2.1. Primer Design, Initial Set up and Validation
The F5 exon 10 segment was amplified with the primers described in
Table 1. The PCR fragment also carries a mandatory additional MnlI site used as an internal digestion control.
Table 1 footer: The mutagenic base in the lower F2 primer is presented with underlined bold characters. As primers were chosen to generate fragments with markedly different sizes, many fluorochrome combinations are possible. The one reported in this article uses a combination of PET, VIC FAM, and NED, but ATTO 565, Yakima Yellow, FAM and ATTO 550 (respectively red, green, blue and black with the Applied G5 filter) would be an equivalent combination.
The HindIII site specific to the F2 G20210A variant was created by a mutagenic primer as previously described [
6,
7]. The upper primer was repositioned to avoid amplification of an MnlI site. Because the putative HindIII restriction site is unique in the F2 amplified fragment, undigested fragments might result from accidentally undigested fragments rather than from the absence of restriction sites.
For HindIII control fragment amplification, the primers described in [
7] could not be used because of the presence of an MnlI site in the PCR product. An FGA intron 1 fragment carrying a mandatory HindIII site and no MnlI site was added to the multiplex PCR as a HindIII digestion control.
The FGA and SE33 STRs (GenBank accession M64982 and V00481) are complex tetranucleotide repeats initially described in DNA fingerprinting applications. Primers were as described in
https://strbase-archive.nist.gov/str_SE33.htm and
https://strbase-archive.nist.gov/str_fga.htm, last accessed on October 18, 2024. With the chosen primers, MnlI cuts the STR of FGA. To overcome this difficulty, the fluorescent labeling was located on the upper primer, still allowing fluorescent STR analysis after MnlI digestion.
Amplifications (MasterMix PCR AmpliTaq Gold360 from Thermofisher Scientific, Waltham, MA) were performed in a final volume of 10µL. Each primer pair was tested in individual reactions on control DNAs before being mixed in multiplex reactions. After testing the initial mix, we concluded that it was necessary to adjust individual primer pair concentrations to limit unbalanced amplification (see
Table 1). Co-digestions were performed with MnlI and HindIII (1 and 3UI/µL final, respectively) from 2 hours to overnight at 37°C in the universal buffer provided by the manufacturer.
2.3. Sensitivity
DNA concentration was determined photometrically. DNA from two individuals were diluted stepwise in sterile water, to concentrations corresponding to 2, 4, 8, 16, 32, 64, 128, and 256ng in the 10µL PCR. Under these conditions, with as low as 2ng of input DNA, all peaks were in the 250-32,000 relative fluorescent units (RFU) range, 32,000 RFU being the saturation limit and 250 RFU the arbitrary detection threshold. When DNAs were in the 15-70ng range per 10µL reaction, which is what is usually achieved with 2µL of DNA extracted using fast extraction protocols, such as the QuickGene Blood DNA extraction kit from Qiagen, Hilden, Germany (expected value in the 25ng/µL range), or the Maxwell RSC whole Blood DNA kit (expected values in the 30-70ng/µL range), satisfactory signal height and peak balance were obtained for 28 cycle amplifications (see
Table 1 for amplification conditions). Thus, after initial set up experiments, DNA concentration was no longer measured and DNA input was set to 2µL, regardless of the actual concentration. A representative electrophoresis is presented in
Figure 1.
2.5. Contamination Detection and Stutter Peaks
FGA and SE33 STRs were chosen because of their high heterozygosity percentage and high allelic heterogeneity rate. For FGA, there are over 28 known alleles ranging from 12.2 to 51.2 repeats and over 76 known alleles ranging from 7 to 39.2 repeats for SE33. The random match probability for the combination of the most frequent genotypes is in the 0.0005 range with these two STRs. Inclusion of such STRs in the multiplex mix enabled quality control and identity verifications. Moreover, duplicate distribution of a given sample instead of two different samples would be detected. STR co-amplification allowed contamination detection by highlighting any additional STR peak (
Figure 1C and ID), regardless of the mechanism involved, whereas traditional methods would only detect reagent contaminations through signal presence in the blank control reaction.
This method amplifies STR-generated artifact stutter peaks in strict repetitive patterns to a size equivalent to alleles one repeat shorter than the respective allelic types. They result from polymerase slippage during template amplification [
8]. In a series of 30 individuals randomly selected in six different assays, stutter heights were 7.6±1.8% and 7.6±1.6% of the allele height for FGA and SE33, respectively (mean ± standard deviation [SD]). Thus, in ongoing assays, any stutter peak above the mean +2SD (i.e., 11%) values will be considered as resulting from contamination.
To assess the potential for contamination detection, two DNA samples were mixed (1:4; 1:10, and 1:20) and analyzed. Amplifications were performed with 25ng of DNA mix. Mixed DNAs at 1:20 ratio allowed contamination identification either by additional STR alleles (
Figure 1C and 1D) or by a stutter size above the expected value. Further dilutions generate peaks below the interpretation limit (250 RFU).
2.2.
