Submitted:
17 November 2025
Posted:
19 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Analysis of Spent Culture Media from Day 5 Blastocysts by Quantitative RT-PCR
2.2. Analysis of Spent Culture Media from Day 6 Blastocysts by Quantitative RT-PCR
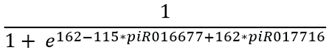 (1)
(1)
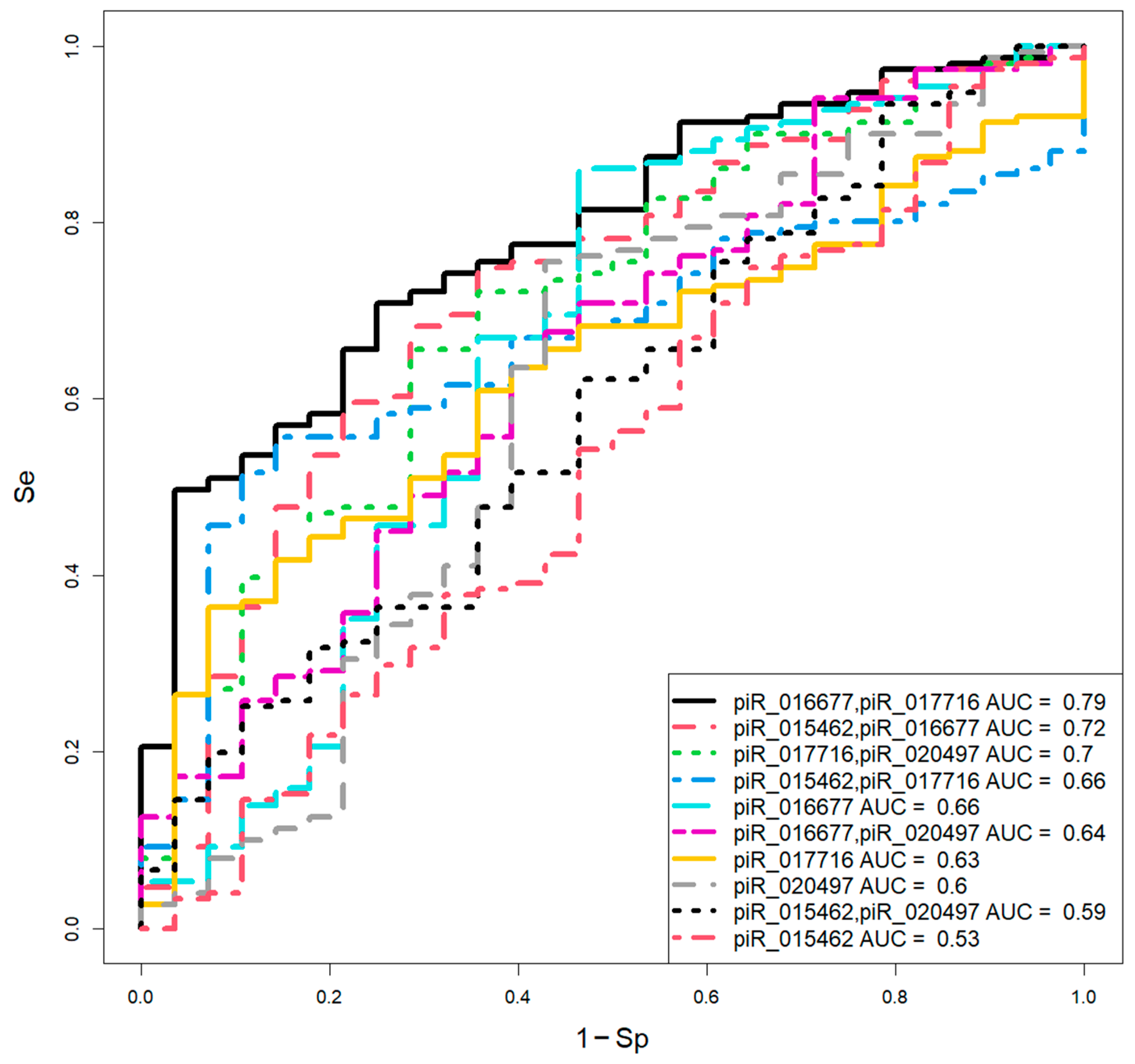
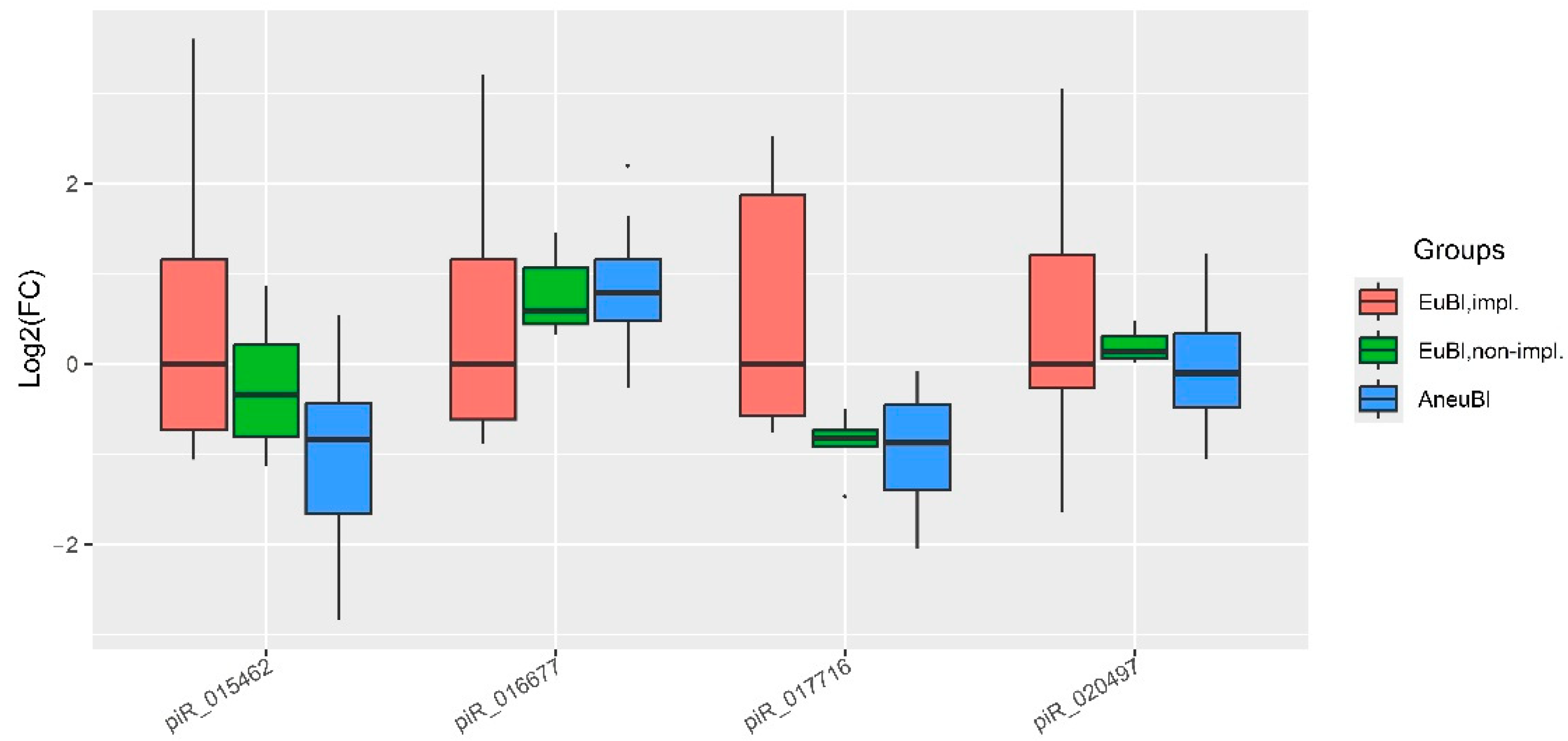
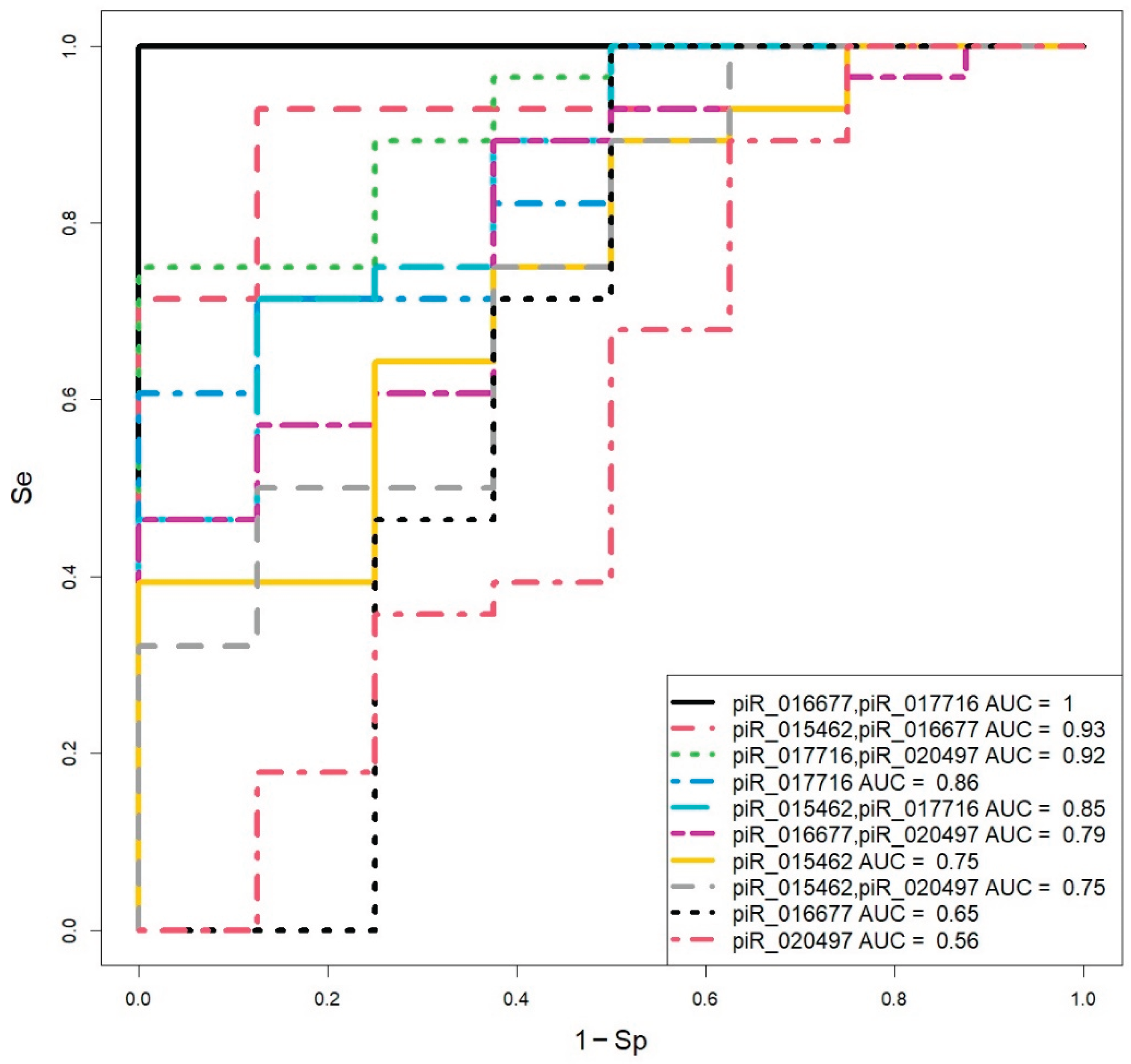
2.3. Assessment of Blastocyst Quality Irrespective of Developmental Rate
| piR_015462 | piR_016677 | piR_017716 | piR_020497 | ||
|---|---|---|---|---|---|
| 5 day EuBl, impl | |||||
| −∆∆Сt | −0.030 | 1.186 | −1.194 | 0.434 | |
| Fold Change* | 0.979 | 2.276 | 0.437 | 1.351 | |
| 6 day EuBl, impl | |||||
| −∆∆Сt | −0.488 | 1.068 | −0.882 | 0.294 | |
| Fold Change* | 0.713 | 2.097 | 0.542 | 1.226 | |
| 5+6 day EuBl, impl | |||||
| −∆∆Сt | −0.258 | 1.186 | −1.180 | 0.403 | |
| Fold Change* | 0.836 | 2.276 | 0.441 | 1.322 | |
| Wald | p_value | 95% CI | OR | Threshold | Se | Sp | Coefficients | |
| (Intercept) | 2.801 | 0.005 | 0.743(0.229;1.276) | 2.103(1.257;3.581) | 0.9338 | 0.4967 | 0.9643 | 0.743 |
| piR_016677 | 4.381 | <0.001 | 1.115(0.645;1.653) | 3.049(1.905;5.221) | 1.115 | |||
| piR_017716 | −3.898 | <0.001 | −0.938(−1.451;−0.496) | 0.391(0.234;0.609) | −0.938 | |||
| (Intercept) | 4.148 | <0.001 | 1.046(0.562;1.557) | 2.845(1.754;4.742) | 0.8451 | 0.6821 | 0.7143 | 1.046 |
| piR_015462 | −2.815 | 0.005 | −0.569(−0.981;−0.184) | 0.566(0.375;0.832) | −0.569 | |||
| piR_016677 | 3.523 | <0.001 | 0.758(0.352;1.202) | 2.135(1.423;3.326) | 0.758 | |||
| (Intercept) | 4.085 | <0.001 | 1.088(0.572;1.625) | 2.97(1.772;5.079) | 0.8505 | 0.6556 | 0.7143 | 1.088 |
| piR_017716 | −3.075 | 0.002 | −0.662(−1.111;−0.255) | 0.516(0.329;0.775) | −0.662 | |||
| piR_020497 | 3.117 | 0.002 | 0.913(0.356;1.516) | 2.491(1.428;4.554) | 0.913 | |||
| (Intercept) | 8.136 | <0.001 | 1.685(1.297;2.112) | 5.393(3.657;8.263) | 0.8524 | 0.5563 | 0.8571 | 1.685 |
| piR_015462 | 0.553 | 0.581 | 0.11(−0.286;0.498) | 1.116(0.751;1.645) | 0.11 | |||
| piR_017716 | −1.002 | 0.316 | −0.177(−0.521;0.179) | 0.837(0.594;1.197) | −0.177 | |||
| (Intercept) | 5.86 | <0.001 | 1.368(0.927;1.847) | 3.928(2.527;6.343) | 0.7991 | 0.8609 | 0.5357 | 1.368 |
| piR_016677 | 2.216 | 0.027 | 0.365(0.062;0.708) | 1.44(1.064;2.031) | 0.365 | |||
| (Intercept) | 5.93 | <0.001 | 1.418(0.966;1.908) | 4.128(2.628;6.74) | 0.8311 | 0.6755 | 0.5714 | 1.418 |
| piR_016677 | 2.443 | 0.015 | 0.582(0.125;1.065) | 1.79(1.134;2.902) | 0.582 | |||
| piR_020497 | −1.257 | 0.209 | −0.384(−0.996;0.209) | 0.681(0.369;1.232) | −0.384 | |||
| (Intercept) | 8.163 | <0.001 | 1.689(1.301;2.116) | 5.415(3.673;8.294) | 0.8576 | 0.3642 | 0.9286 | 1.689 |
| piR_017716 | −0.941 | 0.347 | −0.098(−0.297;0.119) | 0.907(0.743;1.126) | −0.098 | |||
| (Intercept) | 6.825 | <0.001 | 1.569(1.138;2.043) | 4.803(3.121;7.717) | 0.8294 | 0.755 | 0.5714 | 1.569 |
| piR_020497 | 0.978 | 0.328 | 0.177(−0.145;0.566) | 1.193(0.865;1.761) | 0.177 | |||
| (Intercept) | 5.553 | <0.001 | 1.37(0.904;1.877) | 3.935(2.468;6.536) | 0.8399 | 0.6225 | 0.5357 | 1.37 |
| piR_015462 | −1.778 | 0.075 | −0.368(−0.79;0.027) | 0.692(0.454;1.027) | −0.368 | |||
| piR_020497 | 1.994 | 0.046 | 0.516(0.023;1.044) | 1.674(1.023;2.84) | 0.516 | |||
| (Intercept) | 8.183 | <0.001 | 1.689(1.302;2.114) | 5.412(3.675;8.279) | 0.8403 | 0.7483 | 0.3571 | 1.689 |
| piR_015462 | −0.411 | 0.681 | −0.05(−0.28;0.199) | 0.951(0.756;1.22) | −0.05 |
3. Discussion
4. Materials and Methods
4.1. Patient Cohort
