Submitted:
12 September 2025
Posted:
15 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Molecular Detection and Sequencing
2.2. Archaeal Culture Methods
2.3. In Vitro Co-Culture and Microbial Interaction Studies
2.4. Animal Models and Gnotobiotic Systems
2.5. Bioinformatics and Systems Biology Approaches
3. Discussion
3.1. Evidence Linking Archaea to Disease
3.1.1. Inflammatory Bowel Disease (IBD)
3.1.2. Periodontal Disease
3.1.3. Obesity and Metabolic Disorders
3.1.4. Other Associations
3.2. Potential Mechanisms of Archaeal Involvement in Disease
3.2.1. Metabolic Cross-Feeding
3.2.2. Immune Modulation
3.2.3. Indirect Effects on Host Physiology
3.3. Barriers to Establishing Archaeal Pathogenicity
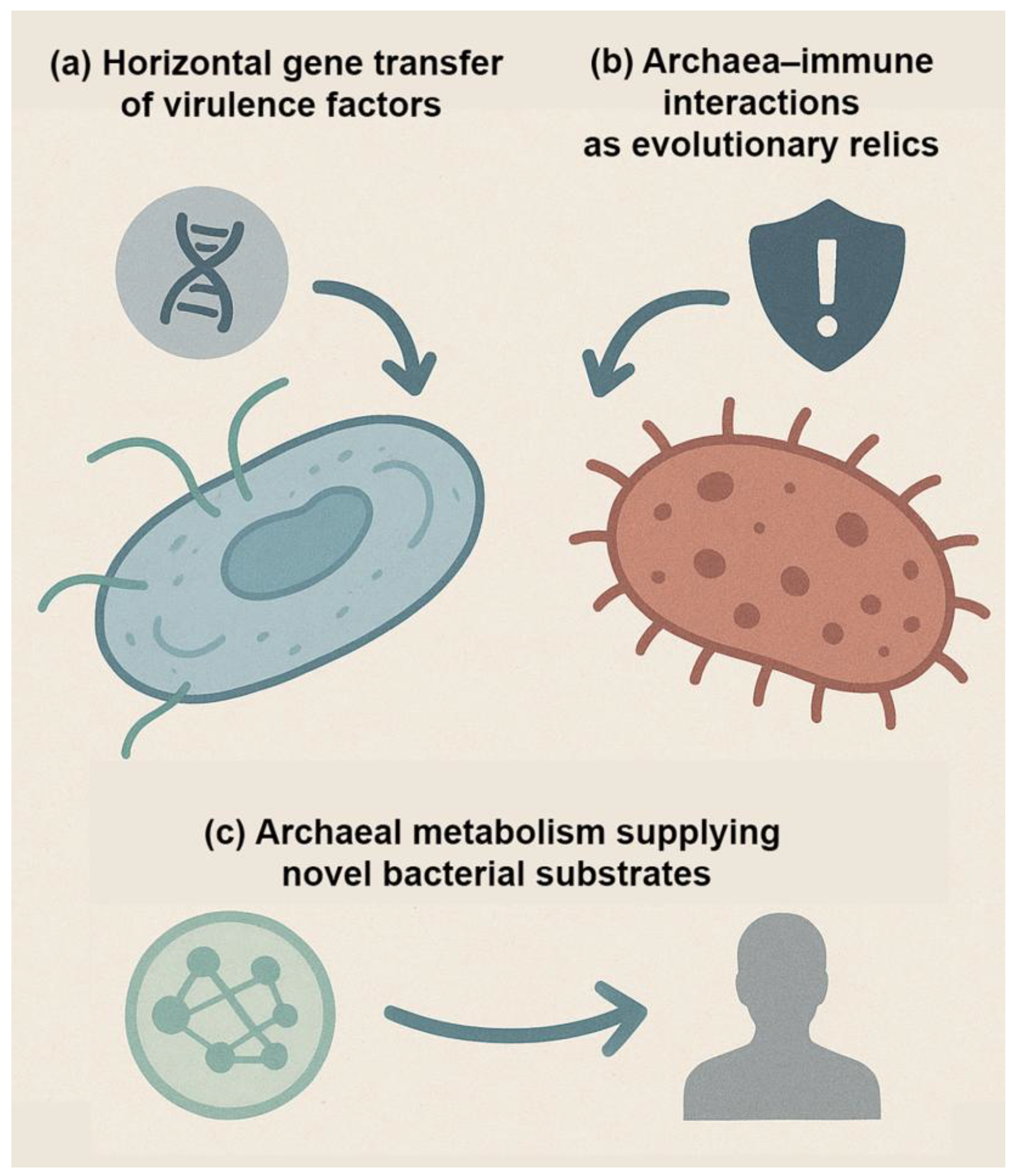
3.3.1. Lack of Virulence Factors
3.3.2. Methodological Challenges
3.3.3. Co-Occurrence with Bacteria
3.3.4. Limited Genetic Tools
3.4. Conceptual Implications
3.5. Evolutionary Perspectives
3.6. Ethical and Practical Considerations
4. Conclusions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| LGT | Lower gestational tract |
| UAT | Upper aerodigestive tract |
| IBS | Irritable bowel syndrome |
| IBD | Irritable bowel disease |
| SCFA | Short chain fatty acids |
| TNF-alpha | Tumor necrosis factor alpha |
| IL-6 | Interleukin 6 |
| LPS | lipopolysaccharide |
| PAMPS | Pathology associated molecular patterns |
| TLR-2,4 | Toll like receptor -2,4 |
| TMAO | Trimethylamine N-oxide |
References
- Woese, C.R. and G.E. Fox, Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc Natl Acad Sci U S A, 1977. 74(11): p. 5088-90.
- Pace, N.R., J. Sapp, and N. Goldenfeld, Phylogeny and beyond: Scientific, historical, and conceptual significance of the first tree of life. Proc Natl Acad Sci U S A, 2012. 109(4): p. 1011-8.
