Submitted:
07 October 2025
Posted:
07 October 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample collecting and Fungal Isolate
2.2. DNA extraction and genome sequencing
2.3. Genome assembly
2.4. Gene prediction
2.5. Functional annotation
2.6. Subcellular localization analysis
2.7. Whole-genome phylogenetic analysis
2.8. Comparative analysis
3. Results and Discussion
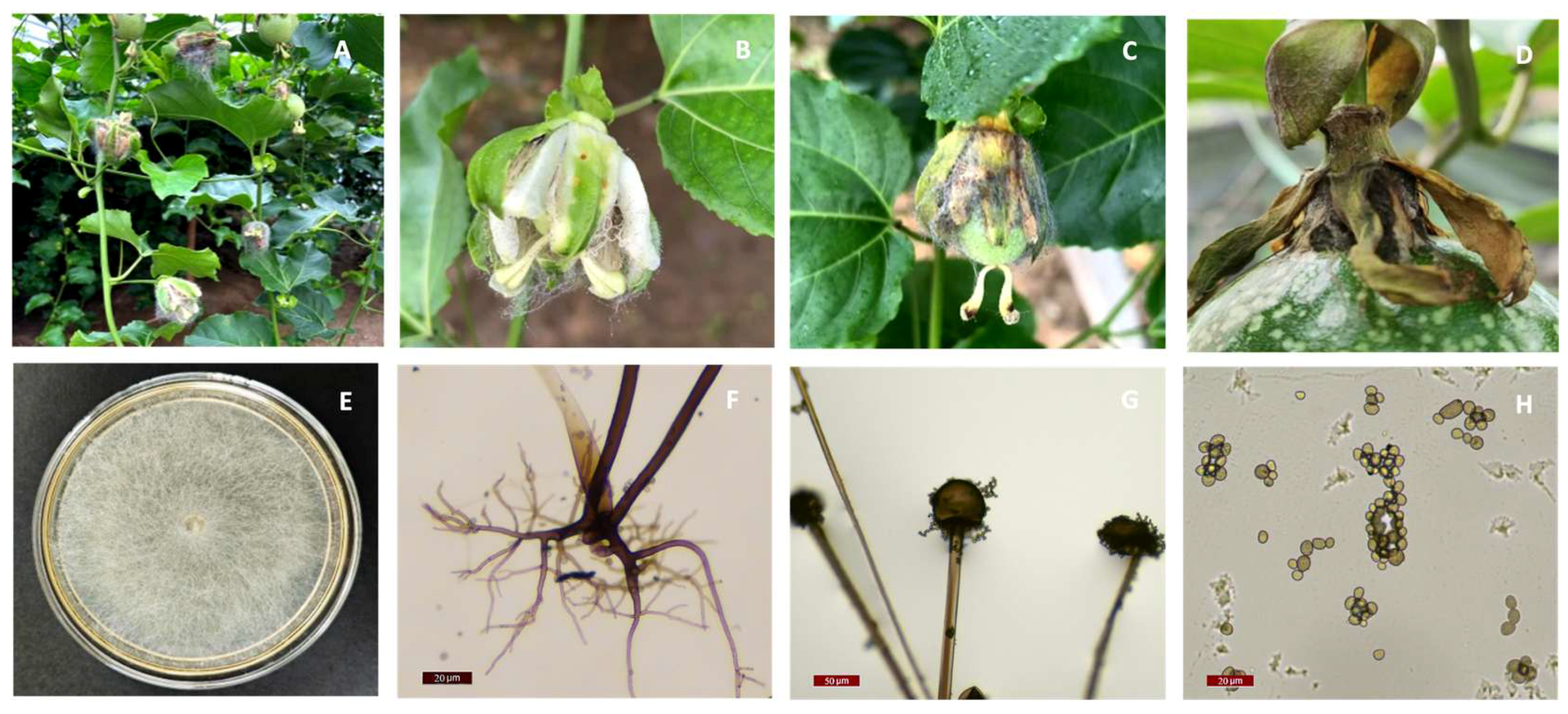
3.1. Pathogen isolation
3.2. Genome assembly
3.3. Functional annotations
3.4. Effector Annotation
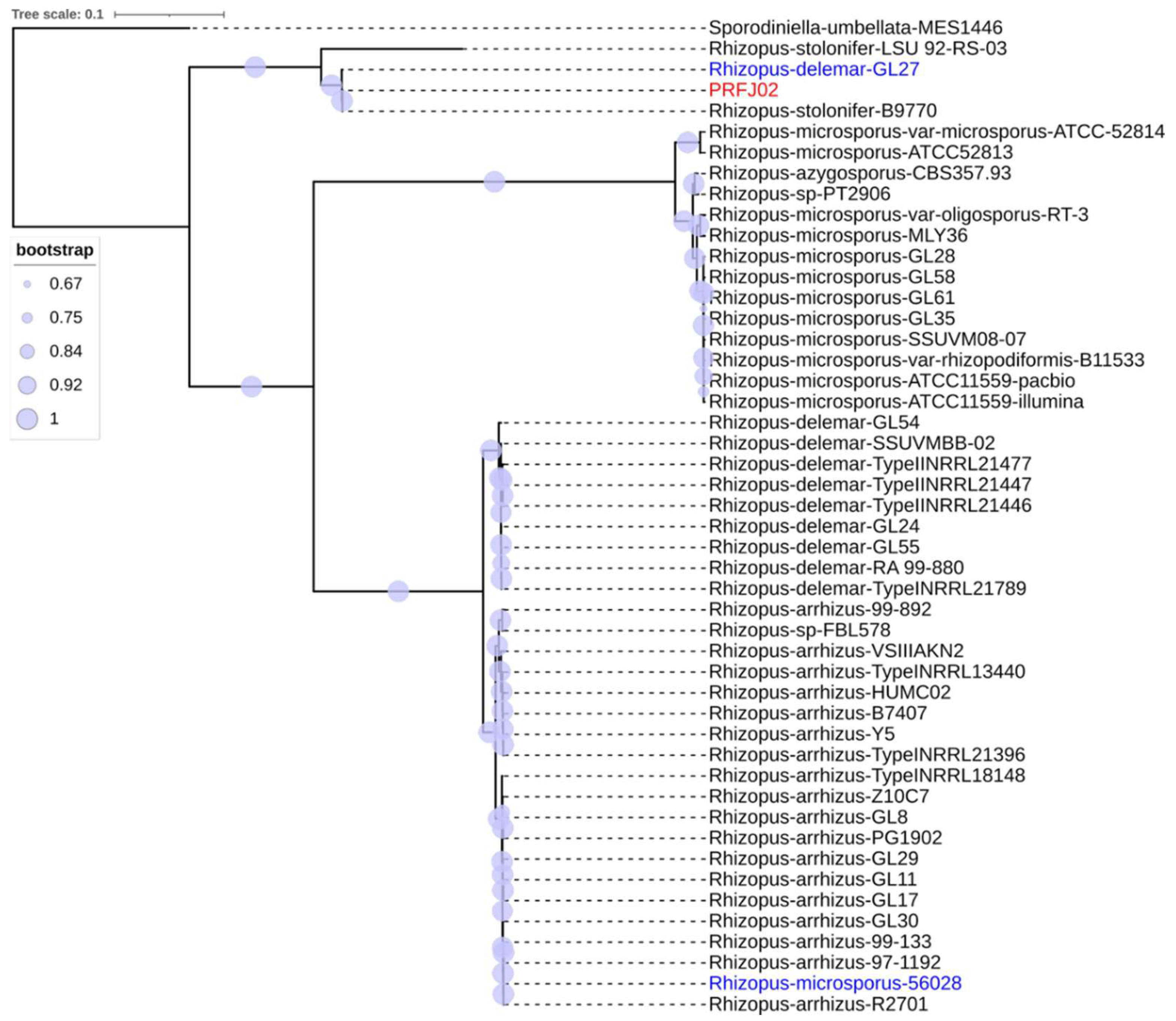
3.5. Whole-genome phylogenetic analysis
3.6. Comparative study
4. Conclusion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Santos-Jiménez, J.L.; Montebianco, C.d.B.; Olivares, F.L.; Canellas, L. P.; Barreto-Bergter, E.; Rosa, R.C.C.; Vaslin, M.F.S. Passion fruit plants treated with biostimulants induce defense-related and phytohormone-associated genes. Plant Gene 2022, 30, 100357. [Google Scholar] [CrossRef]