4.2. Collection of Spent Embryo Culture Medium Samples
4.3. RNA Isolation from Culture Medium Samples
4.4. Reverse Transcription
4.5. Quantitative Real-Time PCR
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Sun, H.; Gong, T.-T.; Jiang, Y.-T.; Zhang, S.; Zhao, Y.-H.; Wu, Q.-J. Global, Regional, and National Prevalence and Disability-Adjusted Life-Years for Infertility in 195 Countries and Territories, 1990-2017: Results from a Global Burden of Disease Study, 2017. Aging (Albany. NY). 2019, 11, 10952–10991. [Google Scholar] [CrossRef] [PubMed]
- Mani, S.; Mainigi, M. Embryo Culture Conditions and the Epigenome. Semin. Reprod. Med. 2018, 36, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Simopoulou, M.; Sfakianoudis, K.; Rapani, A.; Giannelou, P.; Anifandis, G.; Bolaris, S.; Pantou, A.; Lambropoulou, M.; Pappas, A.; Deligeoroglou, E.; et al. Considerations Regarding Embryo Culture Conditions: From Media to Epigenetics. In Vivo 2018, 32, 451–460. [Google Scholar] [CrossRef]
- Gardner, D.K.; Balaban, B. Assessment of Human Embryo Development Using Morphological Criteria in an Era of Time-Lapse, Algorithms and “OMICS”: Is Looking Good Still Important? Mol. Hum. Reprod. 2016, 22, 704–718. [Google Scholar] [CrossRef]
- Simopoulou, M.; Sfakianoudis, K.; Tsioulou, P.; Rapani, A.; Maziotis, E.; Giannelou, P.; Grigoriadis, S.; Pantou, A.; Nikolettos, K.; Vlahos, N.; et al. Should the Flexibility Enabled by Performing a Day-4 Embryo Transfer Remain as a Valid Option in the IVF Laboratory? A Systematic Review and Network Meta-Analysis. J. Assist. Reprod. Genet. 2019, 36, 1049–1061. [Google Scholar] [CrossRef]
- Li, Y.-X.; Wang, J.; Sun, T.-Z.; Lv, M.-Q.; Ge, P.; Li, H.-N.; Zhou, D.-X. Pregnancy Outcomes after Day 5 versus Day 6 Blastocyst-Stage Embryo Transfer: A Systematic Review and Meta-Analysis. J. Obstet. Gynaecol. Res. 2020, 46, 595–605. [Google Scholar] [CrossRef]
- Korsak, Vladislav S; Smirnova, Anna A; Shurygina, O. V. ART Register of RAHR. Russ. J. Hum. Reprod. 2021, 29, 25–40. [Google Scholar] [CrossRef]
- Fesahat, F.; Montazeri, F.; Hoseini, S.M. Preimplantation Genetic Testing in Assisted Reproduction Technology. J. Gynecol. Obstet. Hum. Reprod. 2020, 49, 101723. [Google Scholar] [CrossRef]
- Franasiak, J.M.; Forman, E.J.; Hong, K.H.; Werner, M.D.; Upham, K.M.; Treff, N.R.; Scott, R.T.J. The Nature of Aneuploidy with Increasing Age of the Female Partner: A Review of 15,169 Consecutive Trophectoderm Biopsies Evaluated with Comprehensive Chromosomal Screening. Fertil. Steril. 2014, 101, 656–663.e1. [Google Scholar] [CrossRef]
- Ubaldi, F.M.; Cimadomo, D.; Capalbo, A.; Vaiarelli, A.; Buffo, L.; Trabucco, E.; Ferrero, S.; Albani, E.; Rienzi, L.; Levi Setti, P.E. Preimplantation Genetic Diagnosis for Aneuploidy Testing in Women Older than 44 Years: A Multicenter Experience. Fertil. Steril. 2017, 107, 1173–1180. [Google Scholar] [CrossRef]
- Homer, H.A. Preimplantation Genetic Testing for Aneuploidy (PGT-A): The Biology, the Technology and the Clinical Outcomes. Aust. N. Z. J. Obstet. Gynaecol. 2019, 59, 317–324. [Google Scholar] [CrossRef] [PubMed]
- Domingo-Muelas, A.; Skory, R.M.; Moverley, A.A.; Ardestani, G.; Pomp, O.; Rubio, C.; Tetlak, P.; Hernandez, B.; Rhon-Calderon, E.A.; Navarro-Sánchez, L.; et al. Human Embryo Live Imaging Reveals Nuclear DNA Shedding during Blastocyst Expansion and Biopsy. Cell 2023, 186, 3166–3181.e18. [Google Scholar] [CrossRef] [PubMed]
