Submitted:
17 September 2025
Posted:
18 September 2025
You are already at the latest version
Abstract
Keywords:
Introduction
1. Rapid Adaptive Evolution in a New Host Species
2. Adaptive Radiation and Selective Sweeps
3. The Molecular Nature of Adaptation
4. The Constraints of Adaptation
4.1. Deleterious Pleotropic Effects
4.2. The Order of Mutation Fixation and Diminishing Returns
4.3. Path Exclusion
4.4. “Yoyo Mutations”: Recycling the Same Tricks
5. Punctuated “Equilibrium”
6. Muller’s Ratchet
6.1. Accumulation of Mutations in Nonstructural and Accessory Genes: Weak Selection, Drift, and Hitchhiking
6.2. The Mistigri Hypothesis on Gene Loss
6.3. Deoptimization of Codon Usage
6.4. Loss of Proofreading Function
7. Preservation in Immunocompromised Patients?
Conclusions
References
- Manrubia SC, Lázaro E. Viral evolution. Phys Life Rev. [CrossRef]
- Khare S, Gurry C, Freitas L, et al. GISAID’s Role in Pandemic Response. China CDC Weekly, 2021, Vol 3, Issue 49, Pages: 1049-1051, 1049. [CrossRef]
- Hodcroft, EB. CoVariants: SARS-CoV-2 Mutations and Variants of Interest. https://covariants.org/.
- Mukherjee V, Postelnicu R, Parker C, et al. COVID-19 Across Pandemic Variant Periods: The Severe Acute Respiratory Infection-Preparedness (SARI-PREP) Study. Crit Care Explor. [CrossRef]
- Endo H, Lee K, Ohnuma T, Watanabe S, Fushimi K. Temporal trends in clinical characteristics and in-hospital mortality among patients with COVID-19 in Japan for waves 1, 2, and 3: A retrospective cohort study. Journal of Infection and Chemotherapy, 1393. [CrossRef]
- Flisiak R, Zarębska-Michaluk D, Dobrowolska K, et al. Change in the Clinical Picture of Hospitalized Patients with COVID-19 between the Early and Late Period of Dominance of the Omicron SARS-CoV-2 Variant. Journal of Clinical Medicine 2023, Vol 12, Page 5572, 5572. [CrossRef]
- Kojima N, Adams K, Self WH, et al. Changing Severity and Epidemiology of Adults Hospitalized With Coronavirus Disease 2019 (COVID-19) in the United States After Introduction of COVID-19 Vaccines, March 2021–August 2022. Clinical Infectious Diseases, 20 March. [CrossRef]
- Liu, Y. Is SARS-CoV-2 facing constraints in its adaptive evolution? Biomolecules and Biomedicine, 9 June 2025. [Google Scholar] [CrossRef]
- Hadfield J, Megill C, Bell SM, et al. Nextstrain: real-time tracking of pathogen evolution. Bioinformatics, 4121. [CrossRef]
- Virgin HW, Wherry EJ, Ahmed R. Redefining Chronic Viral Infection. Cell. [CrossRef]
- Keeble, SA. B VIRUS INFECTION IN MONKEYS. Ann N Y Acad Sci. [CrossRef]
- Silvestri G, Paiardini M, Pandrea I, Lederman MM, Sodora DL. Understanding the benign nature of SIV infection in natural hosts. J Clin Invest, 3148. [CrossRef]
- Watanabe S, Masangkay JS, Nagata N, et al. Bat Coronaviruses and Experimental Infection of Bats, the Philippines. Emerg Infect Dis, 1217. [CrossRef]
- Hemnani M, da Silva PG, Thompson G, Poeta P, Rebelo H, Mesquita JR. Detection and Prevalence of Coronaviruses in European Bats: A Systematic Review. Ecohealth. 2024 Dec;21(2-4):125-140. [CrossRef]