- Lurie-Weinberger, M.N. and U. Gophna, Archaea in and on the Human Body: Health Implications and Future Directions. PLoS Pathog, 2015. 11(6): p. e1004833.
- Kuehnast, T., et al., Exploring the human archaeome: its relevance for health and disease, and its complex interplay with the human immune system. FEBS J, 2025. 292(6): p. 1316-1329.
- Gaci, N., et al., Archaea and the human gut: new beginning of an old story. World J Gastroenterol, 2014. 20(43): p. 16062-78.
- Volmer, J.G., H. McRae, and M. Morrison, The evolving role of methanogenic archaea in mammalian microbiomes. Front Microbiol, 2023. 14: p. 1268451.
- Horz, H.P. and G. Conrads, Methanogenic Archaea and oral infections - ways to unravel the black box. J Oral Microbiol, 2011. 3.
- Pilliol, V., et al., Methanobrevibacter oralis: a comprehensive review. J Oral Microbiol, 2024. 16(1): p. 2415734.
- Umbach, A.K., A.A. Stegelmeier, and J.D. Neufeld, Archaea Are Rare and Uncommon Members of the Mammalian Skin Microbiome. mSystems, 2021. 6(4): p. e0064221.
- Bang, C. and R.A. Schmitz, Archaea associated with human surfaces: not to be underestimated. FEMS Microbiol Rev, 2015. 39(5): p. 631-48.
- Gill, E.E. and F.S. Brinkman, The proportional lack of archaeal pathogens: Do viruses/phages hold the key? Bioessays, 2011. 33(4): p. 248-54.
- Eckburg, P.B., P.W. Lepp, and D.A. Relman, Archaea and their potential role in human disease. Infect Immun, 2003. 71(2): p. 591-6.
- Hajishengallis, G. and R.J. Lamont, Polymicrobial communities in periodontal disease: Their quasi-organismal nature and dialogue with the host. Periodontol 2000, 2021. 86(1): p. 210-230.
- Cisek, A.A., et al., The Role of Methanogenic Archaea in Inflammatory Bowel Disease-A Review. J Pers Med, 2024. 14(2).
