Submitted:
19 November 2025
Posted:
20 November 2025
You are already at the latest version
Abstract
Based on the genome and transcriptome data of Lonicera japonica Thunb., this study identified six LjDFR gene family members at the genome-wide level. These genes were located on Chr.04 and Chr.09, and the full-length coding sequences of LjDFR1 to LjDFR6 were successfully cloned. The proteins encoded by the cloned genes are all hydrophilic, with secondary structures dominated by α-helices and random coils. The subcellular localization analysis indicated that LjDFRs are primarily localized in the cell membrane and nucleus. Phylogenetic analysis classified the LjDFR proteins into four subfamilies, clustering with DFR homologs from species such as Capsicum annuum and Camellia sinensis, reflecting a high degree of evolutionary conservation. Promoter analysis identified multiple cis-acting elements associated with light response, hormone signaling, and stress-responses. Expression pattern analysis demonstrated that LjDFR genes exhibit tissue-specific and stage-specific expression patterns during flower development in L. japonica varieties with different floral colors. Notably, LjDFR2 expression was significantly higher in the deeply pigmented tissues of Lonicera japonica Thunb. var. chinensis (Wats.) Bak. than in L. japonica. Together with its phylogenetic clustering with the anthocyanin-related CsDFRa and CaDFR5 genes, this finding suggests that LjDFR2 potentially positively correlated with anthocyanin accumulation. Furthermore, the