- Beigel JH, de Jong D, Thi Kim Tien N, et al. Avian Influenza A (H5N1) Infection in Humans. New England Journal of Medicine, 1374. [CrossRef]
- Omrani AS, Al-Tawfiq JA, Memish ZA. Middle East respiratory syndrome coronavirus (MERS-CoV): animal to human interaction. Pathog Glob Health. [CrossRef]
- Parrish CR, Holmes EC, Morens DM, et al. Cross-species virus transmission and the emergence of new epidemic diseases. Microbiol Mol Biol Rev. [CrossRef]
- Maljkovic Berry I, Athreya G, Kothari M, et al. The evolutionary rate dynamically tracks changes in HIV-1 epidemics: Application of a simple method for optimizing the evolutionary rate in phylogenetic trees with longitudinal data. Epidemics. [CrossRef]
- Amicone M, Borges V, Alves MJ, et al. Mutation rate of SARS-CoV-2 and emergence of mutators during experimental evolution. Evol Med Public Health. [CrossRef]
- Sasani TA, Pedersen BS, Gao Z, et al. Large, three-generation human families reveal post-zygotic mosaicism and variability in germline mutation accumulation. Elife. [CrossRef]
- Eymieux S, Rouillé Y, Terrier O, et al. Ultrastructural modifications induced by SARS-CoV-2 in Vero cells: a kinetic analysis of viral factory formation, viral particle morphogenesis and virion release. Cellular and Molecular Life Sciences, 3565. [CrossRef]
- Wang RJ, Al-Saffar SI, Rogers J, Hahn MW. Human generation times across the past 250,000 years. Sci Adv. [CrossRef]
- Plante JA, Liu Y, Liu J, et al. Spike mutation D614G alters SARS-CoV-2 fitness. Nature, 7852. [CrossRef]
- Focosi D, Spezia PG, Maggi F. Fixation and reversion of mutations in the receptor-binding domain of SARS-CoV-2 spike protein. Diagn Microbiol Infect Dis. Elsevier Inc. [CrossRef]
- Ru H, Zhang P, Wu H. Structural gymnastics of RAG-mediated DNA cleavage in V(D)J recombination. Curr Opin Struct Biol. [CrossRef]
- Polimanti R, Carboni C, Baesso I, et al. Genetic variability of glutathione S-transferase enzymes in human populations: Functional inter-ethnic differences in detoxification systems. Gene. [CrossRef]
- Peng L, Zhong X. Epigenetic regulation of drug metabolism and transport. Acta Pharm Sin B. [CrossRef]
- Kaltenbach M, Tokuriki N. Dynamics and constraints of enzyme evolution. J Exp Zool B Mol Dev Evol. [CrossRef]
- Liu K, Tan S, Niu S, et al. Cross-species recognition of SARS-CoV-2 to bat ACE2. Proceedings of the National Academy of Sciences. [CrossRef]
- Aicher SM, Streicher F, Chazal M, et al. Species-Specific Molecular Barriers to SARS-CoV-2 Replication in Bat Cells. J Virol. [CrossRef]
- Bakre A, Sweeney R, Espinoza E, Suarez DL, Kapczynski DR. The ACE2 Receptor from Common Vampire Bat (Desmodus rotundus) and Pallid Bat (Antrozous pallidus) Support Attachment and Limited Infection of SARS-CoV-2 Viruses in Cell Culture. Viruses. [CrossRef]
- Wark PAB, Pathinayake PS, Kaiko G, et al. <scp>ACE2</scp> expression is elevated in airway epithelial cells from older and male healthy individuals but reduced in asthma. Respirology. [CrossRef]