- Santana, P.T., et al., Dysbiosis in Inflammatory Bowel Disease: Pathogenic Role and Potential Therapeutic Targets. Int J Mol Sci, 2022. 23(7).
- Clarke, S.F., et al., The gut microbiota and its relationship to diet and obesity: new insights. Gut Microbes, 2012. 3(3): p. 186-202.
- Murphy, E.F., et al., Composition and energy harvesting capacity of the gut microbiota: relationship to diet, obesity and time in mouse models. Gut, 2010. 59(12): p. 1635-42.
- Kim, N., et al., Genome-resolved metagenomics: a game changer for microbiome medicine. Exp Mol Med, 2024. 56(7): p. 1501-1512.
- Clarridge, J.E., 3rd, Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin Microbiol Rev, 2004. 17(4): p. 840-62, table of contents.
- Koskinen, K., et al., First Insights into the Diverse Human Archaeome: Specific Detection of Archaea in the Gastrointestinal Tract, Lung, and Nose and on Skin. mBio, 2017. 8(6).
- Fernandez-Carro, E., et al., Alternatives Integrating Omics Approaches for the Advancement of Human Skin Models: A Focus on Metagenomics, Metatranscriptomics, and Metaproteomics. Microorganisms, 2025. 13(8).
- Hofer, U., The majority is uncultured. Nat Rev Microbiol, 2018. 16(12): p. 716-717.
- Samuel, B.S. and J.I. Gordon, A humanized gnotobiotic mouse model of host-archaeal-bacterial mutualism. Proc Natl Acad Sci U S A, 2006. 103(26): p. 10011-6.
- Samuel, B.S., et al., Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc Natl Acad Sci U S A, 2007. 104(25): p. 10643-8.
- Franzosa, E.A., et al., Sequencing and beyond: integrating molecular ‘omics’ for microbial community profiling. Nat Rev Microbiol, 2015. 13(6): p. 360-72.
- Sabih Ur Rehman, S., et al., Integrative systems biology approaches for analyzing microbiome dysbiosis and species interactions. Brief Bioinform, 2025. 26(4).
- Blais Lecours, P., et al., Increased prevalence of Methanosphaera stadtmanae in inflammatory bowel diseases. PLoS One, 2014. 9(2): p. e87734.
- Lepp, P.W., et al., Methanogenic Archaea and human periodontal disease. Proc Natl Acad Sci U S A, 2004. 101(16): p. 6176-81.
- Vianna, M.E., et al., Quantitative analysis of three hydrogenotrophic microbial groups, methanogenic archaea, sulfate-reducing bacteria, and acetogenic bacteria, within plaque biofilms associated with human periodontal disease. J Bacteriol, 2008. 190(10): p. 3779-85.
- Mbakwa, C.A., et al., Gut colonization with methanobrevibacter smithii is associated with childhood weight development. Obesity (Silver Spring), 2015. 23(12): p. 2508-16.
- Triantafyllou, K., C. Chang, and M. Pimentel, Methanogens, methane and gastrointestinal motility. J Neurogastroenterol Motil, 2014. 20(1): p. 31-40.
- Ghoshal, U., et al., Irritable Bowel Syndrome, Particularly the Constipation-Predominant Form, Involves an Increase in Methanobrevibacter smithii, Which Is Associated with Higher Methane Production. Gut Liver, 2016. 10(6): p. 932-938.
- Li, T., et al., Multi-Cohort Analysis Reveals Altered Archaea in Colorectal Cancer Fecal Samples Across Populations. Gastroenterology, 2025. 168(3): p. 525-538 e2.
- Mathlouthi, N.E.H., et al., The Archaeome’s Role in Colorectal Cancer: Unveiling the DPANN Group and Investigating Archaeal Functional Signatures. Microorganisms, 2023. 11(11).
- Malat, I., M. Drancourt, and G. Grine, Methanobrevibacter smithii cell variants in human physiology and pathology: A review. Heliyon, 2024. 10(18): p. e36742.
