Submitted:
09 November 2025
Posted:
10 November 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Master Regulators: Definition and Biological Significance
Rationale for Gene Selection: Why These 36 Genes?
- D Markers 14 genes: CD2, CD3G, CD4, CD7, CDS, CD25, CD27, CD28, CD31, CD45, CD58, CD69, CD80, CD86. These surface markers define T-cell subsets, mediate co-stimulation, and regulate T-cell activation and trafficking. [25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96]
- Transcription Factors Governing Cell Fate 16 genes: FOXP3, FOXJ1, FOXM1, BLIMP1, PU.1, OCT4, TAL1, PROX1, GCM1, TFEB, SIM1, CTCF, MITF, HIF1,VEGF. These transcription factors regulate immune cell differentiation, stem cell pluripotency, hypoxia responses, and tissue-specific development. [106,107,108,109,110,111,112,113,114,115,116,117]
Metadichol: A Pleiotropic Bioactive Nanoemulsion
Systems Biology Perspective
|
Gene |
Full Name |
Category |
Cell Type |
Biological Function | Disease Associations |
|
CD3G |
CD3 gamma chain |
CD Marker |
PBMC |
T cell receptor signaling, T cell activation | Immunodeficiency, autoimmune diseases |
|
CD4 |
Cluster of Differentiation 4 |
CD Marker |
PBMC |
T helper cell marker, MHC class II co- receptor |
HIV/AIDS, autoimmune diseases |
|
CD7 |
Cluster of Differentiation 7 |
CD Marker |
PBMC |
T cell and NK cell development, cell adhesion |
T cell acute lymphoblastic leukemia |
|
CD25 |
Interleukin-2 receptor alpha chain IL- 2Rα |
CD Marker |
PBMC |
IL-2 signaling, T cell activation and proliferation |
Autoimmune diseases, immunodeficiency |
|
CD27 |
Cluster of Differentiation 27 |
CD Marker |
PBMC |
T cell activation, memory T cell generation |
Lymphomas, immunodeficiency |
|
CD25 |
Cluster of Differentiation 2S |
CD Marker |
PBMC |
T cell co-stimulation, survival, and proliferation |
Autoimmune diseases, transplant rejection |
|
CD31 |
Platelet endothelial cell adhesion molecule-1 |
CD Marker |
PBMC |
Cell adhesion, leukocyte migration, angiogenesis |
Cardiovascular disease, inflammation |
|
CD45 |
Protein tyrosine phosphatase receptor type C |
CD Marker |
PBMC |
Leukocyte common antigen, signal transduction |
Immunodeficiency, autoimmune diseases |
|
CD58 |
Lymphocyte function- associated antigen 3 |
CD Marker |
PBMC |
T cell adhesion and co-stimulation |
Autoimmune diseases, cancer immune evasion |
|
CD86 |
B7-2 costimulatory molecule |
CD Marker |
PBMC |
T cell co-stimulation, antigen presentation |
Autoimmune diseases, transplant rejection |
|
CD69 |
Early activation antigen CD69 |
CD Marker |
PBMC |
Early activation marker, tissue retention of lymphocytes | Autoimmune diseases, inflammatory disorders |
|
CD8 |
Cluster of Differentiation S |
CD Marker |
PBMC |
Cytotoxic T cell marker, MHC class I co- receptor |
Immunodeficiency, viral infections, cancer |
|
CD80 |
B7-1 costimulatory molecule |
CD Marker |
PBMC |
T cell co-stimulation, immune checkpoint regulation |
Autoimmune diseases, transplant rejection |
|
CD2 |
Cluster of Differentiation 2 |
CD Marker |
PBMC |
T cell and NK cell adhesion, co-stimulation |
Autoimmune diseases, T cell lymphomas |
|
VEGF |
Vascular endothelial growth factor |
Growth Factor |
Fibroblast |
Angiogenesis, vascular permeability | Cancer, diabetic retinopathy, macular degeneration |
|
TAL1 |
T-cell acute lymphocytic leukemia protein 1 |
Transcription Factor |
Fibroblast |
Hematopoiesis, erythroid differentiation |
T-cell acute lymphoblastic leukemia |
|
BLIMP 1 |
B-lymphocyte- induced maturation protein 1 |
Transcription Factor |
Fibroblast |
Plasma cell differentiation, T cell exhaustion |
Lymphomas, autoimmune diseases |
|
FOXP3 |
Forkhead box protein P3 |
Transcription Factor |
Fibroblast |
Regulatory T cell development and function |
IPEX syndrome, autoimmune diseases, cancer |
|
PU.1 |
Purine-rich box- 1 |
Transcription Factor |
Fibroblast |
Myeloid and B cell development |
Leukemias, immunodeficiency |
|
OCT4 ( POU5F1) |
Octamer- binding transcription factor 4 |
Transcription Factor |
Fibroblast |
Pluripotency, stem cell self- renewal |
Germ cell tumors, cancer stem cells |
|
PROX1 |
Prospero homeobox protein 1 |
Transcription Factor |
Fibroblast |
Lymphatic vessel development, cell fate determination |
Lymphedema, cancer metastasis |
|
GCM1 |
Glial cells missing homolog 1 |
Transcription Factor |
Fibroblast |
Placental development, parathyroid gland development |
Preeclampsia, hypoparathyroidism |
|
PGC1a |
Peroxisome proliferator- activated receptor gamma coactivator 1- alpha |
Transcriptional Coactivator |
Fibroblast |
Mitochondrial biogenesis, oxidative metabolism |
Diabetes, obesity, neurodegenerative diseases, cancer |
|
Nrf2 |
Nuclear factor erythroid 2- related factor 2 |
Transcription Factor |
Fibroblast |
Antioxidant response, detoxification |
Cancer, neurodegenerative diseases, inflammatory diseases |
|
FOXJ1 |
Forkhead box protein J1 |
Transcription Factor |
Fibroblast |
Cilio-genesis, motile cilia development | Primary ciliary dyskinesia, respiratory infections |
|
TFEB |
Transcription factor EB |
Transcription Factor |
Fibroblast |
Autophagy, lysosomal biogenesis | Neurodegenerative diseases, lysosomal storage disorders |
|
cMyc |
MYC proto- oncogene |
Transcription Factor |
Fibroblast |
Cell proliferation, growth, apoptosis, metabolism |
Multiple cancers Burkitt lymphoma, breast, colon, lung |
| p53 | Tumor protein p53 | Transcription Factor | Fibroblast | Tumor suppression, cell cycle arrest, apoptosis | Li-Fraumeni syndrome, multiple cancers >50% of human cancers |
|
p63 |
Tumor protein p63 |
Transcription Factor |
Fibroblast |
Epithelial development, stem cell maintenance |
Ectodermal dysplasia syndromes, squamous cell carcinomas |
|
SIM1 |
Single-minded homolog 1 |
Transcription Factor |
Fibroblast |
Hypothalamic development, energy homeostasis |
Obesity, Prader-Willi- like syndrome |
|
FOXM1 |