expression of LjDFR2 and LjDFR4 was significantly induced under both drought and salt stress, indicating their involvement in abiotic stress responses. This study provides a foundation for further functional characterization of LjDFR genes in anthocyanin metabolism and stress resistance, and offers valuable candidate genes for molecular breeding of L. japonica.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Materials and Stress Treatments
2.2. Identification and Chromosomal Distribution of the LjDFR Gene Family
2.3. Cloning, Physicochemical Properties, and Structural Characteristics Analysis of LjDFR Genes
2.4. Subcellular Localization Analysis
2.5. Multiple Sequence Alignment and Phylogenetic Analysis
2.6. Conserved Motif and Gene Structure Analysis
2.7. Collinearity Analysis and Identification of Cis-Acting Elements in Promoters of the LjDFR Gene Family
2.8. RNA Extraction, cDNA Synthesis, and qRT-PCR Analysis
3. Results
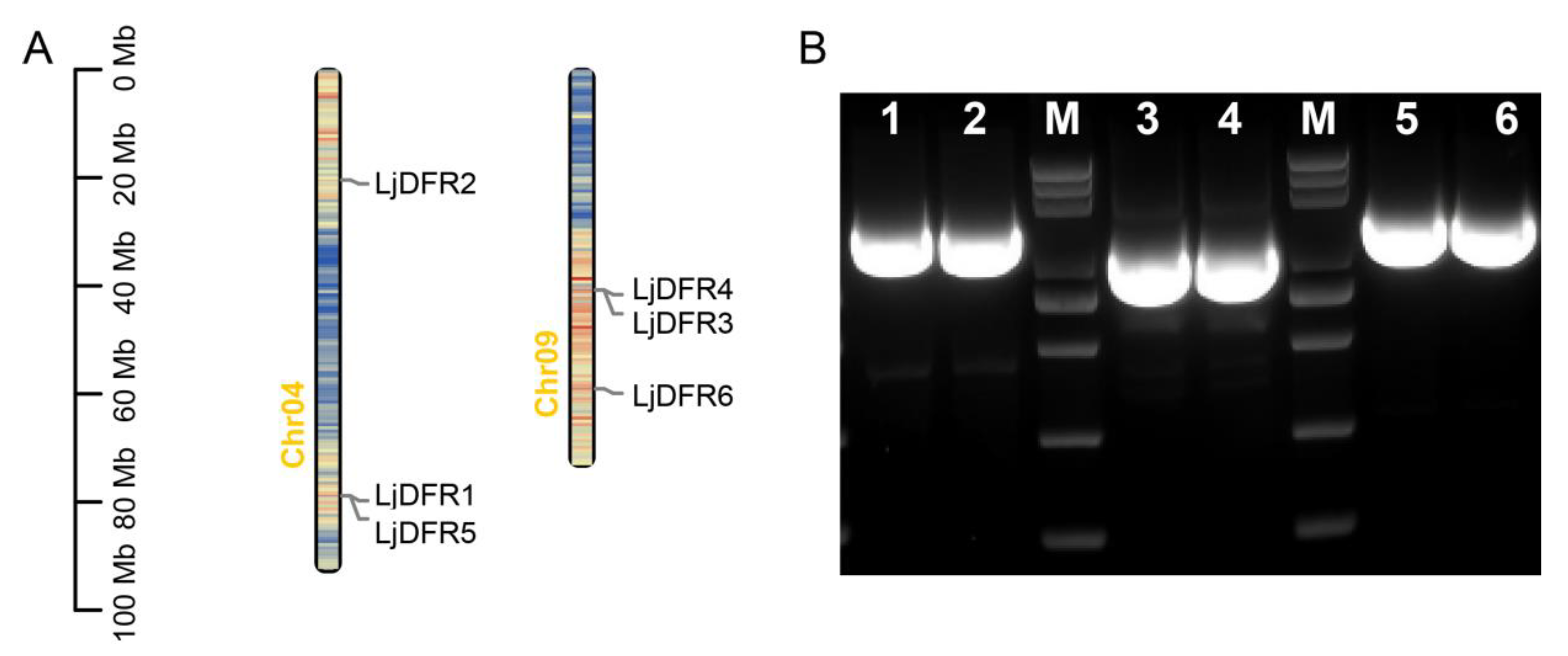
3.1. Genome-Wide Identification, Full-length Cloning, and Chromosomal Distribution of LjDFR Gene Family
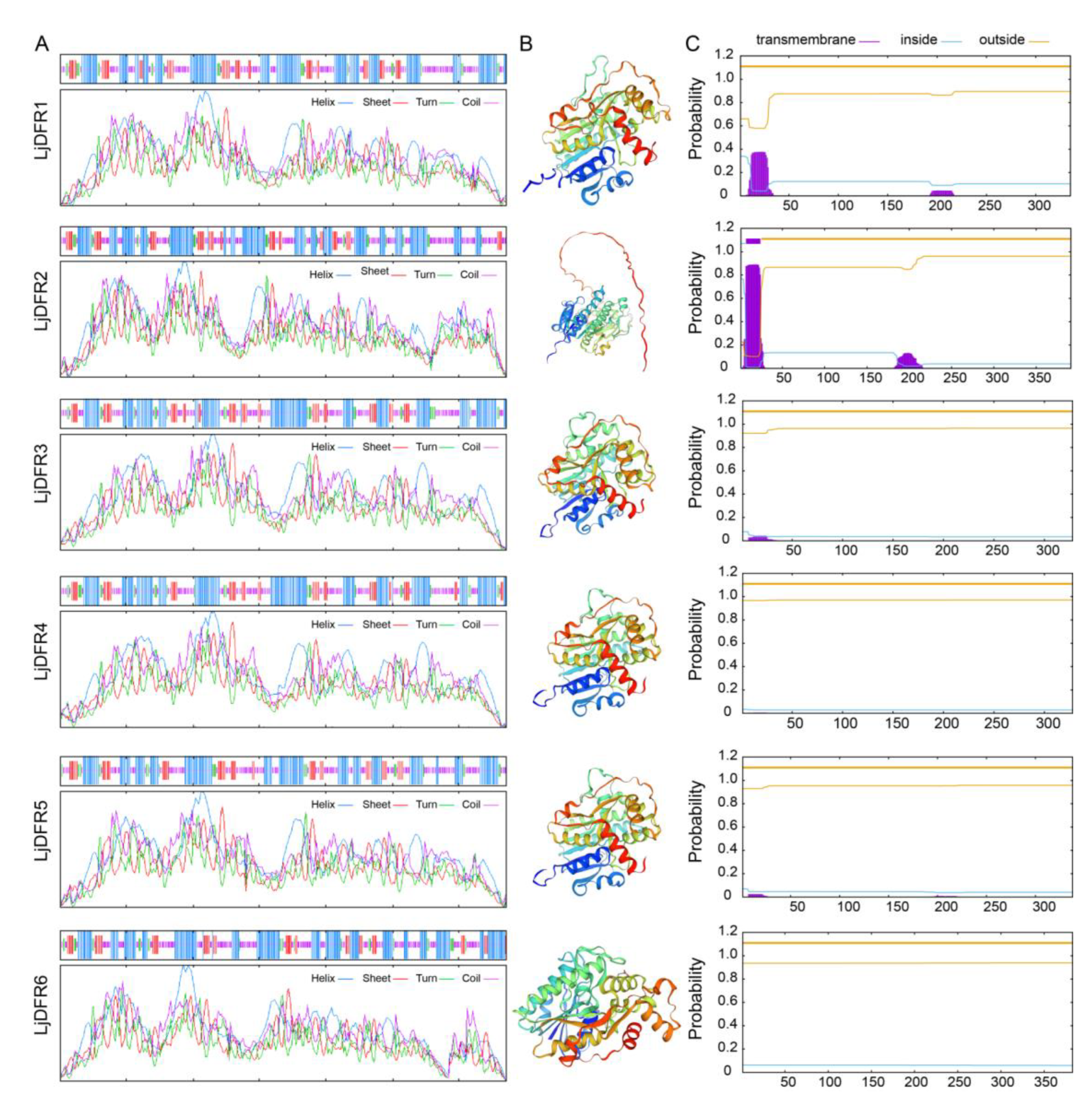
3.2. Protein Physicochemical Properties and Structural Analysis of the LjDFR Gene Family
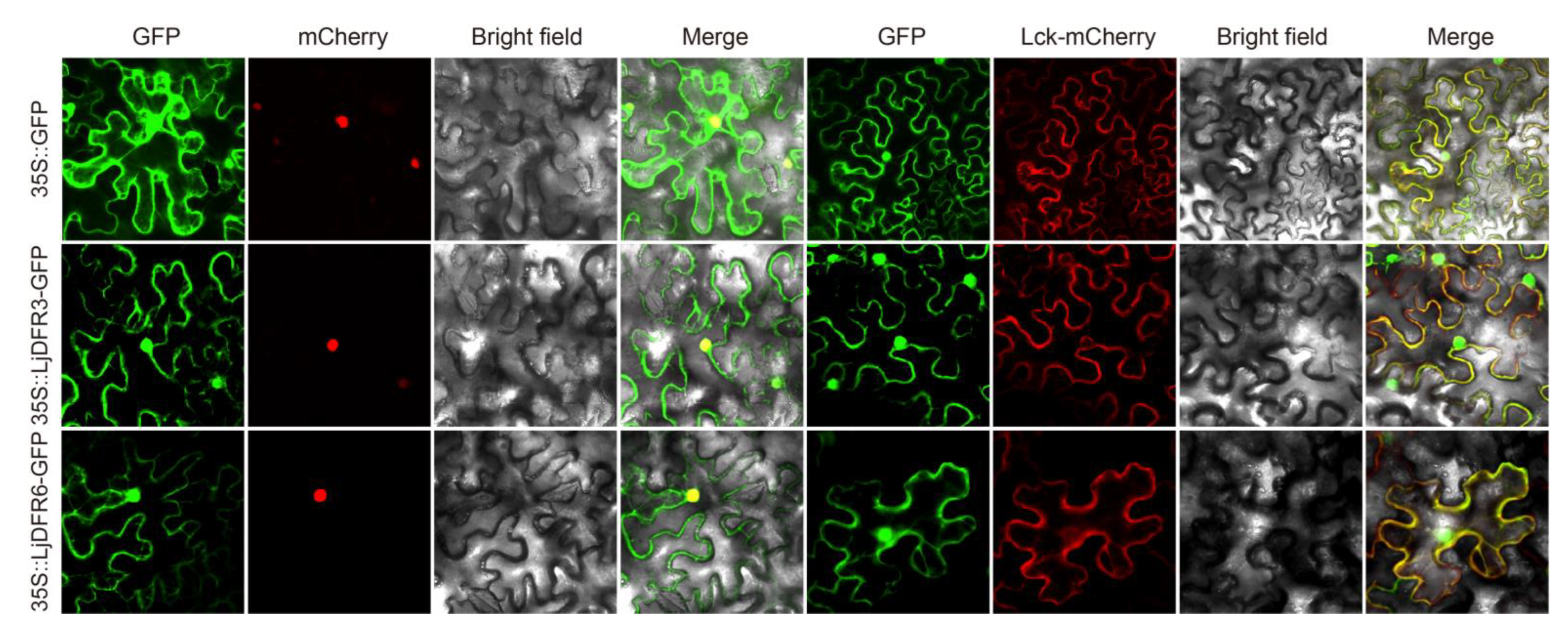
3.3. Subcellular Localization Analysis of LjDFR Proteins
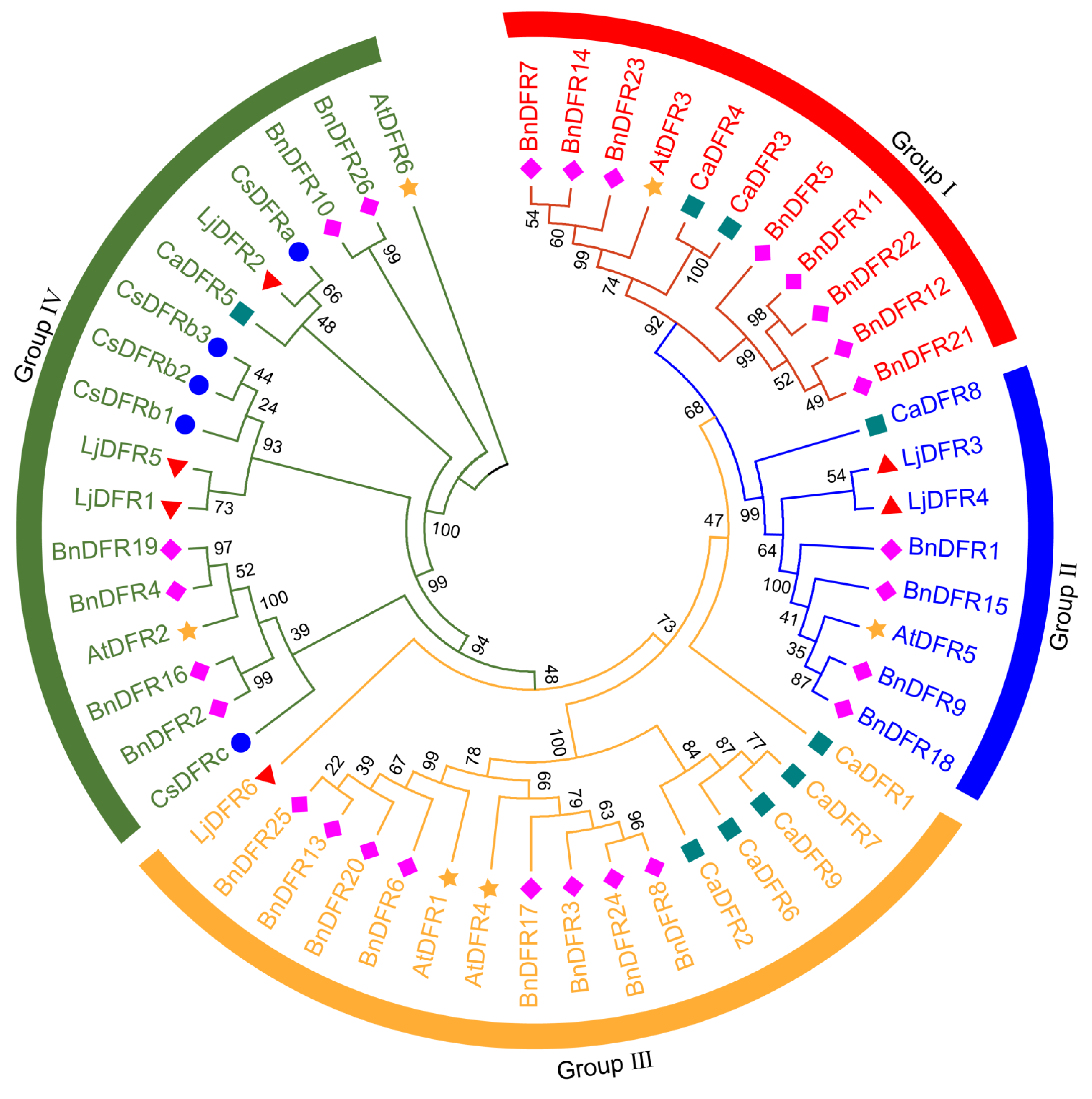
3.4. Phylogenetic Analysis of the LjDFR Family
3.5. Gene Structure, Conserved Motifs, and Synteny Analysis of the LjDFR Gene Family
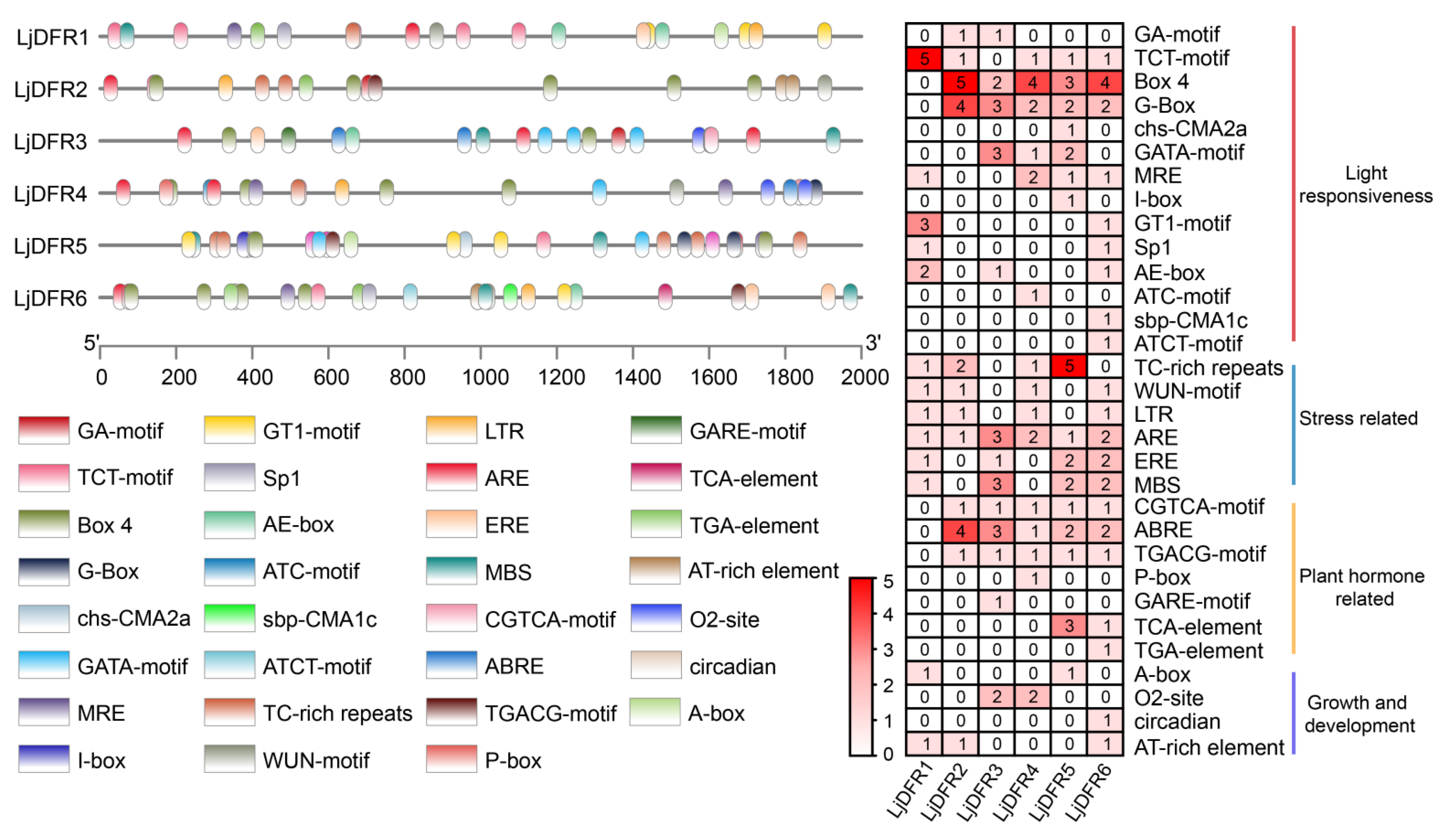