- Hassani, Y., et al., Detection of Methanobrevobacter smithii and Methanobrevibacter oralis in Lower Respiratory Tract Microbiota. Microorganisms, 2020. 8(12).
- Culp, E.J. and A.L. Goodman, Cross-feeding in the gut microbiome: Ecology and mechanisms. Cell Host Microbe, 2023. 31(4): p. 485-499.
- Goyal, A., et al., Ecology-guided prediction of cross-feeding interactions in the human gut microbiome. Nat Commun, 2021. 12(1): p. 1335.
- Houshyar, Y., et al., Going Beyond Bacteria: Uncovering the Role of Archaeome and Mycobiome in Inflammatory Bowel Disease. Front Physiol, 2021. 12: p. 783295.
- Vierbuchen, T., et al., The Human-Associated Archaeon Methanosphaera stadtmanae Is Recognized through Its RNA and Induces TLR8-Dependent NLRP3 Inflammasome Activation. Front Immunol, 2017. 8: p. 1535.
- Campbell, A., et al., H(2) generated by fermentation in the human gut microbiome influences metabolism and competitive fitness of gut butyrate producers. Microbiome, 2023. 11(1): p. 133.
- Duranti, S., et al., Obesity and microbiota: an example of an intricate relationship. Genes Nutr, 2017. 12: p. 18.
- Li, R., J. Li, and X. Zhou, Lung microbiome: new insights into the pathogenesis of respiratory diseases. Signal Transduct Target Ther, 2024. 9(1): p. 19.
- Kim, Y.B., et al., In-depth metataxonomic investigation reveals low richness, high intervariability, and diverse phylotype candidates of archaea in the human urogenital tract. Sci Rep, 2023. 13(1): p. 11746.
- Pimentel, M., et al., Methane, a gas produced by enteric bacteria, slows intestinal transit and augments small intestinal contractile activity. Am J Physiol Gastrointest Liver Physiol, 2006. 290(6): p. G1089-95.
- van de Pol, J.A., et al., Gut Colonization by Methanogenic Archaea Is Associated with Organic Dairy Consumption in Children. Front Microbiol, 2017. 8: p. 355.
- Radaic, A. and Y.L. Kapila, The oralome and its dysbiosis: New insights into oral microbiome-host interactions. Comput Struct Biotechnol J, 2021. 19: p. 1335-1360.
- Wang, P., et al., Gut microbiome-derived ammonia modulates stress vulnerability in the host. Nat Metab, 2023. 5(11): p. 1986-2001.
- Matchado, M.S., et al., Network analysis methods for studying microbial communities: A mini review. Comput Struct Biotechnol J, 2021. 19: p. 2687-2698.
- Liu, Z., et al., Network analyses in microbiome based on high-throughput multi-omics data. Brief Bioinform, 2021. 22(2): p. 1639-1655.
- Abram, F., Systems-based approaches to unravel multi-species microbial community functioning. Comput Struct Biotechnol J, 2015. 13: p. 24-32.
- Papageorgiou, A.C. and P.S. Adam, Archaea appropriate weaponry to breach the bacterial cell wall. PLoS Biol, 2025. 23(8): p. e3003323.
- Takiishi, T., C.I.M. Fenero, and N.O.S. Camara, Intestinal barrier and gut microbiota: Shaping our immune responses throughout life. Tissue Barriers, 2017. 5(4): p. e1373208.
- van Wolferen, M., et al., The cell biology of archaea. Nat Microbiol, 2022. 7(11): p. 1744-1755.
- Kerr, M., et al., Beyond the Gut: Unveiling Methane’s Role in Broader Physiological Systems. FASEB Bioadv, 2025. 7(8): p. e70048.
- Brugere, J.F., et al., Archaebiotics: proposed therapeutic use of archaea to prevent trimethylaminuria and cardiovascular disease. Gut Microbes, 2014. 5(1): p. 5-10.
- Jung, M.Y., et al., Ammonia-oxidizing archaea possess a wide range of cellular ammonia affinities. ISME J, 2022. 16(1): p. 272-283.