Forkhead box protein M1 |
Transcription Factor |
Fibroblast |
Cell cycle progression, proliferation, DNA repair |
Multiple cancers breast, lung, liver, prostate |
|
CTCF |
CCCTC-binding factor |
Transcription Factor |
Fibroblast |
Chromatin organization, insulator function |
Cancer, developmental disorders |
|
MITF |
Microphthalmia- associated transcription factor |
Transcription Factor |
Fibroblast |
Melanocyte development, pigmentation |
Melanoma, Waardenburg syndrome |
|
HIF |
Hypoxia- inducible factor |
Transcription Factor |
Fibroblast |
Hypoxia response, angiogenesis, metabolic adaptation |
Cancer, ischemic diseases, pulmonary hypertension |
|
MYOD1 |
Myogenic differentiation 1 |
Transcription Factor |
Fibroblast |
Muscle cell differentiation, myogenesis |
Rhabdomyosarcoma, muscular dystrophy |
|
PPARg |
Peroxisome proliferator- activated receptor gamma |
Nuclear Receptor |
Fibroblast |
Adipogenesis, glucose metabolism, anti- inflammation |
Type 2 diabetes, obesity, metabolic syndrome, cancer |
Research Rationale and Objectives
- Comprehensively characterize the dose-dependent effects of Metadichol on all 36 genes in human PBMC and Fibroblasts.
- Identify optimal concentrations for maximal transcriptional effects.
- Analyze the relationship between the 36 Master gene regulated and Metadichol's known effects on nuclear receptors, sirtuins, TLRs, and circadian networks; and
- Provide mechanistic insights into the synergistic transcriptional networks underlying Metadichol's pleiotropic biological activities.
Experimental
Materials and Methods
Cell Isolation and Culture
Material Treatment
| Cell lines | Sample |
Treatment details |
|
Human, PBMC, NHDF |
Metadichol |
Control |
| 1 pg/ml | ||
| 100 pg/ml | ||
| 1 ng/ml | ||
| 100ng/ml |
Sample Préparation and RNA Isolation
| Test concentrations | |||||
|---|---|---|---|---|---|
|
RNA yield ng/µl |
0 |
1 pg/ ml |
100 pg/ ml |
1 ng/ ml |
100 ng/ ml |
| Human PBMC's | 619.520 | 317.880 | 681.840 | 700.160 | 667.400 |
| Test concentrations | |||||
| RNA yield ng/µl | 0 | 1 pg/ ml | 100 pg/ ml | 1 ng/ ml | 100 ng/ ml |
| NHDF cells | 147.5 | 593.7 | 311.6 | 392 | 295.4 |
cDNA Synthesis
Primers and qPCR Analysis
| Primer Name | Sequence | Amplicon | Annealing temperature | |
| GAPDH | F | GTCTCCTCTGACTTCAACAGCG | 186 | 60 |
| R | ACCACCCTGTTGCTGTAGCCAA | |||
| CD3G | F | GCATTTTCGTCCTTGCTGTTGGG | 134 | 50 |
| R | GGTCATCTTCTCGATCCTTGAGG | |||
| CD4 | F | CCTCCTGCTTTTCATTGGGCTAG | 125 | 65 |
| R | TGAGGACACTGGCAGGTCTTCT | |||
| CD7 | F | TGTCGGACACTGGCACCTACAC | 114 | 57 |
| R | TCCGAGCATCTGTGCCATCCTT | |||
| CD25 | F | GAGACTTCCTGCCTCGTCACAA | 125 | 53 |
| R | GATCAGCAGGAAAACACAGCCG | |||
| CD27 | F | TCAGCAACTGGGCACAGAAA | 180 | 65 |
| R | TTCCTGGCTCACACATCTGG | |||
| CD28 | F | GAGAAGAGCAATGGAACCATTATC | 121 | 57 |
| R | TAGCAAGCCAGGACTCCACCAA | |||
| CD31 | F | ATTACCTGACCAGCGCCAC | 171 | 57 |
| R | AGAGTGAAGACTGCAGGCAC | |||
| CD45 | F | CTTCAGTGGTCCCATTGTGGTG | 106 | 61 |
| R | CCACTTTGTTCTCGGCTTCCAG | |||
| CD58 | F | AACCTGTATCCCAAGCAGCG | 173 | 65 |
| R | TGCTGTTGTCTTCATCTTCTGT | |||
| CD86 | F | AAGCAAGAGCACTGTCCCTG | 196 | 67 |
| R | TAAGCACAGCAGCATTCCCA | |||
| CD69 | F | CAGAGGTCAGCAGCATGGAA | 138 | 49 |
| R | AGAGCAGCATCCACTGACAC | |||
| CD8 | F | ATGGCCTTACCAGTGACCG | 104 | 49 |
| R | AGGTTCCAGGTCCGATCCAG | |||
| CD80 | F | CTCTTGGTGCTGGCTGGTCTTT | 135 | 49 |
| R | GCCAGTAGATGCGAGTTTGTGC | |||
| CD2 | F | GTCAGCAAGGAATCCAGTGTCG | 197 | 62 |
| R | AACGAGCAGTGCCACAAAGACC | |||
| PU.1SPI1 | F | GACACGGATCTATACCAACGCC | 144 | 67 |
| R | CCGTGAAGTTGTTCTCGGCGAA | |||
| FOXP3 | F | GGCACAATGTCTCCTCCAGAGA | 127 | 53 |
| R | CAGATGAAGCCTTGGTCAGTGC | |||
| PPAR gamma | F | ACGCACCGAAATTCTCCCTT | 171 | 65 |
| R | TCTGCCTCTCCCTTTGCTTG | |||
| MYOD1 | F | CTCCAACTGCTCCGACGGCAT | 148 | 65 |
| R | ACAGGCAGTCTAGGCTCGACAC | |||
| ProX1 | F | AGCGGTCTCTCTAGTACAGGC | 92 | 65 |
| R | AAAGGGGAAAGACACTCTGGG | |||
| SIM1 | F | GACTCTGTACCACCATGTGCAC | 113 | 59 |
| R | GTGTTTCGCCAGGAACCTGTAG | |||
| GCM1 | F | AGTGAACACAGCACCTTCCTCC | 127 | 65 |
| R | TTGGACGCCTTCCTGGAAAGAC | |||
| FOXM1 | F | ATACGTGGATTGAGGACCACT | 175 | 67 |
| R | TCCAATGTCAAGTAGCGGTTG | |||
| HIF-1ALPHA | F | TATGAGCCAGAAGAACTTTTAGGC | 144 | 65 |
| R | CACCTCTTTTGGCAAGCATCCTG | |||
| CTCF | F | GACCACACAAGTGCCATCTCTG | 111 | 59 |
| R | ATGTCGCAGTCTGGGCACTTGT | |||
| FOXJ1 | F | ACTCGTATGCCACGCTCATCTG | 152 | 62 |
| R | GAGACAGGTTGTGGCGGATTGA | |||
| p63 | F | CAGGAAGACAGAGTGTGCTGGT | 121 | 58 |
| R | AATTGGACGGCGGTTCATCCCT | |||
| Nrf2 | F | CACATCCAGTCAGAAACCAGTGG | 111 | 65 |
| R | GGAATGTCTGCGCCAAAAGCTG | |||
| MITF | F | GGCTTGATGGATCCTGCTTTGC | 129 | 62 |
| R | GAAGGTTGGCTGGACAGGAGTT | |||
| TFEB | F | ACCTGTCCGAGACCTATGGG | 222 | 65 |
| R | CGTCCAGACGCATAATGTTGTC | |||
| PGC-1 ALPHA | F | CCAAAGGATGCGCTCTCGTTCA | 146 | 67 |
| R | CGGTGTCTGTAGTGGCTTGACT | |||
| TAL-1 | F | CCACCAACAATCGAGTGAAGAGG | 127 | 67 |
| R | GTTCACATTCTGCTGCCGCCAT | |||
| VEGF | F | GGAACCTCACTATCCGCAGAGT | 131 | 65 |
| R | CCAAGTTCGTCTTTTCCTGGGC | |||
| C-Myc | F | CCTGGTGCTCCATGAGGAGAC | 106 | 62 |
| R | CAGACTCTGACCTTTTGCCAGG | |||
| BLIMP1PRDM1 | F | CAGTTCCTAAGAACGCCAACAGG | 122 | 65 |
| R | GTGCTGGATTCACATAGCGCATC | |||
| Oct4POU5F1 | F | GTAGTCCCTTCGCAAGCCCT | ||
| R | AGGTCCGAGGATCAACCCAG | 163 | 65 | |
| p53 | F | CCTCAGCATCTTATCCGAGTGG | 163 | 58 |
| R | TGGATGGTGGTACAGTCAGAGC |
Results
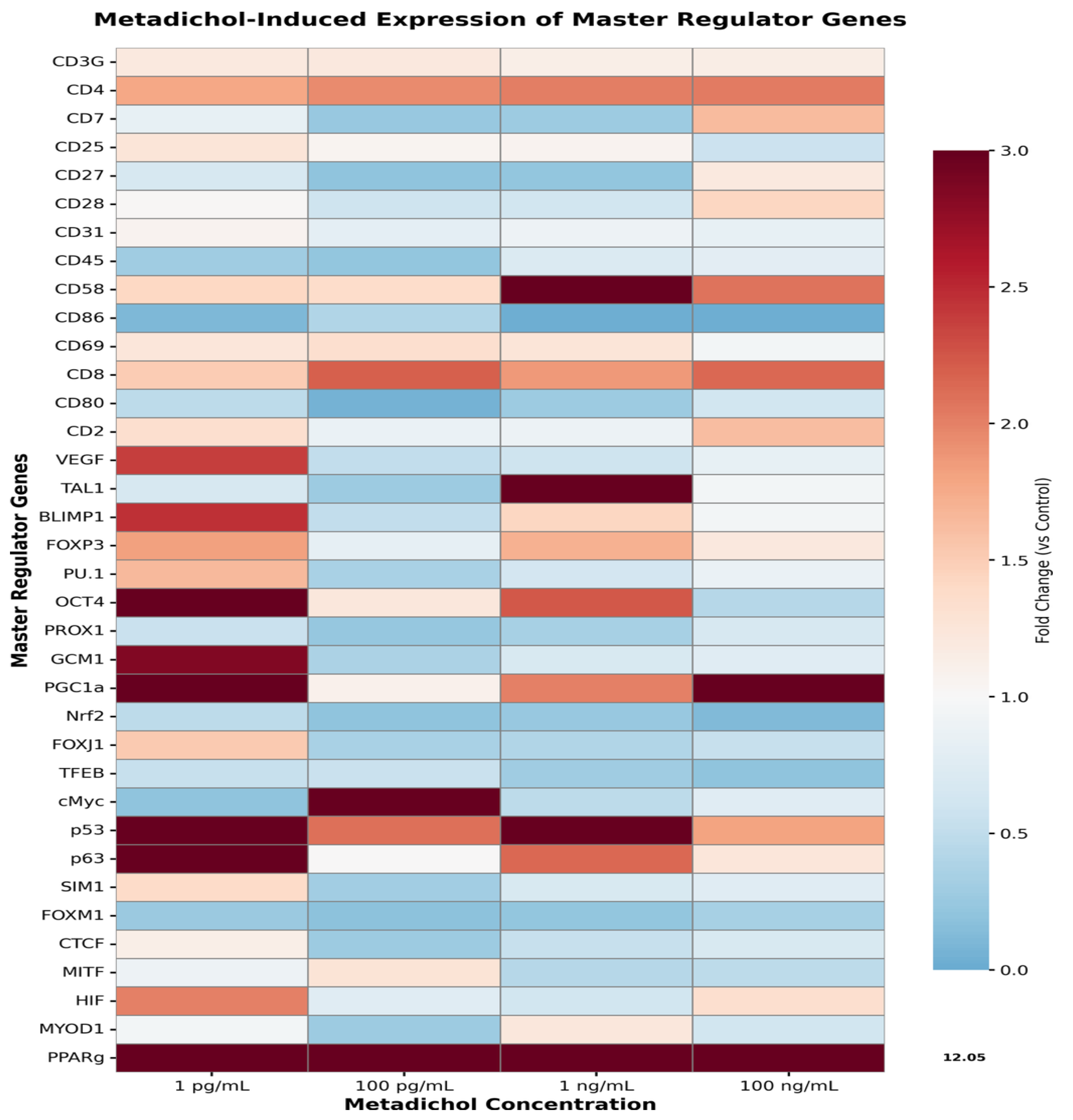
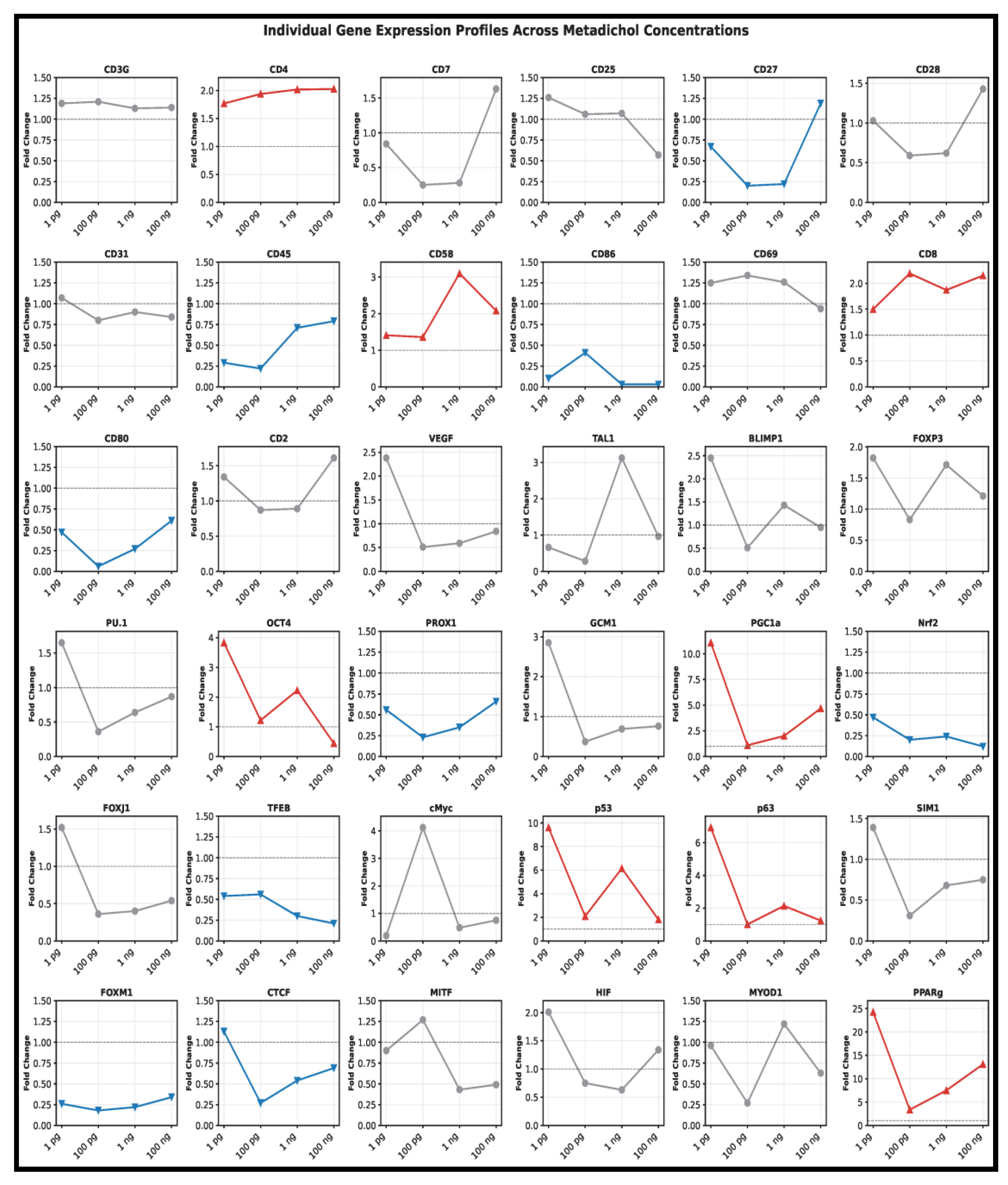
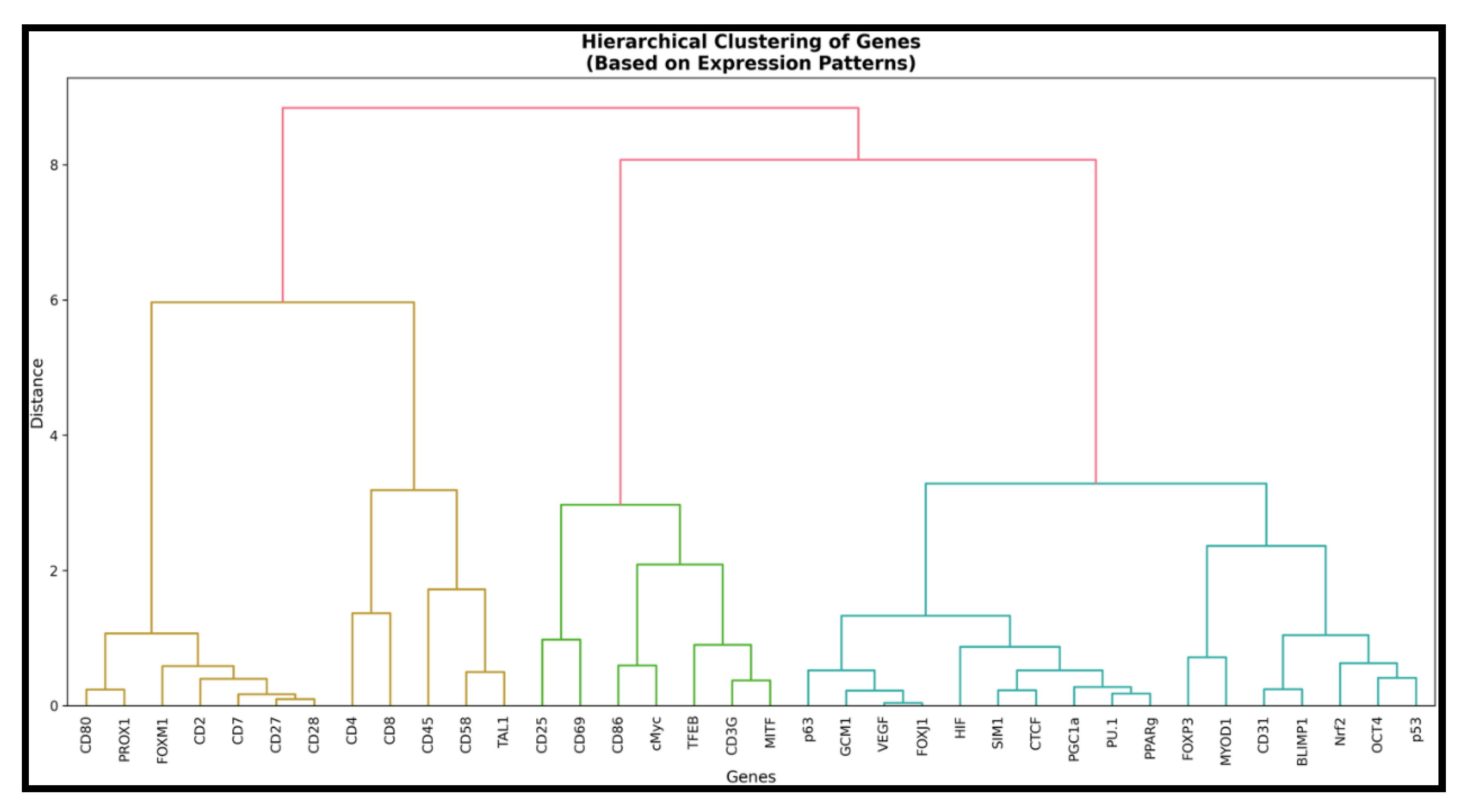
Overview of Metadichol-Induced Gene Expression Changes
Dose-Response Patterns and Functional Classification
- Low-dose responders optimal at 1 pg/mL including PPARγ, PGC1α, and p53.
- Intermediate-dose responders optimal at 100 pg/mL - 1 ng/mL.
- High-dose responders optimal at 100 ng/mL.
- Dose-independent responders showing consistent expression across concentrations.