3.6. Analysis of Cis-Acting Elements in the Promoters of LjDFR Genes
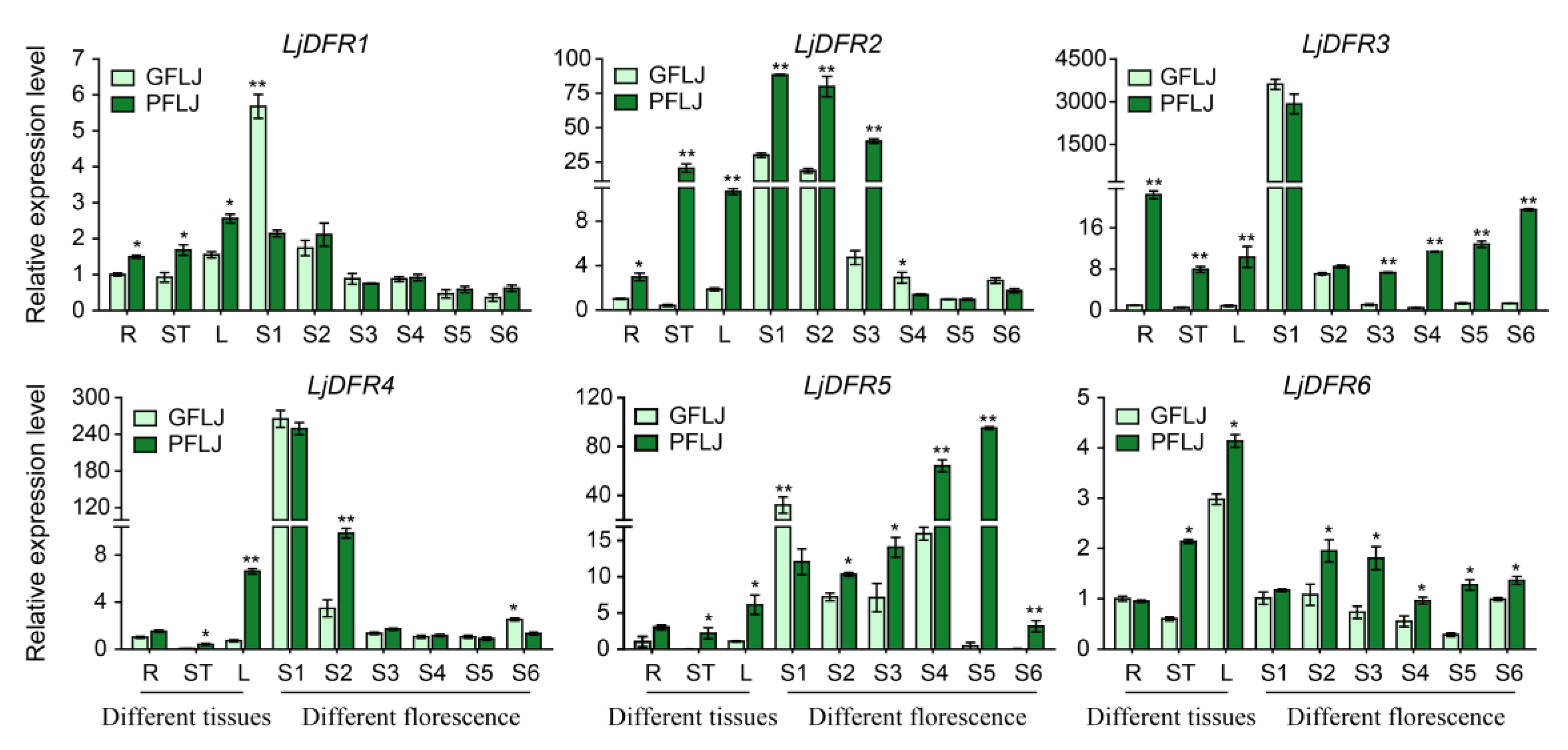
3.7. Tissue Expression Patterns of the LjDFR Gene Family
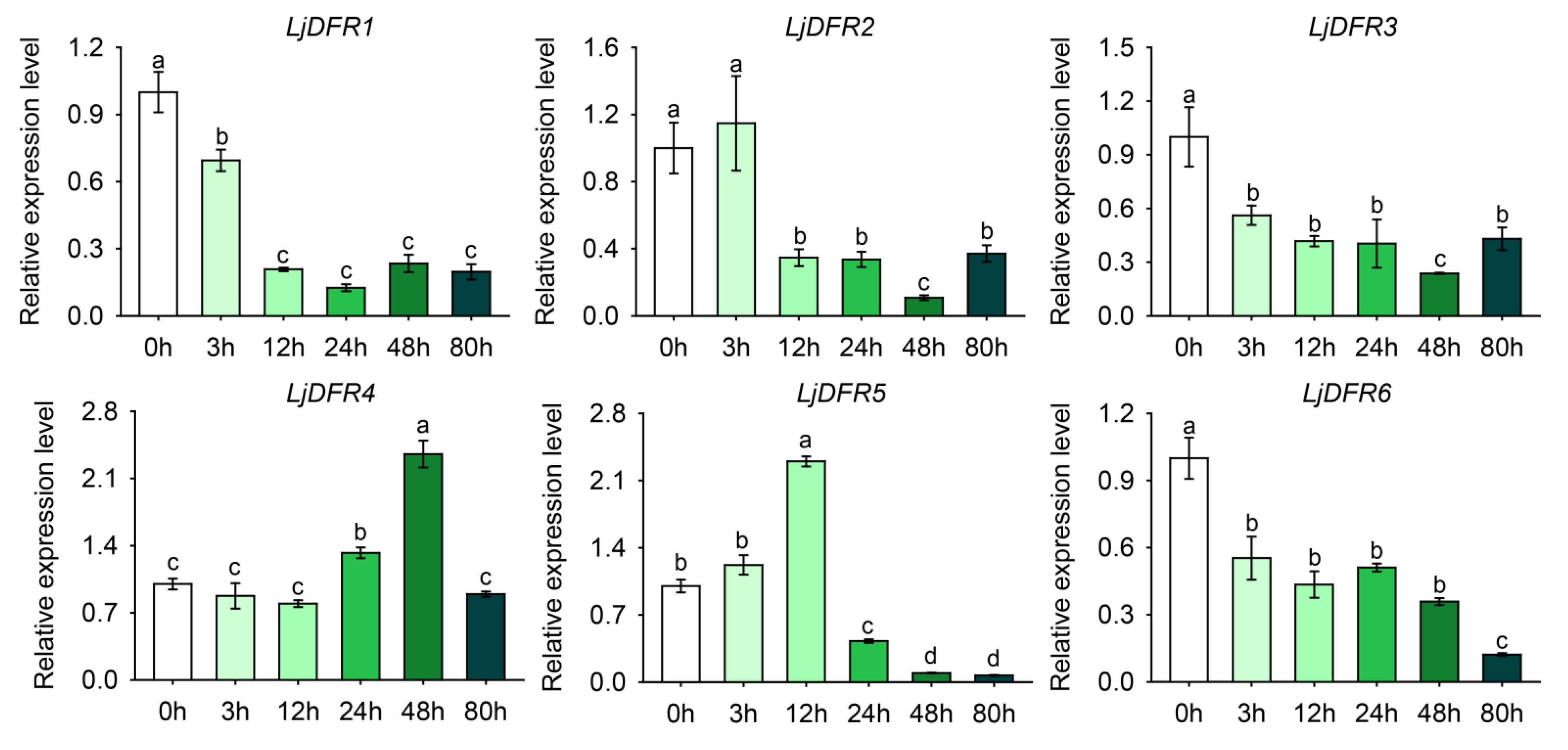
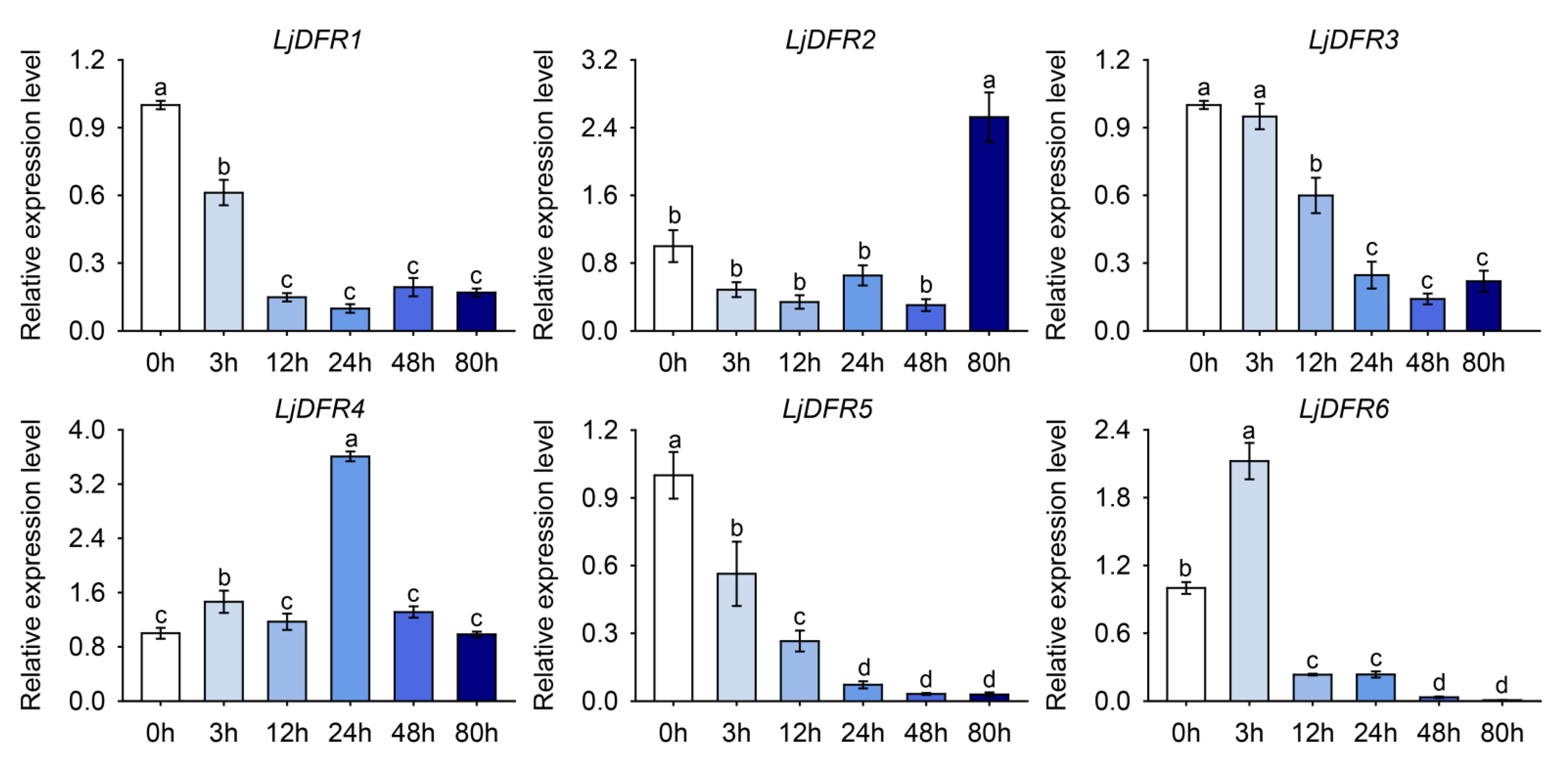
3.8. Expression Patterns of the LjDFR Gene Family under Drought and Salt Stress
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Karanjalker, G.R.; Ravishankar, K.V.; Shivashankara, K.S.; Dinesh, M.R.; Roy, T.K.; Sudhakar Rao, D.V. A Study on the expression of genes involved in carotenoids and anthocyanins during ripening in fruit peel of green, yellow, and red colored mango cultivars. Appl. Biochem. Biotechnol. 2018, 184, 140-154. [CrossRef]
- Shang, Y.; Venail, J.; Mackay, S.; Bailey, P.C.; Schwinn, K.E.; Jameson, P.E.; Martin, C.R.; Davies, K.M. The molecular basis for venation patterning of pigmentation and its effect on pollinator attraction in flowers of Antirrhinum. New Phytol. 2011, 189, 602-615. [CrossRef]
- Zhang, Y.; Butelli, E.; Martin, C. Engineering anthocyanin biosynthesis in plants. Curr. Opin. Plant Biol. 2014, 19, 81-90. [CrossRef]
- Xie, S.; Zhao, T.; Zhang, Z.W.; Meng, J.F. Reduction of dihydrokaempferol by Vitis vinfera dihydroflavonol 4-reductase to produce orange Pelargonidin-Type anthocyanins. J. Agric. Food Chem. 2018, 66, 3524-3532. [CrossRef]
- Jaakola, L. New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant Sci. 2013, 18, 477-483. [CrossRef]
- LaFountain, A.M.; Yuan, Y.W. Repressors of anthocyanin biosynthesis. New Phytol. 2021, 231, 933-949. [CrossRef]