- Green, E.R. and J. Mecsas, Bacterial Secretion Systems: An Overview. Microbiol Spectr, 2016. 4(1).
- Duller, S. and C. Moissl-Eichinger, Archaea in the Human Microbiome and Potential Effects on Human Infectious Disease. Emerg Infect Dis, 2024. 30(8): p. 1505-13.
- Karavaeva, V. and F.L. Sousa, Navigating the archaeal frontier: insights and projections from bioinformatic pipelines. Front Microbiol, 2024. 15: p. 1433224.
- Guerra, A., Human associated Archaea: a neglected microbiome worth investigating. World J Microbiol Biotechnol, 2024. 40(2): p. 60.
- Rafiq, M., et al., Challenges and Approaches of Culturing the Unculturable Archaea. Biology (Basel), 2023. 12(12).
- Kioukis, A., et al., Global Archaeal Diversity Revealed Through Massive Data Integration: Uncovering Just Tip of Iceberg. Microorganisms, 2025. 13(3).
- Atomi, H., T. Imanaka, and T. Fukui, Overview of the genetic tools in the Archaea. Front Microbiol, 2012. 3: p. 337.
- Eren, A.M. and J.F. Banfield, Modern microbiology: Embracing complexity through integration across scales. Cell, 2024. 187(19): p. 5151-5170.
- Bahram, M., et al., Newly designed 16S rRNA metabarcoding primers amplify diverse and novel archaeal taxa from the environment. Environ Microbiol Rep, 2019. 11(4): p. 487-494.
- Chibani, C.M., et al., A catalogue of 1,167 genomes from the human gut archaeome. Nat Microbiol, 2022. 7(1): p. 48-61.
- Kapoore, R.V., et al., Co-culturing microbial consortia: approaches for applications in biomanufacturing and bioprocessing. Crit Rev Biotechnol, 2022. 42(1): p. 46-72.
- Pohlschroder, M., et al., Haloferax volcanii: a versatile model for studying archaeal biology. J Bacteriol, 2025. 207(6): p. e0006225.
- Bai, D., J. Peng, and C. Yi, Advances in single-cell multi-omics profiling. RSC Chem Biol, 2021. 2(2): p. 441-449.
- Lim, J., et al., Advances in single-cell omics and multiomics for high-resolution molecular profiling. Exp Mol Med, 2024. 56(3): p. 515-526.
- Gonzalez, A., A. Fullaondo, and A. Odriozola, Why Are Long-Read Sequencing Methods Revolutionizing Microbiome Analysis? Microorganisms, 2025. 13(8).
- Wen, L. and F. Tang, Single-cell omics sequencing technologies: the long-read generation. Trends Genet, 2025.
- Tronel, A., et al., Untargeted and semi-targeted metabolomics approach for profiling small intestinal and fecal metabolome using high-resolution mass spectrometry. Metabolomics, 2025. 21(4): p. 84.
- Buan, N.R., Methanogens: pushing the boundaries of biology. Emerg Top Life Sci, 2018. 2(4): p. 629-646.
- Aggarwal, N., et al., Microbiome and Human Health: Current Understanding, Engineering, and Enabling Technologies. Chem Rev, 2023. 123(1): p. 31-72.
- Vento, J.M., N. Crook, and C.L. Beisel, Barriers to genome editing with CRISPR in bacteria. J Ind Microbiol Biotechnol, 2019. 46(9-10): p. 1327-1341.
- De Lise, F., et al., Archaea as a Model System for Molecular Biology and Biotechnology. Biomolecules, 2023. 13(1).
- Sleytr, U.B., et al., S-layers: principles and applications. FEMS Microbiol Rev, 2014. 38(5): p. 823-64.
- Cowan, D.A., et al., Extremophiles in a changing world. Extremophiles, 2024. 28(2): p. 26.
- Zhu, P., et al., CRISPR/Cas12a toolbox for genome editing in Methanosarcina acetivorans. Front Microbiol, 2023. 14: p. 1235616.
- Wani, A.K., et al., Discovering untapped microbial communities through metagenomics for microplastic remediation: recent advances, challenges, and way forward. Environ Sci Pollut Res Int, 2023. 30(34): p. 81450-81473.
