Submitted:
12 November 2025
Posted:
13 November 2025
You are already at the latest version
Abstract
Sorghum, the fifth major global cereal, has potential as a source crop in temperate regions. To completely use sorghum bagasse, the ideal enzyme cocktail aims to identify and select the contributed enzymatic system. This study investigated the enzymatic system of Clostridium cellulovorans cellulosome and noncellulosomal enzymes using SDS-PAGE and LC-MS/MS. Enzyme solutions from treated and untreated sorghum bagasse were prepared and compared based on CMCase activity. As a result, the enzyme solution derived from untreated sorghum bagasse had the highest activity. Protein bands from each C. cellulovorans culture showed distinct patterns on SDS-PAGE examination: three enzyme fractions, including culture supernatants, crystalline cellulose (Avicel) bound, and unbound fractions. These results suggested that untreated sorghum bagasse induced a variety of cellulosomal and uncellulosomal proteins. On the other hand, 5% or 10% sorghum supernatants could not induce Avicel-bound proteins, including the cellulosome, although even 5% sorghum juice induced three major bands: 180-kDa, 100-kDa, and 70-kDa, respectively. In contrast, cellobiose induced three major bands, while the total number of all isolated proteins from the cellobiose medium was the most limited among all culture media. More intriguingly, our investigation detected one cellulosomal protein, HbpA and three noncellulosomal enzymes, indicating that glycosidase GH130 was identified as a biomass-induced enzyme in good accord with previously published proteomic studies. Therefore, the development of an enzyme cocktail for degrading target biomass using artificial intelligence and machine learning might be possible in the near future.
Keywords:
1. Introduction
2. Results
2.1. Evaluation of Chemical Pretreatment of Sorghum Bagasse Based on CMCase Activity
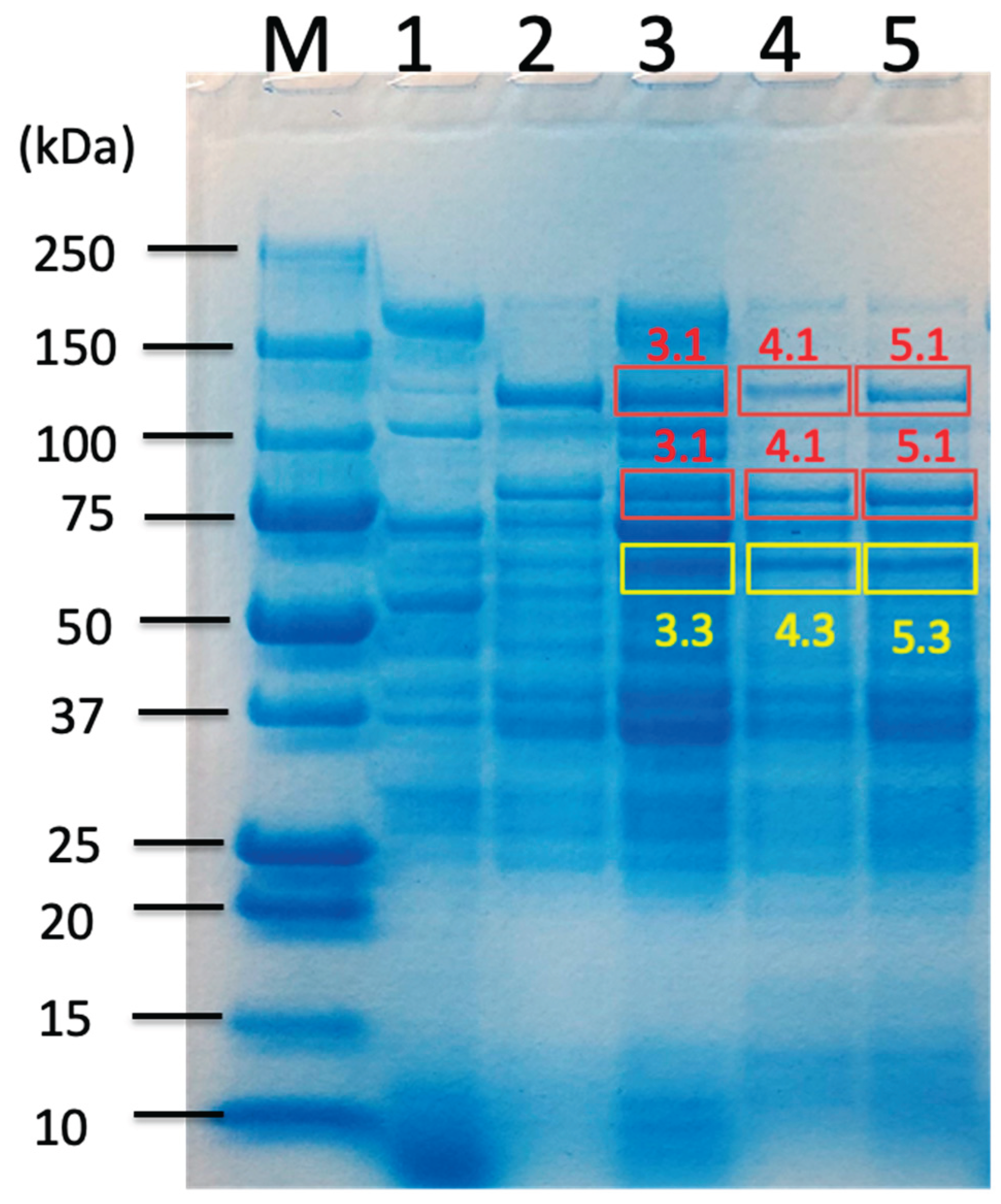
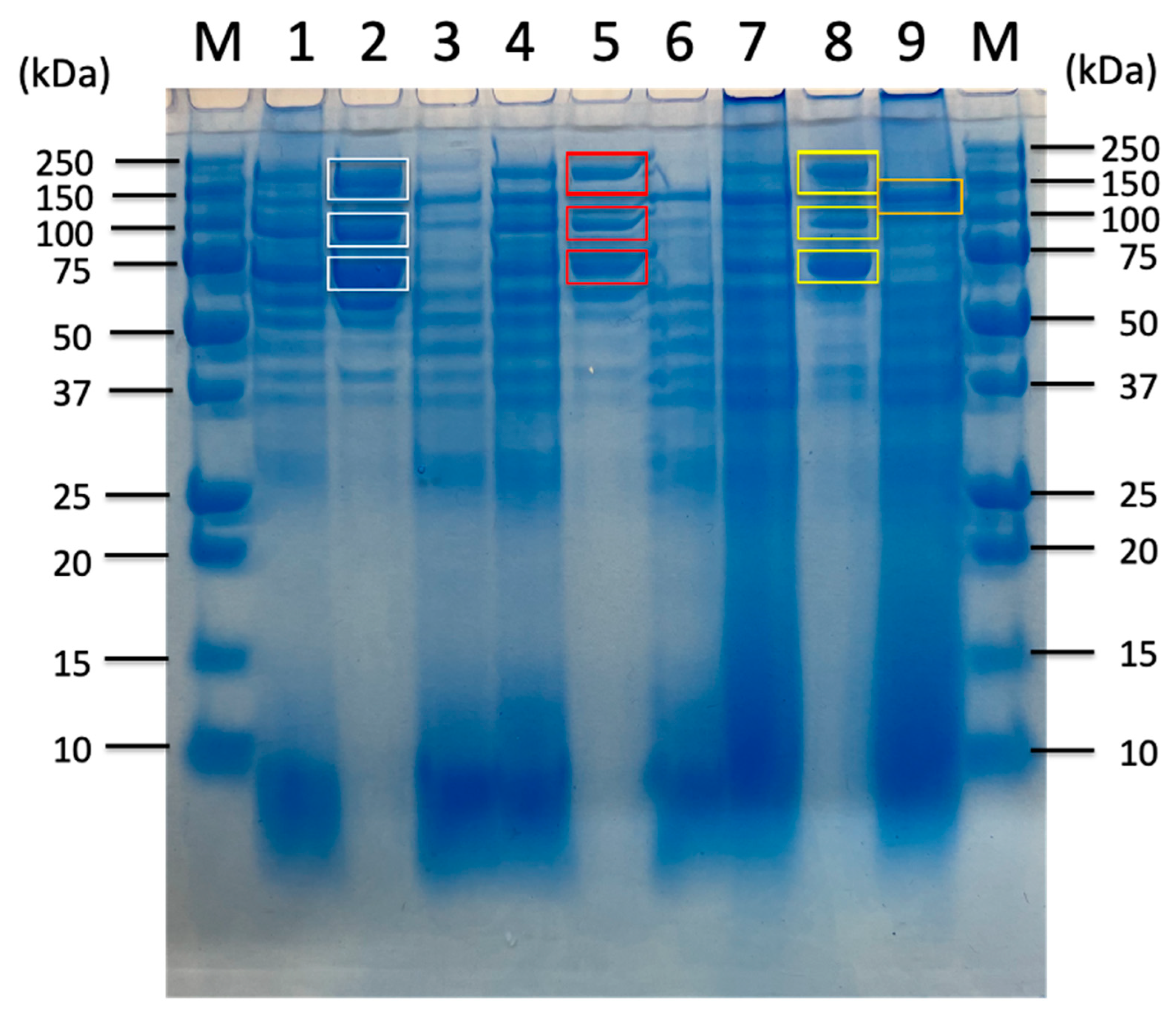
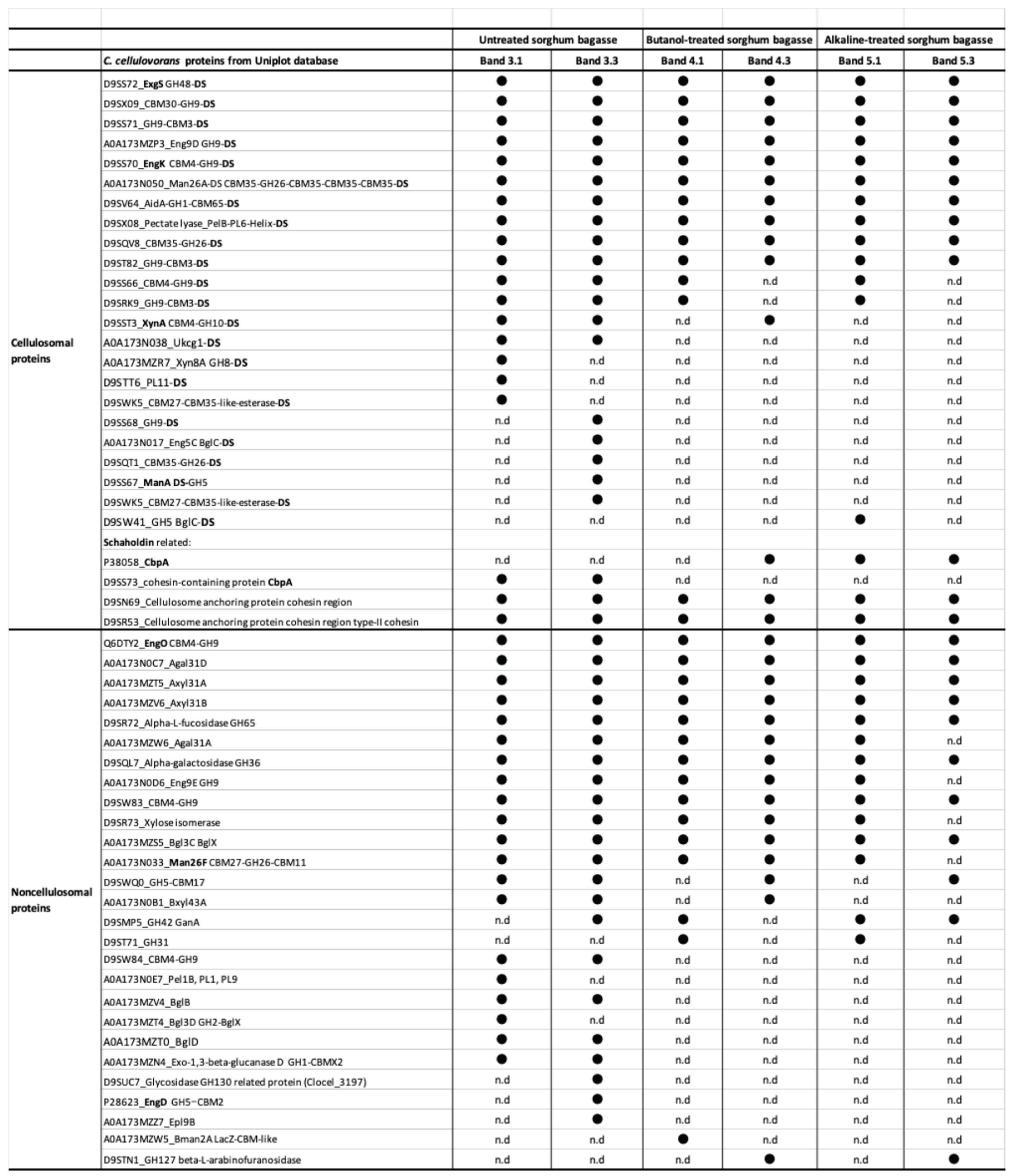
2.2. Proteomic Analysis of Carbohydrate-Related Proteins and Their Comparison Based on Sorghum Bagasse with or Without Chemical Pretreatments
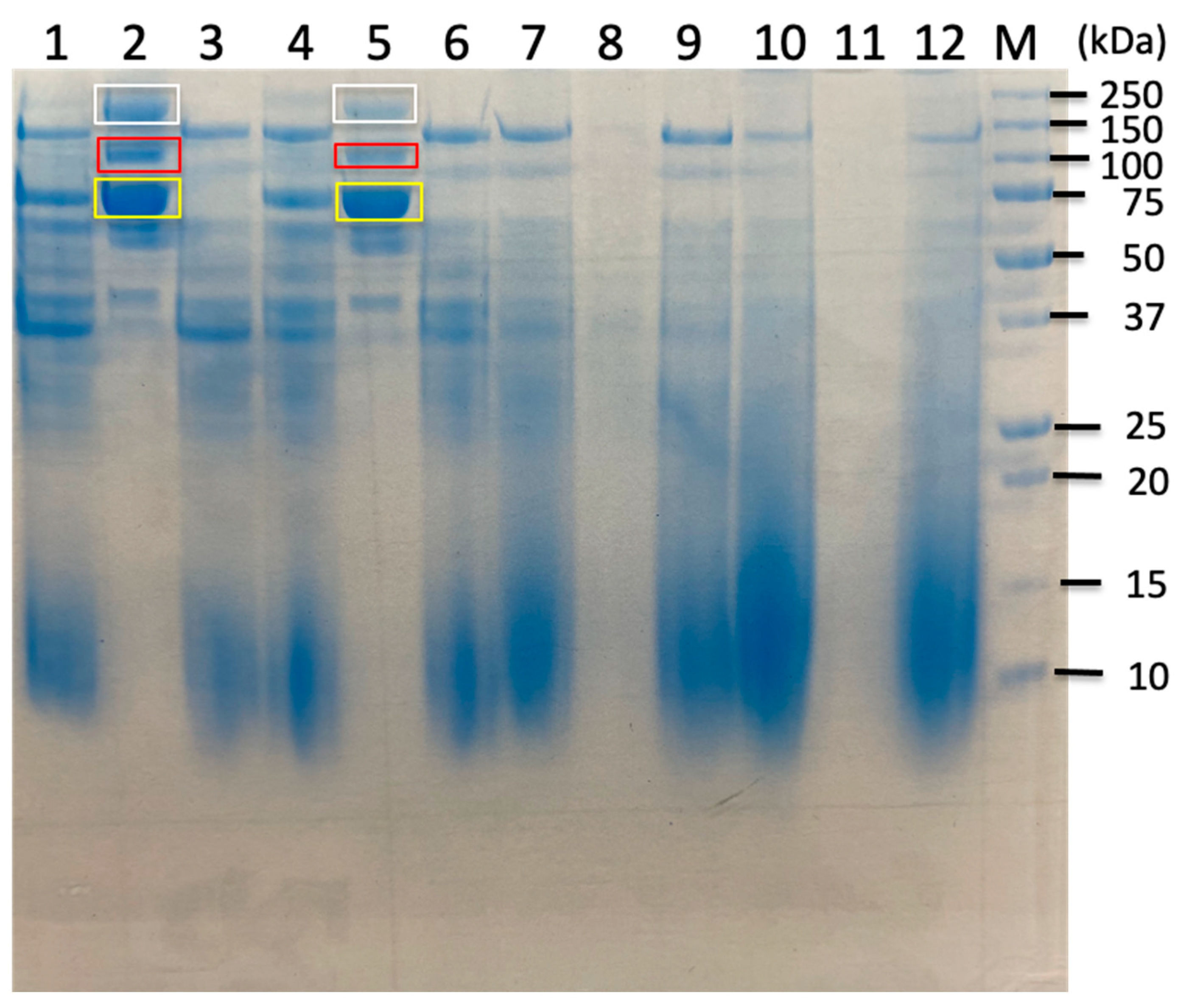
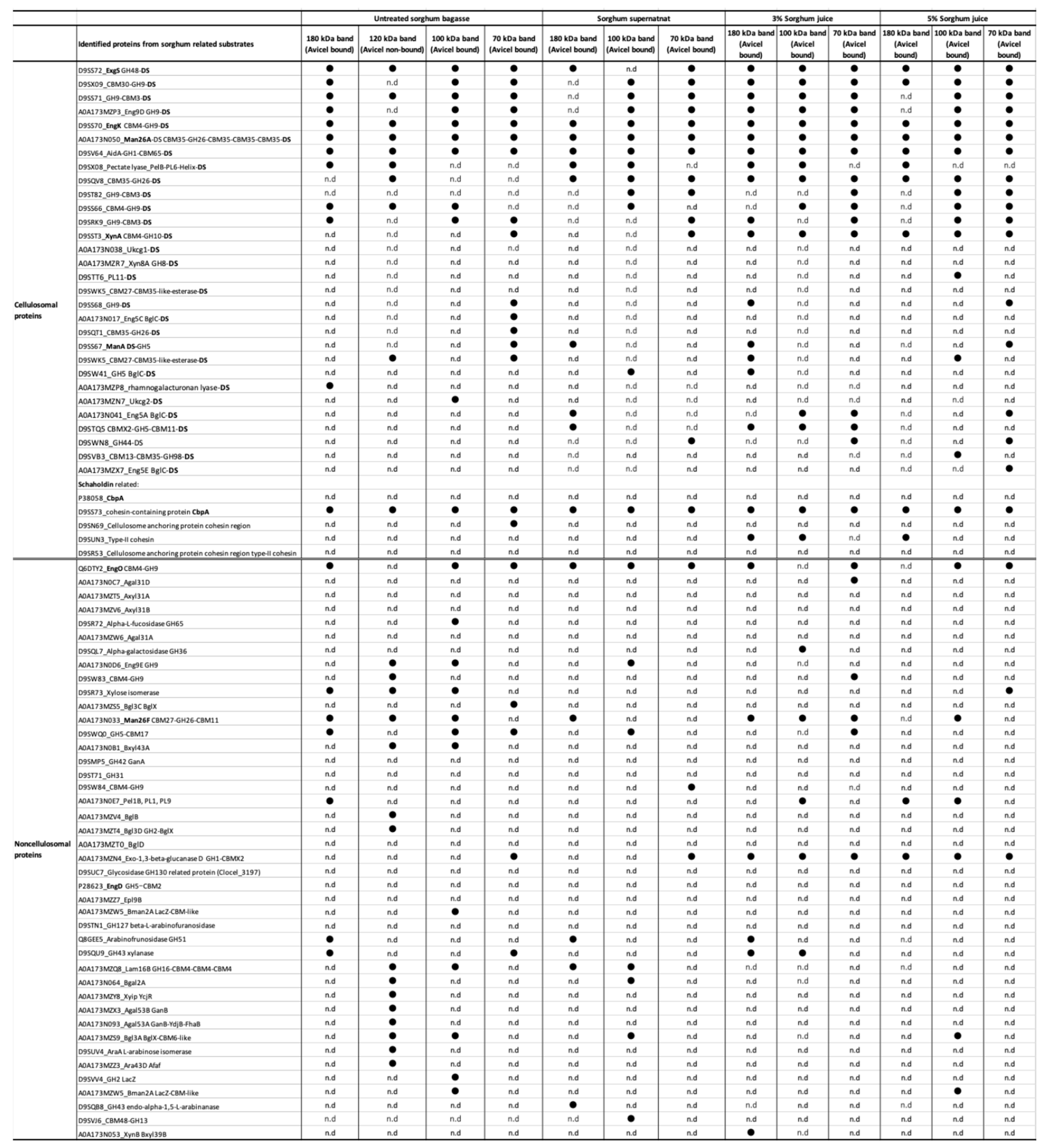
2.3. Proteomic Analysis of Sorghum-Related Proteins and Their Comparison Based on Untreated Sorghum Bagasse and Its Supernatants, and Sorghum Juice
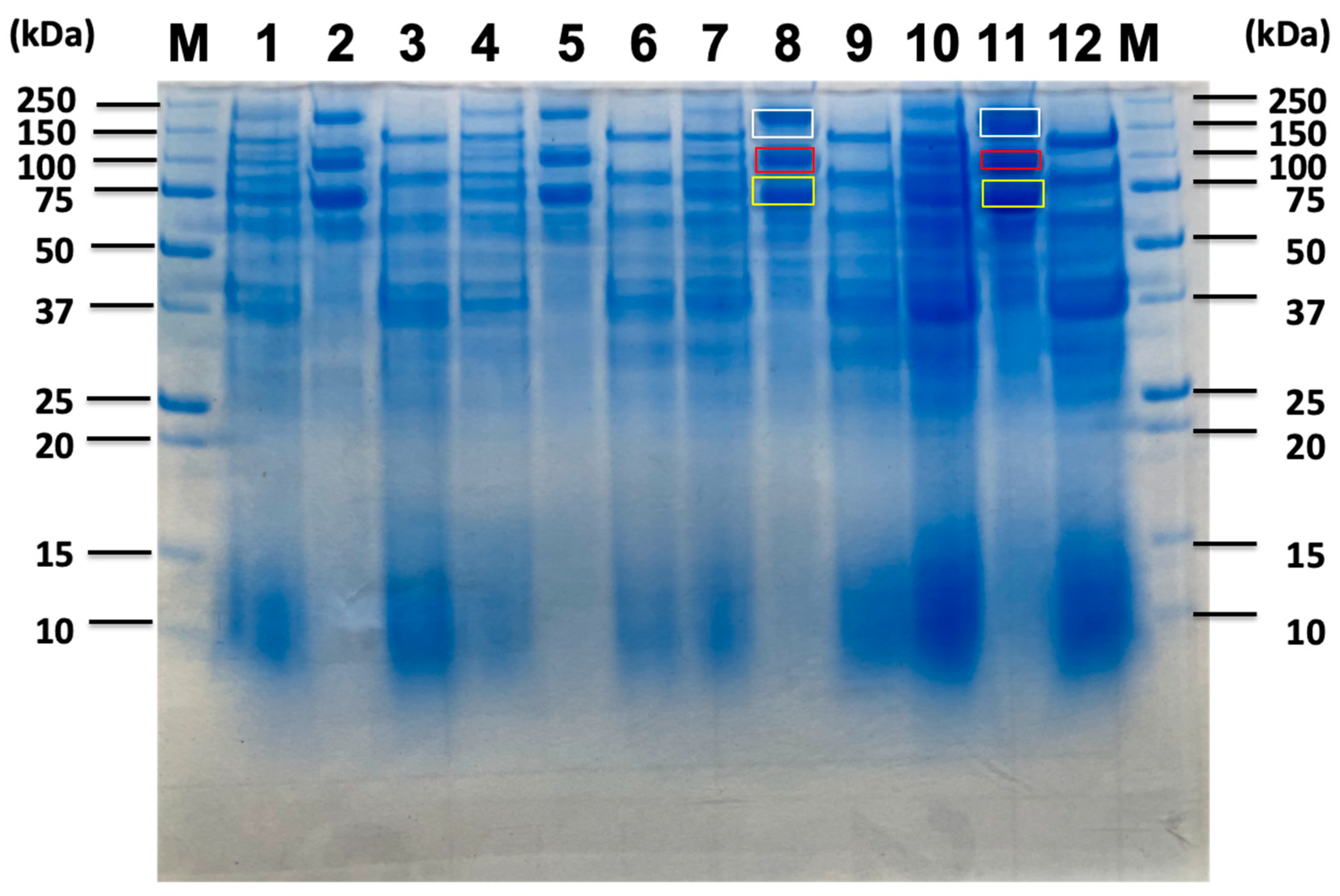
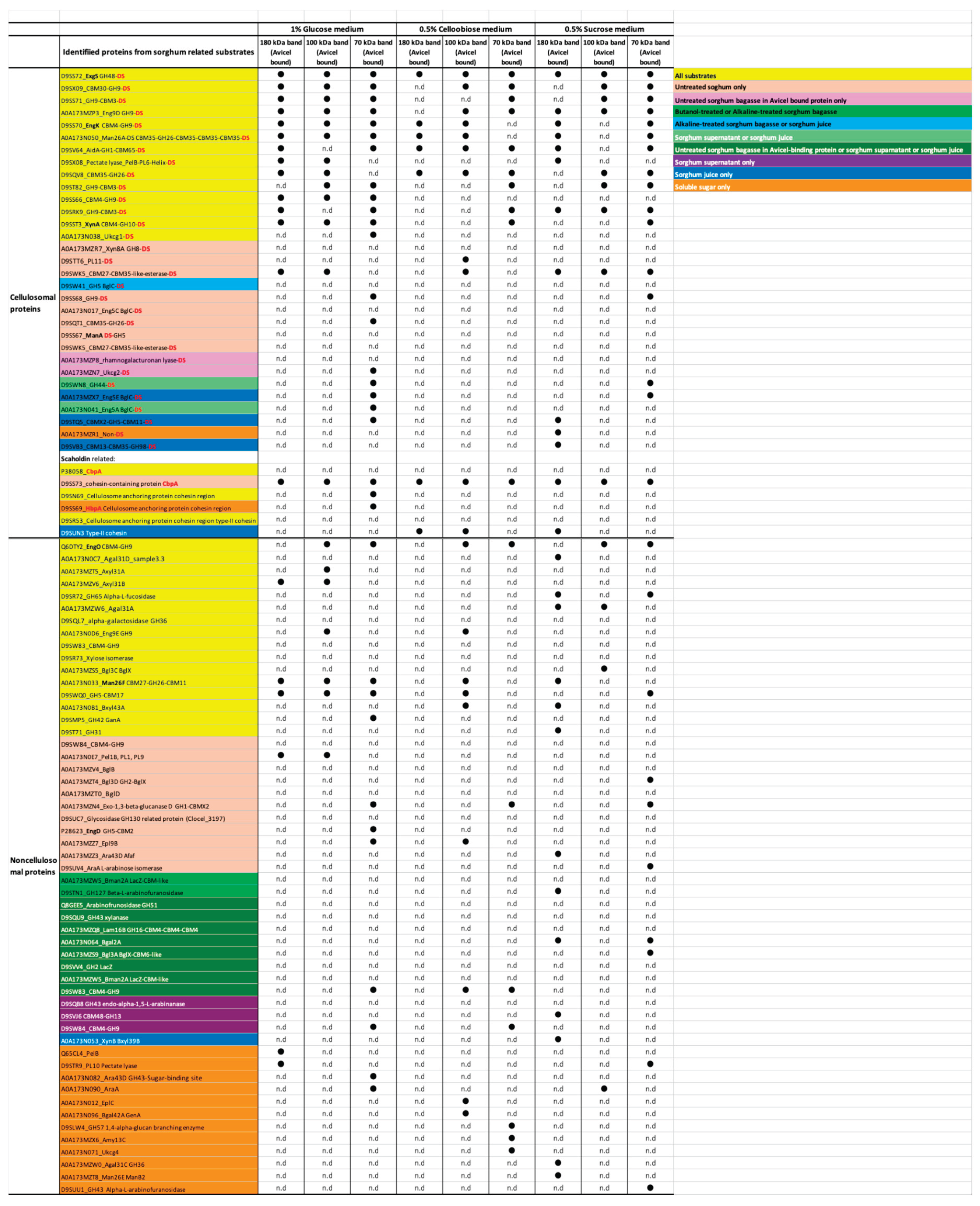
2.4. Proteomic Analysis of Soluble Sugar-Related Proteins and Their Comparison Based on Untreated Sorghum Bagasse and Its Supernatants, and Sorghum Juice