- Zhao H, Lu L, Peng Z, et al. SARS-CoV-2 Omicron variant shows less efficient replication and fusion activity when compared with Delta variant in TMPRSS2-expressed cells. Emerg Microbes Infect. [CrossRef]
- Sakurai Y, Okada S, Ozeki T, Yoshikawa R, Kinoshita T, Yasuda J. SARS-CoV-2 Omicron subvariants progressively adapt to human cells with altered host cell entry. mSphere. [CrossRef]
- Liu Y, Zhao X, Shi J, et al. Lineage-specific pathogenicity, immune evasion, and virological features of SARS-CoV-2 BA.2.86/JN.1 and EG.5.1/HK.3. Nature Communications. [CrossRef]
- Yuan M, Liu H, Wu NC, Wilson IA. Recognition of the SARS-CoV-2 receptor binding domain by neutralizing antibodies. Biochem Biophys Res Commun. [CrossRef]
- Chen EC, Gilchuk P, Zost SJ, et al. Convergent antibody responses to the SARS-CoV-2 spike protein in convalescent and vaccinated individuals. Cell Rep, 1096. [CrossRef]
- Starr TN, Greaney AJ, Addetia A, et al. Prospective mapping of viral mutations that escape antibodies used to treat COVID-19. Science (1979), 6531. [CrossRef]
- Greaney AJ, Loes AN, Crawford KHD, et al. Comprehensive mapping of mutations in the SARS-CoV-2 receptor-binding domain that affect recognition by polyclonal human plasma antibodies. Cell Host Microbe. [CrossRef]
- Zhang L, Jackson CB, Mou H, et al. SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nat Commun. [CrossRef]
- Zhang J, Cai Y, Xiao T, et al. Structural impact on SARS-CoV-2 spike protein by D614G substitution. Science (1979), 6541. [CrossRef]
- Xu X, Wu Y, Kummer AG, et al. Assessing changes in incubation period, serial interval, and generation time of SARS-CoV-2 variants of concern: a systematic review and meta-analysis. BMC Med. [CrossRef]
- Yao Z, Zhang L, Duan Y, Tang X, Lu J. Molecular insights into the adaptive evolution of SARS-CoV-2 spike protein. Journal of Infection, 1061. [CrossRef]
- McGuigan K, Collet JM, Allen SL, Chenoweth SF, Blows MW. Pleiotropic Mutations Are Subject to Strong Stabilizing Selection. Genetics, 1051. [CrossRef]
- Otto, SP. Two steps forward, one step back: The pleiotropic effects of favoured alleles. Proceedings of the Royal Society B: Biological Sciences, 1540. [Google Scholar] [CrossRef]
- Li W, Xu Z, Niu T, et al. Key mechanistic features of the trade-off between antibody escape and host cell binding in the SARS-CoV-2 Omicron variant spike proteins. EMBO J, 1484. [CrossRef]
- Raisinghani N, Alshahrani M, Gupta G, Verkhivker G. AlphaFold2 Modeling and Molecular Dynamics Simulations of the Conformational Ensembles for the SARS-CoV-2 Spike Omicron JN.1, KP.2 and KP.3 Variants: Mutational Profiling of Binding Energetics Reveals Epistatic Drivers of the ACE2 Affinity and Escape Hotspots of Antibody Resistance. Viruses, 1458. [CrossRef]
- Zahradník J, Marciano S, Shemesh M, et al. SARS-CoV-2 variant prediction and antiviral drug design are enabled by RBD in vitro evolution. Nat Microbiol, 1188. [CrossRef]
- Liu H, Wei P, Kappler JW, Marrack P, Zhang G. SARS-CoV-2 Variants of Concern and Variants of Interest Receptor Binding Domain Mutations and Virus Infectivity. Front Immunol. [CrossRef]