- Miyagawa, N.; Miyahara, T.; Okamoto, M.; Hirose, Y.; Sakaguchi, K.; Hatano, S.; Ozeki, Y. Dihydroflavonol 4-reductase activity is associated with the intensity of flower colors in delphinium. Plant Biotechnol. 2015, 32, 249-255. [CrossRef]
- Ni, J.; Ruan, R.J.; Wang, L.J.; Jiang, Z.F.; Gu, X.J.; Chen, L.S.; Xu, M.J. Functional and correlation analyses of dihydroflavonol-4-reductase genes indicate their roles in regulating anthocyanin changes in Ginkgo biloba. Ind. Crops Prod. 2020, 152, 112546. [CrossRef]
- Johnson, E.T.; Ryu, S.; Yi, H.; Shin, B.; Cheong, H.; Choi, G. Alteration of a single amino acid changes the substrate specificity of dihydroflavonol 4-reductase. Plant J. 2001, 25, 325-333. [CrossRef]
- Liu, H.L.; Lou, Q.; Ma, J.R.; Su,B.B.; Gao, Z.Z.; Liu, Y.L. Cloning and functional characterization of dihydroflavonol 4-reductase gene involved in anthocyanidin biosynthesis of Grape Hyacinth. Int. J. Mol. Sci. 2019, 20, 4743. [CrossRef]
- Vainio, J.; Mattila1, S.; Abdou, S.M.; Sipari, N.; Teeri T.H. Petunia dihydroflavonol 4-reductase is only a few amino acids away from producing orange pelargonidinbased anthocyanins. Front. Plant Sci. 2023, 14, 1227219. [CrossRef]
- Shang, X.F.; Pan, H.; Li, M.X.; Miao, X.L.; Ding, H. Lonicera japonica Thunb.: ethnopharmacology, phytochemistry and pharmacology of an important traditional Chinese medicine. J. Ethnopharmacol. 2001, 138, 1-21. [CrossRef]
