Submitted:
14 August 2025
Posted:
19 August 2025
Read the latest preprint version here
Abstract

Keywords:
Introduction
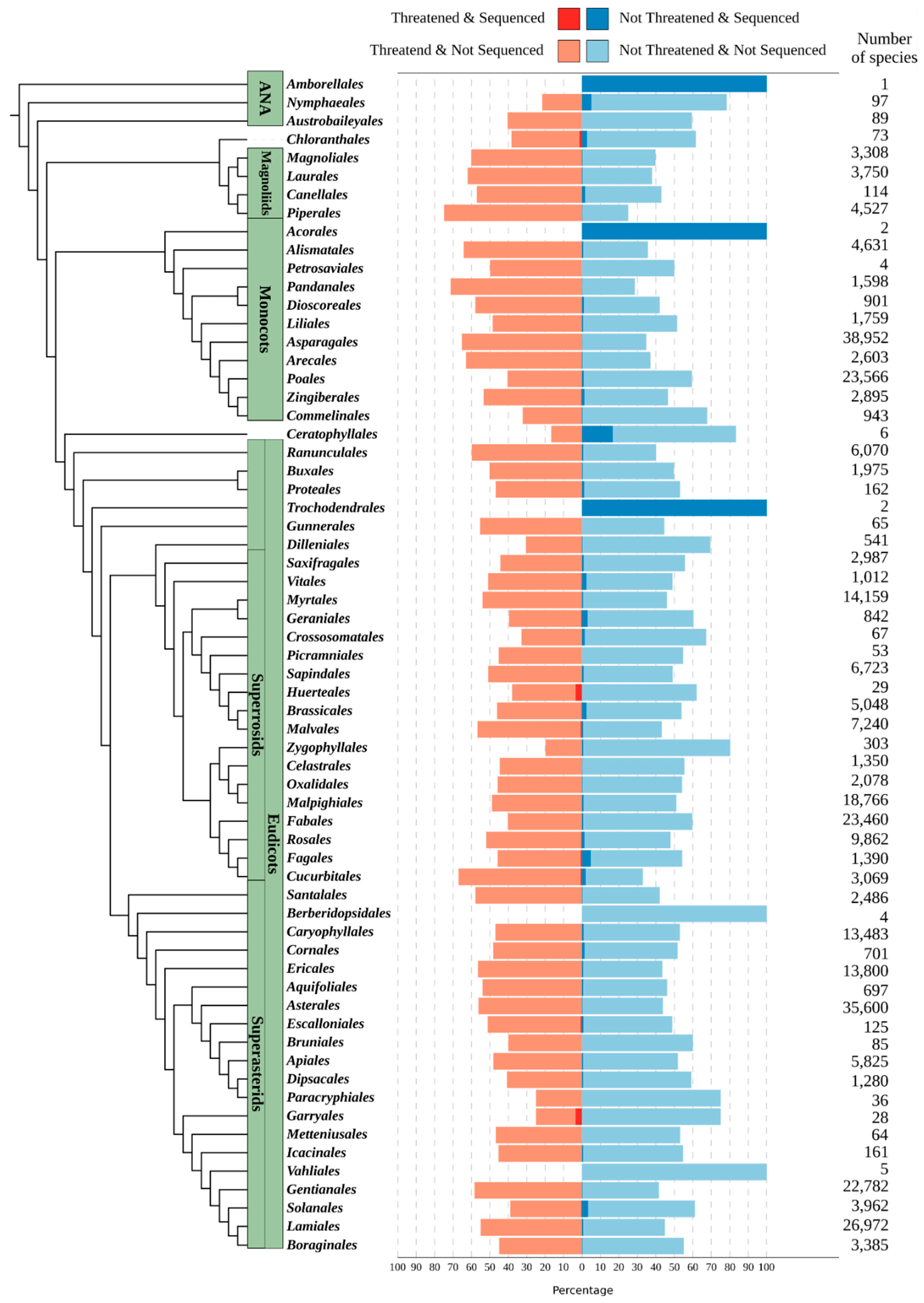

Step 1: Preparations for a Plant Genomics Project
Importance of Botanical Gardens as Source of Material for Sequencing Projects
Evaluating Existing Genomic Data
Resource Estimation Utilising the Plant DNA C-Values Database
Step 2: DNA Extraction Protocols
Step 3: Library Preparation and Sequencing
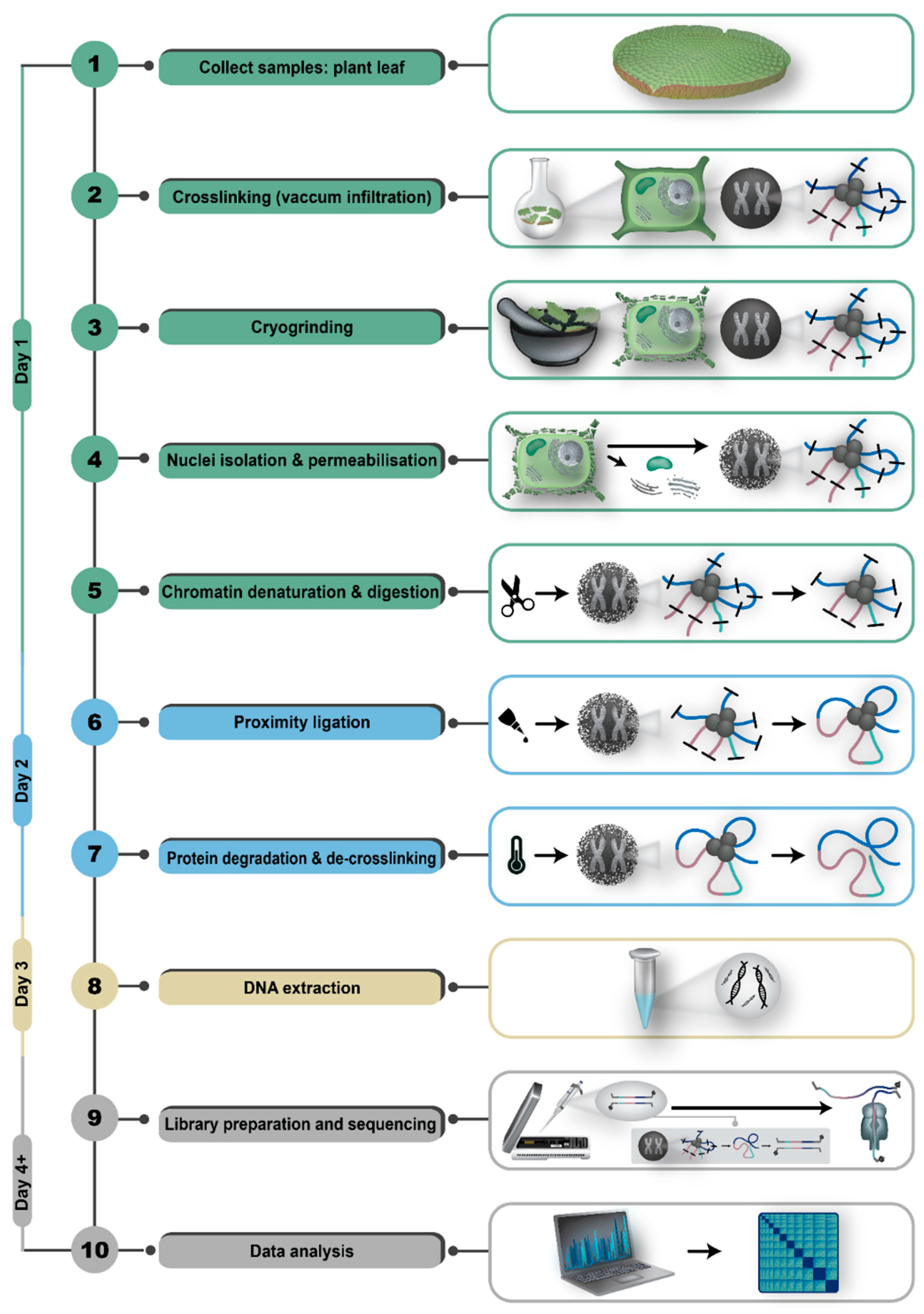
Step 4: Pore-C
Step 5: Basecalling
Step 6: Read Correction
Step 7: Genome Sequence Assembly
Step 8: Assembly Evaluation
Step 9: Scaffolding with Pore-C Data
Step 10: Structural Annotation
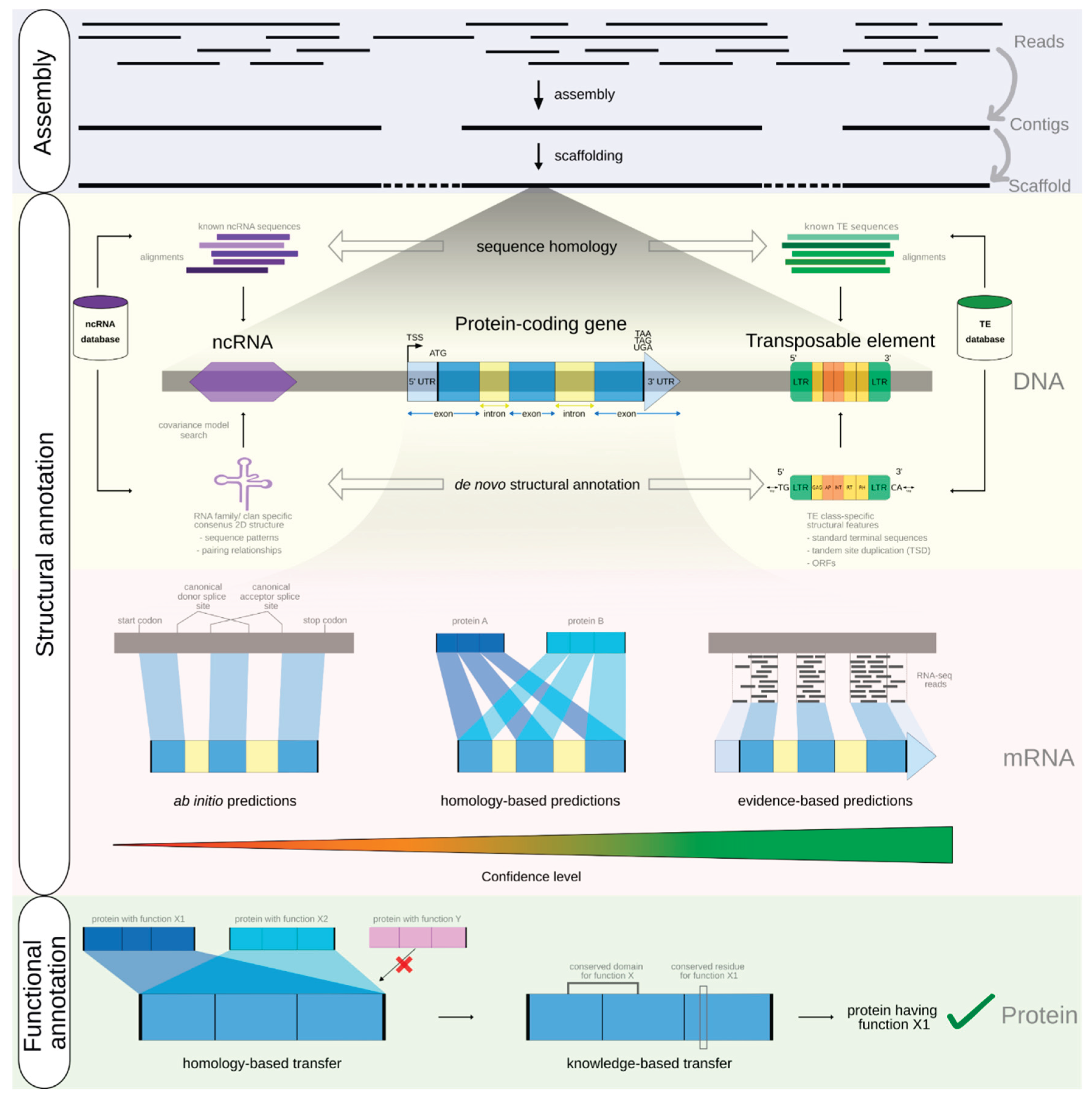
Step 11: Functional Annotation
Step 12: Data Submission
Summary
Supplementary Materials
Data Availability Statement
Additional information
Author Contributions
Acknowledgments
References
- The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature. 2000;408:796.
- Michael TP, Jupe F, Bemm F, Motley ST, Sandoval JP, Lanz C, et al. High contiguity Arabidopsis thaliana genome assembly with a single nanopore flow cell. Nat Commun. 2018;9:541. [CrossRef]
