Submitted:
24 September 2025
Posted:
29 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Artificial Intelligence and Health Systems Technologies
2. Artificial Intelligence in Biochemistry Laboratories
2.1. What is Biochemistry?
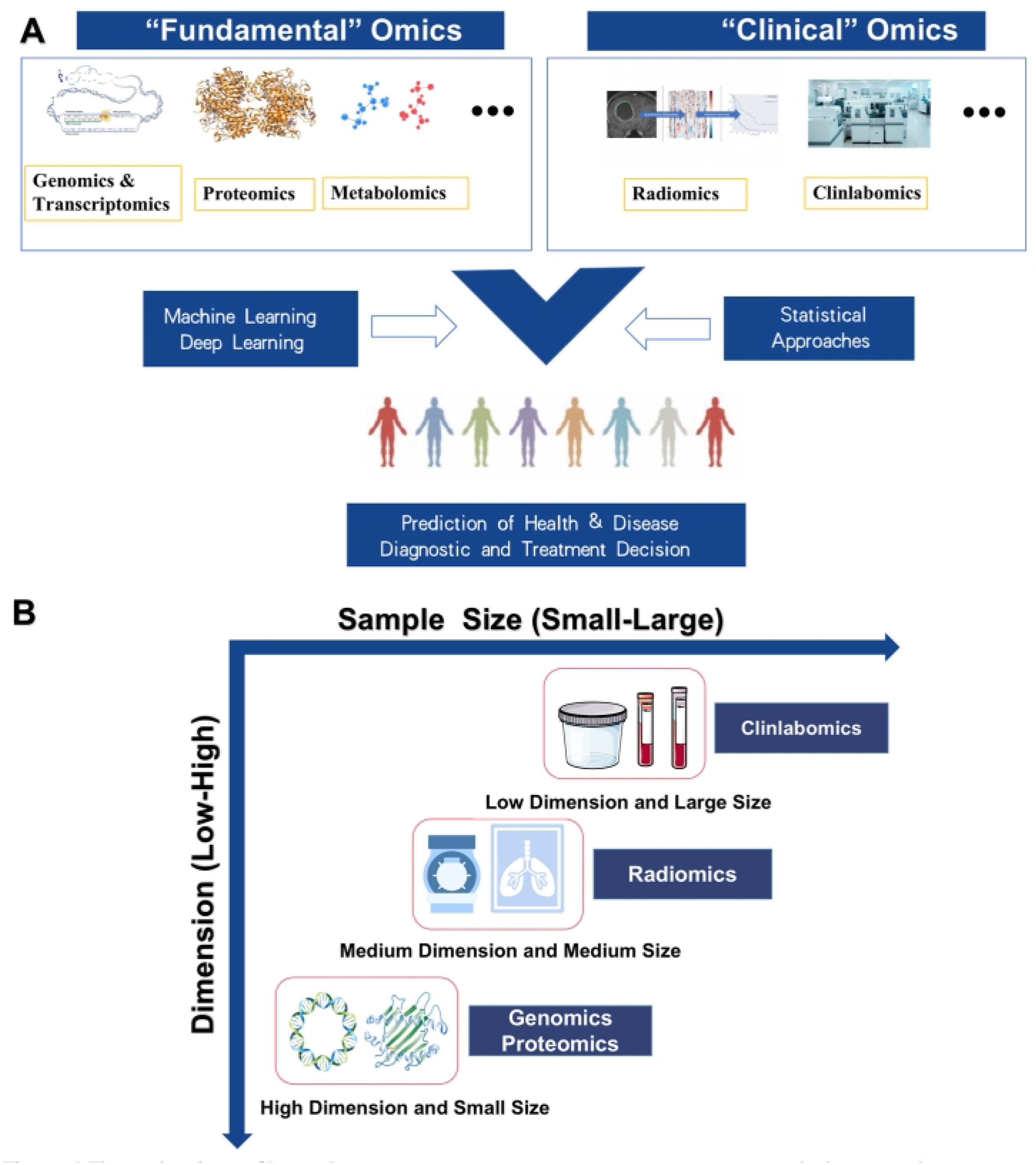
2.2. Biochemistry and Clinlabomics
2.3. AI in Clinical Applications: Case Examples
- Smoking and Aging: Models have been developed that predict smoking status and the rate of aging from blood tests.
- COVID-19: The presence of COVID-19 disease and the risk of death were predicted with high success using blood tests alone.
- Cancer Diagnosis: Powerful prediction models for brain, lung and colorectal cancers were created using simple blood values.
- Laboratory Management: It has been shown that many processes such as detection of mislabeled samples, separation of clotted sera, and automatic determination of reference ranges can be performed with AI.
2.4. Challenges and Future Perspectives of AI in Clinical Biochemistry
- Data Privacy Concerns: Protecting sensitive patient biochemical data from breaches is vital. Because AI systems work with large datasets, ensuring the privacy and security of this data is essential to earn patient trust and comply with regulatory requirements.
- Regulatory Compliance: Ensuring that AI-based diagnostics meet regulatory standards like the FDA and EU is a complex process. This requires rigorous protocols for validating, clinically proving, and continuously monitoring AI models.
- AI Interpretability: Understanding how AI models derive their conclusions from biochemical data is a significant challenge, often referred to as the "black box" problem. It is critical for clinicians and researchers to understand why AI reaches a particular conclusion so they can confidently use it in clinical decision-making.
- Integration with Healthcare Systems: Adapting AI solutions to existing, often complex, hospital and laboratory workflows presents a significant hurdle. This includes not only technological compatibility but also staff training, management of organizational change, and interoperability with legacy systems [7].