- Seo, O.N.; Kim, G.S.; Park, S.; Lee, J.H.; Kim, Y.H.; Lee, W.S.; Lee, S.J.; Kim, C.Y.; Jin, J.S.; Choi, S.K.; Shin, S.C. Determination of polyphenol components of Lonicera japonica Thunb. using liquid chromatography-tandem mass spectrometry: contribution to the overall antioxidant activity. Food Chem. 2012, 134, 572-577. [CrossRef]
- Yuan, Y.; Yang, J.; Yu, X.D.; Huang, L.Q.; Lin, S.F. Anthocyanins from buds of Lonicera japonica Thunb. var. chinensis (Wats.) Bak.. Food Res. Int. 2014, 62, 812-818. [CrossRef]
- Li, J.; Lian, X.; Ye, C.; Wang, L. Analysis of flower color variations at different developmental stages in two L. japonica (Lonicera japonica Thunb.) cultivars. HortScience 2019, 54, 779-782. [CrossRef]
- An, Y.; Li, N.; Zhang, R.L.; Wang, S.; Wang, J.N. Identification and characterization of DFR gene family and cloning of candidate genes for anthocyanin biosynthesis in pepper (Capsicum annuum L.). BMC Plant Biol. 2025, 25, 830. [CrossRef]
- Ruan, H.X.; Shi, X.X.; Gao, L.P.; Rashid, A.; Li, Y.; Lei, T.; Dai, X.L.; Xia, T.; Wang, Y.S. Functional analysis of the dihydroflavonol 4-reductase family of Camellia sinensis: exploiting key amino acids to reconstruct reduction activity. Hortic. Res. 2022, 9, uhac098. [CrossRef]
- Pu, X.D.; Li, Z.; Tian, Y.; Gao, R.R.; Hao, L.J.; Hu, Y.T.; He, C.N.; Sun, W.; Xu, M.M; Peters, R.J.; et al. The honeysuckle genome provides insight into the molecular mechanism of carotenoid metabolism underlying dynamic flower coloration. New phytol. 2020, 227, 930-943. [CrossRef]
- Lu, D.D.; Tan, Z.W.; Yu, Y.L.; Li, L.; Xu, L.J.; Yang, H.Q.; Yang, Q.; Dong, W.; An, S.F.; Liang, H.Z. Cloning, structure and expression profile analysis of CtANR2 and CtANR3 genes from Carthamus tinctorius L.. Acta Agric. Boreali-Sin. 2023, 38, 84-93. [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol Plant. 2020, 13, 1194-1202. [CrossRef]