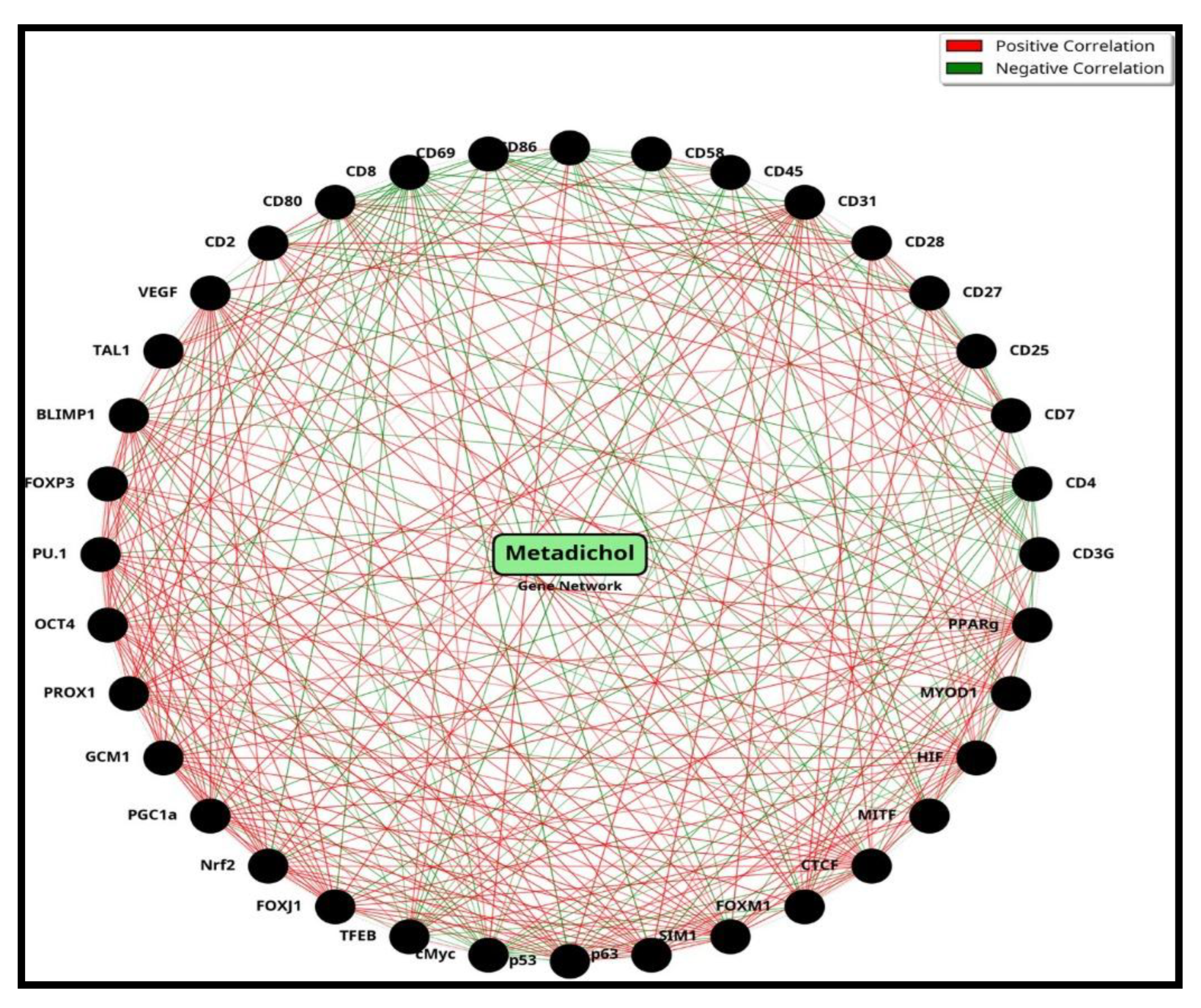
Discussion
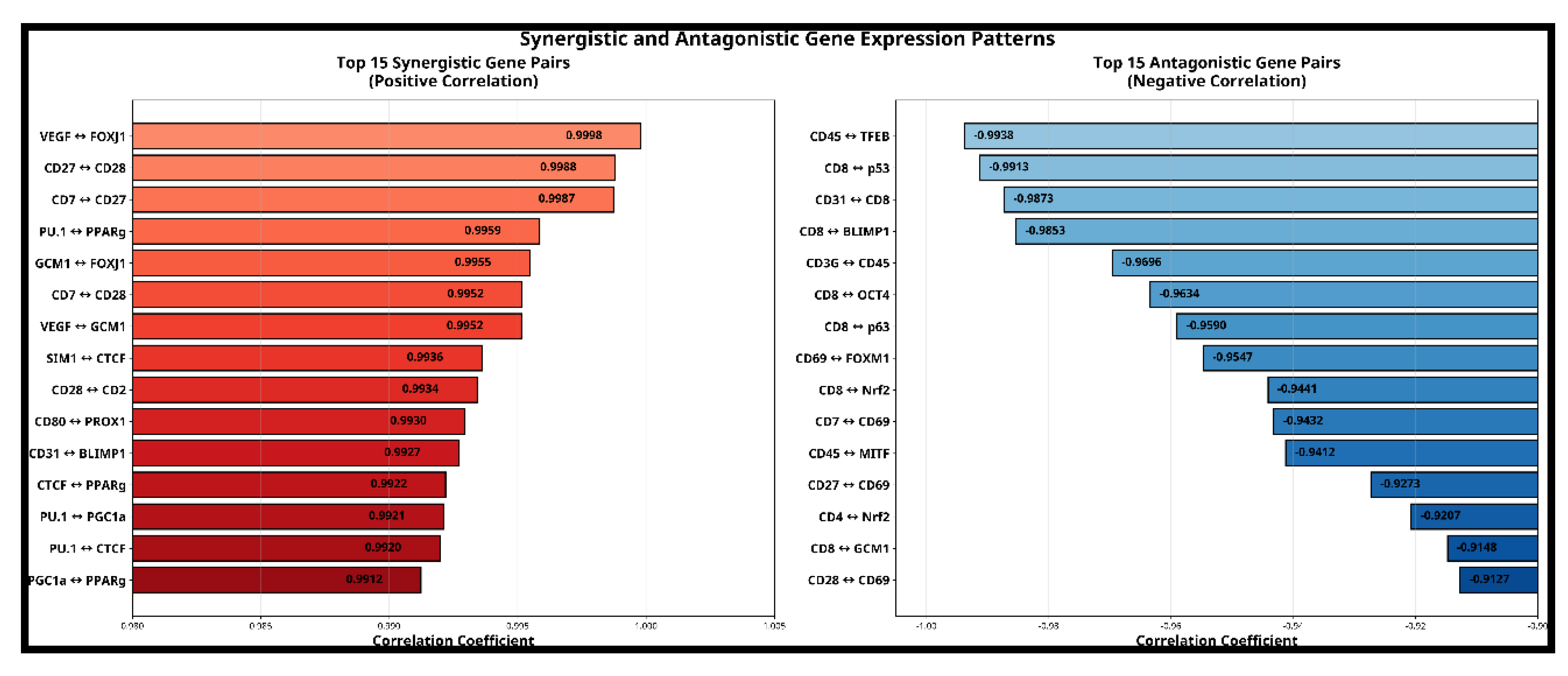
- Synergistic activation of metabolic master regulators creates coordinated enhancement of T-cell fitness,
- Antagonistic regulation of immune cell subsets optimizes the balance between effector and suppressor functions,
- Differential cell- type regulation, exemplified by the CDS- p53 antagonism, suggests selective protection of immune cells while targeting cancer cells.
Metabolic Reprogramming as a Foundation for Anti-Tumor Immunity
The p53 Paradox: Differential Regulation Across Cell Types
Immune Balance Optimization: Shifting from Suppression to Effector Function
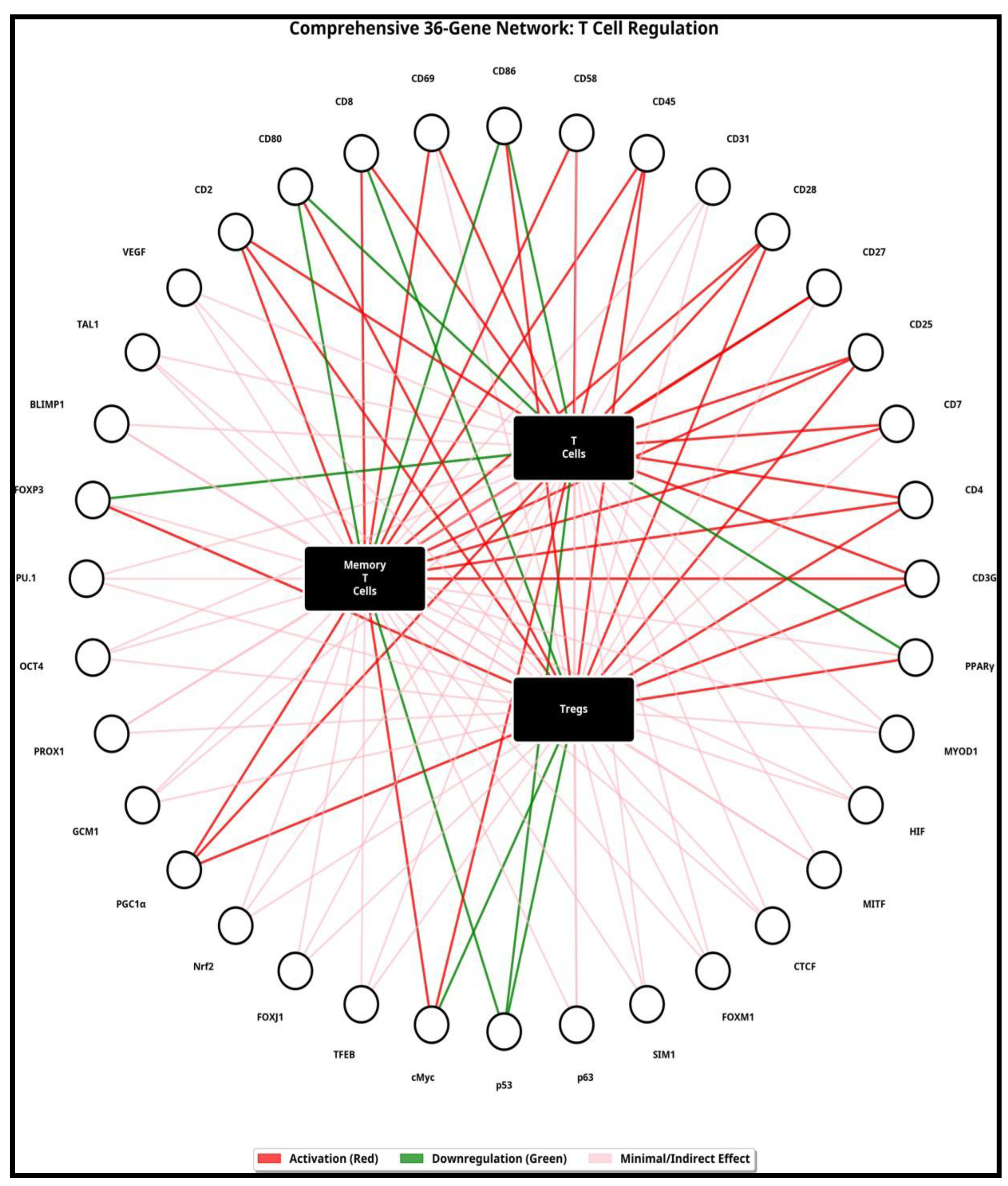
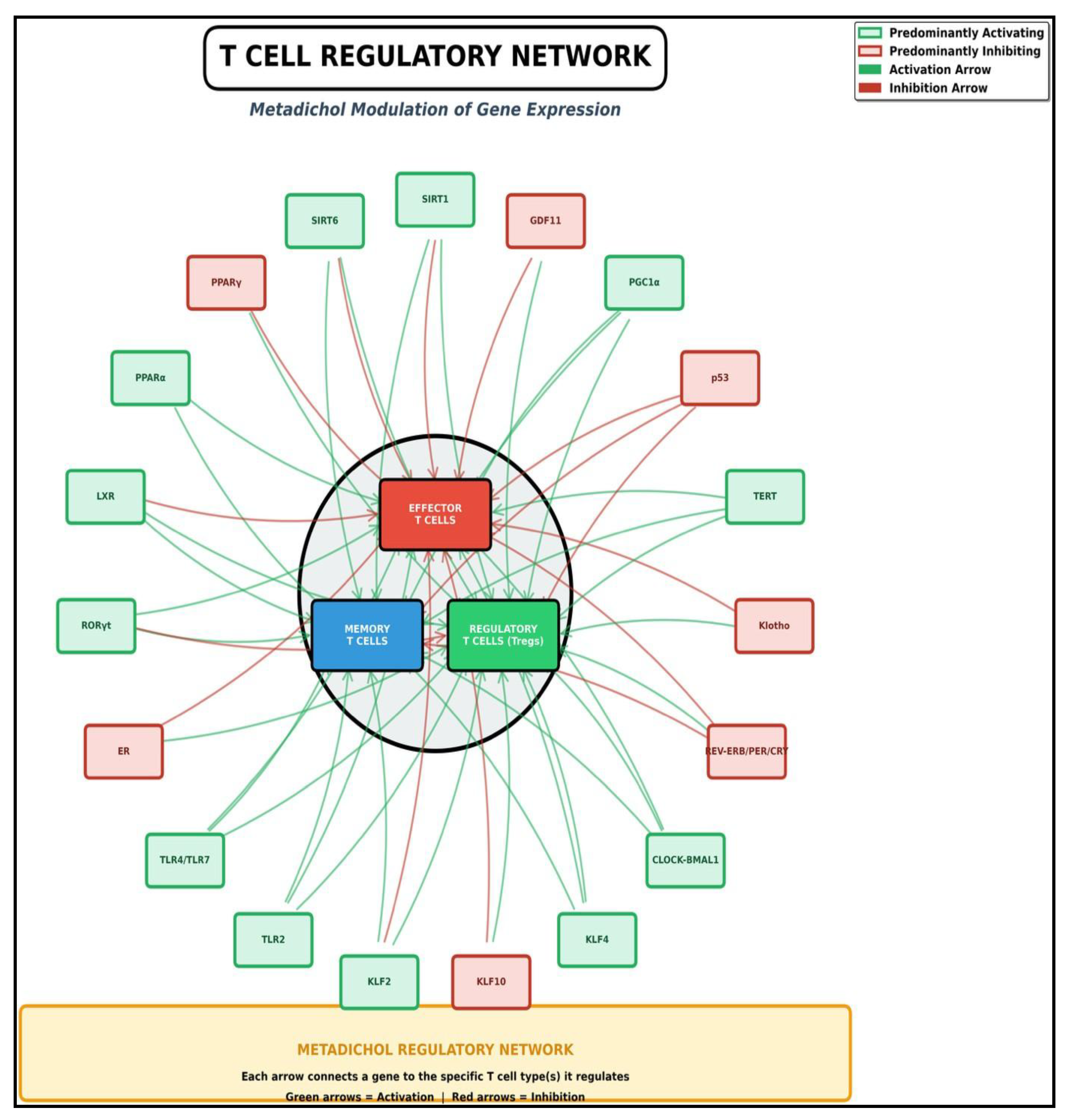
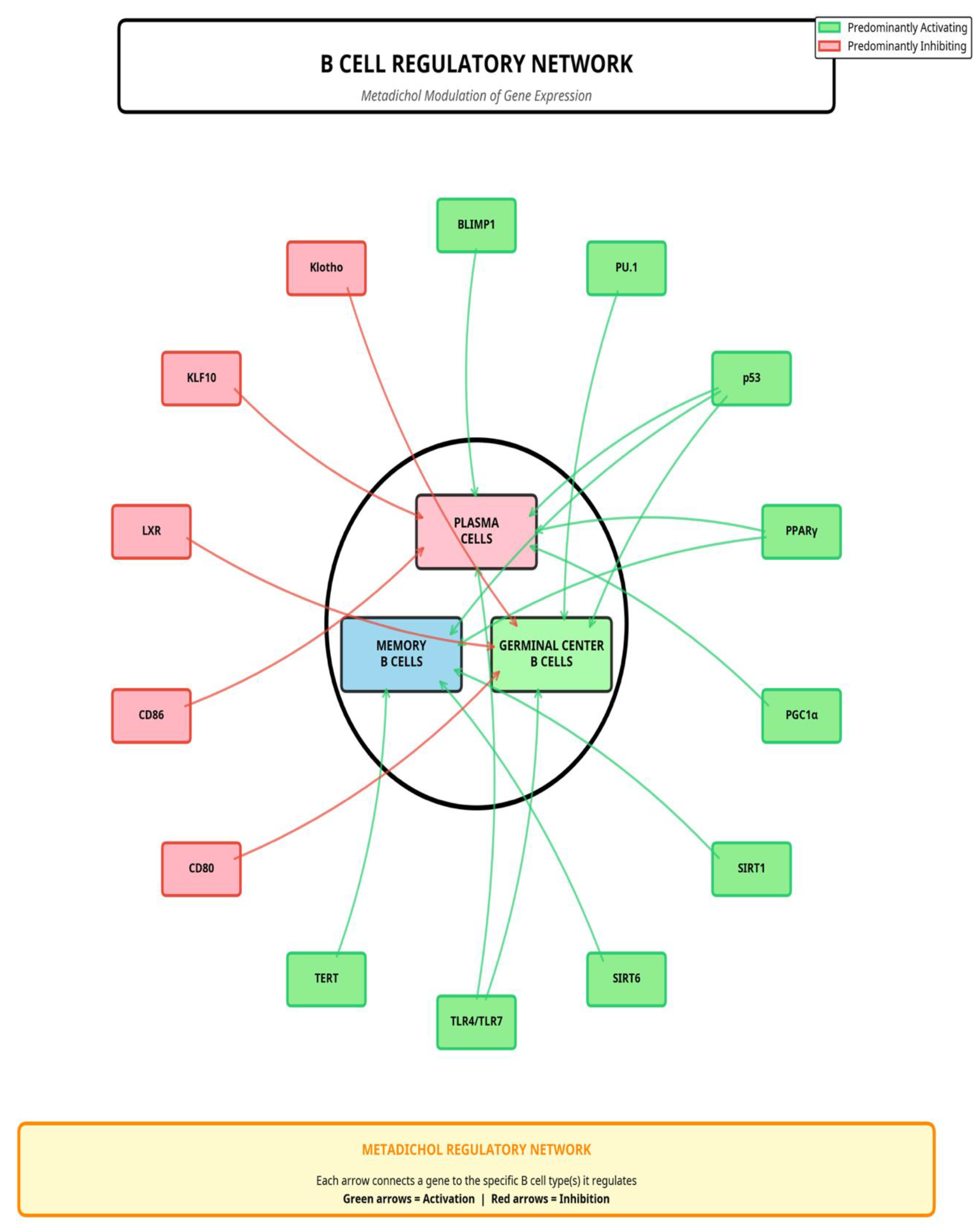
Modulation of T, B, and NK Cell Subsets
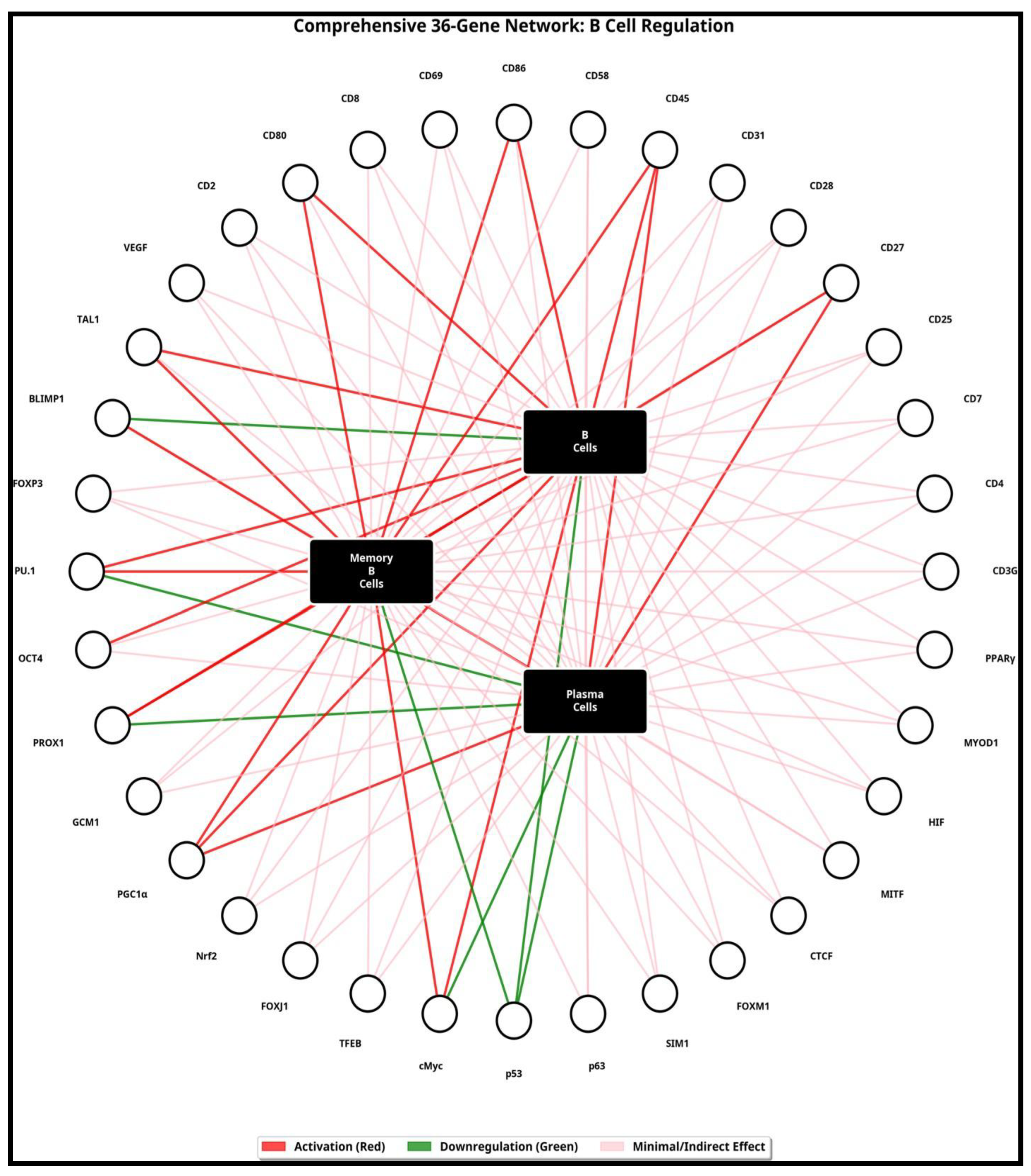
| Gene | Regulation | Effect on B Cells | Effect on Memory B Cells | Effect on Plasma Cells |
| CD3G | ↑ | Minimal effect T cell marker | Minimal effect | Minimal effect |
| CD4 | ↑ | Minimal effect primarily T cell marker | Minimal effect | Minimal effect |
| CD7 | ↓ | Minimal effect primarily T/NK marker | Minimal effect | Minimal effect |
| CD8 | ↑ | Minimal effect T cell marker | Minimal effect | Minimal effect |
| CD25 | ↑/↓ | Modulates B cell activation | May affect memory B cell survival | Minimal effect |
| CD27 | ↓ | Reduced B cell activation | Impaired memory B cell survival and recall | Reduced plasma cell ifferentiation |
| CD28 | ↑/↓ | Minimal direct effect | Minimal effect | Minimal effect |
| CD31 | ↓ | Reduced B cell migration | May affect memory B cell trafficking | Minimal effect |
| CD45 | ↓ | Altered BCR signaling threshold | May affect memory B cell activation | Affects plasma cell function |
| CD58 | ↑ | Enhanced B cell-T cell interactions | Supports memory B cell formation | Minimal effect |
| CD69 | ↑ | Early B cell activation marker | May affect memory B cell retention | Minimal effect |
| CD80 | ↓ | Reduced B cell costimulation | May impair memory B cell function | Reduced plasma cell activation |
| CD86 | ↓ | Reduced B cell costimulation | May impair memory B cell function | Reduced plasma cell activation |
| CD2 | ↑ | Minimal direct effect on B cells | Minimal effect | Minimal effect |
| VEGF | ↑ | Promotes B cell survival and migration | Supports memory B cell niches | Supports plasma cell survival n bone marrow |
| TAL1 | ↓ | Affects early B cell development | May affect memory B cell precursors | Minimal effect |
| BLIMP1 | ↑ | Promotes plasma cell differentiation | Inhibits memory B cell formation | Essential for plasma cell development and antibody secretion |
| FOXP3 | ↑ | Minimal direct effect | Minimal effect | Minimal effect |
| PU.1 | ↑ | Critical for B cell development and function | Supports memory B cell formation | Regulates plasma cell gene expression |
| POU5F1 | ↑ | Promotes B cell stemness | May enhance memory B cell potential | Minimal effect |
| PROX1 | ↓ | Minimal direct effect | Minimal effect | Minimal effect |
| GCM1 | ↑ | Minimal direct effect on B cells | Minimal effect | Minimal effect |
| PGC1α | ↑↑ | Enhances B cell metabolic fitness | Critical for memory B cell longevity | Supports plasma cell antibody production |
| Nrf2 | ↓ | Reduced antioxidant response in B cells | May affect memory B cell survival | May affect plasma cell longevity |
| FOXJ1 | ↑ | Minimal direct effect | Minimal effect | Minimal effect |
| TFEB | ↓ | Reduced autophagy affects B cell metabolism | May affect memory B cell survival | Affects plasma cell protein folding capacity |
| cMyc | ↑ | Promotes B cell proliferation | Inhibits memory B cell formation | Drives plasma cell idfferentiation |
| p53 | ↑↑ | Regulates B cell survival and apoptosis | May affect memory B cell quality | Regulates plasma cell survival |