- Munné, S.; Kaplan, B.; Frattarelli, J.L.; Child, T.; Nakhuda, G.; Shamma, F.N.; Silverberg, K.; Kalista, T.; Handyside, A.H.; Katz-Jaffe, M.; et al. Preimplantation Genetic Testing for Aneuploidy versus Morphology as Selection Criteria for Single Frozen-Thawed Embryo Transfer in Good-Prognosis Patients: A Multicenter Randomized Clinical Trial. Fertil. Steril. 2019, 112, 1071–1079.e7. [Google Scholar] [CrossRef] [PubMed]
- Salmasi, S.; Heidar, M.S.; Khaksary Mahabady, M.; Rashidi, B.; Mirzaei, H. MicroRNAs, Endometrial Receptivity and Molecular Pathways. Reprod. Biol. Endocrinol. 2024, 22, 139. [Google Scholar] [CrossRef]
- Yang, J.; Yang, L.; Zhou, Y.; Cao, F.; Fang, H.; Ma, H.; Ren, J.; Huang, C.; Diao, L.; Li, Q.; et al. Molecular Subtype of Recurrent Implantation Failure Reveals Distinct Endometrial Etiology of Female Infertility. J. Transl. Med. 2025, 23, 792. [Google Scholar] [CrossRef]
- Gleicher, N.; Patrizio, P.; Mochizuki, L.; Barad, D.H. Previously Reported and Here Added Cases Demonstrate Euploid Pregnancies Followed by PGT-A as “Mosaic” as Well as “Aneuploid” Designated Embryos. Reprod. Biol. Endocrinol. 2023, 21, 25. [Google Scholar] [CrossRef]
- Greco, E.; Minasi, M.G.; Fiorentino, F. Healthy Babies after Intrauterine Transfer of Mosaic Aneuploid Blastocysts. N. Engl. J. Med. 2015, 373, 2089–2090. [Google Scholar] [CrossRef]
- Liu, Y.-L.; Yu, T.-N.; Chen, C.-H.; Wang, P.-H.; Chen, C.-H.; Tzeng, C.-R. Healthy Live Births after Mosaic Blastocyst Transfers with the Use of Next-Generation Sequencing. Taiwan. J. Obstet. Gynecol. 2019, 58, 872–876. [Google Scholar] [CrossRef]
- Toporcerová, S.; Badovská, Z.; Kriváková, E.; Mikulová, V.; Mareková, M.; Altmäe, S.; Rabajdová, M. Embryo Secretome in Predicting Embryo Quality and IVF Treatment Outcome. Reprod. Biomed. Online 2025, 51, 104825. [Google Scholar] [CrossRef]
- Onishi, R.; Yamanaka, S.; Siomi, M.C. PiRNA- and SiRNA-Mediated Transcriptional Repression in Drosophila, Mice, and Yeast: New Insights and Biodiversity. EMBO Rep. 2021, 22, e53062. [Google Scholar] [CrossRef]
- Siomi, M.C.; Sato, K.; Pezic, D.; Aravin, A.A. PIWI-Interacting Small RNAs: The Vanguard of Genome Defence. Nat. Rev. Mol. Cell Biol. 2011, 12, 246–258. [Google Scholar] [CrossRef]
- He, C.; Wang, K.; Gao, Y.; Wang, C.; Li, L.; Liao, Y.; Hu, K.; Liang, M. Roles of Noncoding RNA in Reproduction. Front. Genet. 2021, 12, 777510. [Google Scholar] [CrossRef] [PubMed]
- Cox, D.N.; Chao, A.; Lin, H. Piwi Encodes a Nucleoplasmic Factor Whose Activity Modulates the Number and Division Rate of Germline Stem Cells. Development 2000, 127, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Timofeeva, A. V; Fedorov, I.S.; Shamina, M.A.; Chagovets, V. V; Makarova, N.P.; Kalinina, E.A.; Nazarenko, T.A.; Sukhikh, G.T. Clinical Relevance of Secreted Small Noncoding RNAs in an Embryo Implantation Potential Prediction at Morula and Blastocyst Development Stages. Life (Basel, Switzerland) 2021, 11. [Google Scholar] [CrossRef] [PubMed]
- Timofeeva, A.; Drapkina, Y.; Fedorov, I.; Chagovets, V.; Makarova, N.; Shamina, M.; Kalinina, E.; Sukhikh, G. Small Noncoding RNA Signatures for Determining the Developmental Potential of an Embryo at the Morula Stage. Int. J. Mol. Sci. 2020, 21. [Google Scholar] [CrossRef]
- Timofeeva, A. V.; Chagovets, V. V.; Drapkina, Y.S.; Makarova, N.P.; Kalinina, E.A.; Sukhikh, G.T. Cell-Free, Embryo-Specific SncRNA as a Molecular Biological Bridge between Patient Fertility and IVF Efficiency. Int. J. Mol. Sci. 2019, 20, 2912. [Google Scholar] [CrossRef]