- Ojala, T., et al., Current concepts, advances, and challenges in deciphering the human microbiota with metatranscriptomics. Trends Genet, 2023. 39(9): p. 686-702.
- Wolf, M., et al., Advances in the clinical use of metaproteomics. Expert Rev Proteomics, 2023. 20(4-6): p. 71-86.
- Ruden, D.M., TIGR-Tas and the Expanding Universe of RNA-Guided Genome Editing Systems: A New Era Beyond CRISPR-Cas. Genes (Basel), 2025. 16(8).
- Pfeifer, E. and E.P.C. Rocha, Phage-plasmids promote recombination and emergence of phages and plasmids. Nat Commun, 2024. 15(1): p. 1545.
- Yin, Q., et al., Potential interactions between syntrophic bacteria and methanogens via type IV pili and quorum-sensing systems. Environ Int, 2020. 138: p. 105650.
- Ghaly, T.M., et al., Discovery of integrons in Archaea: Platforms for cross-domain gene transfer. Sci Adv, 2022. 8(46): p. eabq6376.
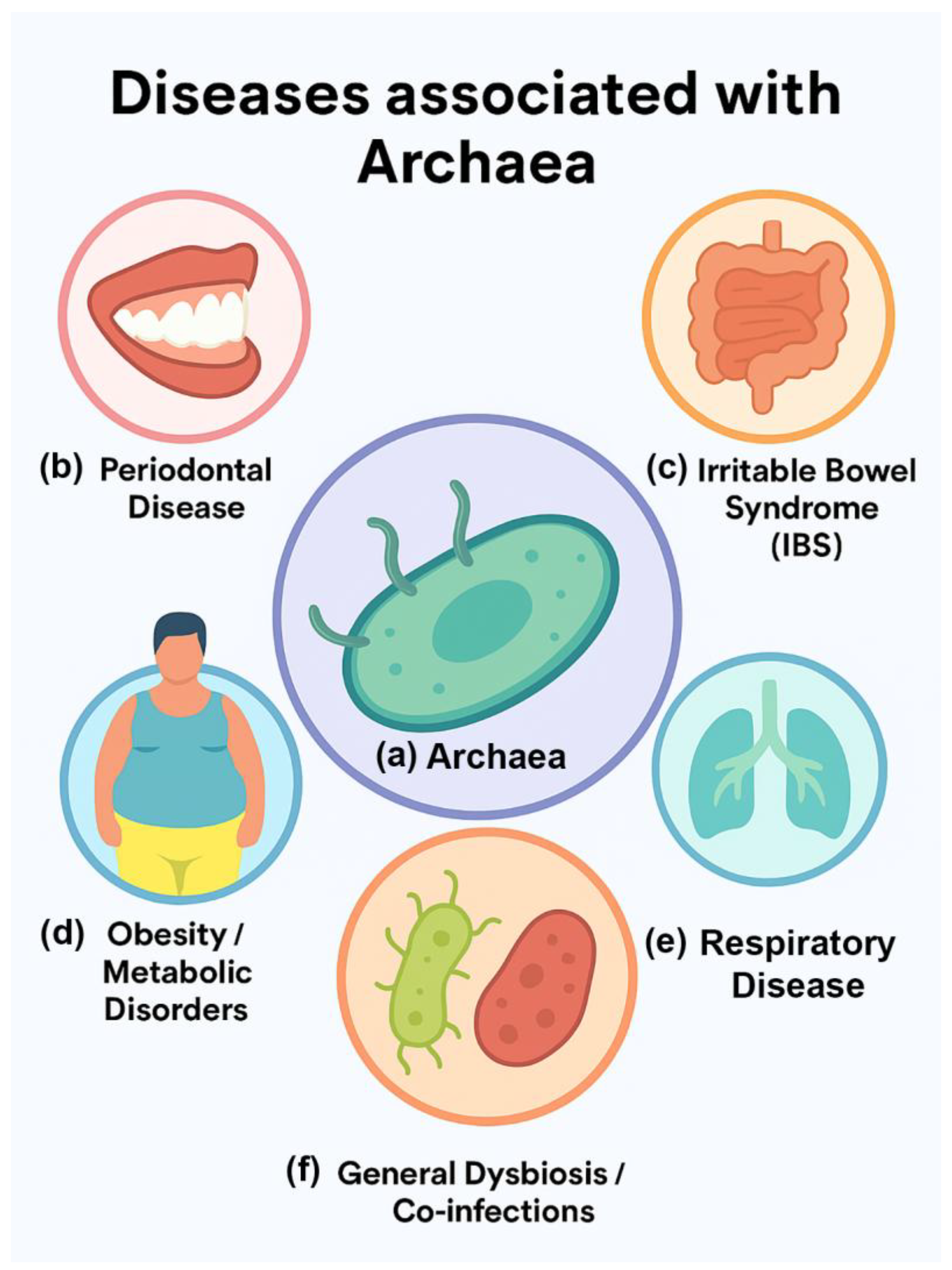
| Disease/Condition | Archaeal Taxa Involved | Proposed Role/Mechanism | Evidence Strength | References (PMIDs) |
|---|---|---|---|---|
| Inflammatory Bowel Disease (IBD) | Methanosphaera stadtmanae, Methanobrevibacter smithii | Induces pro-inflammatory responses; increased abundance in some IBD patients | Moderate | [14,27] |