- Wang, Y.P.; Tang, H.B.; Debarry, J.D.; Tan, X.; Li, J.P.; Wang, X.Y.; Lee, T.H.; Jin, H.Z.; Marler, B.; Guo, H.; Kissinger, J.C.; Paterson, A.H. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [CrossRef]
- Liu, X.Y.; Chen, L.; Qiao, Y.G.; Rong, Y.; Wang J.S. Selection of reference genes by qRT-PCR inflower organ of Lonicera japonica Thunb.. J Shanxi Agri Sci. 2017, 45, 514-517. [CrossRef]
- Meyer, P.; Heidmann, I.; Forkmann, G.; Saedler, H. A new Petunia flower colour generated by transformation of a mutant with a maize gene. Nature 1987, 330, 677-678. [CrossRef]
- Qin, S.; Liu, Y.T.; Cui, B.Q.; Cheng, J.L.; Liu, S.Y.; Liu, H.Z. Isolation and functional diversification of dihydroflavonol 4-Reductase gene HvDFR from Hosta ventricosa indicate its role in driving anthocyanin accumulation. Plant signal. Behav. 2022, 17, e2010389. [CrossRef]
- Sun, W.; Zhou, N.N.; Feng, C.; Sun, S.Y.; Tang, M.; Tang, X.X.; Ju, Z.G.; Yi, Y. Functional analysis of a dihydroflavonol 4-reductase gene in Ophiorrhiza japonica (OjDFR1) reveals its role in the regulation of anthocyanin. Peer J 2021, 9, e12323. [CrossRef]
- Sun, W.; Zhou, N.N.; Wang, Y.H.; Sun, S.Y.; Zhang, Y.; Zhigang Ju, Z.G.; Yi, Y. Characterization and functional analysis of RdDFR1 regulation on flower color formation in Rhododendron delavayi. Plant Physiol. Biochem. 2021, 169: 203-210. [CrossRef]