2.5. Biochemistry and AI Conclusion
3. Artificial Intelligence in Microbiology Laboratories
3.1. What Does Artificial Intelligence Mean in Microbiology?
3.2. Examples of AI Applied in Medical and Clinical Microbiology
3.2.1. Bacteria/Virus Diagnosis and Microbiome/Pathogen Analysis
- a)
-
Bacterial diagnosis system with coherent imaging and DLWaterborne diseases, which affect more than 2 billion people worldwide each year, are a significant economic burden and public health problem. Rapid and accurate detection of pathogens such as E. coli and coliform bacteria in drinking water is crucial. Traditional bacterial detection methods typically rely on counting colonies in petri dishes. However, this process can take 24 to 48 hours for colonies to reach visible size, and interpreting the results requires expertise. In addition to being slow, these methods can struggle to process large sample volumes.To overcome these challenges, Hongda Wang and his team developed a novel system that combines time-lapse coherent imaging and deep learning technologies. This computational system uses a lensless holographic microscopy platform to periodically image bacterial growth within a 60 mm diameter agar plate. The system’s core is two distinct deep neural networks (DNNs) that analyze the spatiotemporal properties of bacteria during their growth phase.Step 1: Detection Network: The first network is used to detect bacterial growth at the earliest possible stage. In this stage, images of the agar plate taken over time are compared and changing objects (bacterial colonies) are identified. Step 2: Classification Network: The second network classifies the species of detected colonies (e.g., E. coli, Klebsiella aerogenes, and Klebsiella pneumoniae). By capturing the unique growth characteristics of bacteria at the microscale, this network can successfully distinguish even species that cannot be distinguished by traditional methods. This developed platform offers significant advantages over traditional methods. Test results showed that the system could detect bacteria in less than 12 hours compared to EPA-approved methods. It achieved a sensitivity of approximately 1 colony-forming unit per liter (CFU/L) in just 9 hours of total testing time. The system also maintains a high precision rate of 99.2%-100% after just 7 hours of incubation, detecting over 95% of all tested bacteria within 12 hours. Its classification capability is also impressive, reaching 97.2% accuracy for E. coli, 98.5% for K. pneumoniae, and 84% for K. aerogenes after 12 hours of incubation. The platform is extremely cost-effective, costing just 0.6 per dollar test, including all testing steps. It also boasts a high-capacity system capable of scanning the entire plate surface at a speed of 