- Pucker B, Kleinbölting N, Weisshaar B. Large scale genomic rearrangements in selected Arabidopsis thaliana T-DNA lines are caused by T-DNA insertion mutagenesis. BMC Genomics. 2021;22:599.
- Pucker B, Irisarri I, Vries J de, Xu B. Plant genome sequence assembly in the era of long reads: Progress, challenges and future directions. Quant Plant Biol. 2022;3:e5. [CrossRef]
- Marks RA, Hotaling S, Frandsen PB, VanBuren R. Representation and participation across 20 years of plant genome sequencing. Nat Plants. 2021;7:1571. [CrossRef]
- Schwacke R, Bolger ME, Usadel B. PubPlant -a continuously updated online resource for sequenced and published plant genomes. Front Plant Sci. 2025;16. [CrossRef]
- Saha D, Panda AK, Datta S. Critical considerations and computational tools in plant genome editing. Heliyon. 2025;11:e41135. [CrossRef]
- Hyun JC, Monk JM, Palsson BO. Comparative pangenomics: analysis of 12 microbial pathogen pangenomes reveals conserved global structures of genetic and functional diversity. BMC Genomics. 2022;23:7. [CrossRef]
- Liao WW, Asri M, Ebler J, Doerr D, Haukness M, Hickey G, et al. A draft human pangenome reference. Nature. 2023;617:312. [CrossRef]
- Meng Q, Xie P, Xu Z, Tang J, Hui L, Gu J, et al. Pangenome analysis reveals yield- and fiber-related diversity and interspecific gene flow in Gossypium barbadense L. Nat Commun. 2025;16:4995. [CrossRef]