- Yang H, Guo H, Wang A, et al. Structural basis for the evolution and antibody evasion of SARS-CoV-2 BA.2.86 and JN.1 subvariants. Nature Communications. [CrossRef]
- Yang S, Yu Y, Xu Y, et al. Fast evolution of SARS-CoV-2 BA.2.86 to JN.1 under heavy immune pressure. Lancet Infect Dis. Elsevier Ltd. [CrossRef]
- Sugano A, Murakami J, Kataguchi H, et al. In silico binding affinity of the spike protein with ACE2 and the relative evolutionary distance of S gene may be potential factors rapidly obtained for the initial risk of SARS-CoV-2. Microb Risk Anal, 1002. [CrossRef]
- Gangavarapu K, Latif AA, Mullen JL, et al. Outbreak.info genomic reports: scalable and dynamic surveillance of SARS-CoV-2 variants and mutations. Nat Methods. [CrossRef]
- Chou HH, Chiu HC, Delaney NF, Segrè D, Marx CJ. Diminishing Returns Epistasis Among Beneficial Mutations Decelerates Adaptation. Science (1979), 6034. [CrossRef]
- Khan AI, Dinh DM, Schneider D, Lenski RE, Cooper TF. Negative Epistasis Between Beneficial Mutations in an Evolving Bacterial Population. Science (1979), 6034. [CrossRef]
- Weinreich DM, Delaney NF, DePristo MA, Hartl DL. Darwinian Evolution Can Follow Only Very Few Mutational Paths to Fitter Proteins. Science (1979), 5770. [CrossRef]
- Jangra S, Ye C, Rathnasinghe R, et al. SARS-CoV-2 spike E484K mutation reduces antibody neutralisation. Lancet Microbe. [CrossRef]
- Barton MI, Macgowan S, Kutuzov M, Dushek O, Barton GJ, Anton Van Der Merwe P. Effects of common mutations in the sars-cov-2 spike rbd and its ligand the human ace2 receptor on binding affinity and kinetics. Elife. [CrossRef]
- Kamble P, Daulatabad V, Singhal A, et al. JN.1 variant in enduring COVID-19 pandemic: is it a variety of interest (VoI) or variety of concern (VoC)? Horm Mol Biol Clin Investig. Walter de Gruyter GmbH. [CrossRef]
- Starr TN, Greaney AJ, Hannon WW, et al. Shifting mutational constraints in the SARS-CoV-2 receptor-binding domain during viral evolution. Science (1979), 6604. [CrossRef]
- Annavajhala MK, Mohri H, Wang P, et al. Emergence and expansion of SARS-CoV-2 B.1.526 after identification in New York. Nature, 7878. [CrossRef]
- Qu P, Evans JP, Zheng YM, et al. Evasion of neutralizing antibody responses by the SARS-CoV-2 BA.2.75 variant. Cell Host Microbe, 1518. [CrossRef]
- Nguyen HL, Nguyen TQ, Li MS. SARS-CoV-2 Omicron Subvariants Do Not Differ Much in Binding Affinity to Human ACE2: A Molecular Dynamics Study. J Phys Chem B, 3340. [CrossRef]
- Cao Y, Song W, Wang L, et al. Characterization of the enhanced infectivity and antibody evasion of Omicron BA.2.75. Cell Host Microbe, 1527. [CrossRef]
- Huo J, Dijokaite-Guraliuc A, Liu C, et al. A delicate balance between antibody evasion and ACE2 affinity for Omicron BA.2.75. Cell Rep, 1119. [CrossRef]
- Mannar D, Saville JW, Zhu X, et al. Structural analysis of receptor binding domain mutations in SARS-CoV-2 variants of concern that modulate ACE2 and antibody binding. Cell Rep, 1101. [CrossRef]