24 cm² per minute. A conventional microscope with a lens would take approximately 128 minutes to scan a plate of the same diameter. In conclusion, this automated and cost-effective deep learning-powered live bacteria detection platform has the potential to be a game-changer in microbiology. It not only significantly reduces testing time but also automates the colony identification process, eliminating the need for expert guidance.
- b)
-
Clinical Viral Metagenomic Studies and Virome Analysis In recent years, the human microbiome—the community of trillions of microbes living within our bodies—has become one of the most exciting topics in science. However, this microscopic ecosystem also has an overlooked, even more complex component: the virome. Defined as the collection of all viruses found in an organism or the environment, the virome has a profound impact on many areas, from human health to ocean ecosystems. So, how do we analyze this invisible world? Virome analysis is a combination of molecular biology and computational techniques that aims to understand the identity, diversity, and functions of viruses by examining their genetic material (DNA or RNA). Studies in this field typically begin with a method called metagenomics, which is the bulk analysis of all genetic material in an environmental sample (e.g., soil, water, or human intestines). The basic steps are: 1. Isolation of Virus Particles: Before sampling, virus-like particles are separated from cellular components such as bacteria. This is typically accomplished through techniques such as filtration, density gradient centrifugation, and enzyme treatments. 2. Obtaining and Sequencing Genetic Material: The nucleic acids (DNA or RNA) of the isolated viruses are extracted. The genetic codes of this material are then read using next-generation sequencing technologies.3. Bioinformatics Analysis: The resulting sequencing data is analyzed using bioinformatics tools and machine learning algorithms. In this phase, unknown viral genes are identified, viral species are classified, and genetic diversity maps are created.Virome analysis not only detects the presence of viruses but also reveals their complex interactions with their hosts. Research on the human virome helps us understand the role of viruses in the immune system, metabolism, and even some chronic diseases (such as diabetes and inflammatory bowel disease). For example, viruses called bacteriophages, which infect only bacteria, play a critical role in maintaining the balance of the gut microbiome.In the context of ecosystems, virome analysis is used to understand viral diversity in seawater, soil, or other environmental niches. Projects such as the Global Ocean Viromes (GOV), published in 2017, have revealed the incredible