| p63 | ↑ | Minimal direct effect | Minimal effect | Minimal effect |
| SIM1 | ↑ | Minimal direct effect | Minimal effect | Minimal effect |
| FOXM1 | ↓ | Reduced B cell proliferation | May affect memory B cell formation | Minimal effect |
| CTCF | ↓ | Altered immunoglobulin gene regulation | May affect memory B cell epigenetics | Affects plasma cell antibody class switching |
| MITF | ↓/↑ | Minimal direct effect | Minimal effect | Minimal effect |
| HIF | ↑ | Enhances B cell function in hypoxia | May affect memory B cell metabolism | Supports plasma cell survival in bone marrow |
| MYOD1 | ↓ | Minimal direct effect on B cells | Minimal effect | Minimal effect |
| PPARγ | ↑↑↑ | Modulates B cell metabolism and function | Supports memory B cell metabolism | Critical for plasma cell survival and antibody production |
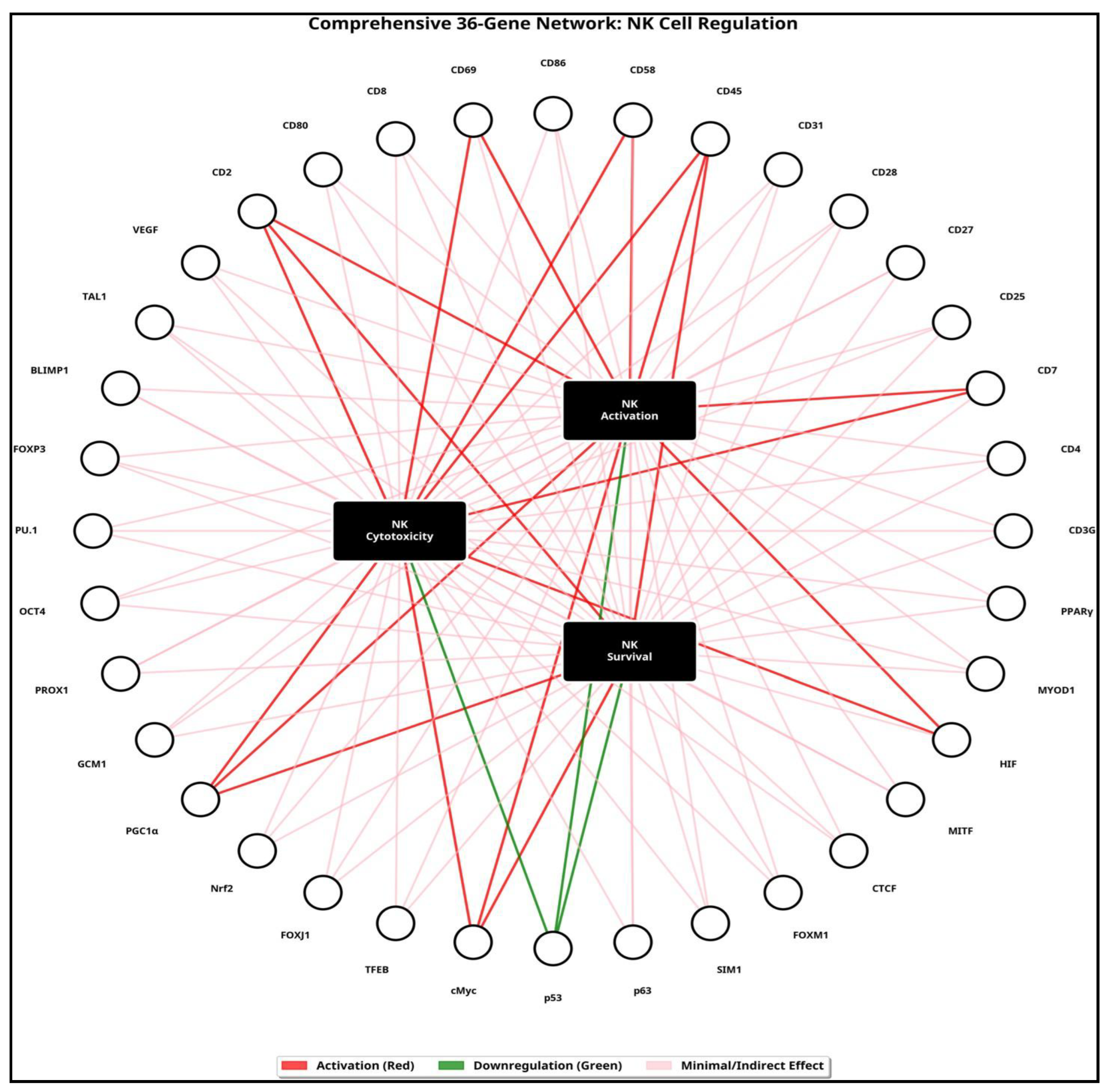
|
Gene |
Regulation |
Effect on NK Cell Activation |
Effect on NK Cell Cytotoxicity |
Effect on NK Cell Survival |
|
CD3G |
↑ |
Minimal effect T cell marker |
Minimal effect |
Minimal effect |
|
CD4 |
↑ |
Minimal effect T cell marker |
Minimal effect |
Minimal effect |
|
CD7 |
↓ |
Reduced NK cell activation |
May affect cytotoxicity |
May affect survival |
|
CD8 |
↑ |
Enhanced NK cell activation |
May enhance cytotoxicity |
Supports survival |
|
CD25 |
↑/↓ |
Modulates IL-2 responsiveness and activation |
Affects IL-2-driven cytotoxicity |
Critical for IL-2-mediated survival |
|
CD27 |
↓ |
Reduced NK cell activation |
May impair cytotoxicity |
Reduced survival signals |
|
CD28 |
↑/↓ |
Modulates NK cell costimulation |
May affect cytotoxicity |
Affects survival signals |
|
CD31 |
↓ |
Reduced NK cell migration |
May affect target cell engagement |
Minimal effect |
|
CD45 |
↓ |
Altered NK cell signaling threshold |
May affect cytotoxic signaling |
Affects survival signaling |
|
CD58 |
↑ |
Enhanced NK cell-target cell interactions | Promotes cytotoxic synapse formation |
Supports survival |
|
CD69 |
↑ |
Early NK cell activation marker |
Promotes cytotoxicity |
Supports tissue retention and survival |
|
CD80 |
↓ |
Reduced costimulatory signaling |
May impair cytotoxicity |
Reduced survival signals |
| CD86 | ↓ | Reduced costimulatory signaling | May impair cytotoxicity | Reduced survival signals |
|
CD2 |
↑ |
Enhanced NK cell activation |
Promotes cytotoxic function |
Supports survival |
|
VEGF |
↑ |
Promotes NK cell recruitment |
May enhance cytotoxicity |
Supports NK cell survival |
|
TAL1 |
↓ |
Affects NK cell development |
May affect cytotoxic potential |
May affect survival |
|
BLIMP1 |
↑ |
Promotes NK cell maturation |
Enhances cytotoxic function |
Supports survival |
|
FOXP3 |
↑ |
May suppress NK cell activation |
May reduce cytotoxicity |
Variable effect |
|
PU.1 |
↑ |
Critical for NK cell development |
Regulates cytotoxic gene expression |
Supports survival |
|
OCT4 |
↑ |
Promotes NK cell stemness |
May affect cytotoxic potential |
Enhances survival |
|
PROX1 |
↓ |
Minimal direct effect |
Minimal effect |
Minimal effect |
|
GCM1 |
↑ |
Minimal direct effect |
Minimal effect |
Minimal effect |
|
PGC1α |
↑↑ |
Enhances NK cell metabolic fitness and activation | Critical for sustained cytotoxicity | Essential for NK cell longevity |
|
Nrf2 |
↓ |
Reduced antioxidant response |
May impair cytotoxicity under oxidative stress |
May reduce survival |
|
FOXJ1 |
↑ |
Minimal direct effect |
Minimal effect |
Minimal effect |
|
TFEB |
↓ |
Reduced autophagy affects NK cell metabolism | May affect cytotoxic granule biogenesis | May reduce survival |
|
cMyc |
↑ |
Promotes NK cell proliferation and activation |
Enhances cytotoxic function |
Supports survival and expansion |
Hierarchical Regulatory Cascade
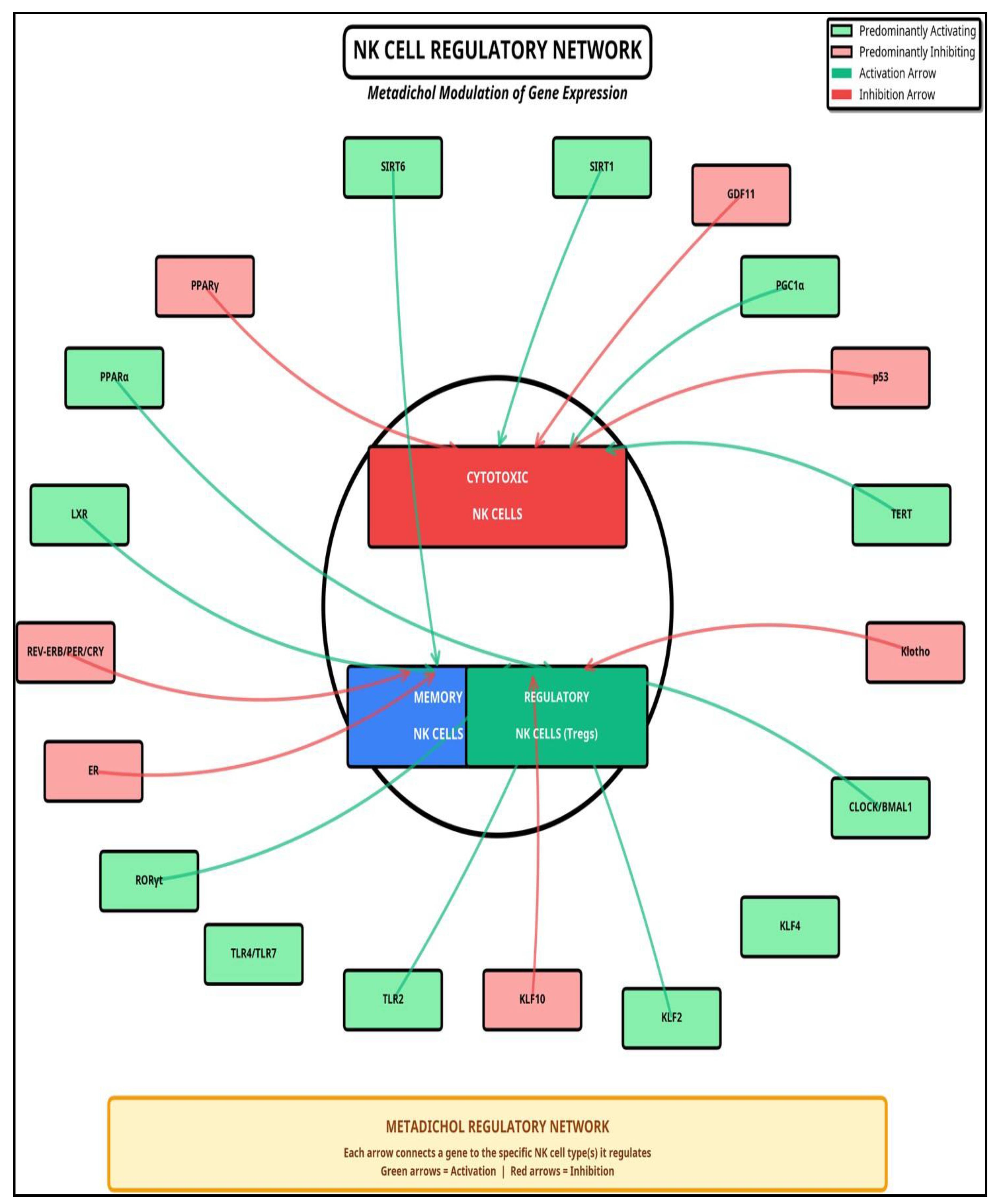