- Timofeeva, Angelika V; Fedorov, Ivan S; Savostina, Guzel V; Ekimov, Alexey N; Perminova, S. G. Quantitative Analysis of PiwiRNAs in the Culture Medium of Euploid and Aneuploid Blastocysts as an Additional Method of Selecting a High-Quality Embryo for Transfer to the Uterine Cavity in Assisted Reproductive Technology Programs. Obstet. Gynecol. 2023, 11, 115–130. [Google Scholar] [CrossRef]
- Gholamy, Afshin; Kreinovich, Vladik; and Kosheleva, O. Why 70/30 or 80/20 Relation Between Training and Testing Sets: A Pedagogical Explanation. Dep. Tech. Reports 2018, 1209.
- Kuznyetsov, V.; Madjunkova, S.; Antes, R.; Abramov, R.; Motamedi, G.; Ibarrientos, Z.; Librach, C. Evaluation of a Novel Non-Invasive Preimplantation Genetic Screening Approach. PLoS One 2018, 13, e0197262. [Google Scholar] [CrossRef]
- Handayani, N.; Aubry, D.; Boediono, A.; Wiweko, B.; Sirait, B.; Sini, I.; Polim, A.A.; Dwiranti, A.; Bowolaksono, A. The Origin and Possible Mechanism of Embryonic Cell-Free DNA Release in Spent Embryo Culture Media: A Review. J. Assist. Reprod. Genet. 2023, 40, 1231–1242. [Google Scholar] [CrossRef]
- Del Collado, M.; Andrade, G.M.; Gonçalves, N.J.N.; Fortini, S.; Perecin, F.; Carriero, M.M. The Embryo Non-Invasive Pre-Implantation Diagnosis Era: How Far Are We? Anim. Reprod. 2023, 20, e20230069. [Google Scholar] [CrossRef]
- Voros, C.; Darlas, M.; Athanasiou, D.; Athanasiou, A.; Athanasiou, A.; Bananis, K.; Papadimas, G.; Tsimpoukelis, C.; Gkirgkinoudis, A.; Sapantzoglou, I.; et al. Evaluation of the Effectiveness and Accuracy of Non-Invasive Preimplantation Genetic Testing (NiPGT) Compared to Invasive Embryo Biopsy. Biomedicines 2025, 13. [Google Scholar] [CrossRef]
- Kobayashi, M.; Kobayashi, J.; Shirasuna, K.; Iwata, H. Abundance of Cell-Free Mitochondrial DNA in Spent Culture Medium Associated with Morphokinetics and Blastocyst Collapse of Expanded Blastocysts. Reprod. Med. Biol. 2020, 19, 404–414. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Ji, H.; Shi, J.; Wang, L.; Ding, L.; Jiang, Y.; Huang, X.; Qiu, P.; Li, P. Digital PCR Detection of MtDNA/GDNA Ratio in Embryo Culture Medium for Prediction of Embryo Development Potential. Pharmgenomics. Pers. Med. 2021, 14, 521–531. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Sun, Y.; Dong, X.; Zhou, J.; Sun, F.; Han, T.; Lei, P.; Mao, R.; Guo, X.; Wang, Q.; et al. Mitochondrial DNA and Genomic DNA Ratio in Embryo Culture Medium Is Not a Reliable Predictor for in Vitro Fertilization Outcome. Sci. Rep. 2019, 9, 5378. [Google Scholar] [CrossRef]
- Wyroba, J.; Kochan, J.; Barszcz, M.; Mirocki, G.; Kordowitzki, P. Reproductive Aging, Preimplantation Genetic Testing for Aneuploidy, and the Diameter of Blastocysts: Does Size Matter? Aging (Albany. NY). 2025, 17, 630–642. [Google Scholar] [CrossRef]
- Chen, X.; Chen, H.; Huang, G.; Lin, R.; Zhou, H.; Fan, X.; Wu, Z.; Xu, K.; Shu, J.; Wang, C. Morphokinetic Parameter S2 Predicts Early Embryo Developmental Potential and Associated Metabolic Pathways. J. Assist. Reprod. Genet. 2025. [Google Scholar] [CrossRef]
- Carvalho, B.R. de; Aires, L.S.C.; Lima, M.F.A.; Bittar, M.V.; Cabral, Í. de O.; Pinheiro Junior, L.C.; Silva, A.A. Prediction of Human Embryo Ploidy Based on Morphokinetic Parameters: Investigating the Use of Time-Lapse Imaging for Non-Invasive Embryo Selection. JBRA Assist. Reprod. 2025, 29, 488–492. [Google Scholar] [CrossRef]
- Liu, X.; Lou, H.; Zhang, J.; Du, M.; Du, Y.; Wu, S.; Guan, Y.; Liu, J. Clinical Outcome Analysis of Frozen-Thawed Embryo Transfer on Day 7. Front. Endocrinol. (Lausanne). 2022, 13, 1082597. [Google Scholar] [CrossRef]