- GBIF Secretariat. GBIF Backbone Taxonomy. 2023;10.15468/39omei. [CrossRef]
- Bachman SP, Brown MJM, Leão TCC, Nic Lughadha E, Walker BE. Extinction risk predictions for the world’s flowering plants to support their conservation. New Phytol. 2024;242:797. [CrossRef]
- The Angiosperm Phylogeny Group, Chase MW, Christenhusz MJM, Fay MF, Byng JW, Judd WS, et al. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot J Linn Soc. 2016;181:1.
- Janssens SB, Couvreur TLP, Mertens A, Dauby G, Dagallier LPMJ, Abeele SV, et al. A large-scale species level dated angiosperm phylogeny for evolutionary and ecological analyses. Biodivers Data J. 2020;8:e39677. [CrossRef]
- McTavish EJ, Hinchliff CE, Allman JF, Brown JW, Cranston KA, Holder MT, et al. Phylesystem: a git-based data store for community-curated phylogenetic estimates. Bioinformatics. 2015;31:2794. [CrossRef]
- Chen G, Sun W. The role of botanical gardens in scientific research, conservation, and citizen science. Plant Divers. 2018;40:181. [CrossRef]
- Aleza P, Juárez J, Hernández M, Pina JA, Ollitrault P, Navarro L. Recovery and characterization of a Citrus clementinaHort. ex Tan. ‘Clemenules’ haploid plant selected to establish the reference whole Citrus genome sequence. BMC Plant Biol. 2009;9:110.