Comparison with Other Small Molecules in Cancer Immunotherapy
Clinical Implications and Therapeutic Potential
Summary and Conclusions: Metadichol as a Master Regulator of Immunity
T-Cell Modulation: Balancing Act for Optimal Immunity
B-Cell and NK-Cell Modulation: A Multi-Pronged Attack
Mechanism of Action: Nuclear Receptors as the Central Hub
Beyond a Single Pathway
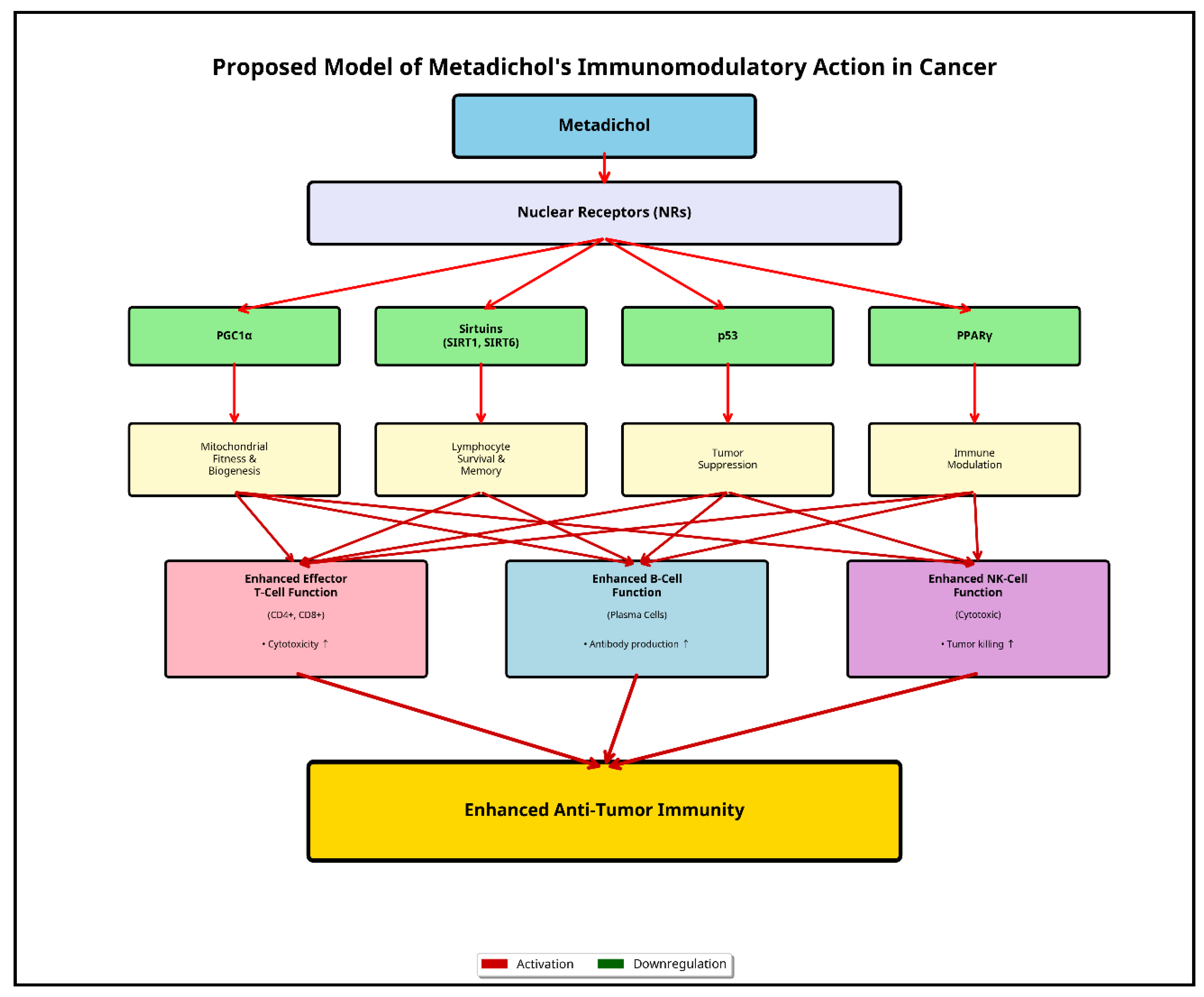
Additional Mechanisms and Safety Profile
Conclusions
Supplementary
Conflicts of Interest
| Glossary of Abbreviations | ||||
| CD3G | CD3 gamma subunit of T-cell receptor complex | FOXJ1 | forkhead box J1 | |
| CD4 | CD4 molecule | TFEB | transcription factor EB | |
| CD7 | CD7 molecule | Myc | MYC proto-oncogene, bHLH transcription factor | |
| CD25 | Interleukin 2 receptor subunit alpha | p53 | tumor protein p53 | |
| CD27 | CD27 molecule | p63 | tumor protein p63 | |
| CD28 | CD28 molecule | SIM1 | SIM bHLH transcription factor 1 | |
| CD31 | platelet and endothelial cell adhesion molecule 1 | FOXM1 | forkhead box M1 | |
| CD45 | PTPRC protein tyrosine phosphatase receptor type C | CTCF | CCCTC-binding factor | |
| CD58 | CD58 molecule | MITF | melanocyte inducing transcription factor | |
| CD86 | CD 86 molecule | HIF1 | hypoxia inducible factor 1 subunit alpha | |
| CD69 | CD69 molecule | MYOD1 | myogenic differentiation 1 | |
| CD8 | CD8 subunit alpha | PPARg | Pperoxisome proliferator activated receptor gamma | |
| CD80 | CD80 molecule | Nrf2 | NFE2 like bZIP transcription factor 2 | |
| CD2 | CD2 molecule | KL | Klotho | |
| VEGF | vascular endothelial growth factor A | PD1 | Programed cell death i | |
| TAL1 | TAL bHLH transcription factor 1, erythroid differentiation factor | PDL1 | Programmed death-ligand 1 | |
| BLIMP1 | PRDM1 PR/SET domain 1 | CTLA | Cytotoxic T-lymphocyte associated protein 4, also known as CD152 | |
| FOXP3 | forkhead box P3 | KLF | Krüppel-like family of transcription factors | |
| PU.1 | SPI1 Spi-1 proto-oncogene | Tert | Telomerase reverse Transcriptase | |
| OCT4 | POU5F1 POU class 5 homeobox 1 | GCM1 | glial cells missing transcription factor 1 | |
| PROX1 | Prospero homeobox 1 | PGC1a | PPARG coactivator 1 alpha | |
References
- Sakaguchi S, Sakaguchi N, Asano M, Itoh M, Toda M. Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains CD25. Breakdown of a single mechanism of self-tolerance causes various autoimmune diseases. J Immunol. 1995;155(3):1151-1164. [CrossRef]
- Brunkow ME, Jeffery EW, Hjerrild KA, et al. Disruption of a new forkhead/winged-helix protein, scurfin, results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat Genet. 2001;27(1):68- 73. [CrossRef]
- Ramsdell F, Ziegler SF. FOXP3 and scurfy: how it all began. Nat Rev Immunol. 2014;14(5):343-349. [CrossRef]
- Sakaguchi S, Yamaguchi T, Nomura T, Ono M. Regulatory T cells and immune tolerance. Cell. 2008;133(5):775-787. [CrossRef]
- Fontenot JD, Gavin MA, Rudensky AY. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat Immunol. 2003;4(4):330-336. [CrossRef]
- Burzyn D, Kuswanto W, Kolodin D, et al. A special population of regulatory T cells potentiates muscle repair. Cell. 2013;155(6):1282-1295. [CrossRef]
- Mathis D, Benoist C. Microbiota and autoimmune disease: the hosted self. Cell Host Microbe. 2011;10(4):297-301. [CrossRef]
- Belkaid Y, Harrison OJ. Homeostatic immunity and the microbiota. Immunity. 2017;46(4):562-576. [CrossRef]
- Bluestone JA, Tang Q. Treg cells-the next frontier of cell therapy. Science. 2018;362(6411):154-155. [CrossRef]
- Sharabi A, Tsokos MG, Ding Y, Malek TR, Klatzmann D, Tsokos GC. Regulatory T cells in the treatment of disease. Nat Rev Drug Discov. 2018;17(11):823-844. [CrossRef]
- Sung H, Ferlay J, Siegel RL, et al. Global Cancer Statistics. [CrossRef]
- 2020. GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71(3):209-249.