3. Discussion
4. Materials and Methods
4.1. Bacterial Strain and Culture Conditions
4.2. Substrates Preparation
4.3. Enzyme Preparation and Concentration
4.4. SDS-PAGE Analysis and Preparation of Crystalline Cellulose Bound and Non-Bound Fractions
4.5. Enzyme Assay
4.6. LC-MS/MS Analysis and Data Acquisition
4.6.1. Chromatographic Conditions
4.6.2. Mass Spectrometry Parameters
4.7. Database Searching and Protein Identification
4.8. Statistics
5. Conclusions
Supplementary Material
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Esaka, K.; Aburaya, S.; Morisaka, H.; Kuroda, K.; Ueda, M. Exoproteome analysis of Clostridium cellulovorans in natural soft-biomass degradation. AMB Express 2015, 5, 1. [Google Scholar] [CrossRef] [PubMed]
- de Morais Cardoso, L.; Pinheiro, S.S.; Martino, H.S.D.; Pinheiro-Sant’Ana, H.M. Sorghum (Sorghum bicolor L.): nutrients, bioactive compounds, and potential impact on human health. Crit. Rev. Food Sci. Nutr. 2017, 57, 372–390. [Google Scholar] [CrossRef]
- Nghiem, N.P.; Montanti, J.; Johnsto, D.B. Sorghum as a renewable feedstock for production of fuels and industrial chemicals. AIMS Bioengin. 2016, 3, 75–91. [Google Scholar] [CrossRef]