- Hirsch CN, Hirsch CD, Brohammer AB, Bowman MJ, Soifer I, Barad O, et al. Draft Assembly of Elite Inbred Line PH207 Provides Insights into Genomic and Transcriptome Diversity in Maize. Plant Cell. 2016;28:2700. [CrossRef]
- Schwartz JC, Farrell CP, Freimanis G, Sewell AK, Phillips JD, Hammond JA. A genome assembly and transcriptome atlas of the inbred Babraham pig to illuminate porcine immunogenetic variation. Immunogenetics. 2024;76:361. [CrossRef]
- Wang B, Jiao Y, Chougule K, Olson A, Huang J, Llaca V, et al. Pan-genome Analysis in Sorghum Highlights the Extent of Genomic Variation and Sugarcane Aphid Resistance Genes. bioRxiv; 2021. p. 2021.01.03.424980.
- Yang N, Liu J, Gao Q, Gui S, Chen L, Yang L, et al. Genome assembly of a tropical maize inbred line provides insights into structural variation and crop improvement. Nat Genet. 2019;51:1052. [CrossRef]
- Ma H, Liu Y, Liu D, Sun W, Liu X, Wan Y, et al. Chromosome-level genome assembly and population genetic analysis of a critically endangered rhododendron provide insights into its conservation. Plant J. 2021;107:1533. [CrossRef]
- Pavese V, Cavalet-Giorsa E, Barchi L, Acquadro A, Torello Marinoni D, Portis E, et al. Whole-genome assembly of Corylus avellana cv “Tonda Gentile delle Langhe” using linked-reads (10X Genomics). G3 GenesGenomesGenetics. 2021;11:jkab152.
- Ben Romdhane W, Ben Saad R, Guiderdoni E, Ali AA mohamed, Tarroum M, Al-Doss A, et al. De novo, high-quality assembly and annotation of the halophyte grass Aeluropus littoralis draft genome and identification of A20/AN1 zinc finger protein family. BMC Plant Biol. 2025;25:556. [CrossRef]
- Liu H, Wei J, Yang T, Mu W, Song B, Yang T, et al. Molecular digitization of a botanical garden: high-depth whole-genome sequencing of 689 vascular plant species from the Ruili Botanical Garden. GigaScience. 2019;8:giz007. [CrossRef]
- Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011;12:323. [CrossRef]
- Li Z, Zhang Z, Yan P, Huang S, Fei Z, Lin K. RNA-Seq improves annotation of protein-coding genes in the cucumber genome. BMC Genomics. 2011;12:540. [CrossRef]
- Arita M, Karsch-Mizrachi I, Cochrane G, on behalf of the International Nucleotide Sequence Database Collaboration. The international nucleotide sequence database collaboration. Nucleic Acids Res. 2021;49:D121.
- Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 2012;40:D1178. [CrossRef]
- CNCB-NGDC Members and Partners. Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2025. Nucleic Acids Res. 2025;53:D30.
- Wani ZA, Bhat A. Figshare: A One-Stop Shop for Research Data Management with Diverse Features and Services. J Inf Knowl. 2022;59:391. [CrossRef]
- Vision T. The Dryad Digital Repository: Published evolutionary data as part of the greater data ecosystem. Nat Preced. 2010;1.
- Arend D, Lange M, Chen J, Colmsee C, Flemming S, Hecht D, et al. e!DAL - a framework to store, share and publish research data. BMC Bioinformatics. 2014;15:214. [CrossRef]
- Arend D, Junker A, Scholz U, Schüler D, Wylie J, Lange M. PGP repository: a plant phenomics and genomics data publication infrastructure. Database. 2016;2016:baw033. [CrossRef]
- Pellicer J, Leitch IJ. The Plant DNA C-values database (release 7.1): an updated online repository of plant genome size data for comparative studies. New Phytol. 2020;226:301. [CrossRef]
- Dohm JC, Minoche AE, Holtgräwe D, Capella-Gutiérrez S, Zakrzewski F, Tafer H, et al. The genome of the recently domesticated crop plant sugar beet (Beta vulgaris). Nature. 2014;505:546. [CrossRef]
- Shi X, Cao S, Wang X, Huang S, Wang Y, Liu Z, et al. The complete reference genome for grapevine (Vitis vinifera L.) genetics and breeding. Hortic Res. 2023;10:uhad061.
- Heslop-Harrison JS (Pat), Schwarzacher T, Liu Q. Polyploidy: its consequences and enabling role in plant diversification and evolution. Ann Bot. 2023;131:1. [CrossRef]
- Duan H, Jones AW, Hewitt T, Mackenzie A, Hu Y, Sharp A, et al. Physical separation of haplotypes in dikaryons allows benchmarking of phasing accuracy in Nanopore and HiFi assemblies with Hi-C data. Genome Biol. 2022;23:84. [CrossRef]
- Sarashetti P, Lipovac J, Tomas F, Šikić M, Liu J. Evaluating data requirements for high-quality haplotype-resolved genomes for creating robust pangenome references. Genome Biol. 2024;25:312. [CrossRef]
- Pellicer J, Kelly LJ, Magdalena C, Leitch IJ. Insights into the dynamics of genome size and chromosome evolution in the early diverging angiosperm lineage Nymphaeales (water lilies). Genome. 2013;56:437. [CrossRef]
- Espinosa E, Bautista R, Larrosa R, Plata O. Advancements in long-read genome sequencing technologies and algorithms. Genomics. 2024;116:110842. [CrossRef]
- Russo A, Mayjonade B, Frei D, Potente G, Kellenberger RT, Frachon L, et al. Low-Input High-Molecular-Weight DNA Extraction for Long-Read Sequencing From Plants of Diverse Families. Front Plant Sci. 2022;13. [CrossRef]
- Friar EA. Isolation of DNA from Plants with Large Amounts of Secondary Metabolites. In: Methods in Enzymology. Academic Press; 2005. p. 1. (Molecular Evolution: Producing the Biochemical Data; vol. 395).