- Lane, S.L.; Reed, L.; Schoolcraft, W.B.; Katz-Jaffe, M.G. Euploid Day 7 Blastocysts of Infertility Patients with Only Slow Embryo Development Have Reduced Implantation Potential. Reprod. Biomed. Online 2022, 44, 858–865. [Google Scholar] [CrossRef]
- Wang, X.; Ramat, A.; Simonelig, M.; Liu, M.-F. Emerging Roles and Functional Mechanisms of PIWI-Interacting RNAs. Nat. Rev. Mol. Cell Biol. 2023, 24, 123–141. [Google Scholar] [CrossRef]
- Hancks, D.C.; Kazazian, H.H. Roles for Retrotransposon Insertions in Human Disease. Mob. DNA 2016, 7, 9. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, R.; Yu, J. New Understanding of the Relevant Role of LINE-1 Retrotransposition in Human Disease and Immune Modulation. Front. cell Dev. Biol. 2020, 8, 657. [Google Scholar] [CrossRef]
- Malki, S.; van der Heijden, G.W.; O’Donnell, K.A.; Martin, S.L.; Bortvin, A. A Role for Retrotransposon LINE-1 in Fetal Oocyte Attrition in Mice. Dev. Cell 2014, 29, 521–533. [Google Scholar] [CrossRef]
- Protasova, M.S.; Andreeva, T.V.; Rogaev, E.I. Factors Regulating the Activity of LINE1 Retrotransposons. Genes (Basel). 2021, 12. [Google Scholar] [CrossRef]
- Gou, L.-T.; Dai, P.; Yang, J.-H.; Xue, Y.; Hu, Y.-P.; Zhou, Y.; Kang, J.-Y.; Wang, X.; Li, H.; Hua, M.-M.; et al. Pachytene PiRNAs Instruct Massive MRNA Elimination during Late Spermiogenesis. Cell Res. 2014, 24, 680–700. [Google Scholar] [CrossRef] [PubMed]
- Goh, W.S.S.; Falciatori, I.; Tam, O.H.; Burgess, R.; Meikar, O.; Kotaja, N.; Hammell, M.; Hannon, G.J. PiRNA-Directed Cleavage of Meiotic Transcripts Regulates Spermatogenesis. Genes Dev. 2015, 29, 1032–1044. [Google Scholar] [CrossRef] [PubMed]
- Zitouni, S.; Nabais, C.; Jana, S.C.; Guerrero, A.; Bettencourt-Dias, M. Polo-like Kinases: Structural Variations Lead to Multiple Functions. Nat. Rev. Mol. Cell Biol. 2014, 15, 433–452. [Google Scholar] [CrossRef] [PubMed]
- Remo, A.; Li, X.; Schiebel, E.; Pancione, M. The Centrosome Linker and Its Role in Cancer and Genetic Disorders. Trends Mol. Med. 2020, 26, 380–393. [Google Scholar] [CrossRef]
- Cunningham, C.E.; Li, S.; Vizeacoumar, F.S.; Bhanumathy, K.K.; Lee, J.S.; Parameswaran, S.; Furber, L.; Abuhussein, O.; Paul, J.M.; McDonald, M.; et al. Therapeutic Relevance of the Protein Phosphatase 2A in Cancer. Oncotarget 2016, 7, 61544–61561. [Google Scholar] [CrossRef]
- Wigley, W.; Fabunmi, R.; Lee, M.G.; Marino, C.; Muallem, S.; DeMartino, G.; Thomas, P. Dynamic Association of Proteasomal Machinery with the Centrosome. J. Cell Biol. 1999, 145, 481–490. [Google Scholar] [CrossRef]
- Lilienbaum, A. Relationship between the Proteasomal System and Autophagy. Int. J. Biochem. Mol. Biol. 2013, 4, 1–26. [Google Scholar]
- Gerhardt, C.; Lier, J.M.; Burmühl, S.; Struchtrup, A.; Deutschmann, K.; Vetter, M.; Leu, T.; Reeg, S.; Grune, T.; Rüther, U. The Transition Zone Protein Rpgrip1l Regulates Proteasomal Activity at the Primary Cilium. J. Cell Biol. 2015, 210, 115–133. [Google Scholar] [CrossRef]
- Holt, J.E.; Tran, S.M.-T.; Stewart, J.L.; Minahan, K.; García-Higuera, I.; Moreno, S.; Jones, K.T. The APC/C Activator FZR1 Coordinates the Timing of Meiotic Resumption during Prophase I Arrest in Mammalian Oocytes. Development 2011, 138, 905–913. [Google Scholar] [CrossRef] [PubMed]
- Rattani, A.; Ballesteros Mejia, R.; Roberts, K.; Roig, M.B.; Godwin, J.; Hopkins, M.; Eguren, M.; Sanchez-Pulido, L.; Okaz, E.; Ogushi, S.; et al. APC/C(Cdh1) Enables Removal of Shugoshin-2 from the Arms of Bivalent Chromosomes by Moderating Cyclin-Dependent Kinase Activity. Curr. Biol. 2017, 27, 1462–1476.e5. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, L.; Ma, Y.; Wang, Y.; Liu, H.; Liu, M.; Qin, L.; Li, J.; Jiang, C.; Zhang, X.; et al. CEP128 Is Involved in Spermatogenesis in Humans and Mice. Nat. Commun. 2022, 13, 1395. [Google Scholar] [CrossRef] [PubMed]