diversity of virus populations in the world’s oceans and the impact these viruses have on our planet’s biochemical cycles.In conclusion, virome analysis is a rapidly developing field that is ushering in a new era in microbiology and ecology. Rather than viewing viruses solely as pathogens, it allows us to understand them as integral components of ecosystems and organisms. Thanks to developing technology, the discovery of this mysterious world of viruses will pave the way for groundbreaking innovations in fields such as personalized medicine and environmental management in the future.[11]
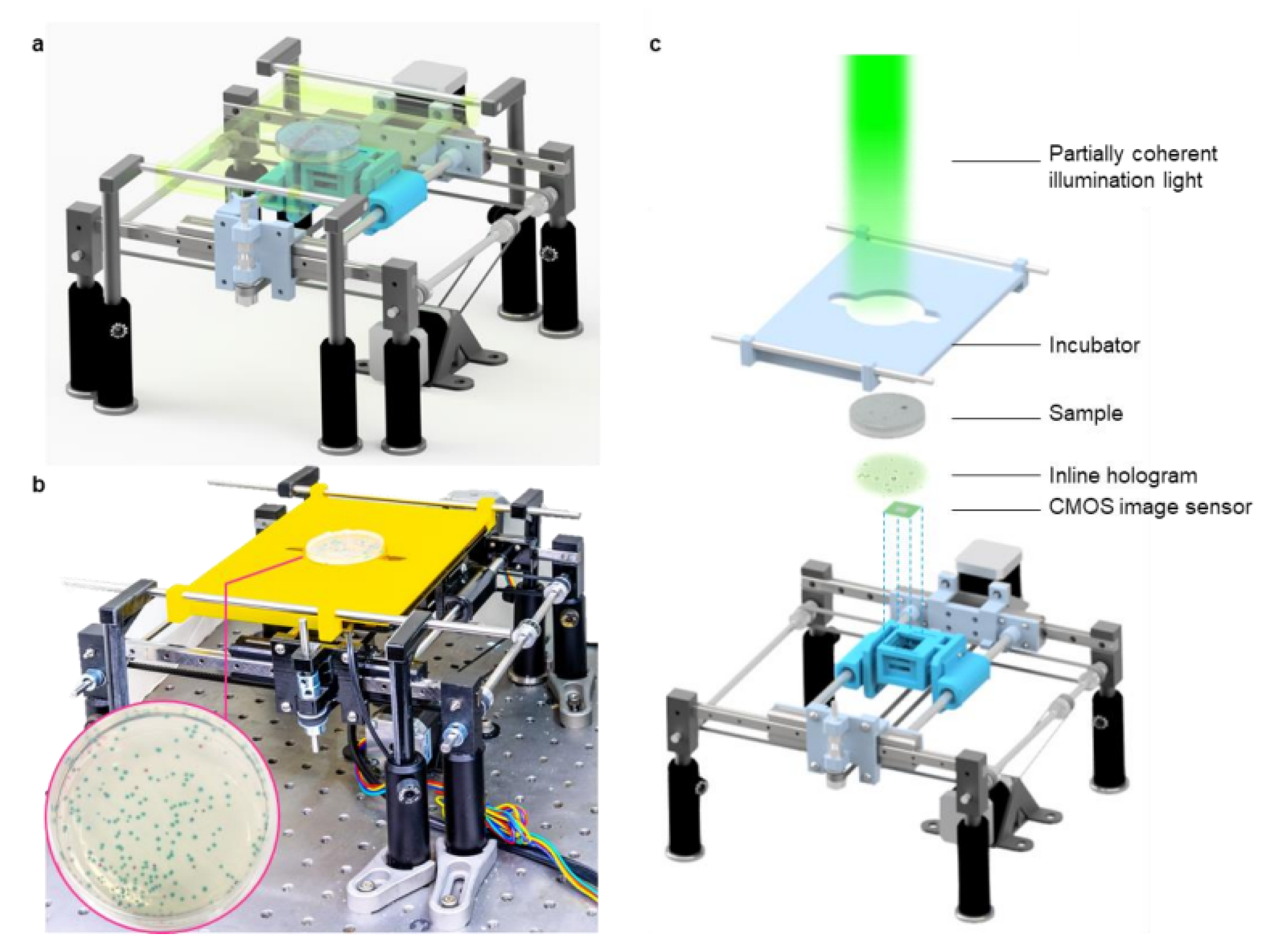
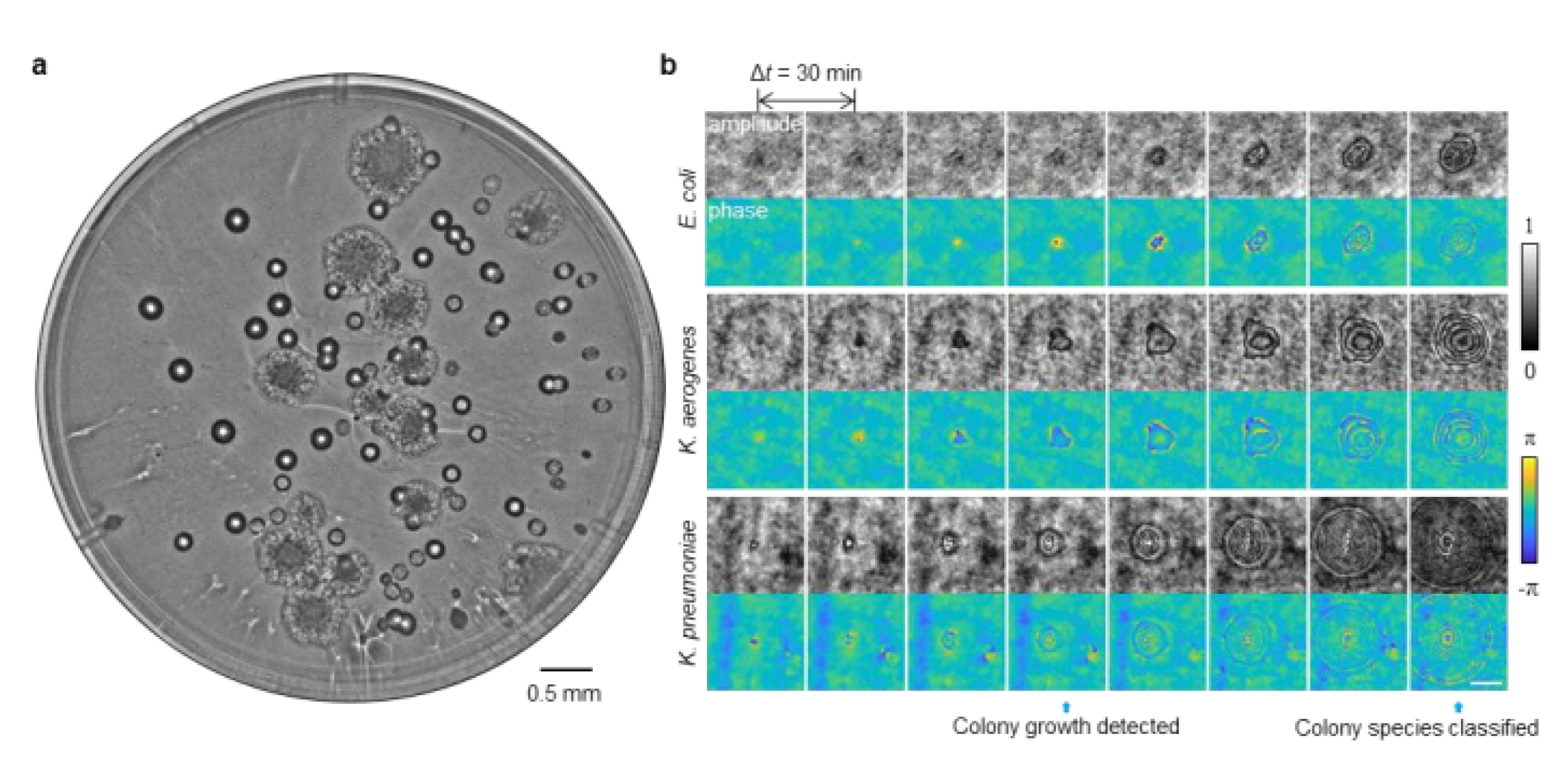
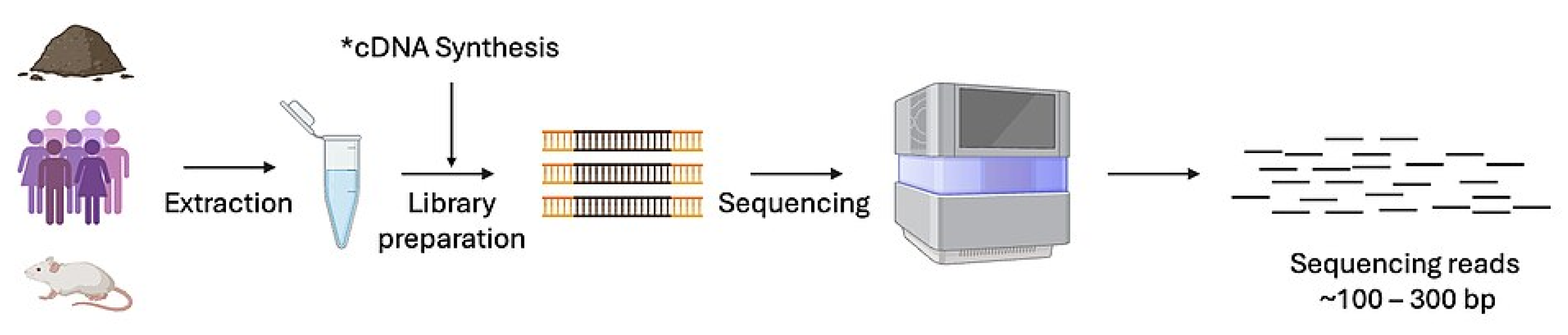
3.2.2. Clinical Diagnosis, AST - Clinical Decision
- a)
-
GPT-4 based EUCAST-GPT-expertToday, antimicrobial resistance (AMR) has become one of the biggest threats to public health. Rapid and accurate detection of beta-lactamase-producing Gram-negative bacteria, in particular, is vital for patients to access the right treatment. Because traditional diagnostic methods are time-consuming and require expertise, artificial intelligence (AI) technologies stand out as a promising new solution in this area.A study on this topic evaluated the ability of "EUCAST-GPT-expert," a ChatGPT-4-based AI agent, to detect potential beta-lactamase mechanisms in clinical diagnostic processes. The study highlights the strengths and current limitations of this new technology.The researchers examined 225 laboratory-collected Gram-negative bacterial isolates. Their antibiotic resistance profiles were determined using standard methods, such as the Kirby-Bauer disk diffusion test. The test results (images and measured inhibition zone diameters) were then presented in three different groups:1-EUCAST-GPT-expert: A GPT-agent specially trained in the European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines and rules2-Microbiologists: Three expert microbiologists with at least five years of experience.3-Uncustomized ChatGPT-4: A standard version of GPT-4 that has not undergone any special training.All groups were asked to determine one of four resistance categories (none, ESBL, AmpC, or carbapenemase) for each isolate and to explain their decision with a brief rationale. The study compared the performance of the groups in detecting resistance mechanisms.
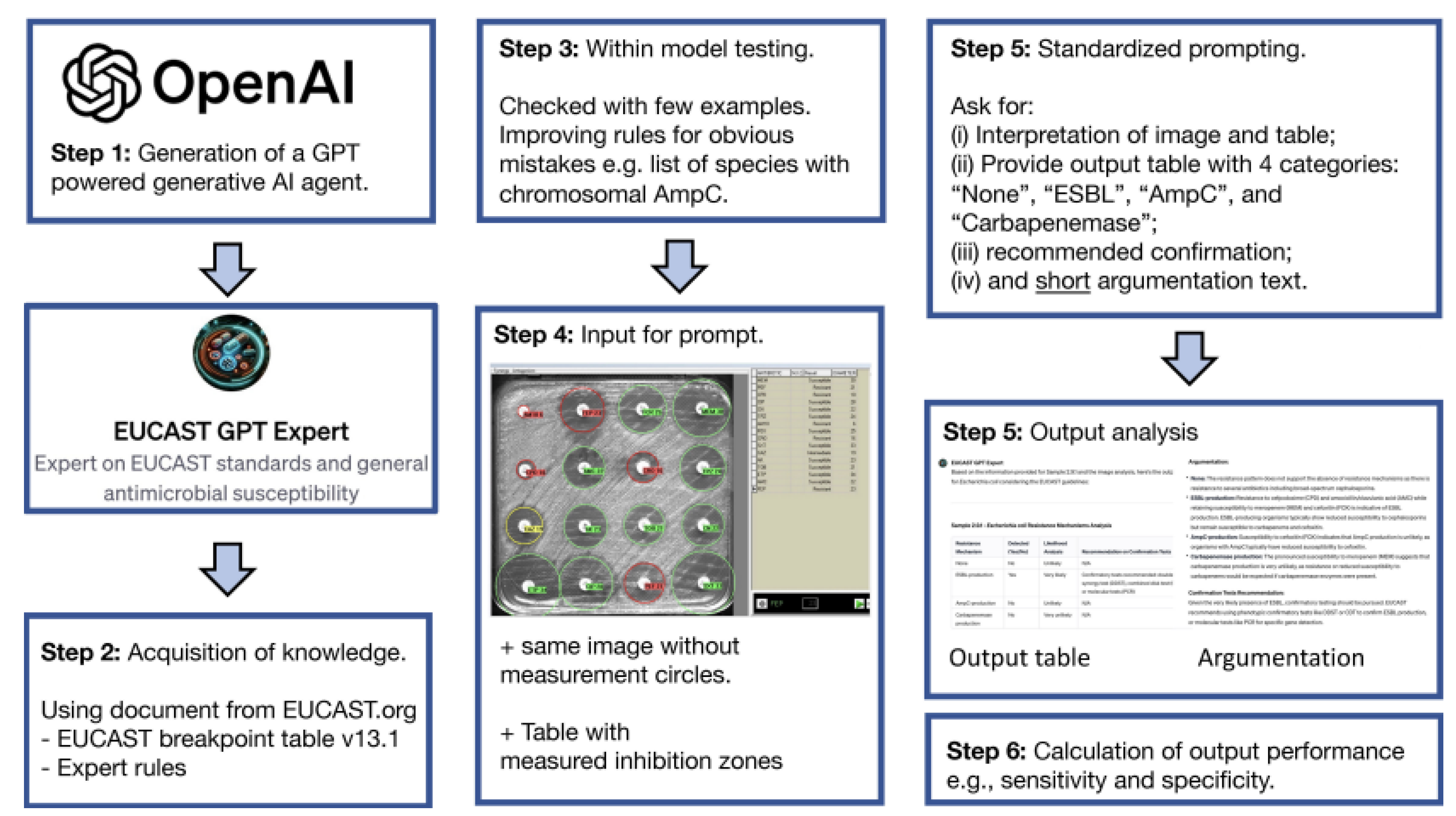
- Sensitivity and Specificity: EUCAST GPT expert performed comparable to human experts in terms of sensitivity for correctly identifying resistant strains. However, the specificity rates for correctly identifying non-resistant strains as absent were lower, particularly for ESBL and AmpC. This suggests that the AI agent tends to over-flag potential resistance.