- Rosso MG, Li Y, Strizhov N, Reiss B, Dekker K, Weisshaar B. An Arabidopsis thaliana T-DNA mutagenized population (GABI-Kat) for flanking sequence tag-based reverse genetics. Plant Mol Biol. 2003;53:247. [CrossRef]
- Doyle JJ, Doyle JL. Isolation of plant DNA from fresh tissue. Focus. 1990;12:13–5.
- Siadjeu C, Pucker B, Viehöver P, Albach DC, Weisshaar B. High Contiguity de novo Genome Sequence Assembly of Trifoliate Yam (Dioscorea dumetorum) Using Long Read Sequencing. Genes. 2020;11:274. [CrossRef]
- Recinos MFM, Winnier S, Lagerhausen K, Ajayi B, Wolff K, Friedhoff R, et al. Cacao genome sequence reveals insights into the flavonoid biosynthesis. bioRxiv; 2024. p. 2024.11.23.624982.
- Pucker B. Plant DNA extraction and preparation for ONT sequencing. 2020;
- Zhang Y, Zhang Y, Burke JM, Gleitsman K, Friedrich SM, Liu KJ, et al. A Simple Thermoplastic Substrate Containing Hierarchical Silica Lamellae for High Molecular Weight DNA Extraction. Adv Mater Deerfield Beach Fla. 2016;28:10630. [CrossRef]
- Butto T, Mungikar K, Baumann P, Winter J, Lutz B, Gerber S. Nuclei on the Rise: When Nuclei-Based Methods Meet Next-Generation Sequencing. Cells. 2023;12:1051. [CrossRef]
- Kang M, Chanderbali A, Lee S, Soltis DE, Soltis PS, Kim S. High-molecular-weight DNA extraction for long-read sequencing of plant genomes: An optimization of standard methods. Appl Plant Sci. 2023;11:e11528. [CrossRef]
- Ling G, Waxman DJ. Isolation of Nuclei for use in Genome-wide DNase Hypersensitivity Assays to Probe Chromatin Structure. Methods Mol Biol Clifton NJ. 2013;977:13.
- Zerpa-Catanho D, Zhang X, Song J, Hernandez AG, Ming R. Ultra-long DNA molecule isolation from plant nuclei for ultra-long read genome sequencing. STAR Protoc. 2021;2:100343. [CrossRef]
- Nowak MS, Pucker B. Pore-C Protocol for Plant Samples. 2025;
- Zhang M, Zhang Y, Scheuring CF, Wu CC, Dong JJ, Zhang HB. Preparation of megabase-sized DNA from a variety of organisms using the nuclei method for advanced genomics research. Nat Protoc. 2012;7:467. [CrossRef]
- Workman R, Fedak R, Kilburn D, Hao S, Liu K, Timp W. High Molecular Weight DNA Extraction from Recalcitrant Plant Species for Third Generation Sequencing. 2019;
- Li Z, Parris S, Saski CA. A simple plant high-molecular-weight DNA extraction method suitable for single-molecule technologies. Plant Methods. 2020;16:38. [CrossRef]
- Mayjonade B, Gouzy ,Jérôme, Donnadieu ,Cécile, Pouilly ,Nicolas, Marande ,William, Callot ,Caroline, et al. Extraction of High-Molecular-Weight Genomic DNA for Long-Read Sequencing of Single Molecules. BioTechniques. 2016;61:203. [CrossRef]
- Jones A, Torkel C, Stanley D, Nasim J, Borevitz J, Schwessinger B. Scalable high-molecular weight DNA extraction for long-read sequencing. 2020;
- Sauvage T, Cormier A, Delphine P. A comparison of Oxford nanopore library strategies for bacterial genomics. BMC Genomics. 2023;24:627. [CrossRef]
- Jain M, Koren S, Miga KH, Quick J, Rand AC, Sasani TA, et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat Biotechnol. 2018;36:338. [CrossRef]
- Dekker J, Rippe K, Dekker M, Kleckner N. Capturing Chromosome Conformation. Science. 2002;295:1306. [CrossRef]
- Deshpande AS, Ulahannan N, Pendleton M, Dai X, Ly L, Behr JM, et al. Identifying synergistic high-order 3D chromatin conformations from genome-scale nanopore concatemer sequencing. Nat Biotechnol. 2022;40:1488. [CrossRef]
- Sati S, Cavalli G. Chromosome conformation capture technologies and their impact in understanding genome function. Chromosoma. 2017;126:33. [CrossRef]
- Hoffman EA, Frey BL, Smith LM, Auble DT. Formaldehyde Crosslinking: A Tool for the Study of Chromatin Complexes. J Biol Chem. 2015;290:26404. [CrossRef]
- Sikorskaite S, Rajamäki ML, Baniulis D, Stanys V, Valkonen JP. Protocol: Optimised methodology for isolation of nuclei from leaves of species in the Solanaceae and Rosaceae families. Plant Methods. 2013;9:31. [CrossRef]
- Belton JM, McCord RP, Gibcus JH, Naumova N, Zhan Y, Dekker J. Hi–C: A comprehensive technique to capture the conformation of genomes. Methods. 2012;58:268. [CrossRef]