| Periodontal Disease | Methanobrevibacter oralis | Hydrogen consumption supports anaerobic bacterial growth in oral biofilms | Strong association, unclear causality | [8,27,28,29] |
| Obesity / Metabolic Disorders | Methanobrevibacter smithii | Enhances bacterial fermentation efficiency; promotes increased host energy harvest | Moderate (mouse models, variable human data) | [16,17,30] |
| Irritable Bowel Syndrome (IBS) | Methanogens (general) | Methane linked to slowed intestinal motility; may exacerbate constipation-type IBS | Moderate | [31,32] |
| Colorectal Cancer | Methanobrevibacter smithii, other methanogens | Detected at higher abundance in some tumor microbiome studies; role unclear | Weak/Correlative | [33,34] |
| Respiratory Disease (e.g., cystic fibrosis lungs) | Methanobrevibacter spp. | Found alongside bacterial pathogens in lung samples; ecological rather than causal role | Weak/Incidental | [35,36] |
| General Dysbiosis/Co-infections | Multiple methanogens | Synergistic effects with bacterial pathogens via metabolic cross-feeding | Strong concept, limited direct proof | [37,38,39] |
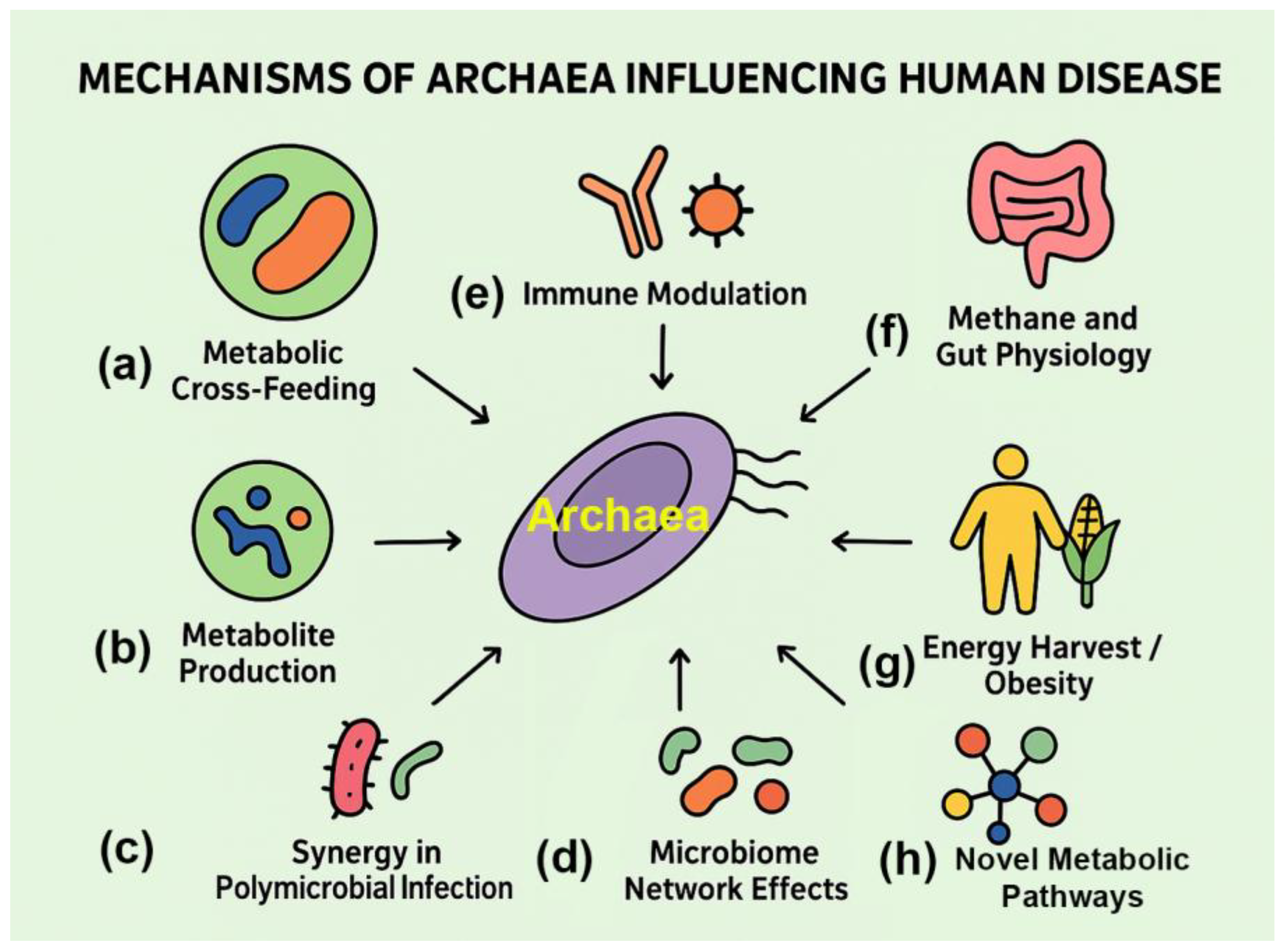
| Mechanism | Archaeal Taxa Involved | Description | Evidence Type | Key References (PMIDs) |
|---|---|---|---|---|
| Metabolic Cross-Feeding | Methanobrevibacter smithii, Methanobrevibacter oralis | Consumption of hydrogen by methanogens enhances bacterial fermentation, supporting pathogen growth | Co-culture, microbiome studies | [3,7,8] |
| Immune Modulation | Methanosphaera stadtmanae | Activation of dendritic cells and cytokine production; unique archaeal cell wall components may act as immune triggers | In vitro immunology | [4,40] |
| Methane and Gut Physiology | General methanogens | Methane slows intestinal motility, potentially contributing to constipation and IBS | Breath methane studies, animal models | [31,45] |
| Energy Harvest / Obesity | Methanobrevibacter smithii | Hydrogen removal improves bacterial breakdown of polysaccharides, increasing caloric extraction | Gnotobiotic mouse models, human studies | [16,17,30,42,46] |
| Synergy in Polymicrobial Infection | Methanobrevibacter oralis, others | Presence of archaea creates favorable niches for anaerobic pathogens (e.g., periodontal bacteria) | Oral biofilm studies | [7,8,47] |
| Metabolite Production | Multiple methanogens | Methane and ammonia production may alter mucosal environment and host tissue function | Metabolomics, breath tests | [9,48] |
| Microbiome Network Effects | Multiple lineages | Archaea may act as keystone species influencing bacterial community composition and stability | Systems biology, network modeling | [49,50,51] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