- Justification: While human experts used concise, average eight words, to explain their decisions, the EUCAST-GPT-expert provided much more detailed justifications, averaging 158 words. While these detailed explanations are useful for educational purposes, they may not be practical in a routine laboratory setting.
- Decision Consistency: Three human microbiologists showed a high agreement of 94.4% on phenomenological categories, while the agreement rate between three different query outputs of EUCAST-GPT-expert was at 81.9%, indicating that the human experts’ decisions were more consistent.
- Importance of Customization: The study found that EUCAST-GPT-expert performed significantly better than uncustomized GPT-4. Standard GPT-4 was only able to interpret 19.6% of cases.
3.2.3. Antibiotic Discovery - Resistance Detection
- a)
-
Abaucin – against Acinetobacter baumannii subspeciesAbaucin (RS-102895, MLJS-21001) is a spirocyclic phenethylamine derivative that exhibits potent activity as a narrow-spectrum antibiotic. It is particularly effective against Acinetobacter baumannii, which the World Health Organization has identified as a "critical threat." Its AI-assisted discovery was carried out by researchers from MIT and McMaster University. Its mechanism of action is based on inhibiting bacterial lipoprotein transport. Interestingly, although the molecule was previously known as a CCR2 chemokine receptor antagonist, its antibiotic property was discovered in 2023. Abaucin was screened among thousands of compounds using graph neural networks (GNN) and machine learning models. Its narrow spectrum (targeting only A. baumannii) carries the potential to reduce the risk of antibiotic resistance. Its efficacy has been confirmed in skin infections and sepsis in mouse models. This discovery is a prime example of how AI can revolutionize antibiotic development [13].
- b)
-
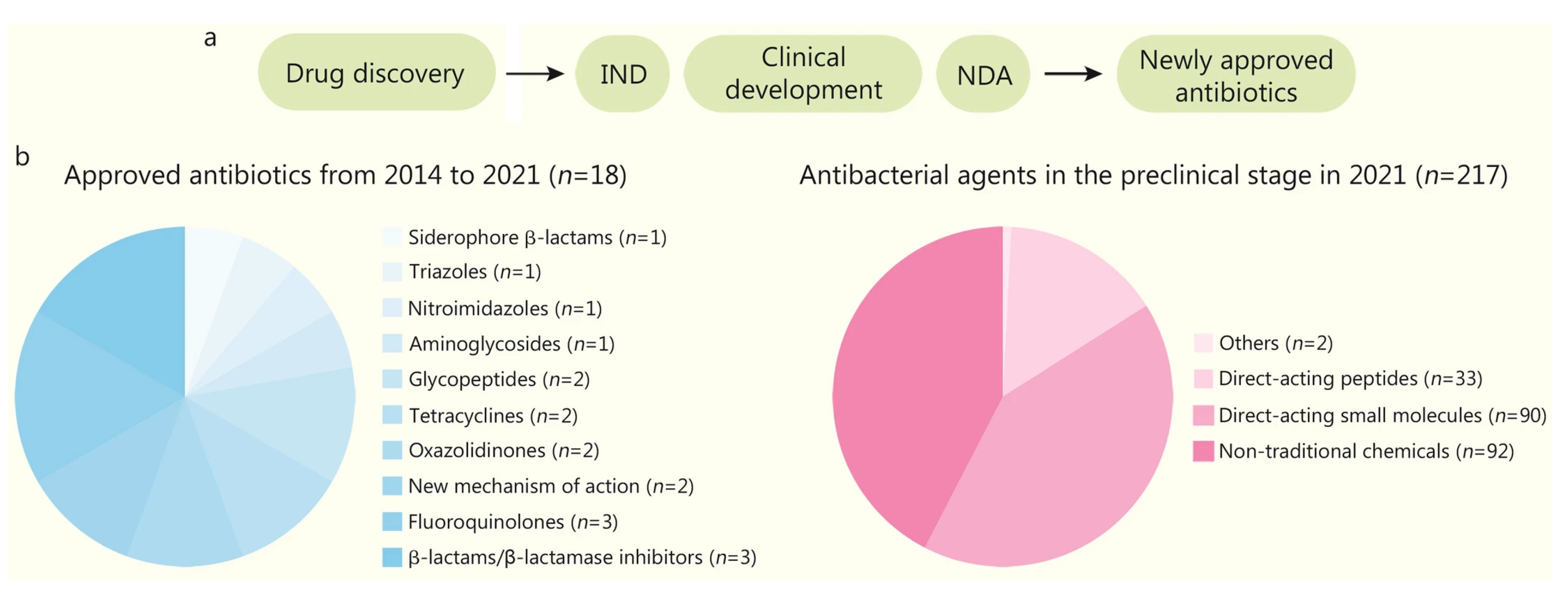
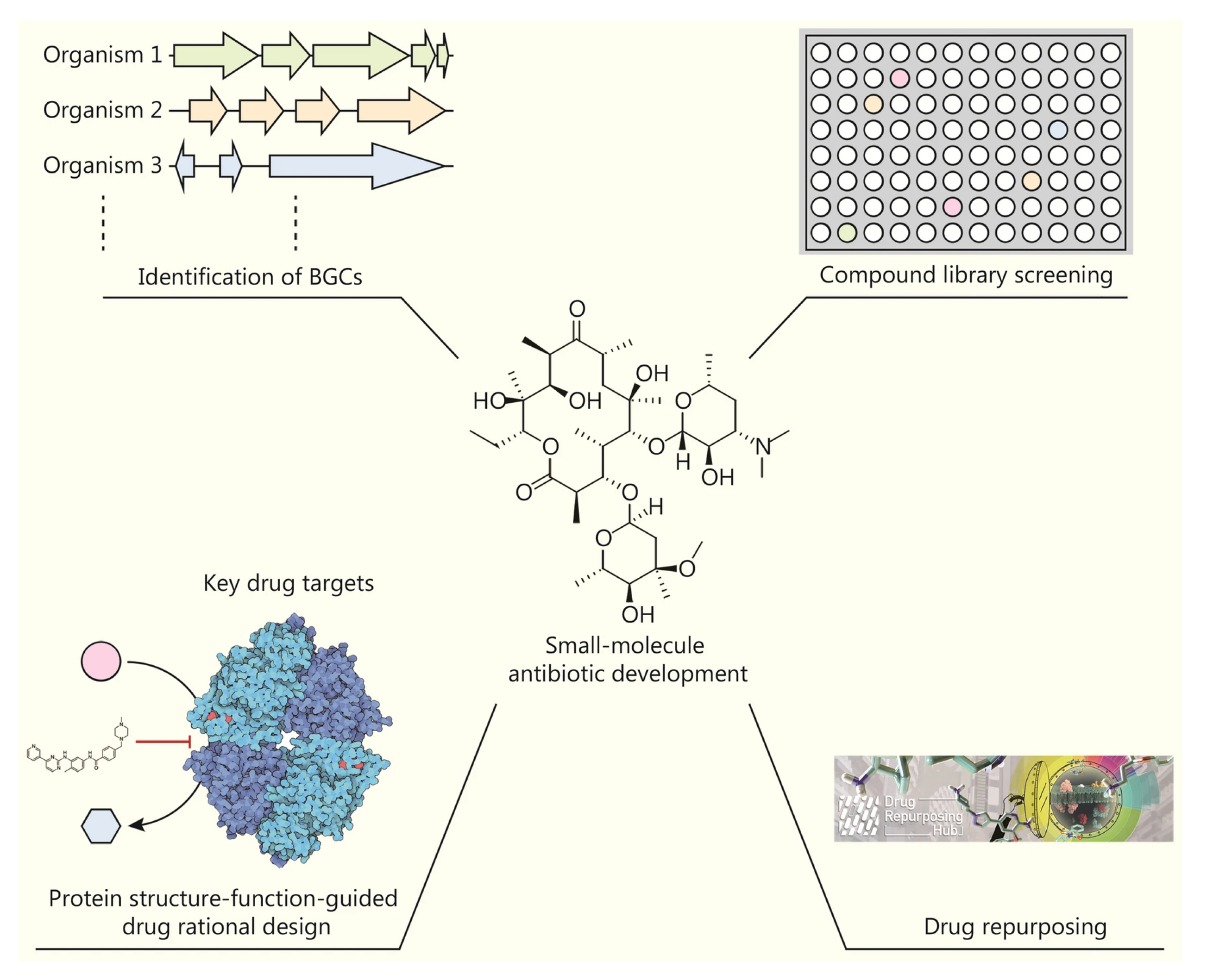
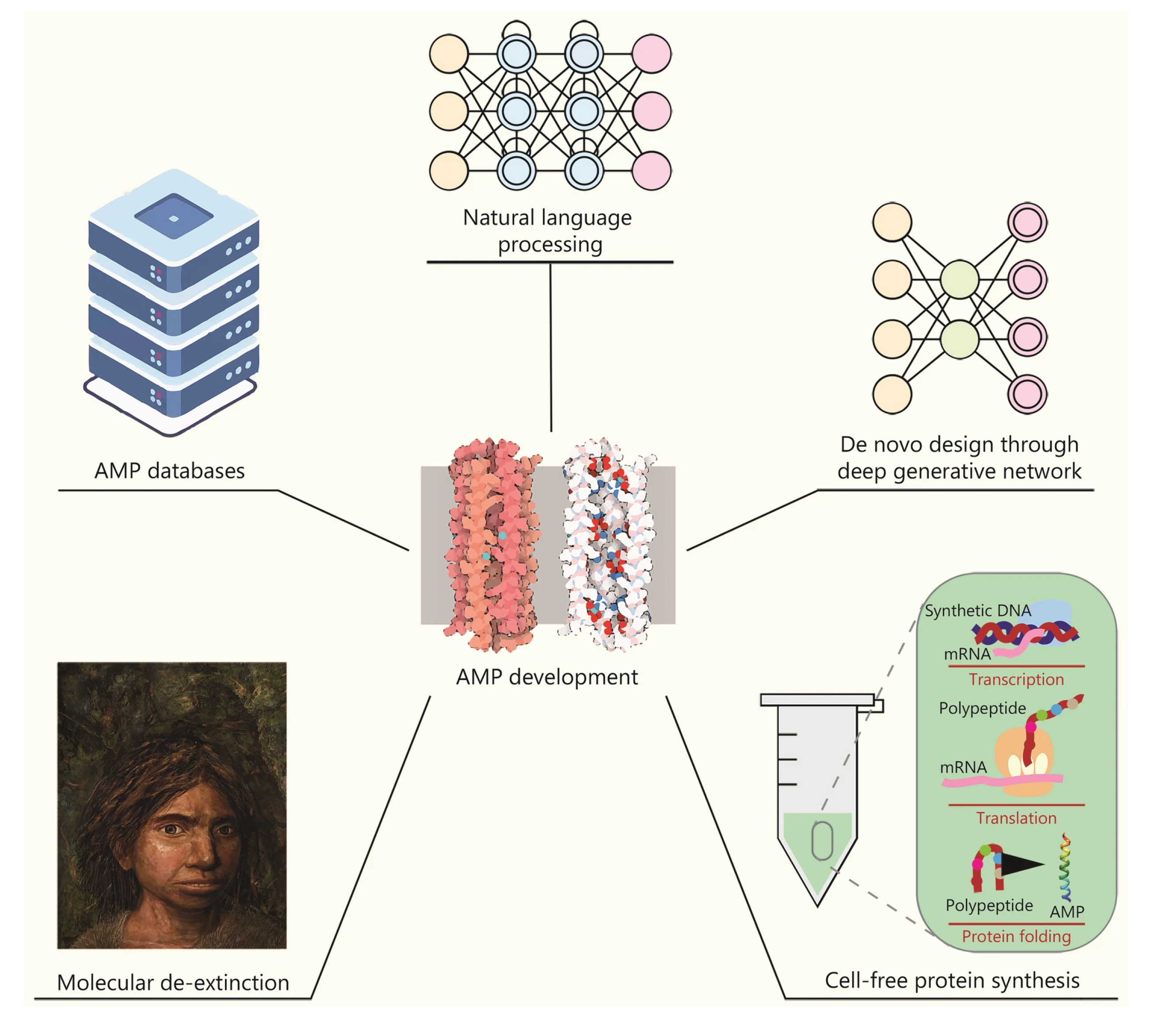
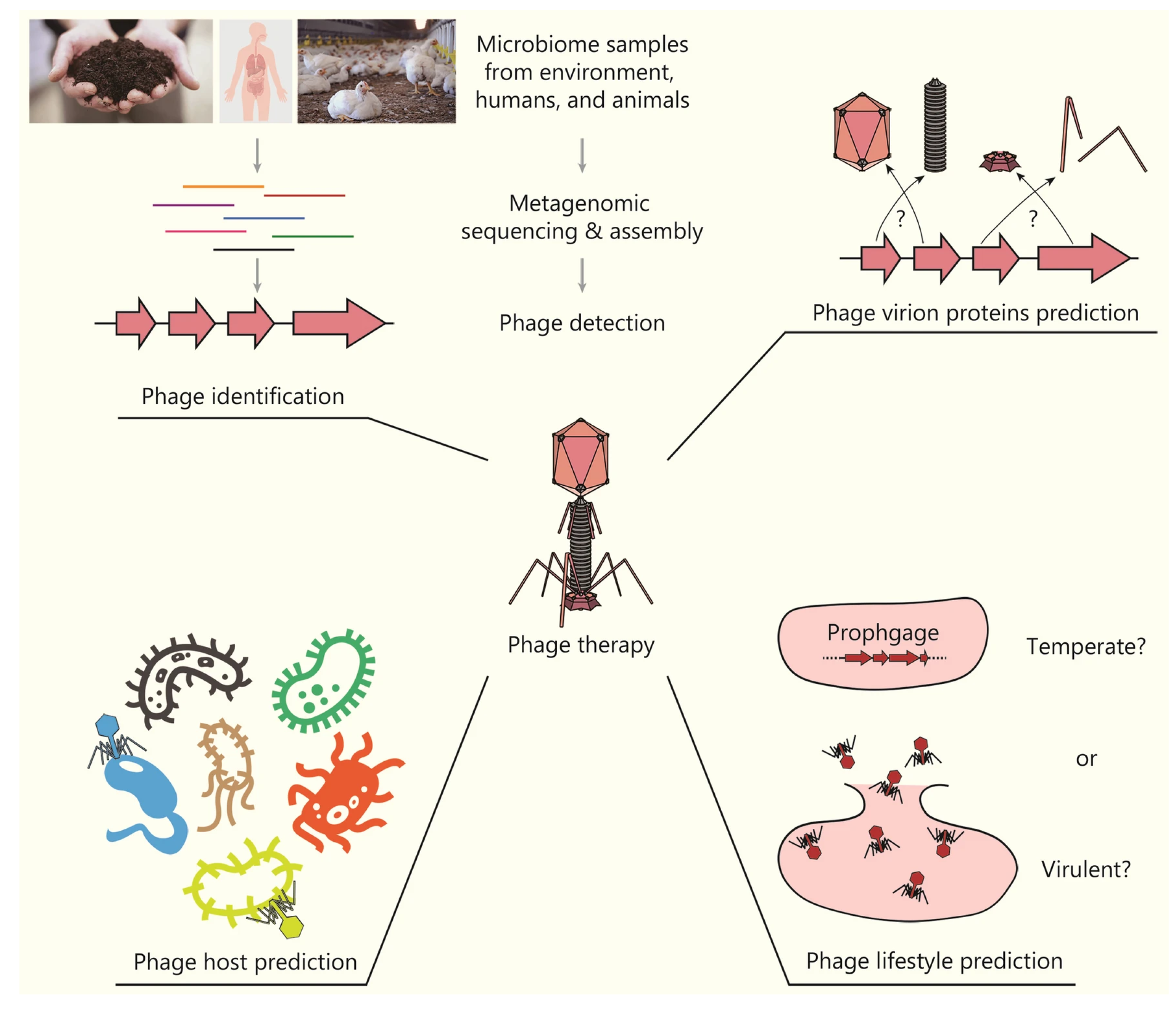