- Weber K, Kuter DJ. Reversible Denaturation of Enzymes by Sodium Dodecyl Sulfate. J Biol Chem. 1971;246:4504. [CrossRef]
- Zhong JY, Niu L, Lin ZB, Bai X, Chen Y, Luo F, et al. High-throughput Pore-C reveals the single-allele topology and cell type-specificity of 3D genome folding. Nat Commun. 2023;14:1250. [CrossRef]
- Schalamun M, Schwessinger B. DNA size selection (>1kb) and clean up using an optimized SPRI beads mixture. 2017;
- Schalamun M, Nagar R, Kainer D, Beavan E, Eccles D, Rathjen JP, et al. Harnessing the MinION: An example of how to establish long-read sequencing in a laboratory using challenging plant tissue from Eucalyptus pauciflora. Mol Ecol Resour. 2019;19:77. [CrossRef]
- DeAngelis MM, Wang DG, Hawkins TL. Solid-phase reversible immobilization for the isolation of PCR products. Nucleic Acids Res. 1995;23:4742. [CrossRef]
- Lis JT, Schleif R. Size fractionation of double-stranded DNA by precipitation with polyethylene glycol. Nucleic Acids Res. 1975;2:383. [CrossRef]
- He Z, Zhu Y, Gu H. A new method for the determination of critical polyethylene glycol concentration for selective precipitation of DNA fragments. Appl Microbiol Biotechnol. 2013;97:9175. [CrossRef]
- Oxford Nanopore Technologies [Internet]. 2019 [cited 2025 July 23]. Restriction enzyme Pore-C info sheet. Available from: https://nanoporetech.com/document/restriction-enzyme-pore-c.
- Oxford Nanopore Technologies [Internet]. 2022 [cited 2025 July 23]. Ligation sequencing DNA V14 (SQK-LSK114). Available from: https://nanoporetech.com/document/genomic-dna-by-ligation-sqk-lsk114.
- Pagès-Gallego M, de Ridder J. Comprehensive benchmark and architectural analysis of deep learning models for nanopore sequencing basecalling. Genome Biol. 2023;24:71.
- Dorado Documentation [Internet]. [cited 2025 July 23]. Available from: https://dorado-docs.readthedocs.io/en/latest/.
- Dorado [Internet]. Oxford Nanopore Technologies; [cited 2025 July 23]. Available from: https://github.com/nanoporetech/dorado.
- Koren S, Bao Z, Guarracino A, Ou S, Goodwin S, Jenike KM, et al. Gapless assembly of complete human and plant chromosomes using only nanopore sequencing. Genome Res. 2024;34:1919. [CrossRef]
- Nowak MS, Harder B, Meckoni SN, Friedhoff R, Wolff K, Pucker B. Genome sequence and RNA-seq analysis reveal genetic basis of flower coloration in the giant water lily Victoria cruziana. bioRxiv; 2024. p. 2024.06.15.599162.
- Krawczyk K, Szablińska-Piernik J, Paukszto Ł, Maździarz M, Sulima P, Przyborowski JA, et al. Chromosome-scale telomere to telomere genome assembly of common crystalwort (Riccia sorocarpa Bisch.). Sci Data. 2025;12:77. [CrossRef]
- ELIXIR [Internet]. [cited 2025 July 23]. Available from: https://elixir-europe.org/.
- CyVerse [Internet]. [cited 2025 July 23]. Available from: https://cyverse.org/.
- Koren S, Walenz BP, Berlin K, Miller JR, Bergman NH, Phillippy AM. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017;27:722. [CrossRef]
- Hu J, Wang Z, Sun Z, Hu B, Ayoola AO, Liang F, et al. NextDenovo: an efficient error correction and accurate assembly tool for noisy long reads. Genome Biol. 2024;25:107. [CrossRef]
- Nurk S, Walenz BP, Rhie A, Vollger MR, Logsdon GA, Grothe R, et al. HiCanu: accurate assembly of segmental duplications, satellites, and allelic variants from high-fidelity long reads. Genome Res. 2020;30:1291. [CrossRef]
- HERRO [Internet]. Šikić lab; [cited 2025 July 23]. Available from: https://github.com/lbcb-sci/herro.
- Stanojevic D, Lin D, Nurk S, Florez De Sessions P, Sikic M. Telomere-to-Telomere Phased Genome Assembly Using HERRO-Corrected Simplex Nanopore Reads. bioRxiv; 2024.
- Shafin K, Pesout T, Lorig-Roach R, Haukness M, Olsen HE, Bosworth C, et al. Nanopore sequencing and the Shasta toolkit enable efficient de novo assembly of eleven human genomes. Nat Biotechnol. 2020;38:1044. [CrossRef]
- Antipov D, Rautiainen M, Nurk S, Walenz BP, Solar SJ, Phillippy AM, et al. Verkko2: Integrating proximity ligation data with long-read De Bruijn graphs for efficient telomere-to-telomere genome assembly, phasing, and scaffolding. bioRxiv; 2024.