- Chen, G.; Xu, Y.; Gui, J.; Huang, Y.; Ma, F.; Wu, W.; Han, T.; Qiu, W.; Yang, L.; Song, S. Characterization of Dof transcription factors and the heat-tolerant function of PeDof-11 in passion fruit (Passiflora edulis). Int. J. Mol. Sci. 2023, 24, 12091. [Google Scholar] [CrossRef]
- Sun, J. M.; Chen, G.; Huang, Y.C.; Yang, L.; Zhang, J.Z.; Qin, J.F. First Report of Rhizopus stolonifer causing flower rot of yellow passion fruit (Passiflora edulis f. flavicarpa) in China. Plant disease 2024, 108, 4. [Google Scholar] [CrossRef]
- Wu, Y.; Shi, G.; Zhou, J.; Tian, Q.; Liu, J.; Huang, W.; Xia, X.; Mou, H.; Yang, X. Identification and validation of stem rot disease resistance genes in passion fruit (Passiflora edulis). Hort. Sci. (Prague) 2025, 52, 67–80. [Google Scholar] [CrossRef]
- Chen, Y.H.; Sung, K.Y.; Tuan, S.J.; Huang, J.W.; Lin, Y.H.; Huang, T.P. A streptomyces agent for biocontrol of phytophthora blight and its modulation of rhizosphere microbiomes in passion fruit. Plant disease 2025. [CrossRef] [PubMed]
- Santos-Jiménez, J.L.; Montebianco, C.d.B.; Bernardino, M.C.; Barreto-Bergter, E.; Rosa, R.C.C.; Vaslin, M.F.S. Enhancing passion fruit resilience: the role of hariman in mitigating viral damage and boosting productivity in organic farming systems. Int. J. Mol. Sci. 2025, 26, 2177. [Google Scholar] [CrossRef]
- Choi, M.K. Integrating viral infection and correlation analysis in Passiflora edulis and surrounding weeds to enhance sustainable agriculture in Republic of Korea. Viruses 2025, 17, 383. [Google Scholar] [CrossRef]
- Manicom, B.; et al. 2003. Diseases of passion fruit. Page 413 in: Diseases of Tropical Fruit Crops. R. C. Ploetz, ed. CABI Publishing, Wallingford, U.K.