- Pitaval, A.; Senger, F.; Letort, G.; Gidrol, X.; Guyon, L.; Sillibourne, J.; Théry, M. Microtubule Stabilization Drives 3D Centrosome Migration to Initiate Primary Ciliogenesis. J. Cell Biol. 2017, 216, 3713–3728. [Google Scholar] [CrossRef]
- Yang, Z.; Gallicano, G.I.; Yu, Q.C.; Fuchs, E. An Unexpected Localization of Basonuclin in the Centrosome, Mitochondria, and Acrosome of Developing Spermatids. J. Cell Biol. 1997, 137, 657–669. [Google Scholar] [CrossRef]
- Zhang, X.; Chou, W.; Haig-Ladewig, L.; Zeng, W.; Cao, W.; Gerton, G.; Dobrinski, I.; Tseng, H. BNC1 Is Required for Maintaining Mouse Spermatogenesis. Genesis 2012, 50, 517–524. [Google Scholar] [CrossRef]
- Ma, J.; Zeng, F.; Schultz, R.M.; Tseng, H. Basonuclin: A Novel Mammalian Maternal-Effect Gene. Development 2006, 133, 2053–2062. [Google Scholar] [CrossRef]
- Mao, B.-P.; Ge, R.; Cheng, C.Y. Role of Microtubule +TIPs and -TIPs in Spermatogenesis - Insights from Studies of Toxicant Models. Reprod. Toxicol. 2020, 91, 43–52. [Google Scholar] [CrossRef]
- Dubois, F.; Bergot, E.; Zalcman, G.; Levallet, G. RASSF1A, Puppeteer of Cellular Homeostasis, Fights Tumorigenesis, and Metastasis-an Updated Review. Cell Death Dis. 2019, 10, 928. [Google Scholar] [CrossRef]
- Lim, H.Y.G.; Plachta, N. Cytoskeletal Control of Early Mammalian Development. Nat. Rev. Mol. Cell Biol. 2021, 22, 548–562. [Google Scholar] [CrossRef] [PubMed]
- Herreros, L.; Rodríguez-Fernandez, J.L.; Brown, M.C.; Alonso-Lebrero, J.L.; Cabañas, C.; Sánchez-Madrid, F.; Longo, N.; Turner, C.E.; Sánchez-Mateos, P. Paxillin Localizes to the Lymphocyte Microtubule Organizing Center and Associates with the Microtubule Cytoskeleton. J. Biol. Chem. 2000, 275, 26436–26440. [Google Scholar] [CrossRef] [PubMed]
- Ezoe, K.; Miki, T.; Ohata, K.; Fujiwara, N.; Yabuuchi, A.; Kobayashi, T.; Kato, K. Prolactin Receptor Expression and Its Role in Trophoblast Outgrowth in Human Embryos. Reprod. Biomed. Online 2021, 42, 699–707. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, S.; Mui, V.J.; Rosenberg, S.K.; Homma, K.; Cheatham, M.A.; Zheng, J. Cadherin 23-C Regulates Microtubule Networks by Modifying CAMSAP3’s Function. Sci. Rep. 2016, 6, 28706. [Google Scholar] [CrossRef]
- Meng, W.; Mushika, Y.; Ichii, T.; Takeichi, M. Anchorage of Microtubule Minus Ends to Adherens Junctions Regulates Epithelial Cell-Cell Contacts. Cell 2008, 135, 948–959. [Google Scholar] [CrossRef]
- Embryology, A.S. in R.M. and E.S.I.G. of The Istanbul Consensus Workshop on Embryo Assessment: Proceedings of an Expert Meeting†. Hum. Reprod. 2011, 26, 1270–1283. [Google Scholar] [CrossRef]
- Team, R.C. A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria Available online:. Available online: https://www.r-project.org (accessed on 10 October 2025).
- Team, Rs. RStudio: Integrated Development for R. Available online: http://www.rstudio.com/. (accessed on 10 October 2025).
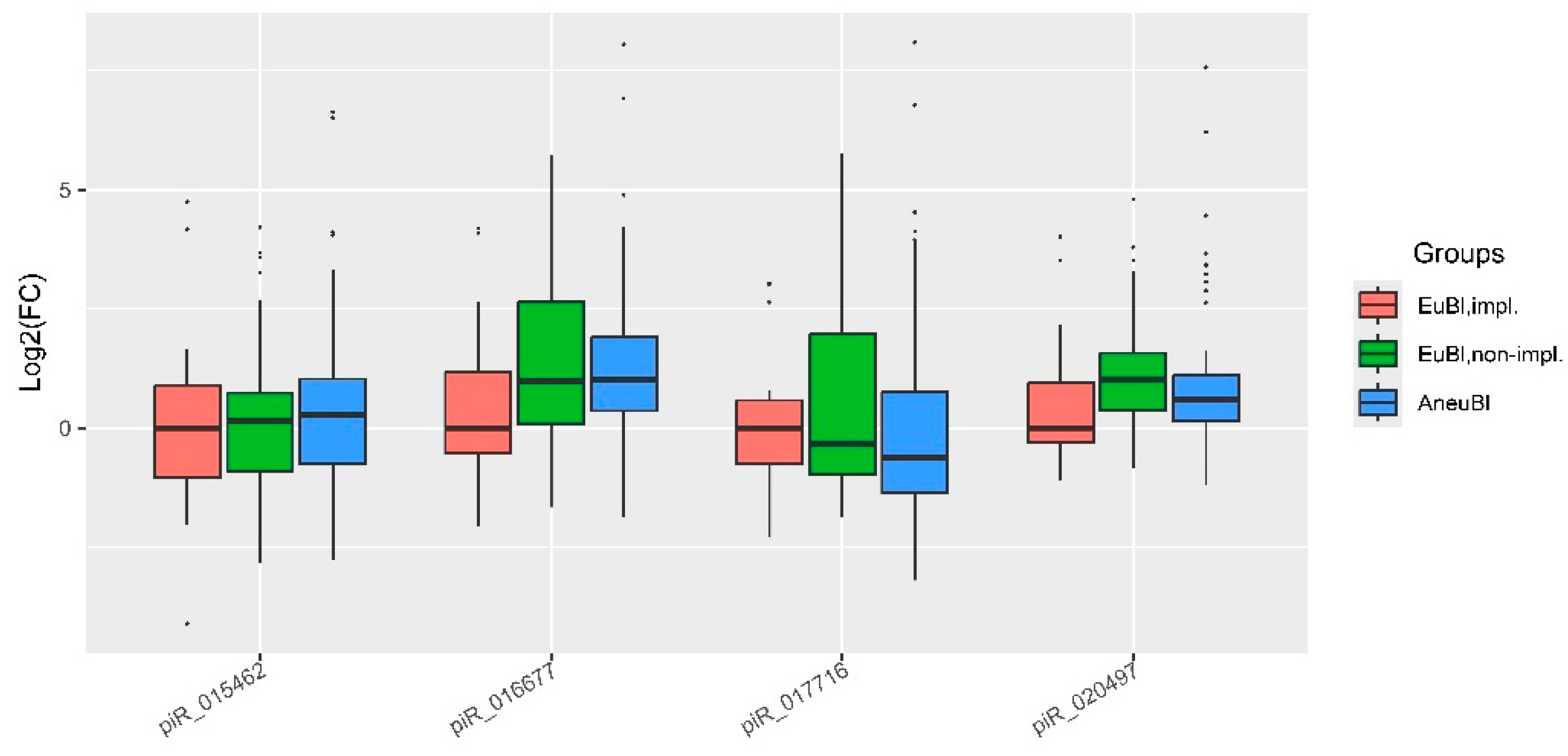
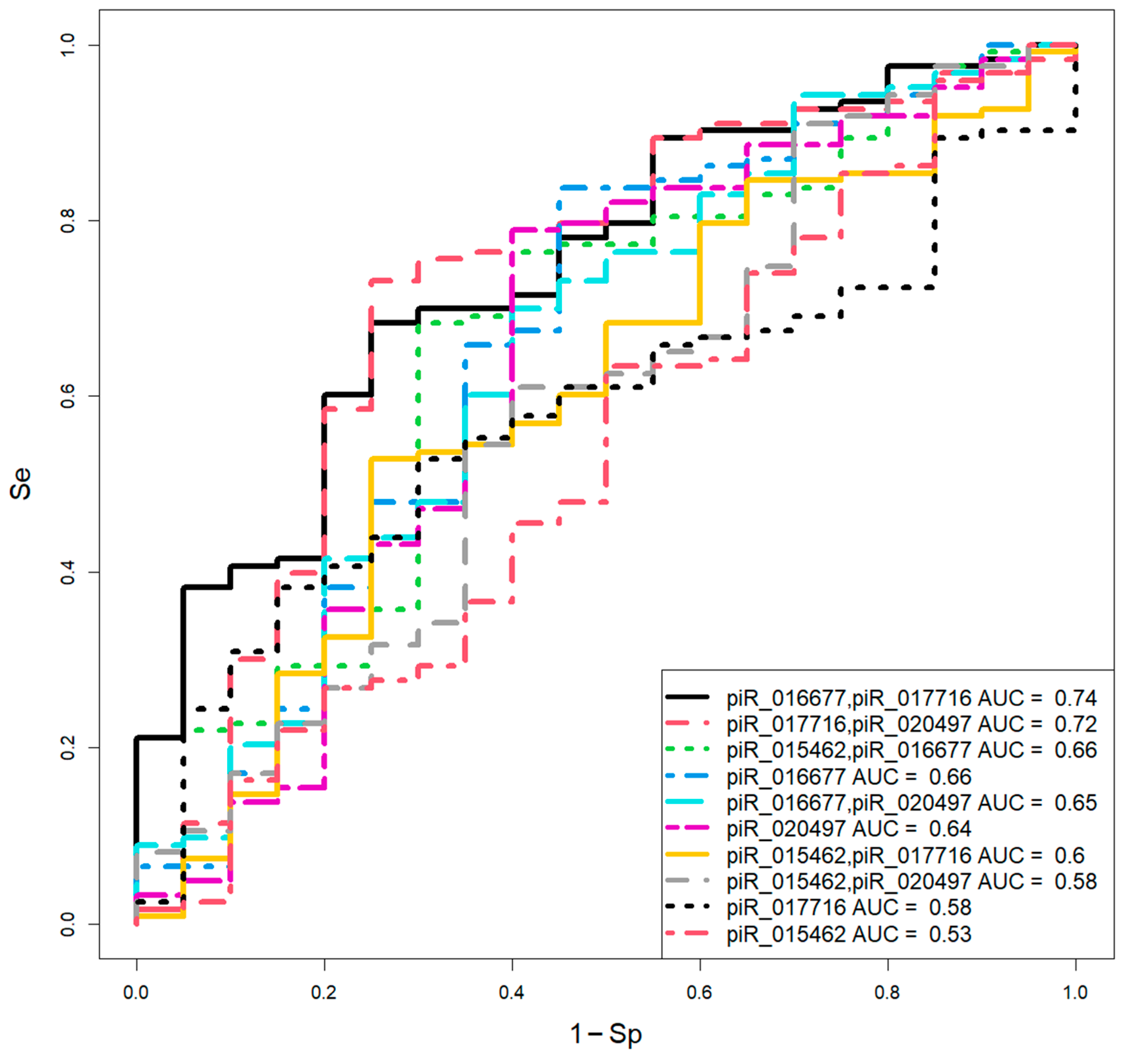
| piwiRNA | Group | Me(Q1;Q3)* | EuBl,impl** | EuBl,non-impl** | AneuBl** |
|---|---|---|---|---|---|
| piR_016677 | EuBl,impl | 0(−0.54;1.17) | 1 | 0.04 | 0.02 |
| EuBl,non-impl | 0.99(0.1;2.65) | 0.04 | 1 | 0.9 | |
| AneuBl | 1.01(0.36;1.9) | 0.02 | 0.9 | 1 | |
| piR_017716 | EuBl,impl | 0(−0.74;0.58) | 1 | 0.78 | 0.16 |
| EuBl,non-impl | −0.34(−0.98;1.98) | 0.78 | 1 | 0.15 | |