- Uddin, M.R.; Kee Woong Park, K.W.; Kim, Y.K.; Sang Un Park, S.U.; Pyon, J.Y. Enhancing sorgoleone levels in grain sorghum root exudates. J. Chem. Ecol. 2010, 36, 914–922. [Google Scholar] [CrossRef]
- Tamaru, Y.; Miyake, H.; Kuroda, K.; Nakanishi, A.; Kawade, Y.; Yamamoto, K.; Uemura, M.; Fujita, Y.; Doi, R.H.; Ueda, M. Genome sequence of the cellulosome-producing mesophilic organism Clostridium cellulovorans 743B. J. Bacteriol. 2010, 192, 901–902. [Google Scholar] [CrossRef]
- Tamaru, Y.; Miyake, H.; Kuroda, K.; Nakanishi, A.; Matsushima, C.; Doi, R.H.; Ueda, M. Comparison of the mesophilic cellulosome-producing Clostridium cellulovorans genome with other cellulosome-related clostridial genomes. Microbial Biotechnol. 2011, 4, 64–73. [Google Scholar] [CrossRef]
- Morisaka, H.; Matsui, K.; Tatsukami, Y.; Kuroda, K.; Miyake, H.; Tamaru, Y.; Ueda, M. Profile of native cellulosomal proteins of Clostridium cellulovorans adapted to various carbon sources. AMB Express 2012, 2, 37. [Google Scholar] [CrossRef]
- Aburaya, S.; Aoki, W.; Kuroda, K.; Minakuchi, H.; Ueda, M. Temporal proteome dynamics of Clostridium cellulovorans cultured with major plant cell wall polysaccharides. BMC Microbiol. 2019, 19, 118. [Google Scholar] [CrossRef] [PubMed]
- Usai, G.; Cirrincione, S.; Re, A.; Manfredi, M.; Pagnani, A.; Pessione, E.; Mazzoli, R. Clostridium cellulovorans metabolism of cellulose as studied by comparative proteomic approach. J. Proteomics 2020, 216, 103667. [Google Scholar] [CrossRef] [PubMed]