PROSPECT / A. baumannii sample One of the most serious problems threatening public health worldwide is the development of drug resistance by microorganisms, particularly bacteria. This phenomenon, known as Antimicrobial Resistance (AMR), renders previously effective treatments ineffective, prolongs illnesses, increases mortality rates, and escalates healthcare costs. According to the World Health Organization (WHO), 1.27 million deaths were directly attributed to bacterial AMR in 2019 alone. It is estimated that this number could reach 10 million per year by 2050. The discovery and development of new antibiotics to address this emergency is both time-consuming and costly using traditional methods. At this point, artificial intelligence (AI) technology emerges as the greatest beacon of hope in the face of this global crisis by accelerating drug discovery processes and offering new solutions.[14]Artificial intelligence, particularly machine learning (ML) and deep learning (DL) algorithms, analyze massive amounts of biological data to develop systems that mimic human intelligence. These technologies are being integrated into drug development processes, accelerating the discovery of new antibiotics. Traditionally, discovering a new drug involved lengthy and laborious steps, such as synthesizing thousands of chemicals derived from existing drugs and testing them for activity and toxicity. AI is transforming this process with multi-dimensional approaches:1-Development of Small Molecule Antibiotics:Discovery of Biosynthetic Gene Clusters (BGCs): In nature, many microorganisms harbor BGCs, which encode secondary metabolites with the potential to be developed into new antibiotics. However, most of these genes belong to organisms that cannot be cultured in vitro. To address this challenge, AI uses deep learning algorithms such as DeepBGC and DeepRiPP to predict new BGC classes and the chemical activity of their products from genetic data. Screening of Compound Libraries: Unlike traditional screening methods, machine learning-based screening can explore broad chemical spaces in silico. This significantly increases the likelihood of discovering structurally and functionally novel compounds with potent antibacterial properties. For example, a message-passing deep neural network trained on growth inhibition data from over 7,500 FDA-approved compounds discovered aubacin, a narrow-spectrum antibiotic that targets lipoprotein trafficking and has a novel mechanism of action (MOA).