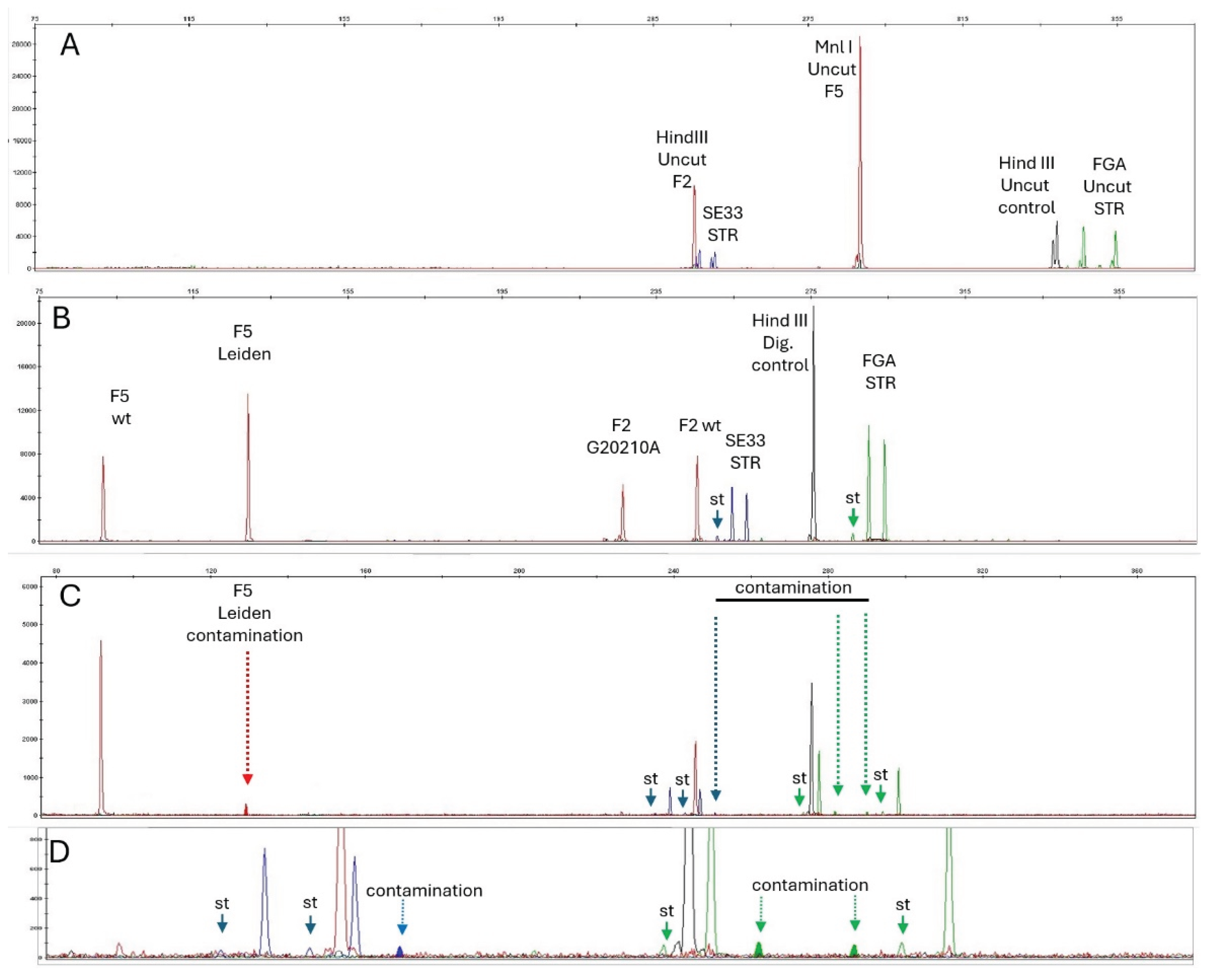
Figure 1 Undigested sample, double heterozygous sample, and contaminated sample.
Figure 1.
Undigested sample, double heterozygous sample, and contaminated sample.
Figure 1.
Undigested sample, double heterozygous sample, and contaminated sample.
One mL of polymerase chain reaction (PCR) product was added to 9μL Hi-Di formamide (Applied Biosystems) and 0.5μL size standard (GS™-LIZ® 600, Applied Biosystems), and then loaded onto a capillary electrophoresis apparatus (Applied Biosystems 3500 xL Dx).
Panel A. Undigested PCR sample: the undigested HindIII NED-labeled control fragment is present (Hind III Uncut control, in black), as well as the MnlI undigested FV fragment (MnlI Uncut F5, in red).
Panel B. Electropherogram of a double heterozygous sample (FV Leiden +/- and F2 G20210A +/-). A: PET-labeled F2 and F5 fragments (red) digested with Mnl I and Hind III, respectively, and NED-labeled FGA intron I (green) for Hind III digestion control (expected size 275 bp for digested PCR fragment). Uncut FGA intron 1 fragment would be 339 pb and uncut FV fragment would be 288 pb, they are both absent. The different size for VIC-labeled PCR products (FGA STR alleles, green) and 6-FAM-labeled PCR products (SE33 STR alleles, blue) in panels A to C reflects the different sample genotypes. Stutters (st) are indicated by plain arrows.
Panel C: Experimental dilution 1:20 of a heterozygous FV Leiden individual's DNA into a wild-type DNA. The DNA sample used for experimental contamination was deliberately chosen to carry the FV-Leiden variant. The FV Leiden+/- and F2 G20210A -/- was mixed (1:20 ratio) in a FV-Leiden-/- and F2 G20210A -/- DNA sample. Stutters are indicated by plain arrows. Dotted arrows show peaks resulting from contamination.
Panel D: Zoom in on contamination and stutter peaks. Definitive contamination proof is provided by panel D analysis: contamination is detected by the presence of extra FGA and SE33 peaks (plain line arrows). Extra STR alleles are present (long arrow) and correspond to the FGA and SE33 alleles from the contaminating DNA. Of note, any contaminating allele the size of the stutter (in base pair) would be superimposed on the stutter peak. It would be detected because of the size above the expected stutter size (11% of main peak).