- Wang Y, Liu C, Zhang C, et al. Structural basis for SARS-CoV-2 Delta variant recognition of ACE2 receptor and broadly neutralizing antibodies. Nat Commun. [CrossRef]
- Neher, RA. Contributions of adaptation and purifying selection to SARS-CoV-2 evolution. Virus Evol. [CrossRef]
- Tchesnokova V, Kulasekara H, Larson L, et al. Acquisition of the L452R Mutation in the ACE2-Binding Interface of Spike Protein Triggers Recent Massive Expansion of SARS-CoV-2 Variants. J Clin Microbiol. [CrossRef]
- Chan KC, Song Y, Xu Z, Shang C, Zhou R. SARS-CoV-2 Delta Variant: Interplay between Individual Mutations and Their Allosteric Synergy. Biomolecules, 1742. [CrossRef]
- Innocenti G, Obara M, Costa B, et al. Real-time identification of epistatic interactions in SARS-CoV-2 from large genome collections. Genome Biol. [CrossRef]
- Philip AM, Ahmed WS, Biswas KH. Reversal of the unique Q493R mutation increases the affinity of Omicron S1-RBD for ACE2. Comput Struct Biotechnol J, 1966. [CrossRef]
- Wang Q, Guo Y, Iketani S, et al. Antibody evasion by SARS-CoV-2 Omicron subvariants BA.2.12.1, BA.4 and BA.5. Nature, 7923. [CrossRef]
- Taylor AL, Starr TN. Deep mutational scanning of SARS-CoV-2 Omicron BA.2.86 and epistatic emergence of the KP.3 variant. Virus Evol. [CrossRef]
- Feng L, Sun Z, Zhang Y, et al. Structural and molecular basis of the epistasis effect in enhanced affinity between SARS-CoV-2 KP.3 and ACE2. Cell Discov. [CrossRef]
- Jian F, Feng L, Yang S, et al. Convergent evolution of SARS-CoV-2 XBB lineages on receptor-binding domain 455–456 synergistically enhances antibody evasion and ACE2 binding. PLoS Pathog, 1011. [CrossRef]
- Hulo C, De Castro E, Masson P, et al. ViralZone: a knowledge resource to understand virus diversity. Nucleic Acids Res. [CrossRef]
- Dejnirattisai W, Huo J, Zhou D, et al. SARS-CoV-2 Omicron-B.1.1.529 leads to widespread escape from neutralizing antibody responses. Cell. [CrossRef]
- Wang Q, Mellis IA, Ho J, et al. Recurrent SARS-CoV-2 spike mutations confer growth advantages to select JN.1 sublineages. Emerg Microbes Infect. Taylor and Francis Ltd. [CrossRef]
- Duarte CM, Ketcheson DI, Eguíluz VM, et al. Rapid evolution of SARS-CoV-2 challenges human defenses. Sci Rep, 6457. [CrossRef]
- Muller, HJ. The relation of recombination to mutational advance. Mutation Research - Fundamental and Molecular Mechanisms of Mutagenesis. [CrossRef]
- Chao, L. Fitness of RNA virus decreased by Muller’s ratchet. Nature, 6300. [Google Scholar] [CrossRef]
- Andersson DI, Hughes D. Muller’s ratchet decreases fitness of a DNA-based microbe. Proc Natl Acad Sci U S A. [CrossRef]
- Pandit A, Sinha S. Differential Trends in the Codon Usage Patterns in HIV-1 Genes. PLoS One, 2888. [CrossRef]
- Carter RW, Sanford JC. A new look at an old virus: patterns of mutation accumulation in the human H1N1 influenza virus since 1918. Theor Biol Med Model. [CrossRef]
- Chernyaeva EN, Ayginin AA, Kosenkov A V., et al. SARS-CoV-2 Recombination and Coinfection Events Identified in Clinical Samples in Russia. Viruses, 1660. [CrossRef]
- Bloom JD, Neher RA. Fitness effects of mutations to SARS-CoV-2 proteins. Virus Evol. [CrossRef]