| AneuBl | −0.62(−1.35;0.75) | 0.16 | 0.15 | 1 | |
| piR_020497 | EuBl,impl | 0(−0.29;0.95) | 1 | 0.02 | 0.1 |
| EuBl,non-impl. | 1.02(0.37;1.57) | 0.02 | 1 | 0.04 | |
| AneuBl | 0.6(0.15;1.1) | 0.1 | 0.04 | 1 | |
| piR_015462 | EuBl,impl | 0(−1.04;0.9) | 1 | 0.9 | 0.5 |
| EuBl,non-impl | 0.16(−0.91;0.74) | 0.9 | 1 | 0.48 | |
| AneuBl | 0.27(−0.76;1.03) | 0.5 | 0.48 | 1 |
| Wald | p_value | 95%CI | OR | Threshold | Se | Sp | Coefficients | |
|---|---|---|---|---|---|---|---|---|
| (Intercept) | 3.594 | <0.001 | 1.055(0.494;1.655) | 2.873(1.639;5.232) | 0.84 | 0.69 | 0.75 | 1.055 |
| piR_016677 | 3.364 | <0.001 | 0.967(0.441;1.586) | 2.63(1.554;4.883) | 0.967 | |||
| piR_017716 | −2.743 | 0.006 | −0.769(−1.376;−0.259) | 0.464(0.253;0.772) | −0.769 | |||
| (Intercept) | 3.658 | <0.001 | 1.157(0.546;1.801) | 3.179(1.727;6.053) | 0.85 | 0.73 | 0.75 | 1.157 |
| piR_017716 | −2.43 | 0.015 | −0.578(−1.074;−0.121) | 0.561(0.341;0.886) | −0.578 | |||
| piR_020497 | 2.782 | 0.005 | 0.955(0.302;1.667) | 2.598(1.353;5.298) | 0.955 | |||
| (Intercept) | 5.02 | <0.001 | 1.388(0.868;1.962) | 4.009(2.383;7.112) | 0.85 | 0.68 | 0.71 | 1.388 |
| piR_015462 | −1.211 | 0.226 | −0.269(−0.719;0.16) | 0.764(0.487;1.174) | −0.269 | |||
| piR_016677 | 2.344 | 0.019 | 0.542(0.103;1.017) | 1.719(1.109;2.764) | 0.542 | |||
| (Intercept) | 5.577 | <0.001 | 1.494(0.992;2.05) | 4.453(2.697;7.767) | 0.82 | 0.84 | 0.55 | 1.494 |
| piR_016677 | 2.027 | 0.043 | 0.364(0.037;0.742) | 1.439(1.037;2.101) | 0.364 | |||
| (Intercept) | 5.414 | <0.001 | 1.526(0.999;2.111) | 4.599(2.717;8.261) | 0.84 | 0.71 | 0.61 | 1.526 |
| piR_016677 | 1.638 | 0.101 | 0.442(−0.073;0.993) | 1.557(0.929;2.698) | 0.443 | |||
| piR_020497 | −0.39 | 0.697 | −0.144(−0.878;0.576) | 0.866(0.416;1.778) | −0.144 | |||
| (Intercept) | 5.752 | <0.001 | 1.589(1.074;2.164) | 4.899(2.928;8.709) | 0.83 | 0.79 | 0.61 | 1.589 |
| piR_020497 | 1.35 | 0.177 | 0.308(−0.089;0.805) | 1.36(0.915;2.237) | 0.308 | |||
| (Intercept) | 7.314 | <0.001 | 1.782(1.329;2.289) | 5.94(3.776;9.866) | 0.87 | 0.53 | 0.75 | 1.782 |
| piR_015462 | 1.249 | 0.212 | 0.281(−0.164;0.725) | 1.324(0.849;2.066) | 0.281 | |||
| piR_017716 | −1.1 | 0.271 | −0.222(−0.617;0.184) | 0.801(0.539;1.202) | −0.222 | |||
| (Intercept) | 5.238 | <0.001 | 1.523(0.981;2.131) | 4.584(2.668;8.423) | 0.81 | 0.91 | 0.32 | 1.523 |
| piR_015462 | −0.639 | 0.523 | −0.145(−0.604;0.291) | 0.865(0.547;1.338) | −0.145 | |||
| piR_020497 | 1.444 | 0.149 | 0.438(−0.137;1.061) | 1.549(0.872;2.888) | 0.438 | |||
| (Intercept) | 7.515 | <0.001 | 1.811(1.37;2.325) | 6.17(3.935;10.223) | 0.86 | 0.38 | 0.85 | 1.82 |
| piR_017716 | −0.179 | 0.858 | −0.022(−0.250;0.241) | 0.978(0.778;1.273) | −0.022 | |||
| (Intercept) | 7.406 | <0.001 | 1.795(1.344;2.299) | 6.019(3.835;9.969) | 0.85 | 0.63 | 0.51 | 1.795 |
| piR_015462 | 0.618 | 0.537 | 0.091(−0.182;0.399) | 1.095(0.833;1.490) | 0.091 |
| piwiRNA | group | Me(Q1;Q3)* | EuBl, impl** | EuBl, non-impl** | AneuBl** |
|---|---|---|---|---|---|
| piR_016677 | EuBl, impl | 0(−0.61;1.16) | 1 | 0.5 | 0.2 |
| EuBl, non-impl | 0.58(0.44;1.06) | 0.5 | 1 | 0.67 | |
| AneuBl | 0.78(0.47;1.16) | 0.2 | 0.67 | 1 | |
| piR_017716 | EuBl, impl | 0(−0.57;1.87) | 1 | 0.01 | 0.003 |
| EuBl, non-impl | −0.82(−0.91;−0.74) | 0.01 | 1 | 0.97 | |
| AneuBl | −0.88(−1.4;−0.45) | 0.003 | 0.97 | 1 | |
| piR_020497 | EuBl, impl | 0(−0.27;1.21) | 1 | 0.85 | 0.4 |
| EuBl, non-impl | 0.13(0.06;0.31) | 0.85 | 1 | 0.14 | |
| AneuBl | −0.1(−0.48;0.34) | 0.4 | 0.14 | 1 | |
| piR_015462 | EuBl, impl | 0(−0.73;1.15) | 1 | 0.57 | 0.015 |