- Sharma P, Hu-Lieskovan S, Wargo JA, Ribas A. Primary, Adaptive, and Acquired Resistance to Cancer Immunotherapy. Cell. 2017;168(4):707-723. [CrossRef]
- Ribas A, Wolchok JD. Cancer immunotherapy using checkpoint blockade. Science. 2018;359(6382):1350-1355. [CrossRef]
- Munn DH, Bronte V. Immune suppressive mechanisms in the tumor microenvironment. Curr Opin Immunol. 2016;39:1-6. [CrossRef]
- Ikeda H, Togashi Y. Immune evasion through mitochondrial transfer in the tumor microenvironment. Nature. 2025;638:182-190. [CrossRef]
- Klein K, He K, Younes AI, et al. Role of Mitochondria in Cancer Immune Evasion and Potential Therapeutic Approaches. Front Immunol. 2020;11:573326. [CrossRef]
- Tanaka A, Sakaguchi S. Regulatory T cells in cancer immunotherapy. Cell Res. 2017;27(1):109- 118. [CrossRef]
- Tanaka A, Sakaguchi S. Targeting Treg cells in cancer immunotherapy. Eur J Immunol. 2019;498:1140-1146. [CrossRef]
- Bahrambeigi S, Molaparast M, Sohrabi F, et al. Targeting PPAR ligands as possible approaches for metabolic reprogramming of T cells in cancer immunotherapy. Immunol Lett. 2020;219:12- 22. [CrossRef]
- Cheng HS, Tan WR, Low ZS, et al. PPARs and Tumor Microenvironment: The Emerging Roles of the Metabolic Master Regulators in Tumor Stromal-Epithelial Crosstalk and Carcinogenesis. Cancers Basel. 2021;13(9):2153. [CrossRef]
- Chowdhury PS, Chamoto K, Kumar A, Honjo T. PPAR-Induced Fatty Acid Oxidation in T Cells Increases the Number of Tumor-Reactive CDS+ T Cells and Facilitates Anti- PD-1 Therapy. Cancer Immunol Res. 201S;6(11):1375-13S7. [CrossRef]
- Dumauthioz N, Tschumi B, Wenes M, et al. Enforced PGC-1α expression promotes CDS T cell fitness, memory formation and antitumor immunity. Cell Mol Immunol. 2021;18(7):1761-1771. [CrossRef]
- Hori S, Nomura T, Sakaguchi S. Control of regulatory T cell development by the transcription factor Foxp3. Science. 2003;2995609:1057-1061. [CrossRef]
- Wang Y, Huang Y, Guan F, et al. From metabolism to malignancy: the multifaceted role of PGC1α in cancer. Front Oncol. 2024;14:13S3S09. [CrossRef]
- Binder C, Cvetkovski F, Sellberg F, et al. CD2 Immunobiology. Front Immunol. 2020;11:1090. [CrossRef]
- Li B, Wang X, Hsu C, et al. Cis interactions between CD2 and its ligands on T cells are required for T cell activation. Sci Immunol. 2022;7(74):eabn6373. [CrossRef]
- Moingeon P, Chang HC, Wallner BP, Stebbins C, Frey AZ, Reinherz EL. CD2-mediated adhesion facilitates T lymphocyte antigen recognition. Nature. 1989;339339(6225):544-547. [CrossRef]
- Van der Merwe PA, Barclay AN, Mason DW, Davies EA, Morgan BP, Tonegawa S. Topology of the CD2-CD48 cell-adhesion molecule complex: implications for antigen recognition by T cells. Curr Biol. 1995;5(1):74-84. [CrossRef]
- Killeen N, Stuart SG, Littman DR. Development and function of T cells in mice with a disrupted CD2 gene. EMBO J. 1992;11(12):4329-4336. [CrossRef]
- Rowe JH, Stadinski BD, Henderson LA, et al. Patients with CD3G mutations reveal a role for human CD3γ in Treg diversity and suppressive function. Blood. 2018;131(21):2335-2345. [CrossRef]
- Torres PS, Zapata DA, Pacheco-Castro A, et al. Contribution of CD3γ to TCR regulation and signaling in human mature T lymphocytes. Int Immunol. 2002;14(11):1357-1363. [CrossRef]
- Menon AP, Lizee G, Hwu P. Modulating T Cell Responses by Targeting CD3. Cancers Basel. 2023;15(4):1189. [CrossRef]
- Xin W, Li Y, Wu J, et al. Structures of human γδ T cell receptor--CD3 complex. Nature. 2024;630(8015):222-230. [CrossRef]
- National Center for Biotechnology Information. Gene: CD3G CD3 gamma subunit of T-cell receptor complex. Updated September 9, 2025. Accessed November 2, 2025. https://[www.ncbi.nlm.nih.gov/gene/917].
- Janeway CA Jr. The co-receptor function of CD4. Semin Immunol. 1991;3(3):153-160.
- Rushdi MN, Sethi A, Grewal R, et al. Cooperative binding of T cell receptor and CD4 to peptide-MHC class II complex. Nat Commun. 2022;13(1):6466.
- Mørch AM, Pedersen SF, Holst PJ. Coreceptors and TCR Signaling - the Strong and the Weak of It. Front Cell Dev Biol. 2020;8:597627.
- Swain SL, McKinstry KK, Strutt TM. Expanding roles for CD4+ T cells in immunity to viruses. Nat Rev Immunol. 2012;12(2):136-148. [CrossRef]
- Kitchen SG, Jones-Trower A, Zack JA. The CD4 molecule on CD8+ T lymphocytes directly enhances the immune response to viral immunogen. Proc Natl Acad Sci U S A. 2005;1024(4):15962-15967.
- Sempowski GD, Haynes BF. Structure and function of the CD7 molecule. Immunol Res. 1999;20(2):115-123. [CrossRef]
- Bárcena A, Muench MO, Galy AH, Chen X, Tórre-Cisneros J, Khavari P. CD7 expression in the early stages of T- and myeloid-cell development. Clin Immunol Immunopathol. 1993;68(2):123-132.
- Aandahl EM, Quigley MF, Moretto WJ, Nixon DF, Våbenø J. CD7 is a differentiation marker that.
- Hao QL, Zhu J, Price MA, et al. Human intrathymic lineage commitment is marked by differential CD7 expression: identification of CD7- lympho-myeloid thymic progenitors. Blood. 2008;111(3):1318- 1326. [CrossRef]
- Liu J, Zhou Y, Jiang W, et al. Targeted CD7 CAR T-cells for treatment of T-Lymphocyte Malignancies. Front Immunol. 2023;14:1170968.
- Raskov H, Orhan A, Gaggar S, Gögenur I. Cytotoxic CD8+ T cells in cancer and cancer immunotherapy. Br J Cancer. 2021;124(2):359-367. [CrossRef]
- Daniels MA, Devine L, Miller JD, et al. CD8 binding to MHC class I molecules is influenced by T cell maturation and glycosylation. Immunity. 2001;15(6):1051-1061. [CrossRef]
- Srinivasan S, Purbhoo MA, Tsuchida CA, et al. Structure, function, and immunomodulation of the CD8 co-receptor. Front Immunol. 2024;15:1412513.
- Zhang N, Bevan MJ. CD8+ T cells: foot soldiers of the immune system. Immunity. 2011;35(2):161- 168. [CrossRef]
- Koh CH, Pua CJ, Lee J, et al. CD8 T-cell subsets: heterogeneity, functions, and therapeutic potential. Exp Mol Med. 2023;55(11):2287-2299.
- Rolink A, Grawunder U, Winkler TH, Karasuyama H, Melchers F. IL-2 receptor alpha chain CD25, TAC expression defines a crucial stage in pre-B cell development. Int Immunol. 1994;68:1257-1264. [CrossRef]
- Shatrova AN, Misyurin VA, Misyurin AV, et al. Time-Dependent Regulation of IL-2R α-Chain CD25 Expression by TCR Signal Strength and IL-2-Induced STAT5 Signaling in Activated Human Blood T Lymphocytes. PLoS One. 2016;11(12):e0167215.
- Létourneau S, van Leeuwen PA, Krieg C, et al. IL-2- and CD25-dependent immunoregulatory mechanisms in the homeostasis of T-cell subsets. J Allergy Clin Immunol. 2009;1234:758-762. [CrossRef]
- Roifman, CM. Human IL-2 receptor α chain deficiency. Pediatr Res. 2000;481:6-11. [CrossRef]
- Caudy AA, Reddy ST, Chatila T, Atkinson JP, Verbsky JW. CD25 deficiency causes an immune dysregulation, polyendocrinopathy, enteropathy, X-linked-like syndrome, and defective IL-10 expression from CD4 lymphocytes. J Allergy Clin Immunol. 2007;1192:482-487. [CrossRef]
- Watts TH, Yeung KKM, Yu T, Lee S. TNF/TNFR Superfamily Members in Costimulation of T Cell Responses---Revisited. Annu Rev Immunol. 2025;43. [CrossRef]
- Watts, TH. TNF/TNFR family members in costimulation of T cell responses. Annu Rev Immunol. 2005;23:23-68. [CrossRef]
- Starzer AM, Berghoff AS. New emerging targets in cancer immunotherapy: CD27 TNFRSF7. ESMO Open. 2019;4Suppl 3:e000629. [CrossRef]
- Hendriks J, Xiao Y, Borst J. CD27 promotes survival of activated T cells and complements CD28 in generation and establishment of the effector T cell pool. J Exp Med. 2003;1989:1369-1380. [CrossRef]
- Jaeger-Ruckstuhl CA, Jäeger R, Zurbuchen Y, et al. Signaling via a CD27-TRAF2-SHP-1 axis during naive T cell priming regulates memory CD8+ T cell differentiation. Immunity. 2024;572:283-299.e9.
- Esensten JH, Helou YA, Chopra G, Weiss A, Bluestone JA. CD28 costimulation: from mechanism to therapy. Immunity. 2016;445:973-988.
- Beyersdorf N, Hanke T. CD28 co-stimulation in T-cell homeostasis: a recent perspective. Immunotargets Ther. 2015;4:111-122.
- Porciello N, Tuosto L. CD28 costimulatory signals in T lymphocyte activation: Emerging functions and implications for cancer immunotherapy. Semin Immunol. 2016;281:3-11.
- Schildberg FA, Klein SR, Freeman GJ, Sharpe AH. Coinhibitory pathways in the B7-CD28 ligand- receptor family. Immunity. 2016;445:955-972.
- Diehn M, Alizadeh AA, Rando OJ, et al. Genomic expression programs and the integration of the CD28 costimulatory signal in T cell activation. Proc Natl Acad Sci U S A. 2002;9918:11796-11801. [CrossRef]
- Lertkiatmongkol P, Liao D, Mei H, Hu Y, Newman PJ. Endothelial functions of PECAM-1 CD31. Vascul Pharmacol. 2016;87:3-9.
- Woodfin A, Voisin MB, Nourshargh S. PECAM-1: A Multi-Functional Molecule in Inflammation and Vascular Biology. Arterioscler Thromb Vasc Biol. 2007;2712:2514-2523. [CrossRef]
- Ma L, Mauro C, Marelli-Berg FM. CD31 Exhibits Multiple Roles in Regulating T Lymphocyte Trafficking In Vivo. J Immunol. 2012;1899:4104-4111. [CrossRef]
- DeLisser HM, Baldwin HS, Albelda SM. Platelet endothelial cell adhesion molecule 1 PECAM- 1/CD31: a multifunctional vascular cell adhesion molecule. Trends Cardiovasc Med. 1997;76:203- 210. [CrossRef]
- Caligiuri, G. CD31 as a Therapeutic Target in Atherosclerosis. Circ Res. 2020;1269:1178-1189. [CrossRef]
- Trowbridge IS, Thomas ML. CD45: an emerging role as a protein tyrosine phosphatase required for lymphocyte activation and development. Annu Rev Immunol. 1994;12:85-116. [CrossRef]
- Al Barashdi MA, Vallance TM, Al-Zakwani I, Bailey K, Majid S. Protein tyrosine phosphatase receptor type C PTPRC or CD45. J Clin Pathol. 2021;749:548-552.
- Charbonneau H, Tonks NK, Walsh KA, Fischer EH. The leukocyte common antigen CD45: a putative receptor-linked protein tyrosine phosphatase. Proc Natl Acad Sci U S A. 1988;8519:7182-7186. [CrossRef]
- Tonks NK, Charbonneau H, Diltz CD, Fischer EH, Walsh KA. Demonstration that the leukocyte common antigen CD45 is a protein tyrosine phosphatase. Biochemistry. 1988;2724:8695-8701. [CrossRef]
- Courtney AH, Shvets E, Lu W, et al. CD45 functions as a signaling gatekeeper in T cells. Sci Signal. 2019;12604:eaaw8151. [CrossRef]
- Dustin ML, Olszowy MW, Holdorf AD, et al. Low affinity interaction of human or rat cell adhesion molecule CD2 with its ligand aligns adhering membranes to achieve high physiological affinity. J Biol Chem. 1997;27249:30889-30898.
- Ikemizu S, Sparks MA, van der Merwe PA, Jones EY, Stuart DI. Crystal structure of the CD2-binding domain of CD58 lymphocyte function-associated antigen 3 at 1.8-Å resolution. Proc Natl Acad Sci U S A. 1999;968:4289-4294.
- National Center for Biotechnology Information. Gene: CD58 CD58 molecule. Updated August 19, 2025. Accessed November 2, 2025. https://[www.ncbi.nlm.nih.gov/gene/965].
- Cibrián D, Sánchez-Madrid F. CD69: from activation marker to metabolic gatekeeper. Eur J Immunol. 2017;476:946-953. [CrossRef]
- Ziegler SF, Ramsdell F, Hjerrild KA, et al. The activation antigen CD69. Stem Cells. 1994;125:456- 465. [CrossRef]
- Sancho D, Gómez M, Sánchez-Madrid F. CD69 is an immunoregulatory molecule induced following activation. Trends Immunol. 2005;263:136-140. [CrossRef]
- Yu L, Wang Y, Yang J, et al. CD69 enhances immunosuppressive function of regulatory T-cells and attenuates colitis by prompting IL-10 production. Cell Death Dis. 2018;99:905.
- Marzio R, Mauël J, Corradin SB. CD69 and regulation of the immune function. Immunopharmacol Immunotoxicol. 1999;213:565-582.
- Satoh J, Lee HC, Kim SU, et al. T-cell costimulatory molecules B7-1 CD80 and B7-2 CD86 in the NOD mouse. Diabetes. 1995;4410:1156-1162.
- Butte MJ, Keir ME, Phamduy TB, Sharpe AH, Freeman GJ. Programmed death-1 ligand 1 interacts specifically with the B7-1 costimulatory molecule to inhibit T cell responses. Immunity.2007:27(1):111-122. [CrossRef]
- Ransom, DM. CD28, CTLA-4 and their ligands: who does what and to whom? Immunology. 2000;1012:169-177.
- Collins M, Ling V, Carreno BM. The B7 family of immune-regulatory ligands. Genome Biol. 2005;66:223. [CrossRef]
- Linsley PS, Peach R, Gladstone P, Bajorath J. Extending the B7 CD80 gene family. Protein Sci. 1994;38:1341-1343. [CrossRef]
- Vasilevko V, Ghochikyan A, Holterman MJ, Agadjanyan MG. CD80 B7-1 and 9B7-2 are functionally equivalent in the initiation and maintenance of CD4+ T-cell proliferation after activation with suboptimal doses of PHA. DNA Cell Biol. 2002;213:137-149. [CrossRef]
- Jeannin P, Magistrelli G, Aubry JP, et al. Soluble CD86 is a costimulatory molecule for human T lymphocytes. Immunity. 2000;13(3):303-312. [CrossRef]
- Kennedy A, Waters E, Rowshanravan B, et al. Differences in CD80 and CD86 transendocytosis reveal CD86 as a key target for CTLA-4 immune regulation. Nat Immunol. 2022;239:1365-1378. [CrossRef]
- Halliday N, Williams C, Kennedy A, et al. CD86 is a selective CD28 ligand supporting FoxP3+ regulatory T cell homeostasis in the presence of high levels of CTLA-4. Front Immunol. 2020;11:600000. [CrossRef]
- National Center for Biotechnology Information. Gene: CD86 CD86 molecule. Updated September 5, 2025. Accessed November 2, 2025. https://[www.ncbi.nlm.nih.gov/gene/942.].