- Lippi G, Henry BM. The landscape of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) genomic mutations. J Lab Precis Med. [CrossRef]
- McGrath ME, Xue Y, Dillen C, et al. SARS-CoV-2 variant spike and accessory gene mutations alter pathogenesis. Proceedings of the National Academy of Sciences. [CrossRef]
- Gupta S, Gupta D, Bhatnagar S. Analysis of SARS-CoV-2 genome evolutionary patterns. Microbiol Spectr. [CrossRef]
- Yi K, Kim SY, Bleazard T, Kim T, Youk J, Ju YS. Mutational spectrum of SARS-CoV-2 during the global pandemic. Exp Mol Med, 1229. [CrossRef]
- Colson P, Chaudet H, Delerce J, et al. Role of SARS-CoV-2 mutations in the evolution of the COVID-19 pandemic. Journal of Infection, 1061. [CrossRef]
- Omotoso OE, Babalola AD, Matareek A. Mutational hotspots and conserved domains of SARS-CoV-2 genome in African population. Beni Suef Univ J Basic Appl Sci. [CrossRef]
- Klute S, Nchioua R, Cordsmeier A, et al. Mutation T9I in Envelope confers autophagy resistance to SARS-CoV-2 Omicron. iScience, 1129. [CrossRef]
- Wang W, Qu Y, Wang X, et al. Genetic variety of ORF3a shapes SARS-CoV-2 fitness through modulation of lipid droplet. J Med Virol. [CrossRef]
- Hartmann S, Radochonski L, Ye C, Martinez-Sobrido L, Chen J. SARS-CoV-2 ORF3a drives dynamic dense body formation for optimal viral infectivity. Nat Commun, 4393. [CrossRef]
- López-Ayllón BD, de Lucas-Rius A, Mendoza-García L, et al. SARS-CoV-2 accessory proteins involvement in inflammatory and profibrotic processes through IL11 signaling. Front Immunol. [CrossRef]
- Focosi D, Spezia PG, Maggi F. Subsequent Waves of Convergent Evolution in SARS-CoV-2 Genes and Proteins. Vaccines (Basel). [CrossRef]
- Foster CSP, Bull RA, Tedla N, et al. Persistence of a Frameshifting Deletion in SARS-CoV-2 ORF7a for the Duration of a Major Outbreak. Viruses. [CrossRef]
- Panzera Y, Calleros L, Goñi N, et al. Consecutive deletions in a unique Uruguayan SARS-CoV-2 lineage evidence the genetic variability potential of accessory genes. PLoS One, 0263. [CrossRef]
- Lam JY, Kok KH. The Recurring Loss of ORF8 Secretion in Dominant SARS-CoV-2 Variants. Int J Mol Sci, 5778. [CrossRef]
- Zhang Y, Chen Y, Li Y, et al. The ORF8 protein of SARS-CoV-2 mediates immune evasion through down-regulating MHC-Ι. Proceedings of the National Academy of Sciences. [CrossRef]
- Arduini A, Laprise F, Liang C. SARS-CoV-2 ORF8: A Rapidly Evolving Immune and Viral Modulator in COVID-19. Viruses. [CrossRef]
- Wagner C, Kistler KE, Perchetti GA, et al. Positive selection underlies repeated knockout of ORF8 in SARS-CoV-2 evolution. Nature Communications. [CrossRef]
- Colson P, Delerce J, Fantini J, Pontarotti P, La Scola B, Raoult D. The return of the “Mistigri” (virus adaptative gain by gene loss) through the SARS-CoV-2 XBB.1.5 chimera that predominated in 2023. J Med Virol. [CrossRef]
- Rayati Damavandi A, Dowran R, Al Sharif S, Kashanchi F, Jafari R. Molecular variants of SARS-CoV-2: antigenic properties and current vaccine efficacy. Med Microbiol Immunol. [CrossRef]