- Cheng H, Concepcion GT, Feng X, Zhang H, Li H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat Methods. 2021;18:170. [CrossRef]
- hifiasm [Internet]. 2025 [cited 2025 July 23]. Available from: https://github.com/chhylp123/hifiasm.
- Hakim SE, Choudhary N, Malhotra K, Peng J, Bültemeier A, Arafa A, et al. Phylogenomics and metabolic engineering reveal a conserved gene cluster in Solanaceae plants for withanolide biosynthesis. Nat Commun. 2025;16:6367. [CrossRef]
- Horz JM, Wolff K, Friedhoff R, Pucker B. Genome sequence of the ornamental plant Digitalis purpurea reveals the molecular basis of flower color and morphology variation. bioRxiv; 2024. p. 2024.02.14.580303.
- Manni M, Berkeley MR, Seppey M, Zdobnov EM. BUSCO: Assessing Genomic Data Quality and Beyond. Curr Protoc. 2021;1:e323. [CrossRef]
- Ou S, Chen J, Jiang N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic Acids Res. 2018;46:e126. [CrossRef]
- Natarajan S, Gehrke J, Pucker B. Mapping-based genome size estimation. BMC Genomics. 2025;26:482. [CrossRef]
- Rhie A, Walenz BP, Koren S, Phillippy AM. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 2020;21:245. [CrossRef]
- Gurevich A, Saveliev V, Vyahhi N, Tesler G. QUAST: quality assessment tool for genome assemblies. Bioinformatics. 2013;29:1072. [CrossRef]
- McCord RP, Kaplan N, Giorgetti L. Chromosome Conformation Capture and Beyond: Toward an Integrative View of Chromosome Structure and Function. Mol Cell. 2020;77:688. [CrossRef]
- CPhasing [Internet]. 2025 [cited 2025 July 23]. Available from: https://github.com/wangyibin/CPhasing.
- SRA - NCBI [Internet]. [cited 2025 July 23]. Available from: https://www.ncbi.nlm.nih.gov/sra/.
- Keilwagen J, Hartung F, Grau J. GeMoMa: Homology-Based Gene Prediction Utilizing Intron Position Conservation and RNA-seq Data. In: Kollmar M, editor. Gene Prediction: Methods and Protocols [Internet]. New York, NY: Springer; 2019 [cited 2025 July 23]. p. 161–77. Available from: . [CrossRef]
- Kallenborn F, Chacon A, Hundt C, Sirelkhatim H, Didi K, Dallago C, et al. GPU-accelerated homology search with MMseqs2. bioRxiv; 2024. p. 2024.11.13.623350.
- Gabriel L, Brůna T, Hoff KJ, Ebel M, Lomsadze A, Borodovsky M, et al. BRAKER3: Fully automated genome annotation using RNA-seq and protein evidence with GeneMark-ETP, AUGUSTUS, and TSEBRA. Genome Res. 2024;34:769. [CrossRef]
- Brůna T, Lomsadze A, Borodovsky M. GeneMark-ETP significantly improves the accuracy of automatic annotation of large eukaryotic genomes. Genome Res. 2024;34:757. [CrossRef]
- Stanke M, Diekhans M, Baertsch R, Haussler D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinforma Oxf Engl. 2008;24:637. [CrossRef]
- Palmer JM, Stajich J. Funannotate v1.8.1: Eukaryotic genome annotation. Zenodo; 2020.
- Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114. [CrossRef]
- Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol. 2011;29:644. [CrossRef]
- Haas BJ, Delcher AL, Mount SM, Wortman JR, Smith RK, Hannick LI, et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 2003;31:5654.
- Kim D, Paggi JM, Park C, Bennett C, Salzberg SL. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol. 2019;37:907. [CrossRef]
- Bray NL, Pimentel H, Melsted P, Pachter L. Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol. 2016;34:525. [CrossRef]
- Majoros WH, Pertea M, Salzberg SL. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinforma Oxf Engl. 2004;20:2878. [CrossRef]
- Korf I. Gene finding in novel genomes. BMC Bioinformatics. 2004;5:59. [CrossRef]
- Haas BJ, Salzberg SL, Zhu W, Pertea M, Allen JE, Orvis J, et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 2008;9:R7. [CrossRef]
- Karbstein K, Choudhary N, Xie T, Tomasello S, Wagner ND, Barke BH, et al. Efficient assembly of plant genomes: A case study with evolutionary implications in Ranunculus (Ranunculaceae). bioRxiv; 2024. p. 2023.08.08.552429.