- Ahmadian M, Suh JM, Hah N, et al. PPARγ signaling and metabolism: the good, the bad and the future. Nat Med. 2013;195:557-566. [CrossRef]
- Lehrke M, Lazar MA. The Many Faces of PPARγ. Cell. 2005;1236:993-999.
- Lin J, Handschin C, Spiegelman BM. Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab. 2005;16:361-370. [CrossRef]
- Halling JF, Pilegaard H. PGC-1α-mediated regulation of mitochondrial function and physiological implications. Appl Physiol Nutr Metab. 2020;459:927-936. [CrossRef]
- He F, Ru X, Wen T. NRF2, a Transcription Factor for Stress Response and Beyond. Int J Mol Sci. 2020;2113:4777. [CrossRef]
- Tonelli C, Chio IIC, Tuveson DA. Transcriptional Regulation by Nrf2. Antioxid Redox Signal. 2018;2917:1727-1745.
- Aubrey BJ, Kelly GL, Janic A, Herold MJ, Strasser A. How does p53 induce apoptosis and how does this relate to p53-mediated tumour suppression? Cell Death Differ. 2018;25(1):104-113. [CrossRef]
- Li Y, Giovannini S, Wang T, et al. p63: a crucial player in epithelial stemness regulation. Oncogene. 2023;4247:3435-3444. [CrossRef]
- Dhanasekaran R, Deutzmann A, Mahauad-Fernandez WD, Hansen AS, Gouw AM, Felsher DW. The MYC oncogene - the grand orchestrator of cancer growth and immune evasion. Nat Rev Clin Oncol. 2022;191:23-36. [CrossRef]
- Rudensky, AY. Regulatory T cells and Foxp3. Immunol Rev. 2011;2411:260-268.
- Yu X, Ng CP, Habacher H, Roy S. Foxj1 transcription factors are master regulators of the motile ciliogenic program. Nat Genet. 2008;4012:1445-1453. [CrossRef]
- Liao GB, Li XZ, Zeng S, et al. Regulation of the master regulator FOXM1 in cancer. Cell Commun Signal. 2018;161:57. [CrossRef]
- Shaffer AL, Lin KI, Kuo TC, et al. Blimp-1 orchestrates plasma cell differentiation by extinguishing the mature B cell gene expression program. Immunity. 2002;171:51-62. [CrossRef]
- Rothenberg EV, Hosokawa H, Ungerbäck J. Mechanisms of Action of Hematopoietic Transcription Factor PU.1 in Initiation of T-Cell Development. Front Immunol. 2019;10:228. [CrossRef]
- Jerabek S, Merino F, Schöler HR, Cojocaru V. OCT4: Dynamic DNA binding pioneers stem cell Pluripotency, Biochim.Biophys.Acts. 2014;18393;138-154. [CrossRef]
- Porcher C, Chagraoui H, Kristiansen MS. SCL/TAL1: a multifaceted regulator from blood development to disease. Blood. 2017;12915:2051-2060. [CrossRef]
- Ducoli L, Detmar M. Beyond PROX1: transcriptional, epigenetic, and noncoding RNA regulation of lymphatic identity and function. Dev Cell. 2021;564:406-426. [CrossRef]
- Jeyarajah MJ, Jaju Bhattad G, Kops BF, Renaud SJ. The multifaceted role of GCM1 during trophoblast differentiation in the human placenta. Proc Natl Acad Sci U S A. 2022;11950:e2203071119. [CrossRef]
- Martini-Stoica H, Xu Y, Ballabio A, Zheng H. The autophagy-lysosomal pathway in neurodegeneration: A TFEB perspective. Trends Neurosci. 2016;394:221-234. [CrossRef]
- Michaud JL, Boucher F, Melnyk A, et al. Sim1 haploinsufficiency causes hyperphagia, obesity and reduction of the paraventricular nucleus of the hypothalamus. Hum Mol Genet. 2001;1014:1465- 1473. [CrossRef]
- Phillips-Cremins JE, Corces VG. Chromatin Insulators: Linking genome organization to cellular function. Mol Cell. 2013;504:461-474. [CrossRef]
- Goding CR, Arnheiter H. MITF-the first 25 years. Genes Dev. 2019;3315-16:983-1007.
- Kaelin WG Jr, Ratcliffe PJ. Oxygen sensing by metazoans: the central role of the HIF hydroxylase pathway. Mol Cell. 2008;304:393-402. [CrossRef]
- Hernández-Hernández JM, García-González EG, Brun CE, Rudnicki MA. The myogenic regulatory factors, determinants of muscle development, cell identity and regeneration. Semin Cell Dev Biol. 2017;72:10-18. [CrossRef]
- M. Vascular Endothelial Growth Factor VEGF and Its Receptor VEGFR Signaling in Angiogenesis: A Crucial Target for Anti- and Pro-Angiogenic Therapies. Genes Cancer. 2011;212:1097-1105.
- P.R. Raghavan US patents 8,722,093 2014, 9,034,383 2015, and 9,006,292 2015.
- Raghavan, PR. Metadichol® A Nano Lipid Emulsion that Expresses All 49 Nuclear Receptors in Stem and Somatic Cells. Arch Clin Biomed Res. 2023;76:543-555. [CrossRef]
- Raghavan, PR. Metadichol®-induced expression of Sirtuins 1-7 in somatic and cancer cells. Med Res Arch. 2024;126. [CrossRef]
- Raghavan, PR. Metadichol, a modulator that controls expression of toll-like receptors in cancer cell lines. Br J Cancer Res. 2024;73:720-732. [CrossRef]
- Raghavan, PR. Metadichol induced expression of toll receptor family members in peripheral blood mononuclear cells. Med Res Arch. 2024;128. [CrossRef]
- Raghavan, PR. Metadichol-induced expression of circadian clock transcription factors in human fibroblasts. Med Res Arch. 2024;126. [CrossRef]
- P.R. Raghavan. Metadichol induced CD14 Glycoprotein expression in human embryonic stem cells and fibroblasts. [CrossRef]
- Raghavan, PR. VDR inverse agonism by metadichol enhances VDBP-mediated immunity. Preprints.org. 2025. [CrossRef]
- Raghavan, P. R. Metadichol Stimulates Gene Expression Across Mammalian Species: Dose Response Characterization and Implications for Restoring Vitamin C Biosynthesis. Preprints 2025, 2025081935. [Google Scholar] [CrossRef]
- Raghavan, PR. Metadichol orchestrates pluripotency via nuclear receptors during cellular reprogramming. Med Res Arch. 2025;136. [CrossRef]
- Raghavan P. R.. Beyond Rapamycin: Metadichol Represents a New Class of Multi-Target mTOR Modulators. Medical Research Archives, S.l., v. 13, n. 9, sep. 2025. ISSN 2375-1924. [CrossRef]
- Qian L, Zhu Y, Deng C, et al. Peroxisome proliferator-activated receptor gamma coactivator-1 PGC-1 family in physiological and pathophysiological process and diseases. Signal Transduct Target Ther. 2024;91:50. [CrossRef]
- Reina-Campos M, Scharping NE, Goldrath AW. CD8+ T cell metabolism in infection and cancer. Nat Rev Immunol. 2021;2111:718-738. [CrossRef]
- Scharping NE, Menk AV, Moreci RS, et al. The tumor microenvironment represses T cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity. 2016;452:374-388. [CrossRef]
- Michalek RD, Gerriets VA, Jacobs SR, et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J Immunol. 2011;1866:3299-3303. [CrossRef]
- Chowdhury PS, Chamoto K, Kumar A, Honjo T. PPAR-induced fatty acid oxidation in T cells increases the number of tumor-reactive CD8+ T cells and facilitates anti-PD-1 therapy. Cancer Immunol Res. 2018;6(11):1375-1387. [CrossRef]
- Dumauthioz N, Tschumi B, Wenes M, et al. Enforced PGC-1α expression promotes CD8 T cell fitness, memory formation and antitumor immunity. Cell Mol Immunol. 2021;187:1761-1771. [CrossRef]
- Puigserver P, Spiegelman BM. Peroxisome proliferator-activated receptor-gamma coactivator 1 alpha PGC-1 alpha: transcriptional coactivator and metabolic regulator. Endocr Rev. 2003;241:78-90. [CrossRef]
- Kishton RJ, Sukumar M, Restifo NP. Metabolic regulation of T cell longevity and function in tumor immunotherapy. Cell Metab. 2017;261:94-109. [CrossRef]
- Levine, AJ. p53: 800 million years of evolution and 40 years of discovery. Nat Rev Cancer. 2020;208:471-480. [CrossRef]
- Watanabe M, Moon KD, Vacchio MS, Hathcock KS, Hodes RJ. Downmodulation of tumor suppressor p53 by T cell receptor signaling is critical for antigen-specific CD4+ T cell responses. Immunity. 2014;405:681-691. [CrossRef]
- Kruiswijk F, Labuschagne CF, Vousden KH. p53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat Rev Mol Cell Biol. 2015;167:393-405. [CrossRef]
- Bieging KT, Mello SS, Attardi LD. Unravelling mechanisms of p53-mediated tumour suppression. Nat Rev Cancer. 2014;14(5):359-370. [CrossRef]
- Jiang L, Kon N, Li T, et al. Ferroptosis as a p53-mediated activity during tumour suppression. Nature. 2015;5207545:57-62. [CrossRef]
- Togashi Y, Shitara K, Nishikawa H. Regulatory T cells in cancer immunosuppression - implications for anticancer therapy. Nat Rev Clin Oncol. 2019;166:356-371. [CrossRef]
- Tanaka A, Sakaguchi S. Targeting Treg cells in cancer immunotherapy. Eur J Immunol. 2019;498:1140-1146. [CrossRef]
- Arce Vargas F, Furness AJS, Litchfield K, et al. Fc effector function contributes to the activity of human anti-CTLA-4 antibodies. Cancer Cell. 2018;334:649-663.e4. [CrossRef]
- Sharpe AH, Freeman GJ. The B7-CD28 superfamily. Nat Rev Immunol. 2002;22:116-126. [CrossRef]
- Buck MD, O\'Sullivan D, Klein Geltink RI, et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell. 2016;1661:63-76. [CrossRef]
- Scharping NE, Menk AV, Whetstone RD, Zeng X, Delgoffe GM. Efficacy of PD-1 blockade is potentiated by metformin-induced reduction of tumor hypoxia. Cancer Immunol Res. 2017;5(1):9-16. [CrossRef]
- St-Pierre J, Drori S, Uldry M, Silvaggi JM, Rhee J, Jäger S, Handschin C, Zheng K, Lin J, Yang W, Simon DK, Bachoo R, Spiegelman BM. Suppression of reactive oxygen species and neurodegeneration by the PGC-1 transcriptional coactivators. Cell. 2006 Oct 20;1272:397-408. [CrossRef]
- Bengsch B, Johnson AL, Kurachi M, et al. Bioenergetic insufficiencies due to metabolic alterations regulated by the inhibitory receptor PD-1 are an early driver of CD8+ T cell exhaustion. Immunity. 2016;452:358-373. [CrossRef]
- Levine, AJ. p53, the cellular gatekeeper for growth and division. Cell. 1997;883:323-331.
- Zheng SJ, Lamhamedi-Cherradi SE, Wang P, Xu L, Chen YH. Tumor suppressor p53 inhibits autoimmune inflammation and macrophage function. Diabetes. 2005;545:1423-1428. [CrossRef]
- Lujambio A, Akkari L, Simon J, et al. Non-cell-autonomous tumor suppression by p53. Cell. 2013;1532:449-460. [CrossRef]
- Vousden KH, Prives C. Blinded by the light: the growing complexity of p53. Cell. 2009;1373:413- 431. [CrossRef]
- Kastenhuber ER, Lowe SW. Putting p53 in context. Cell. 2017;1706:1062-1078. [CrossRef]
- Togashi Y, Shitara K, Nishikawa H. Regulatory T cells in cancer immunosuppression - implications for anticancer therapy. Nat Rev Clin Oncol. 2019;166:356-371. [CrossRef]
- Tanaka A, Sakaguchi S. Regulatory T cells in cancer immunotherapy. Cell Res. 2017;27(1):109-118.
- Saleh R, Elkord E. Treg-mediated acquired resistance to immune checkpoint inhibitors. Cancer Lett. 2019;457:168-179. [CrossRef]
- Lenschow DJ, Walunas TL, Bluestone JA. CD28/B7 system of T cell costimulation. Annu Rev Immunol. 1996;14:233-258. [CrossRef]
- Klein Geltink RI, Kyle RL, Pearce EL. Unraveling the complex interplay between T cell metabolism and function. Annu Rev Immunol. 2018;36:461-488.
- Alarcón B, Gil D, Delgado P, Schamel WW. Initiation of TCR signaling: regulation within CD3 dimers. Immunol Rev. 2003;191:38-46. [CrossRef]
- Zaunders JJ, Kelleher AD, Cooper DA. The role of T-cell receptor diversity in HIV-1 infection. Curr Opin Immunol. 2001;134:483-489.
- Rabinowich H, Pricop L, Herberman RB, Whiteside TL. Expression and function of CD7 molecule on human natural killer cells. J Immunol. 1994;1522:517-526. [CrossRef]
- Gao GF, Murata K, Rao Z, et al. Crystal structure of the complex between human CD8αα and HLA-A2. Nature. 1997;387663(3):630-634.
- Létourneau S, van Leeuwen EM, Krieg C, et al. IL-2/anti-IL-2 antibody complexes show strong biological activity by avoiding interaction with IL-2 receptor alpha subunit CD25. Proc Natl Acad Sci USA. 2010;1075:2171-2176.
- Van Gisbergen KP, van der Vlist M, ten Brinke A, et al. The costimulatory molecule CD27 maintains clonally diverse CD8+ T cell responses of low antigen affinity to protect against viral variants. Immunity. 2011;35(1):97-108.
- Linsley PS, Clark EA, Ledbetter JA. T-cell antigen CD28 mediates adhesion with B cells by interacting with activation antigen B7/BB-1. Proc Natl Acad Sci USA. 1990;871(3):5031-5035.
- Hermiston ML, Xu Z, Weiss A. CD45: a critical regulator of signaling thresholds in immune cells. Annu Rev Immunol. 2003;21:107-137. [CrossRef]
- De-la-Fuente C, Pantoja-Uceda D, G-Trevijano ER, et al. The structure and plasticity of the complex formed by the two N-terminal domains of human CD58. J Mol Biol. 2020;4329:3141-3157.
- Davis SJ, van der Merwe PA. The kinetic-segregation model: TCR triggering and beyond. Nat Immunol. 2006;78:803-809. [CrossRef]
- Ferrara N, Gerber HP, LeCouter J. The biology of VEGF and its receptors. Nat Med. 2003;96:669- 676.