- Moran, NA. Microbial Minimalism. Cell. [CrossRef]
- Meyer H, Sutter G, Mayr A. Mapping of deletions in the genome of the highly attenuated vaccinia virus MVA and their influence on virulence. Journal of General Virology, 1031. [CrossRef]
- Haddox HK, Angehrn G, Sesta L, et al. The mutation rate of SARS-CoV-2 is highly variable between sites and is influenced by sequence context, genomic region, and RNA structure. Nucleic Acids Res. [CrossRef]
- Bai H, Ata G, Sun Q, Rahman SU, Tao S. Natural selection pressure exerted on “Silent” mutations during the evolution of SARS-CoV-2: Evidence from codon usage and RNA structure. Virus Res. [CrossRef]
- Fumagalli SE, Padhiar NH, Meyer D, et al. Analysis of 3.5 million SARS-CoV-2 sequences reveals unique mutational trends with consistent nucleotide and codon frequencies. Virol J. [CrossRef]
- Davidson A, Parr M, Totzeck F, et al. Over time analysis of the codon usage of SARS-CoV-2 and its variants. Comput Struct Biotechnol J, 2034. [CrossRef]
- Mogro EG, Bottero D, Lozano MJ. Analysis of SARS-CoV-2 synonymous codon usage evolution throughout the COVID-19 pandemic. Virology. [CrossRef]
- Wu X, Shan K jia, Zan F, Tang X, Qian Z, Lu J. Optimization and Deoptimization of Codons in SARS-CoV-2 and Related Implications for Vaccine Development. Advanced Science. [CrossRef]
- Posani E, Dilucca M, Forcelloni S, Pavlopoulou A, Georgakilas AG, Giansanti A. Temporal evolution and adaptation of SARS-CoV-2 codon usage. Frontiers in Bioscience-Landmark. [CrossRef]
- Huang W, Guo Y, Li N, Feng Y, Xiao L. Codon usage analysis of zoonotic coronaviruses reveals lower adaptation to humans by SARS-CoV-2. Infection, Genetics and Evolution. [CrossRef]
- Padhiar NH, Ghazanchyan T, Fumagalli SE, et al. SARS-CoV-2 CoCoPUTs: analyzing GISAID and NCBI data to obtain codon statistics, mutations, and free energy over a multiyear period. Virus Evol. [CrossRef]
- Takada K, Ueda MT, Shichinohe S, et al. Genomic diversity of SARS-CoV-2 can be accelerated by mutations in the nsp14 gene. iScience, 1062. [CrossRef]
- Eskier D, Suner A, Oktay Y, Karakülah G. Mutations of SARS-CoV-2 nsp14 exhibit strong association with increased genome-wide mutation load. PeerJ, 1018. [CrossRef]
- Hassan SS, Bhattacharya T, Nawn D, et al. SARS-CoV-2 NSP14 governs mutational instability and assists in making new SARS-CoV-2 variants. Comput Biol Med, 1078. [CrossRef]
- Sangare K, Liu S, Selvaraj P, Stauft CB, Starost MF, Wang TT. Combined mutations in nonstructural protein 14, envelope, and membrane proteins mitigate the neuropathogenicity of SARS-CoV-2 Omicron BA.1 in K18-hACE2 mice. mSphere. [CrossRef]
- Ahmed JQ, Maulud SQ, Al-Qadi R, et al. Sequencing and mutations analysis of the first recorded SARS-CoV-2 Omicron variant during the fourth wave of pandemic in Iraq. The Brazilian Journal of Infectious Diseases, 1026. [CrossRef]
- Markov P, V. , Ghafari M, Beer M, et al. The evolution of SARS-CoV-2. Nat Rev Microbiol. Nature Research. [CrossRef]