- Eljonaid, M.Y.; Tomita, H.; Okazaki, F.; Tamaru, Y. Enzymatic characterization of unused biomass degradation using the Clostridium cellulovorans cellulosome. Microorganisms 2022, 10, 2514. [Google Scholar] [CrossRef]
- Wang, F.; Ouyang, D.; Zhou, Z.; Page, S.J.; Liu, D.; Zhao, X. Lignocellulosic biomass as sustainable feedstock and materials for power generation and energy storage. J. Energy Chem. 2021, 57, 247–280. [Google Scholar] [CrossRef]
- Liu, F.; Wodajo, B.; Zhao, K.; Tang, S.; Xie, Q.; Xie, P. Unravelling sorghum functional genomics and molecular breeding: past achievements and future prospects. J. Gen. Genom. 2025, 52, 719–732. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, M.; Sazuka, T.; Oda, Y.; Kawahigashi, H.; Wu, J.; Takanashi, H.; Ohnishi, T.; Yoneda, J.I.; Ishimori, M.; Kajiya-Kanegae, H.; Hibara, K.; Ishizuna, F.; Ebine, K.; Ueda, T.; Tokunaga, T.; Iwata, H.; Matsumoto, T.; Kasuga, S.; Yonemaru, J.; Tsutsumi, N. Transcriptional switch for programmed cell death in pith parenchyma of sorghum stems. Proc. Natl. Acad. Sci. U.S.A. 2018, 115, E8783–E8792. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.M.; Leng, C.Y.; Luo, H.; Wu, X.Y.; Liu, Z.Q.; Zhang, Y.M.; Zhang, H.; Xia, Y.; Shang, L.; Liu, C.M.; Hao, D.Y.; Zhou, Y.H.; Chu, C.C.; Cai, H.W.; Jing, H.J. Sweet sorghum originated through selection of Dry, a plant-specific NAC transcription factor gene. Plant Cell 2018, 30, 2286–2307. [Google Scholar] [CrossRef]
- Kim, J.; Hopfield, J.J.; Winfree, E. Neural network computation by in vitro transcriptional circuits. Adv. Neural Inf. Process. Syst. 2004, 17, 681–688. [Google Scholar]
- Genot, A.J.; Fujii, T.; Rondelez, Y. Scaling down DNA circuits with competitive neural networks. J. R. Soc. Interface 2013, 10, 20130212. [Google Scholar] [CrossRef]
- Okumura, S.; Gines, G.; Lobato-Dauzier, N.; Baccouche, A.; Deteix, R.; Fujii, T.; Rondelez, Y.; Genot, A.J. Nonlinear decision-making with enzymatic neural networks. Nature 2022, 610, 496–501. [Google Scholar] [CrossRef]
- Tomita, H.; Okazaki, F.; Tamaru, Y. Biomethane production from sugar beet pulp under cocultivation with Clostridium cellulovorans and methanogens. AMB Express 2019, 9, 28. [Google Scholar] [CrossRef]
- Tomita, H.; Okazaki, F.; Tamaru, Y. Direct IBE fermentation from mandarin orange wastes by combination of Clostridium cellulovorans and Clostridium beijerinckii. AMB Express 2019, 9, 1. [Google Scholar] [CrossRef]