- Ranallo-Benavidez, T.R.; Jaron, K.S.; Schatz, M.C. GenomeScope 2. 0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat Commun 2020, 11, 1432. [Google Scholar] [CrossRef]
- Servant, N.; Varoquaux, N.; Lajoie, B.R. Viara, E.; Chen, C.; Vert, J.; Heard, E.;Dekker, J.; Barillot, E. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol 2015, 16, 259. [Google Scholar] [CrossRef]
- Zhang, H.; Song, L.; Wang, X.; Tang, M.; Aluru, S.; Cheng, H.; Wang, C.; Meyer, C.A.; Yue, F.; Liu, X.S.; Li, H. Fast alignment and preprocessing of chromatin profiles with Chromap. Nat Commun 2021, 12, 6566. [Google Scholar] [CrossRef]
- Mikheenko, A.; Prjibelski, A.; Saveliev, V.; Antipov, D. Gurevich A. Versatile genome assembly evaluation with QUAST-LG. Bioinformatics 2018, 34, 142–150. [Google Scholar] [CrossRef]
- Manni, M.; Berkeley, M.R.; Seppey, M.; Zdobnov, E.M. BUSCO: Assessing Genomic Data Quality and Beyond. Current Protocols 2021, 1, e323. [Google Scholar] [CrossRef]
- Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 1999, 27, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Flynn, J. M.; Hubley, R.; Goubert, C.; Rosen, J.; Clark, A. G.; Feschotte, C.; Smit, A. F. RepeatModeler2 for automated genomic discovery of transposable element families. PANS 2020, 117(17), 9451–9457. [Google Scholar] [CrossRef]
- Chen, N. Using RepeatMasker to Identify Repetitive Elements in Genomic Sequences. Curr Protoc. [CrossRef]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 1997, 25, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Lagesen, K.; Hallin, P.; Rødland, E.A.; Stærfeldt, H.H.; Rognes, T.; Ussery, D.W. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 2007, 35, 3100–3108. [Google Scholar] [CrossRef] [PubMed]
- Stanke, M.; Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res 2005, 33, W465–W467. [Google Scholar] [CrossRef]
- Majoros, W.H.; Pertea, M.; Salzberg, S.L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 2004, 20, 2878–2879. [Google Scholar] [CrossRef]
- Ter-Hovhannisyan, V.; Lomsadze, A.; Chernoff, Y.O.; Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res 2008, 18, 1979–1990. [Google Scholar] [CrossRef]
- Haas, B.J.; Salzberg, S.L.; Zhu, W.; Pertea, M.; Allen, J.E.; Orvis, J.; White, O.; Buell, C.R.; Wortman, J.R. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Bio. 2008, 9, 1–22. [Google Scholar] [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Huerta-Cepas, J.; Forslund, K.; Coelho, L.P.; Szklarczyk, D.; Jensen, L.J.; Mering, C.; Bork, P. Fast Genome-Wide Functional Annotation through Orthology Assignment by eggNOG-Mapper. Mol. Biol. Evol 2017, 34, 8–2115. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Attwood, T.K.; Babbitt, P.C.; Bateman, A.; Bork, P.; Bridge, A.J.; Chang, H.Y.; Dosztányi, Z.; El-Gebali, S.; Fraser, M.; Gough, J.; Haft, D.; Holliday, G.L.; Huang, H.; Huang, X.; Letunic, I.; Lopez, R.; Lu, S.; Marchler-Bauer, A.; Mi, H.; Mistry, J.; Natale, D.A.; Necci, M.; Nuka, G.; Orengo, C.A.; Park, Y.; Pesseat, S.; Piovesan, D.; Potter, S.C.; Rawlings, N.D.; Redaschi, N.; Richardson, L.; Rivoire, C.; Sangrador-Vegas, A.; Sigrist, C.; Sillitoe, I.; Smithers, B.; Squizzato, S.; Sutton, G.; Thanki, N.; Thomas, P.D.; Tosatto, S.C.; Wu, C.H.; Xenarios, I.; Yeh, L.S.; Young, S.Y.; Mitchell, A.L. InterPro in 2017-beyond protein family and domain annotations. Nucleic Acids Res. 2017, 4: 190-199.
- Almagro Armenteros, J.J.; Tsirigos, K.D.; Sønderby, C.K.; Petersen, T.N.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. [CrossRef]
- Almagro Armenteros, J.J.; Salvatore, M.; Emanuelsson, O.; Winther, O.; von Heijne, G.; Elofsson, A.; Nielsen, H. Detecting sequence signals in targeting peptides using deep learning. Life Sci. Alliance 2019, 2, e201900429. [Google Scholar] [CrossRef] [PubMed]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 2001, 305, 567–580. [Google Scholar] [CrossRef]
- Sperschneider, J.; Dodds, P.N.; Gardiner, D.M.; Singh, K.B.; Taylor, J.M. Improved prediction of fungal effector proteins from secretomes with EffectorP 3.0. Mol. Plant Pathol, 23.