- Herblot S, Steff AM, Hugo P, et al. SCL/TAL1 is a positive regulator of thymocyte maturation. Blood. 2000;956:2025-2032.
- Shapiro-Shelef M, Lin KI, McHeyzer-Williams LJ, et al. Blimp-1 is required for the formation of immunoglobulin secreting plasma cells and pre-plasma memory B cells. Immunity. 2003;194:607- 620. [CrossRef]
- Scott EW, Simon MC, Anastasi J, Singh H. Requirement of transcription factor PU.1 in the development of multiple hematopoietic lineages. Science. 1994;2655178:1573-1577. [CrossRef]
- Nichols J, Zevnik B, Anastassiadis K, et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell. 1998;953:379-391. [CrossRef]
- Wigle JT, Oliver G. Prox1 function is required for the development of the murine lymphatic system. Cell. 1999;986:769-778. [CrossRef]
- Anson-Cartwright L, Dawson K, Holmyard D, et al. The glial cells missing-1 protein is essential for branching morphogenesis in the chorioallantoic placenta. Nat Genet. 2000;253:311-314. [CrossRef]
- Scarpulla, RC. Metabolic control of mitochondrial biogenesis through the PGC-1 family regulatory network. Biochim Biophys Acta. 2011;18137:1269-1278. [CrossRef]
- Itoh K, Chiba T, Takahashi S, et al. An Nrf2/small Maf heterodimer mediates the induction of phase II detoxifying enzyme genes through antioxidant response elements. Biochem Biophys Res Commun. 1997;2362:313-322.
- Watanabe M, Moon KD, Vacchio MS, et al. FOXJ1 is essential for Treg cell development and function. J Exp Med. 2014;2112:233-247.
- Sardiello M, Palmieri M, di Ronza A, et al. A gene network regulating lysosomal biogenesis and function. Science. 2009;3255939:473-477. [CrossRef]
- Dang, CV. MYC on the path to cancer. Cell. 2012;1491:22-35. [CrossRef]
- Mills AA, Zheng B, Wang XJ, et al. p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature. 1999;3986729:708-713. [CrossRef]
- Michaud JL, Boucher F, Melnyk A, et al. Sim1 haploinsufficiency causes hyperphagia, obesity and reduction of the paraventricular nucleus of the hypothalamus. Hum Mol Genet. 2001;1014:1465- 1473. [CrossRef]
- Laoukili J, Kooistra MR, Brás A, et al. FoxM1 is required for execution of the mitotic programme and chromosome stability. Nat Cell Biol. 2005;72:126-136. [CrossRef]
- Bell AC, West AG, Felsenfeld G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell. 1999;983:387-396. [CrossRef]
- Tachibana M, Takeda K, Nobukuni Y, et al. Ectopic expression of MITF, a gene for Waardenburg syndrome type 2, converts fibroblasts to cells with melanocyte characteristics. Nat Genet. 1996;141:50-54. [CrossRef]
- Wang GL, Jiang BH, Rue EA, Semenza GL. Hypoxia-inducible factor1. is a basic-helix-loop-helix- PAS heterodimer regulated by cellular O2 tension. Proc Natl Acad Sci USA. 1995;9212:5510-5514. [CrossRef]
- Davis RL, Weintraub H, Lassar AB. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell. 1987;516:987-1000. [CrossRef]
- Cipolletta D, Feuerer M, Li A, et al. PPAR-γ is a major driver of the accumulation and phenotype of adipose tissue Treg cells. Nature. 2012;4867404:549-553. [CrossRef]
- Alarcón B, Gil D, Delgado P, Schamel WW. Initiation of TCR signaling: regulation within CD3 dimers. Immunol Rev. 2003;191:38-46. [CrossRef]
- Klein U, Rajewsky K, Küppers R. Human immunoglobulin IgM+IgD+ peripheral blood B cells expressing the CD27 cell surface antigen carry somatically mutated variable region genes: CD27 as a general marker for somatically mutated memory B cells. J Exp Med. 1998;1889:1679-1689. [CrossRef]
- Sardiello M, Palmieri M, di Ronza A, et al. A gene network regulating lysosomal biogenesis and function. Science. 2009;3255939:473-477. [CrossRef]
- Tellier J, Shi W, Minnich M, et al. Blimp-1 controls plasma cell function through the regulation of immunoglobulin secretion and the unfolded protein response. Nat Immunol. 2016;173:323-330. [CrossRef]
- Nutt SL, Hodgkin PD, Tarlinton DM, Corcoran LM. The generation of antibody-secreting plasma cells. Nat Rev Immunol. 2015;153:160-171. [CrossRef]
- Lam WY, Becker AM, Kennerly KM, et al. Mitochondrial pyruvate import promotes long-term survival of antibody-secreting plasma cells. Immunity. 2016;45(1):60-73. [CrossRef]
- Vitale M, Della Chiesa M, Carlomagno S, et al. The small subset of CD56brightCD16- natural killer cells is selectively responsible for both cell proliferation and interferon-gamma production upon interaction with dendritic cells. Eur J Immunol. 2004;346:1715-1722.
- Hesslein DG, Palacios EH, Sun JC, et al. Dysregulation of signaling pathways in CD45-deficient NK cells leads to differentially regulated cytotoxicity and cytokine production. Proc Natl Acad Sci USA. 2006;10318:7012-7017.
- De-la-Fuente C, Pantoja-Uceda D, G-Trevijano ER, et al. The structure and plasticity of the complex formed by the two N-terminal domains of human CD58. J. Mol. Biol.; 4329; 3141-3157.
- Bryceson YT, March ME, Ljunggren HG, Long EO. Synergy among receptors on resting NK cells for the activation of natural cytotoxicity and cytokine secretion. Blood. 2006;1071:159-166. [CrossRef]
- Kallies A, Carotta S, Huntington ND, et al. A role for Blimp1 in the transcriptional network controlling natural killer cell maturation. Blood. 2011;1176:1869-1879. [CrossRef]
- Colucci F, Samson SI, DeKoter RP, et al. Differential requirement for the transcription factor PU.1 in the generation of natural killer cells versus B and T cells. Blood. 2001;979:2625-2632. [CrossRef]
- Anson-Cartwright L, Dawson K, Holmyard D, et al. The glial cells missing-1 protein is essential for branching morphogenesis in the chorioallantoic placenta. Nat Genet. 2000;253:311-314. [CrossRef]
- Davis RL, Weintraub H, Lassar AB. Expression of a single transfected cDNA converts fibroblasts to myoblasta. Cell 1987:516; 987-1000. [CrossRef]
- Szanto A, Balint BL, Nagy ZS, et al. STAT6 transcription factor is a facilitator of the nuclear receptor PPARγ-regulated gene expression in macrophages and dendritic cells. Immunity. 2010;335:699-712. [CrossRef]
- Smith MA, Maurin M, Cho HI, et al. PRDM1/Blimp-1 controls effector cytokine production in human NK cells. J Immunol. 2010;18510:6058-6067. [CrossRef]
- Huntington ND, Puthalakath H, Gunn P, et al. Interleukin 15-mediated survival of natural killer cells is determined by interactions among Bim, Noxa and Mcl-1. Nat Immunol. 2007;88:856-863. [CrossRef]
- Vivier E, Tomasello E, Baratin M, et al. Functions of natural killer cells. Nat Immunol. 2008;95:503- 510.
- Chiossone L, Chaix J, Fuseri N, et al. Maturation of mouse NK cells is a 4-stage developmental program. Blood. 2009;11322:5488-5496.
- Wang W, Erbe AK, Hank JA, et al. NK cell-mediated antibody-dependent cellular cytotoxicity in cancer immunotherapy. Front Immunol. 2015;6:368.
- Vachharajani VT, Liu T, Wang X, Hoth JJ, Yoza BK, McCall CE. Sirtuins link inflammation and metabolism. J Immunol Res. 2016;2016:8167273. [CrossRef]
- Hamaidi I, Kim S. Sirtuins are crucial regulators of T cell metabolism and functions. Exp Mol Med. 2022;543:207-215. [CrossRef]
- Li X, Liu L, Li T, et al. Metabolic mechanisms orchestrated by Sirtuin family to modulate inflammatory responses. Front Immunol. 2024;15:1448535.
- Zhao X, Li X, Zhang Y, et al. New insights into the role of Klotho in inflammation and fibrosis: molecular and cellular mechanisms. Front Immunol. 2024;15:1454142.
- Wang Y, Wang K, Bao Y, et al. The serum soluble Klotho alleviates cardiac aging and regulates M2a/M2c macrophage polarization via inhibiting TLR4/Myd88/NF-κB pathway. Tissue Cell. 2022;77:101842. [CrossRef]
- Zhu L, Stein LR, Kim D, et al. Klotho controls the brain--immune system interface in the choroid plexus. Proc Natl Acad Sci U S A. 2018;115(48):E11388-E11396. [CrossRef]
- Bi F, Liu W, Wu Z, et al. Antiaging Factor Klotho Retards the Progress of Intervertebral Disc Degeneration through the Toll-Like Receptor 4-NF-κB Pathway. Int J Cell Biol. 2020;2020:8319516. [CrossRef]
- Lim SW, Jin L, Luo K, et al. Klotho enhances FoxO3-mediated manganese superoxide dismutase expression by negatively regulating PI3K/AKT pathway during tacrolimus-induced oxidative stress. Cell Death Dis. 2017;8(8):e2972. [CrossRef]
- Tai NC, Shih CT, Chen TY, et al. Soluble klotho regulates the function of salivary glands by activating KLF4 pathways. Biomed J. 2019;42(5):319-332.
- Cho H, Zhao X, Hatori M, et al. Regulation of circadian behaviour and metabolism by REV-ERB-α and REV-ERB-β. Nature. 2012;485(7396):123-127. [CrossRef]
- Korbecki J, Bobiński R, Dutka M. Self-regulation of the inflammatory response by peroxisome proliferator-activated receptors. Inflamm Res. 2019;68(6):443-458. [CrossRef]
- Raghavan, PR. Metadichol; An Agonist that Expresses the Anti-aging gene Klotho in Various Cell Lines. Fortune Journal of Health Sciences. 6 2023: 357-362.
- Raghavan, PR. Metadichol and Vitamin C increase in vivo, an open-label study. Preprints.org. 2019. [CrossRef]
- Kishton RJ, Sukumar M, Restifo NP. Metabolic regulation of T cell longevity and function in tumor immunotherapy. Cell Metab. 2017;261:94-109. [CrossRef]
- Alemán, CL, Más Ferreiro, et al. Carcinogenicity of policosanol in Sprague‒Dawley rats: A 24-month study. 1994; Teratog Carcinog Mutagen: 14:239-49.
- Raghavan, PR. Metadichol induces CD14 glycoprotein expression in human embryonic stem cells and fibroblasts. Medical Research Archives, S.l., v. 13, n. 5, may 2025. ISSN 2375-1924. [CrossRef]

| Gene | Regulation | Effect on T Cells Effector | Effect on T Memory Cells | Effect on T Regulatory Cells | |
| CD3G | ↑ | Enhances TCR signaling and T cell activation | Supports memory T cell formation | Minimal effect | |
| CD4 | ↑ | Enhances helper T cell function and activation | Promotes memory CD4+ T cell development | Essential for Treg development and function | |
| CD7 | ↓ | Reduced early T cell activation marker | May affect memory formation | Minimal effect | |
| CD8 | ↑ | Enhances cytotoxic T cell function and killing | Promotes memory CD8+ T cell development | Minimal effect | |
| CD25 | ↑/↓ | Modulates IL-2 responsiveness | Variable effect | Critical for Treg suppressive function | |
| CD27 | ↓ | Reduced co-stimulation | Impaired memory T cell survival | Minimal effect | |
| CD28 | ↑/↓ | Modulates T cell co-stimulation | Affects memory T cell maintenance | Important for Treg homeostasis | |
| CD31 | ↓ | Reduced T cell migration | May affect tissue-resident memory | Minimal effect | |
| CD45 | ↓ | Altered TCR signaling threshold | May affect memory recall | Affects Treg signaling | |
| CD58 | ↑ | Enhanced T cell adhesion and activation | Promotes memory T cell interactions | Minimal effect | |
| CD69 | ↑ | Early activation marker, tissue retention | Promotes tissue-resident memory | Minimal effect | |
| CD80 | ↓ | Reduced costimulatory signaling | May impair memory formation | Affects Treg suppression | |
| CD86 | ↓ | Reduced costimulatory signaling | May impair memory formation | Affects Treg suppression | |
| CD2 | ↑ | Enhanced T cell activation | Supports memory T cell function | Minimal effect | |
| VEGF | ↑ | Promotes T cell survival and migration | May support memory T cell niches | Minimal effect | |
| TAL1 | ↓ | Reduced T cell development | May affect memory precursor formation | Minimal effect | |
| BLIMP1 | ↑ | Promotes effector differentiation | Inhibits memory T cell formation | Promotes Treg suppressive function | |
| FOXP3 | ↑ | Inhibits effector T cell function | Minimal effect | Master regulator of Treg development and function | |
| PU.1 | ↑ | Modulates T cell differentiation | May affect memory formation | Minimal effect | |
| OCT4 | ↑ | Promotes T cell stemness | Enhances memory T cell potential | Minimal effect | |
| PROX1 | ↓ | Affects T cell migration | Regulates memory T cell localization | Minimal effect | |
| GCM1 | ↑ | Modulates T cell function | Minimal effect | Minimal effect | |
| PGC1α | ↑↑ | Enhances metabolic fitness and function | Critical for memory T cell formation and maintenance | Supports Treg metabolic adaptation | |
| Nrf2 | ↓ | Reduced antioxidant response | May affect memory T cell longevity | Minimal effect | |
| FOXJ1 | ↑ | Modulates T cell function | Minimal effect | May affect Treg function | |
| TFEB | ↓ | Reduced autophagy and metabolism | May affect memory T cell survival | Minimal effect | |
| cMyc | ↑ | Promotes T cell proliferation and effector function | Inhibits memory T cell formation | Minimal effect | |
| p53 | ↑↑ | Regulates T cell survival and apoptosis | May affect memory T cell quality | Regulates Treg stability | |
| p63 | ↑ | Modulates T cell development | Minimal effect | Minimal effect | |
| SIM1 | ↑ | Modulates T cell metabolism | Minimal effect | Minimal effect | |
| FOXM1 | ↓ | Reduced T cell proliferation | May affect memory formation | Minimal effect | |
| CTCF | ↓ | Altered gene expression patterns | May affect memory T cell epigenetics | Minimal effect | |
| MITF | ↓/↑ | Modulates T cell function | Minimal effect | Minimal effect | |
| HIF | ↑ | Enhances T cell function in hypoxia | May affect memory T cell metabolism | Supports Treg function in tissues | |
| MYOD1 | ↓ | Minimal direct effect on T cells | Minimal effect | Minimal effect | |
| PPARγ | ↑↑↑ | Promotes metabolic reprogramming | Supports memory T cell metabolism | Critical for Treg function and suppression | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).