| EuBl, non-impl | −0.34(−0.8;0.22) | 0.57 | 1 | 0.1 | |
| AneuBl | −0.84(−1.66;−0.44) | 0.015 | 0.1 | 1 |
| Wald | p_value | 95%CI | OR | Threshold | Se | Sp | Coefficients | |
|---|---|---|---|---|---|---|---|---|
| (Intercept) | −0.001 | 0.999 | −179.213(−82930.486;20089.428) | 1.47E−78(0;Inf) | 0.5 | 1 | 1 | −179.213 |
| piR_016677 | 0.001 | 0.999 | 131.019(−4729.481;4991.519) | 7.96E+56(0;Inf) | 131.019 | |||
| piR_017716 | −0.002 | 0.999 | −177.891(−73828.946;−162647.348) | 5.53E−78(0;0) | −177.891 | |||
| (Intercept) | −0.973 | 0.331 | −0.891(−3.013;0.701) | 0.41(0.049;2.034) | 0.75 | 0.93 | 0.88 | −0.891 |
| piR_015462 | −2.945 | 0.003 | −2.214(−4.068;−1.008) | 0.109(0.017;0.365) | −2.214 | |||
| piR_016677 | 2.467 | 0.014 | 2.537(0.872;5.031) | 12.646(2.392;153.113) | 2.537 | |||
| (Intercept) | −0.877 | 0.381 | −1.072(−4.125;0.898) | 0.342(0.016;2.455) | 0.83 | 0.75 | 1 | −1.072 |
| piR_017716 | −2.027 | 0.043 | −5.369(−12.553;−1.741) | 0.005(3.54E−06;0.175) | −5.369 | |||
| piR_020497 | 1.909 | 0.051 | 2.088(0.193;4.735) | 8.061(1.213;113.859) | 2.088 | |||
| (Intercept) | 0.179 | 0.858 | 0.139(−1.554;1.505) | 1.149(0.211;4.505) | 0.89 | 0.61 | 1 | 0.139 |
| piR_017716 | −2.058 | 0.04 | −2.476(−5.52;−0.847) | 0.084(0.004;0.429) | −2.476 | |||
| (Intercept) | 0.135 | 0.893 | 0.107(−1.609;1.499) | 1.113(0.201;4.479) | 0.86 | 0.71 | 0.88 | 0.107 |
| piR_015462 | 0.431 | 0.667 | 0.407(−1.36;2.496) | 1.502(0.257;12.138) | 0.407 | |||
| piR_017716 | −1.758 | 0.079 | −2.911(−7.128;−0.595) | 0.054(0.001;0.551) | −2.911 | |||
| (Intercept) | 1.247 | 0.212 | 0.683(−0.412;1.797) | 1.981(0.662;6.034) | 0.7 | 0.89 | 0.62 | 0.683 |
| piR_016677 | 1.991 | 0.047 | 1.438(0.159;3.109) | 4.213(1.172;22.402) | 1.438 | |||
| piR_020497 | −2.184 | 0.029 | −1.606(−3.289;−0.307) | 0.201(0.037;0.736) | −1.606 | |||
| (Intercept) | 2.044 | 0.041 | 0.962(0.046;1.937) | 2.618(1.047;6.936) | 0.67 | 0.89 | 0.5 | 0.962 |
| piR_015462 | −2.112 | 0.035 | −0.992(−2.174;−0.25) | 0.371(0.114;0.779) | −0.992 | |||
| (Intercept) | 1.665 | 0.096 | 0.811(−0.145;1.815) | 2.251(0.865;6.139) | 0.68 | 0.89 | 0.5 | 0.811 |
| piR_015462 | −2.227 | 0.026 | −1.394(−2.902;−0.321) | 0.248(0.055;0.726) | −1.394 | |||
| piR_020497 | 1.009 | 0.313 | 0.735(−0.633;2.436) | 2.086(0.531;11.43) | 0.735 | |||
| (Intercept) | 1.991 | 0.046 | 0.987(0.056;2.048) | 2.684(1.058;7.753) | 0.7 | 1 | 0.5 | 0.987 |
| piR_016677 | 0.815 | 0.415 | 0.413(−0.529;1.498) | 1.512(0.589;4.471) | 0.413 | |||
| (Intercept) | 3.179 | 0.002 | 1.377(0.593;2.323) | 3.965(1.809;10.209) | 0.9 | 1 | 0.13 | 1.378 |
| piR_020497 | −1.329 | 0.184 | −0.591(−1.573;0.271) | 0.553(0.207;1.311) | −0.591 |
| Blastocyst quality | piR_016677 | piR_017716 | Model Result (Treshold=0.5) |
|---|---|---|---|
| EuBl, impl | 2.313089 | 2.359285 | 6E−122 |
| EuBl, impl | −0.5407 | 0.318697 | 1E−120 |
| EuBl, non−impl | 0.320295 | −0.89333 | 1 |
| AneuBl | −0.04268 | −2.0471 | 1 |
| AneuBl | 1.12153 | −0.5075 | 1 |
| AneuBl | −0.26189 | −1.36328 | 1 |
| AneuBl | 1.100154 | −0.31455 | 1 |
| AneuBl | 2.194816 | −0.11771 | 1 |
| piwiРНК 1 | Identification number 1 | Nucleotide sequence of the sense primer for PCR, 5′-3′ |
|---|---|---|
| hsa_piR_015462 | DQ591122 | tgtcctgggccagcctgatgatgtcctcctc |
| hsa_piR_016677 | DQ592953 | cccctggtggtctagtggttaggattcggc |
| hsa_piR_020497 | DQ598177 | ggggggtgtagctcagtggtagagcgcgtgct |
| hsa_piR_017716 | DQ594453 | ttccctggtggtctagtggttaggattcggc |
| hsa_piR_022258 | DQ600471 | tactacctgattggtcgggtgtgagc |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