[14]2-Protein Structure Prediction and Rational Drug Design: AlphaFold2 and RoseTTAFold: Understanding the three-dimensional structure of a drug’s target protein is fundamental to rational drug design. Traditional methods are slow and technically challenging. AlphaFold2, developed by DeepMind, has revolutionized this field by predicting protein structure with near-experimental accuracy. RoseTTAFold, on the other hand, can predict protein-protein interactions directly from protein sequences.Problems and Solutions: Despite these powerful tools, accurately predicting how drug candidates bind to their targets remains challenging.Despite the high quality of AlphaFold2 predictions, the performance of docking methods is limited. Therefore, improving docking calculations using machine learning-based scoring functions is critical to overcoming these hurdles.[14]3-Design of Antimicrobial Peptides (AMP):Unique Mechanisms of Action: AMPs are short peptides, generally consisting of 2-50 amino acids, produced by multicellular organisms against pathogens. They act by directly interacting with the bacterial membrane, disrupting physiological processes such as cell wall synthesis, cell division, and membrane permeability. They can also prevent the formation of biofilms resistant to conventional antibiotics. AI-Assisted Design: AI plays a key role in the design and modification of new AMPs. According to the WHO’s 2021 report, 15.2% of 217 preclinical chemicals are direct-acting peptides. This demonstrates the growing interest in biologic drugs.[14]4-Development of Bacteriophage (Phage) Therapy: Phage Discovery and Classification: Phages, viruses that infect bacteria, are a promising alternative to the AMR crisis. AI has become an important tool for studying phages from natural sources. AI-powered tools such as PHACTS and BACPHLIP can classify phages as virulent (non-lysogenic) or temperate (lysogenic), facilitating the selection of the most suitable phages for treatment. These tools have achieved impressive accuracy rates of 99% and 98%, respectively.[14]
3.3. Challenges and Future Perspectives of Microbiology AI
3.4. Microbiology and AI Conclusion
4. Comparative Analysis-Result
4.1. Common and Domain-Specific Challenges
4.2. Solutions and Recommendations
4.3. Future Strategies in Biochemistry and Microbiology
References
- Esteva A, et al. A guide to deep learning in healthcare. Nature Medicine. 2019;25(1):24–29. [CrossRef]