- Stanke, M.; Diekhans, M.; Baertsch, R.; Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 2008, 24, 637–644. [Google Scholar] [CrossRef]
- Stanke, M.; Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 2003, 19, 215–225. [Google Scholar] [CrossRef]
- Keller, O.; Kollmar, M.; Stanke, M.; Waack, S. A novel hybrid gene prediction method employing protein multiple sequence alignments, Bioinformatics 2011, 27, 757–763. 27.
- Shen, W.; Sipos, B.; Zhao, L. SeqKit2: a Swiss army knife for sequence and alignment processing. iMeta 2024, 3, e191. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef]
- Capella-Gutierrez, S.; Silla-Martinez, J.M.; Gabaldon, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Gardiner, D.M.; Aitken, E.A.B.; van Dam, P.; Le, D.P.; Smith, L.J.; Chen, A. De novo long-read assembly and annotation for genomes of two cotton-associated Fusarium oxysporum isolates. Australasian Plant Pathol. 2025, 54, 377–386. [Google Scholar] [CrossRef]
- Edler, D.; Klein, J.; Antonelli, A.; Silvestro, D. raxmlGUI 2.0: A graphical interface and toolkit for phylogenetic analyses using RAxML. Methods Ecol. Evol. 2021, 12, 373–377. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v6: Recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 2024, 52, W78–W82. [Google Scholar] [CrossRef] [PubMed]
- Cantarel, B.L.; Coutinho, P.M.; Rancurel, C.; Bernard, T.; Lombard, V.; Henrissat, B. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res. 2009, 37, D233–D238. [Google Scholar] [CrossRef]
- Le Naour-Vernet, M.L.; Lahfa, M.; Maidment, J.H.R.; Padilla, A.; Roumestand, C.; de Guillen, K.; Kroj, T.; Césari, S. Structure-guided insights into the biology of fungal effectors. New Phytol. 2025, 246, 1460–1477. [Google Scholar] [CrossRef]