- Yang C, Zhao H, Espín E, Tebbutt SJ. Association of SARS-CoV-2 infection and persistence with long COVID. Lancet Respir Med. [CrossRef]
- Machkovech HM, Hahn AM, Garonzik Wang J, et al. Persistent SARS-CoV-2 infection: significance and implications. Lancet Infect Dis. [CrossRef]
| SARS-CoV-2 | Human | |
|---|---|---|
| Mutation rate (per nucleotide per generation) | 1-2 x 10-6 [19] | 1-2 x 10-8 [20] |
| Generation time | 8-24 hours [21] | 26.9 years [22] |
| History | 5.5 years | 7 million years |
| Generations | 2,000 - 6,000 | 260,000 |
| Total mutations per nucleotide | 2 x 10-3 – 1.2 x 10-2 | 2.6 x 10-3 - 5.2 x 10-3 |
| RBD position | 358 | 445 | 446 | 448 | 460 | 468 | 470 | 477 | 478 | 481 | 483 | 484 | 490 | 493 | 494 | 498 | 501 | |
| WT residue | I | V | G | N | N | I | T | S | T | N | V | E | F | Q | S | Q | N | |
| Observed in In Vitro evolution products | F | K | R | S | K | T/V | M | N | S | Y | E | K | Y/S | H | P | R | Y | |
| In vitro product Clone B62 | F | K | K | T | M | N | K | R | Y | |||||||||
| Alpha | Y | |||||||||||||||||
| Beta | K | Y | ||||||||||||||||
| Gamma | K | Y | ||||||||||||||||
| Delta | K | |||||||||||||||||
| Zeta | K | |||||||||||||||||
| Eta | K | |||||||||||||||||
| Theta | K | Y | ||||||||||||||||
| Iota | (N) | (K) | ||||||||||||||||
| Kappa | K | |||||||||||||||||
| Lambda | S | |||||||||||||||||
| Mu | K | Y | ||||||||||||||||
| O m i c r o n |
BA.1 | S | N | K | A | R | R | Y | ||||||||||
| BA.2 | N | K | A | R | R | Y | ||||||||||||
| BA.4 | N | K | A | R | Y | |||||||||||||
| BA.5 | N | K | A | R | Y | |||||||||||||
| BQ.1 | K | N | K | A | R | Y | ||||||||||||
| BA.2.75 | S | K | N | K | A | R | Y | |||||||||||
| XBB.1.5 | P | S | K | N | K | A | S | R | R | Y | ||||||||
| BA2.86 | H | S | K | N | K | K | K | R | Y | |||||||||
| JN.1 | H | S | K | N | K | K | K | R | Y | |||||||||
| KP3.1 | H | S | K | N | K | K | K | E | R | Y | ||||||||
| LP.8.1 | R | S | K | N | K | K | K | E | R | Y | ||||||||
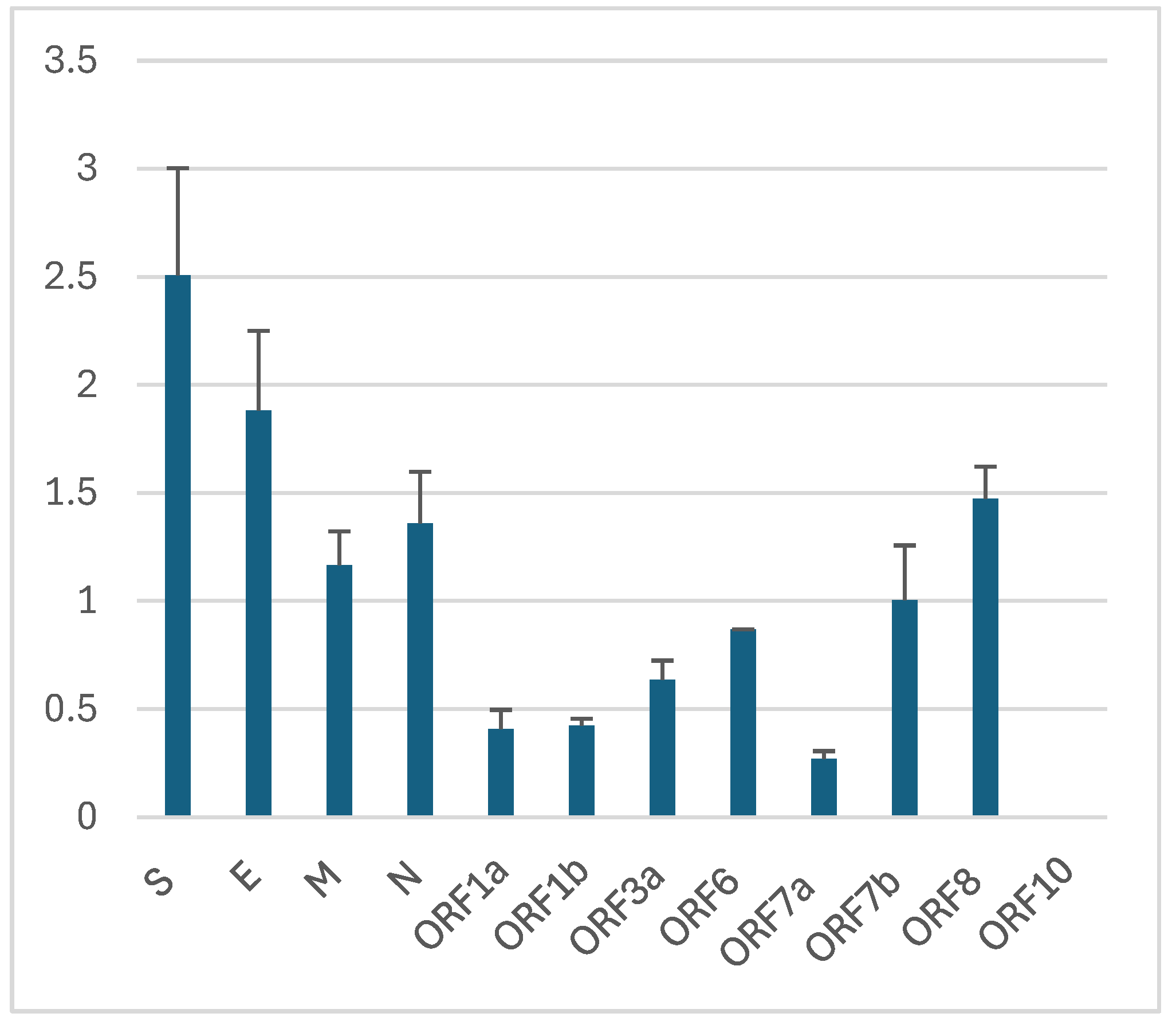
| Protein | Gene size | Number of mutations according to [88] | Number of mutations according to [92] | Fixed by Feb 2022 [88] | Fixed by Oct 2022 [92] | Fixation percent according to [88] | Fixation percent according to [92] | Mean Fixation Rate | SD |
|---|---|---|---|---|---|---|---|---|---|
| Spike | 3821 | 3799 | 2974 | 82 | 85 | 2.2 | 2.9 | 2.5 | 0.5 |
| Envelope | 227 | 185 | 140 | 3 | 3 | 1.6 | 2.1 | 1.9 | 0.4 |
| Membrane | 668 | 568 | 470 | 6 | 6 | 1.1 | 1.3 | 1.2 | 0.2 |
| Nucleocapsid | 1259 | 1512 | 1243 | 18 | 19 | 1.2 | 1.5 | 1.4 | 0.2 |
| ORF1a | 13176 | 12218 | 9358 | 42 | 44 | 0.3 | 0.5 | 0.4 | 0.1 |
| ORF1b | 8111 | 4489 | 4713 | 18 | 21 | 0.4 | 0.4 | 0.4 | 0 |
| ORF3a | 827 | 1225 | 1002 | 7 | 7 | 0.6 | 0.7 | 0.6 | 0.1 |
| ORF6 | 185 | 230 | 230 | 2 | 2 | 0.9 | 0.9 | 0.9 | 0 |
| ORF7a | 365 | 823 | 678 | 2 | 2 | 0.2 | 0.3 | 0.3 | 0 |
| ORF7b | 131 | 242 | 169 | 2 | 2 | 0.8 | 1.2 | 1 | 0.3 |
| ORF8 | 365 | 584 | 507 | 8 | 8 | 1.4 | 1.6 | 1.5 | 0.1 |
| ORF10 | 116 | 130 | 121 | 0 | 0 | 0 | 0 | 0 | 0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).