- Tan, Z.W.; Lu, D.D.; Li, L.; Yu, Y.L.; Xu, L.J.; Dong, W.; Yang, H.Q.; Yang, Q.; Li, C.M.; Liang, H.Z. Cloning and expression analysis of dihydroflavonol 4-reductase gene from Safflower (Carthamus tinctorius L.). Mol Plant Breeding. 2022, 20, 5309-5318. [CrossRef]
- Yang, C.L.; Li, Y.Z.; He, L.L.; Song, Y.H.; Zhang, P.; Liu, Z.X.; Li, P.H.; Liu, S.J. Genome-wide identification and analysis of TPS gene family and functional verification of VvTPS4 in the formation of monoterpenes in Grape. Sci Agric Sin. 2025, 58, 1397-1417. [CrossRef]
- Feng, X.; Zhang, Y.T.; Wang, H.; Tian, Z.D.; Siyao Xin, S.Y.; Zhu, P.F. The dihydrofavonol 4-reductase BoDFR1 drives anthocyanin accumulation in pink-leaved ornamental kale. Theor. Appl. Genet. 2021, 134, 159-169. [CrossRef]
- Shi, Z.; Li, H.; Gao, M.; Guo, C.H.; Guo, D.L.; Bi, Y.D. Cloning of GmDFR gene from Soybean (Glycine max) and identification of its function on resistance to iron deficiency. J. Agric. Biotechnol. 2023, 31, 259-272. [CrossRef]
- Wang, H.J.; Chen, X.J.; Li, T.J.; Luo, J.; Qu, Y. Cloning and expression analysis of DFR gene from Meconopsis with different colors. Acta Agric Boreali Sin. 2024, 39, 88-95. [CrossRef]
- Zheng, H.; Zhang, F.; Jian, Y.; Huang, W.L.; Liang, S.; Jiang, M.; Yuan, Q.; Wang, Q.M.; Sun, B. Cloning and function identification of Dihydroflavonol 4-Reductase gene BoaDFR in Chinese Kale. Acta Hortic. Sin. 2021, 48, 73-82. [CrossRef]
- Zhang, B.Y.; Li, C.H.; Liu, X.; Liao, X.S.; Rong, D.Y. Cloning and subcellular localization analysis of LcDRF1 and LcDRF2 in Loropetalum chinense var. rubrum. J. South. Agric. 2020, 51, 2865-2874. [CrossRef]