- Tomita, H.; Tamaru, Y. The second-generation biomethane from mandarin orange peel under cocultivation with methanogens and the armed Clostridium cellulovorans. Fermentation 2019, 5, 95. [Google Scholar] [CrossRef]
- Koukiekolo, R.; Cho, H.Y.; Kosugi, A.; Inui, M.; Yukawa, H.; Doi, R.H. Degradation of corn fiber by Clostridium cellulovorans cellulases and hemicellulases and contribution of scaffolding protein CbpA. Appl. Environ. Microbiol. 2005, 71, 3504–3511. [Google Scholar] [CrossRef]
- Tamaru, Y.; Ui, S.; Murashima, K.; Kosugi, A.; Chan, H.; Doi, R.H.; Liu, B. Formation of protoplasts from cultured tobacco cells and Arabidopsis thaliana by the action of cellulosomes and pectate lyase from Clostridium cellulovorans. Appl. Environ. Microbiol. 2002, 68, 2614–2618. [Google Scholar] [CrossRef] [PubMed]
- Kosugi, A.; Murashima, K.; Doi, R.H. Xylanase and acetyl xylan esterase activities of XynA, a key Subunit of the Clostridium cellulovorans cellulosome for xylan degradation. Appl. Environ. Microbiol. 2002, 68, 6399–6402. [Google Scholar] [CrossRef] [PubMed]
- Beg, Q.K.; Kapoor, M.; Mahajan, L.; Hoondal, G.S. Microbial xylanases and their industrial applications: a review. Appl. Microbiol. Biotechnol. 2001, 56, 326–338. [Google Scholar] [CrossRef]