- Greenspan H, van Ginneken B, Summers RM. Deep learning in medical imaging: overview and future promise of an exciting new technique. IEEE Transactions on Medical Imaging. 2016;35(5):1153–1159. [CrossRef]
- Jha S, Topol EJ. Adapting to artificial intelligence. JAMA. 2016;316(22):2353–2354. [CrossRef]
- Jiang F, Jiang Y, Zhi H, Dong Y, Li H, Ma S, et al. Artificial intelligence in healthcare: past, present and future. Stroke and Vascular Neurology. 2017;2(4):230–243. [CrossRef]
- Nelson DL, Cox MM. Lehninger Principles of Biochemistry. 8th ed. New York: W.H. Freeman and Company; 2021.
- Jin B, Liu H, Zhang Y, Guo T, Liu Y. Clinlabomics: leveraging clinical laboratory data by data mining and machine learning. BMC Bioinformatics. 2022;23(1):116. [CrossRef]
- Gupta P, Ali I, Awasthi A, Bhardwaj S, Shankhdhar PK, Gangwar V, Kumar A. The role of artificial intelligence in clinical biochemistry: a systematic review. International Journal of Pharmacy and Pharmaceutical Research. 2025;5(2):45–58.
- Smith KP, et al. Applications of artificial intelligence in clinical microbiology diagnostic testing. Clinical Microbiology Newsletter. 2020;42(8):61–70. [CrossRef]
- Iadanza E, et al. Gut microbiota and artificial intelligence approaches: a scoping review. Health and Technology. 2020;10:1–16. [CrossRef]
- Wang H, Koydemir HC, Qiu Y, Bai B, Zhang Y, Jin Y, et al. Early-detection and classification of live bacteria using time-lapse coherent imaging and deep learning. Light: Science & Applications. 2020;9:118. [CrossRef]
- Quince C, Walker AW, Simpson JT, Loman NJ, Segata N. Shotgun metagenomics, from sampling to analysis. Nature Biotechnology. 2017;35(9):833–844. [CrossRef]
- Lescure FX, et al. GPT-4-based AI agents—the new expert system for detection of antimicrobial resistance mechanisms? Journal of Clinical Microbiology. 2024;62(11):e00689-24. [CrossRef]
- Liu G, Catacutan DB, Rathod K, Swanson K, Jin W, Mohammed JC, et al. Deep learning-guided discovery of an antibiotic targeting Acinetobacter baumannii. Nature Chemical Biology. 2023;19(11):1342–1350. [CrossRef]
- Liu GY, Yu D, Fan MM, Zhang X, Jin ZY, Tang C, et al. Antimicrobial resistance crisis: could artificial intelligence be the solution? Military Medical Research. 2024;11:7. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