- Zhu, Y.; Peng, Q.Z.; Li, K.G.; Xie, D.Y. Molecular cloning and functional characterization of a dihydroflavonol 4-reductase from Vitis bellula. Molecules 2018, 23, 861. [CrossRef]
- Zhou, X.Z.; Yao, S.H.; Li, J.W.; Chen, K.Y.; Yu, D. Analyses of gene duplication and synteny in Melampsora larici-populina (Pucciniales, Basidiomycota). Mycosystema 2021, 40, 580-591. [CrossRef]
- Xu, G.X.; Guo, C.C.; Shan, H.Y.; Kong, H.Z. Divergence of duplicate genes in exon-intron structure. Proc. Natl. Acad. Sci. U.S.A. 2012, 109, 1187-1192. [CrossRef]
- Hu, K.; Meng, L.; Han, K.T.; Sun, Y.; Dai, S.L. Isolation and expression analysis of key genes involved in anthocyanin biosynthesis of Cineraria. Acta Hortic. Sin. 2009, 36, 1013-1022. [CrossRef]
- Jiang, B.X.; Wang, Q.H.; Yang, G.X.; Jia, Y.H.; Xie, X.H.; Wu, Y.Y. Cloning and analysis of RhDFR gene in Rhododendron hybridum Hort.. Acta Bot. Boreat. Occident. Sin. 2023, 43, 10-20. [CrossRef]
- Yu, X; Li, Y.H.; Zhang, L.Y.; Liu H.; Luo C.; Cheng X.; Gao K.; Huang C.L.; Chen D.L. Cloning and expression analysis of CmDFRa gene in Chrysanthemum × morifolium. Mol Plant Breeding. 2024, 4, 1-13. https://link.cnki.net/urlid/46.1068.S.20240424.0948.005.
- Lim, S.H.; Park, B.; Kim, D.H.; Park, S.; Yang, J.H.; Jung J.A.; Lee J.M.; Lee J.Y. Cloning and functional characterization of dihydroflavonol 4-reductase gene involved in anthocyanin biosynthesis of Chrysanthemum. Int. J. Mol. Sci. 2020, 21, 7960. [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