- Wang, X.; Xiao, M.; Chen, C.; Qi, K.; Chen, C.; Cui, Q.; Kosugi, A.; Liu, Y.J.; Feng, Y. Effective conversion of lignocellulose to fermentable sugars using engineered Clostridium thermocellum with co-enhanced cellulolytic and xylanolytic activities. Bioresour. Technol. 2025, 440, 133466. [Google Scholar] [CrossRef] [PubMed]
- Teramura, H.; Sasaki, K.; Oshima, T.; Kawaguchi, H. Effective usage of sorghum bagasse: Optimization of organosolv pretreatment using 25% 1-butanol and subsequent nanofiltration membrane separation. Bioresour. Technol. 2017, 252, 157–164. [Google Scholar] [CrossRef]
- Wunna, K.; Nakasaki, K.; Auresenia, J.L.; Abella, L.C.; Gaspillo, P.D. Effect of alkali pretreatment on removal of lignin from sugarcane bagasse. Chem. Engin. Trans. 2017, 56, 1831–1836. [Google Scholar]
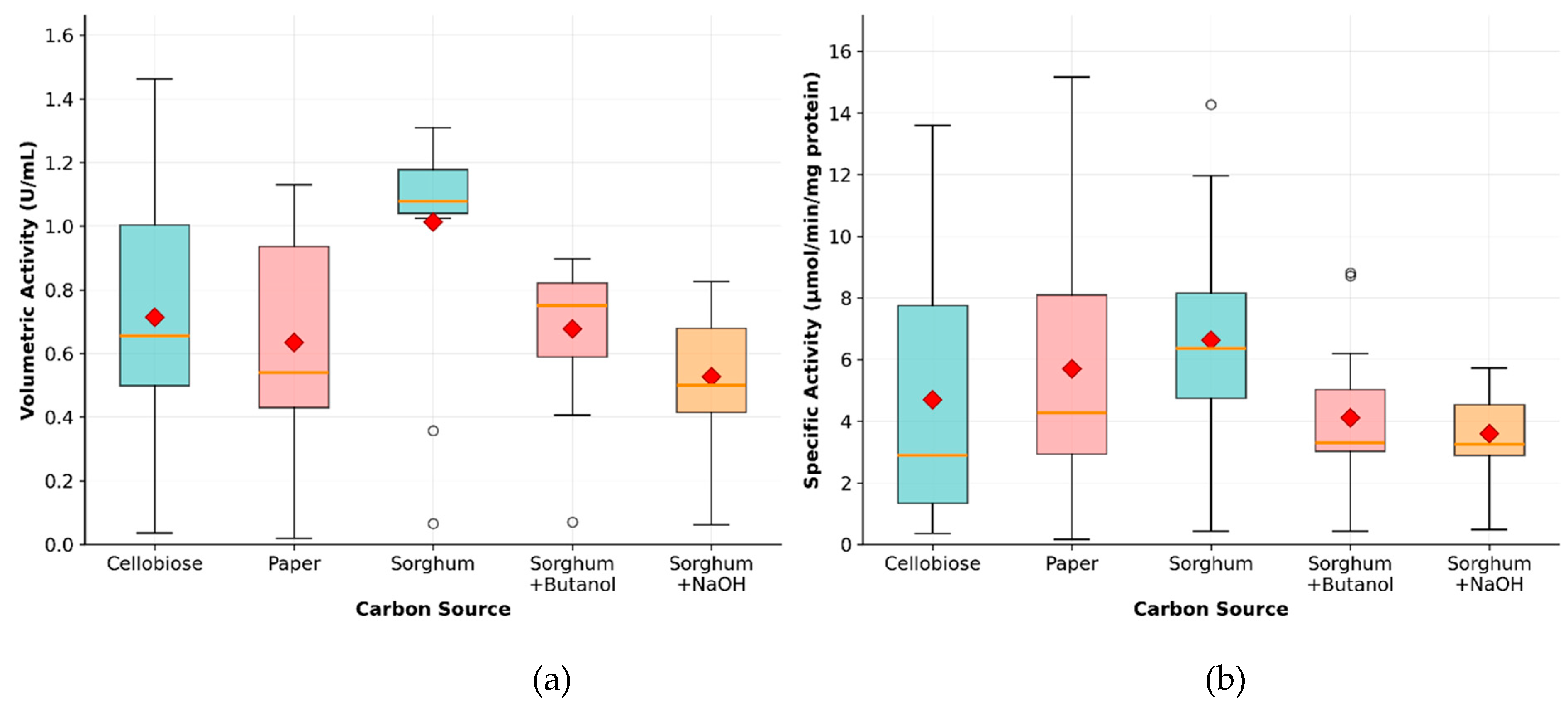

| Carbon Sources | Mean Specific Activity (U/mg) | Peak Specific Activity (U/mg) | Time to Peak (h) | Mean Volumetric Activity (U/mL) | Peak Volumetric Activity (U/mL) | Time to Peak (h) |
|---|---|---|---|---|---|---|
| Cellobiose | 4.70 | 13.60 | 123 | 0.71 | 1.46 | 170 |
| Filter paper | 5.70 | 15.17 | 146 | 0.63 | 1.13 | 146 |
| Untreated sorghum bagasse | 6.63 | 14.27 | 123 | 1.01 | 1.31 | 84 |
| Sorghum bagasse + butanol | 4.11 | 8.81 | 123 | 0.68 | 0.90 | 24 |
| Sorghum bagasse + NaOH | 3.59 | 5.72 | 84 | 0.53 | 0.83 | 24 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