- Testa AC, Hane JK, Ellwood SR, Oliver RP. CodingQuarry: highly accurate hidden Markov model gene prediction in fungal genomes using RNA-seq transcripts. BMC Genomics. 2015;16:170. [CrossRef]
- Vuruputoor VS, Monyak D, Fetter KC, Webster C, Bhattarai A, Shrestha B, et al. Welcome to the big leaves: Best practices for improving genome annotation in non-model plant genomes. Appl Plant Sci. 2023;11:e11533. [CrossRef]
- Woldesemayat AA, Ntushelo K, Modise DM. Identification and characterization of protein coding genes in monsonia (Monsonia burkeana Planch. ex harv) using a combination of approaches. Genes Genomics. 2017;39:245. [CrossRef]
- Robinson JT, Thorvaldsdottir H, Turner D, Mesirov JP. igv.js: an embeddable JavaScript implementation of the Integrative Genomics Viewer (IGV). Bioinformatics. 2023;39:btac830. [CrossRef]
- Ou S, Su W, Liao Y, Chougule K, Agda JRA, Hellinga AJ, et al. Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Genome Biol. 2019;20:275. [CrossRef]
- Nawrocki EP, Eddy SR. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics. 2013;29:2933. [CrossRef]
- Chan PP, Lin BY, Mak AJ, Lowe TM. tRNAscan-SE 2.0: improved detection and functional classification of transfer RNA genes. Nucleic Acids Res. 2021;49:9077. [CrossRef]
- Nawrocki E. Structural RNA Homology Search and Alignment Using Covariance Models. Theses Diss ETDs. 2009;
- Lagesen K, Hallin P, Rødland EA, Staerfeldt HH, Rognes T, Ussery DW. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 2007;35:3100. [CrossRef]
- Pucker B. Functional Annotation – How to Tackle the Bottleneck in Plant Genomics. Preprints; 2024.
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403. [CrossRef]
- Buchfink B, Xie C, Huson DH. Fast and sensitive protein alignment using DIAMOND. Nat Methods. 2015;12:59. [CrossRef]
- Pucker B, Holtgräwe D, Sörensen TR, Stracke R, Viehöver P, Weisshaar B. A De Novo Genome Sequence Assembly of the Arabidopsis thaliana Accession Niederzenz-1 Displays Presence/Absence Variation and Strong Synteny. PLOS ONE. 2016;11:e0164321. [CrossRef]
- Jones P, Binns D, Chang HY, Fraser M, Li W, McAnulla C, et al. InterProScan 5: genome-scale protein function classification. Bioinformatics. 2014;30:1236. [CrossRef]
- Rempel A, Choudhary N, Pucker B. KIPEs3: Automatic annotation of biosynthesis pathways. PLOS ONE. 2023;18:e0294342. [CrossRef]
- Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 2021;49:D412. [CrossRef]
- Hunter S, Apweiler R, Attwood TK, Bairoch A, Bateman A, Binns D, et al. InterPro: the integrative protein signature database. Nucleic Acids Res. 2009;37:D211. [CrossRef]
- Jensen LJ, Julien P, Kuhn M, von Mering C, Muller J, Doerks T, et al. eggNOG: automated construction and annotation of orthologous groups of genes. Nucleic Acids Res. 2008;36:D250.
- The UniProt Consortium. UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 2023;51:D523.
- Rawlings ND, Barrett AJ. MEROPS: the peptidase database. Nucleic Acids Res. 2000;28:323.
- Siva Shanmugam NR, Yin Y. CAZyme3D: A Database of 3D Structures for Carbohydrate-active Enzymes. J Mol Biol. 2025;437:169001.
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene Ontology: tool for the unification of biology. Nat Genet. 2000;25:25. [CrossRef]
- Wilkinson MD, Dumontier M, Aalbersberg IjJ, Appleton G, Axton M, Baak A, et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci Data. 2016;3:160018. [CrossRef]
- Hafner A, DeLeo V, Deng CH, Elsik CG, S Fleming D, Harrison PW, et al. Data reuse in agricultural genomics research: challenges and recommendations. GigaScience. 2025;14:giae106. [CrossRef]
- Sielemann K, Hafner A, Pucker B. The reuse of public datasets in the life sciences: potential risks and rewards. PeerJ. 2020;8:e9954. [CrossRef]
- ENA - European Nucleotide Archive [Internet]. [cited 2025 July 23]. Available from: https://www.ebi.ac.uk/ena/browser/home.
- Sequence Read Archive [Internet]. 2025 [cited 2025 July 23]. Available from: https://www.ddbj.nig.ac.jp/dra/index-e.html.
- Dainat J, Hereñú D, Murray DKD, Davis E, Ugrin I, Crouch K, et al. NBISweden/AGAT: AGAT-v1.4.1. Zenodo; 2024.
- Geib SM, Hall B, Derego T, Bremer FT, Cannoles K, Sim SB. Genome Annotation Generator: a simple tool for generating and correcting WGS annotation tables for NCBI submission. GigaScience. 2018;7:giy018. [CrossRef]
- Norling M, Jareborg N, Dainat J. EMBLmyGFF3: a converter facilitating genome annotation submission to European Nucleotide Archive. BMC Res Notes. 2018;11:584. [CrossRef]
- Submission Portal | NCBI | NLM | NIH [Internet]. [cited 2025 July 23]. Available from: https://submit.ncbi.nlm.nih.gov/.
- Webin command line submission interface (Webin-CLI) [Internet]. European Nucleotide Archive; 2025 [cited 2025 July 23]. Available from: https://github.com/enasequence/webin-cli.
- Pucker B, Choudhary N, Meckoni SN, de Oliveira JAVS. Plant Genomics Guide [Internet]. 2025 [cited 2025 Aug 14]. Available from: https://github.com/bpucker/PlantGenomicsGuide.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



