu, J.; Pompanon, F. Conservation Genetics of Cattle, Sheep, and Goats. C R Biol 2011, 334. [Google Scholar] [CrossRef] [PubMed]

| Population | N | Percentaje (%) |
| Total | 8,653 | 100 |
| Females | 4,867 | 56 |
| Males | 3,786 | 44 |
| Dams | 2,367 | 491 |
| Sires | 223 | 6 2 |
| Individuals without progeny | 6,063 | 70 |
| Reference population | 6,840 | 79 |
| Individuals with one known parent | 67 | 0.8 |
| Individuals with both parents unknown. | 1,763 | 20 |
| Pathways | GI | SE | AB | SE |
| Sire-Son | 8.9 | ± 0.5 | 8.1 | ± 0.08 |
| Sire-Daughter | 7.6 | ± 0.1 | 7.7 | ± 0.07 |
| Dam-Son | 8.3 | ± 0.4 | 8.6 | ± 0.07 |
| Dam-Daughter | 8.0 | ± 0.1 | 8.3 | ± 0.07 |
| Generation |
No. Animals |
Average F % |
No Inbred animals % (Average F %) | AverageAR % | Ne |
| 0 | 1748 | 0.0 | --- | 0.1 | |
| 1 | 1022 | 0.0 | --- | 1.6 | |
| 2 | 1634 | 1.5 | 9.6 (15.7) | 2.2 | 33 |
| 3 | 846 | 3.7 | 28.4 (13.2) | 3.6 | 22 |
| 4 | 1128 | 4.6 | 42.0 (11.1) | 4.2 | 53 |
| 5 | 1137 | 4.8 | 51.1 (9.3) | 3.6 | 387 |
| 6 | 574 | 4.7 | 57.0 (8.2) | 3.7 | |
| 7 | 360 | 5.2 | 81.4 (6.3) | 3.6 | 124 |
| 8 | 196 | 4.8 | 85.2 (5.7) | 3.4 | |
| 9 | 8 | 3.0 | 75.0 (4.0) | 3.6 |
| Population, metrics | N |
| Total | 8,653 |
| Base population | 1,815 |
| Effective number of founders | 57.4 |
| Number of animals in the reference population | 6,838 |
| Number of founders | 1,187 |
| Number of ancestors contributing to the reference population | 1,175 |
| Effective number of ancestors in the reference population | 32 |
| Effective number of founders in the reference population | 37 |
| Number of ancestors explaining 50% of the ancestry | 21 |
| Herd | Calvings | Own sires %1 | Foreign calvings % 2 |
| 1 | 2,817 | 75.9 | 6.8 |
| 2 | 3,872 | 38.1 | 8.2 |
| 3 | 176 | 68.6 | 0.6 |
| 4 | 430 | 68.6 | 1.3 |
| 5 | 1,002 | 83.5 | 2.9 |
| 6 | 356 | 61.0 | 0.0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