- Snelders, N.C.; Rovenich, H.; Thomma, B.P. Microbiota manipulation through the secretion of effector proteins is fundamental to the wealth of lifestyles in the fungal kingdom. Fems Microbiol Rev. 2022, 46, fuac022. [Google Scholar] [CrossRef]
- Wu, J.; Wang, P.; Wang, W.; Hu, H.; Wei, Q.; Bao, C.; Yan, Y. Comprehensive Genomic and Proteomic Analysis Identifies Effectors of Fusarium oxysporum f. sp. melongenae. J. Fungi 2024, 10, 828. [Google Scholar] [CrossRef]
- Sun, J.; Zhang, J.; Gardiner, D.M.; van Dam, P.; Fu, G.; Ferguson, B.J.; Aitken, E.A.B.; Chen, A. Long-read draft genome sequences of two Fusarium oxysporum f. sp. cubense isolates from banana (Musa spp.). J. Fungi. 2025, 11, 421. [Google Scholar] [CrossRef]

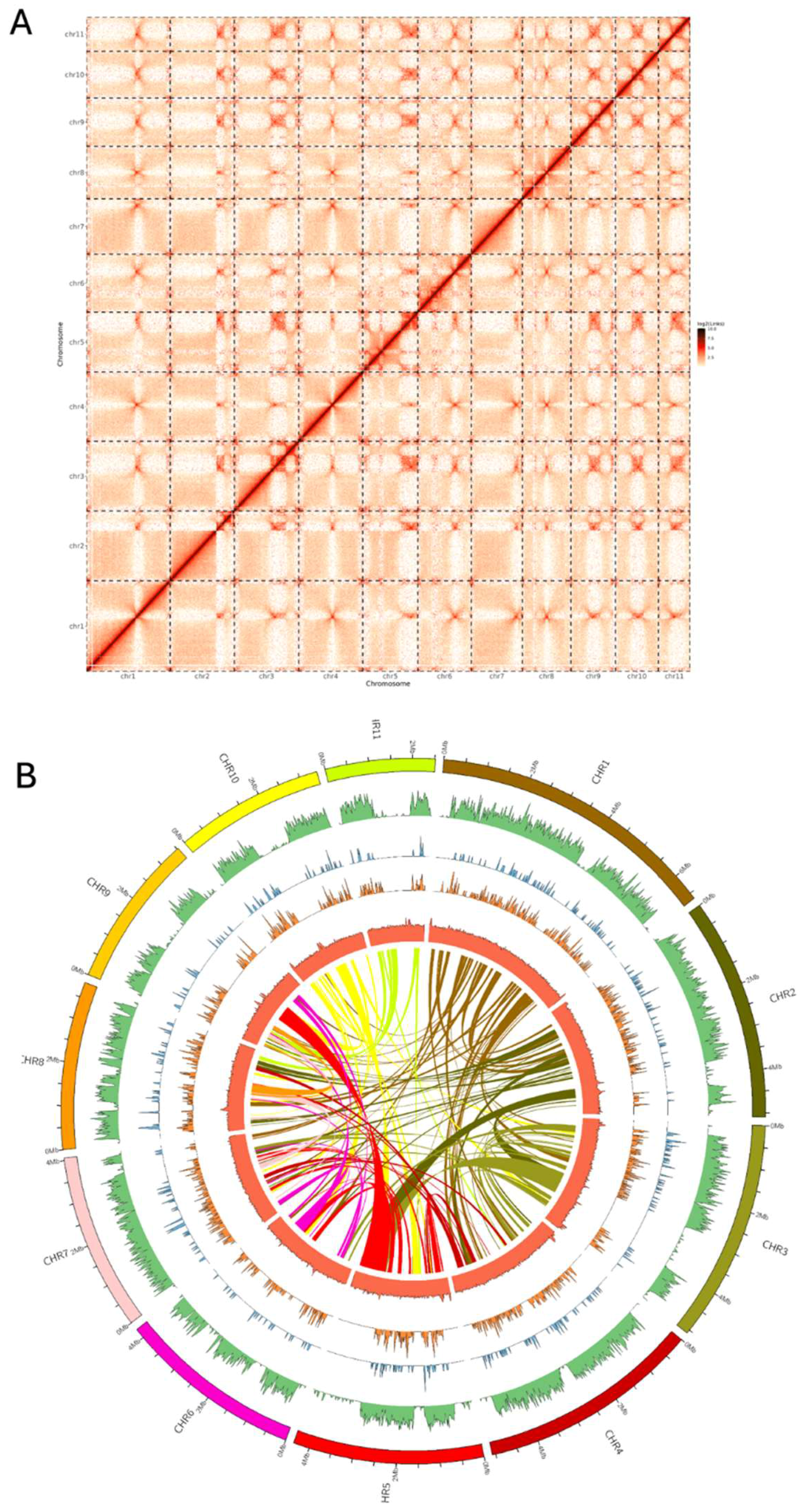
| Assembly statistics | Primary assembly | Hi-C scaffolded assembly |
|---|---|---|
| Total length (Gbp) | 6.6 a, 6.8 b, 4.1 c | 6.6 a, 6.8 b, 4.1 c |
| Coverage (fold) | 138 d | 138 d |
| Assembly size (bp) | 48,208,123 | 48,208,381 |
| No. of contigs (or scaffolds) | 21 | 18 |
| Maximum contig length (bp) | 6,159,681 | 6,651,178 |
| N50 contig (or scaffold) length (bp) | 4,233,866 | 4,366,636 |
| Contig L50 | 5 | 5 |
| Contig L90 | 10 | 10 |
| GC content (%) | 35.9 | 35.9 |
| BUSCO coverage (%) | 95.2 | 95.2 |
| Total no. of BUSCOs | 2,449 | 2,449 |
| No. of duplicate BUSCOs | 104 | 100 |
| No. of fragmented BUSCOs | 3 | 5 |
| No. of missing BUSCOs | 116 | 112 |
| Gene model | ||
| Total no. of genes | NA | 11,885 |
| Total no. of proteins | NA | 11,737 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




