Submitted:
15 August 2025
Posted:
15 August 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
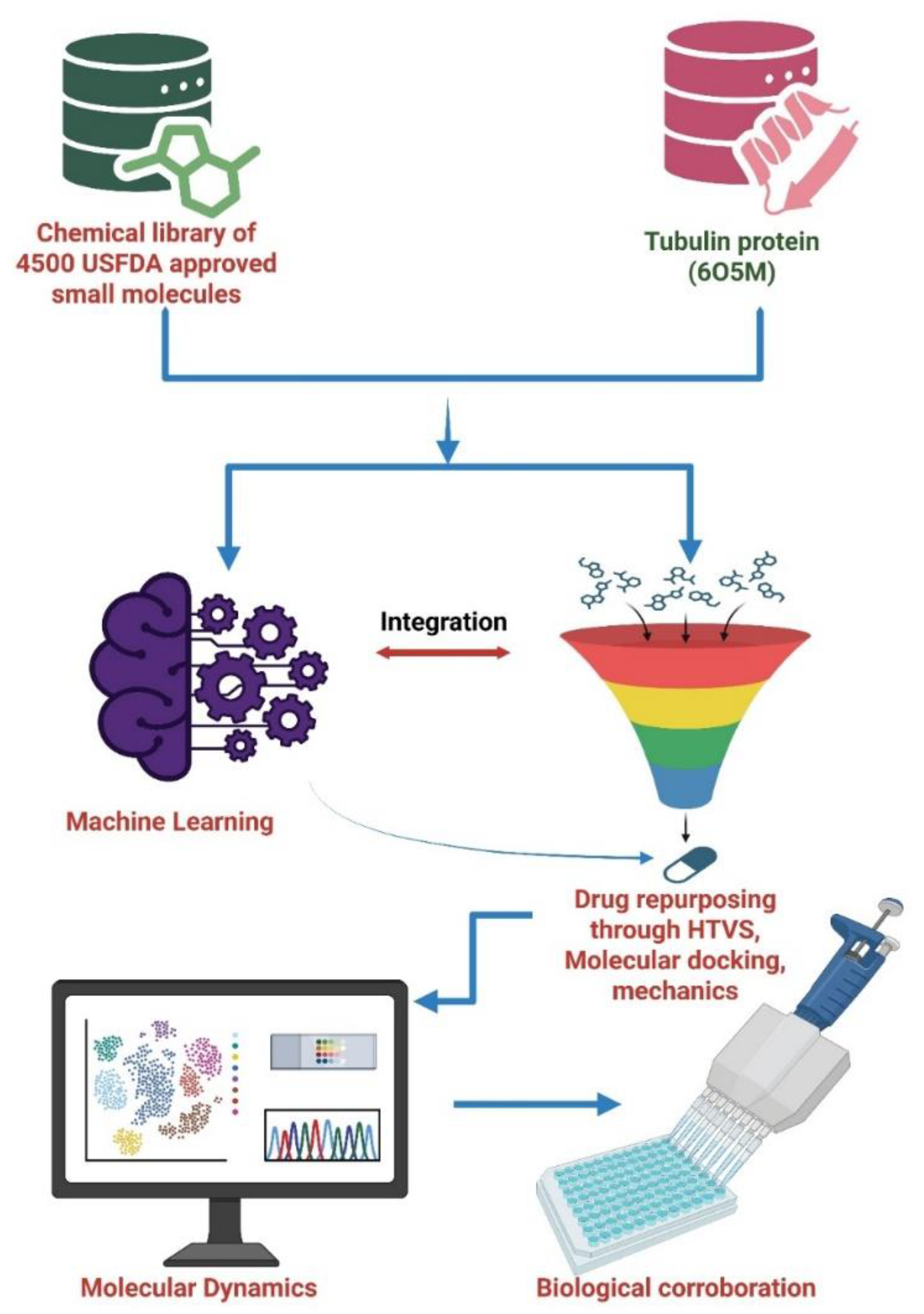
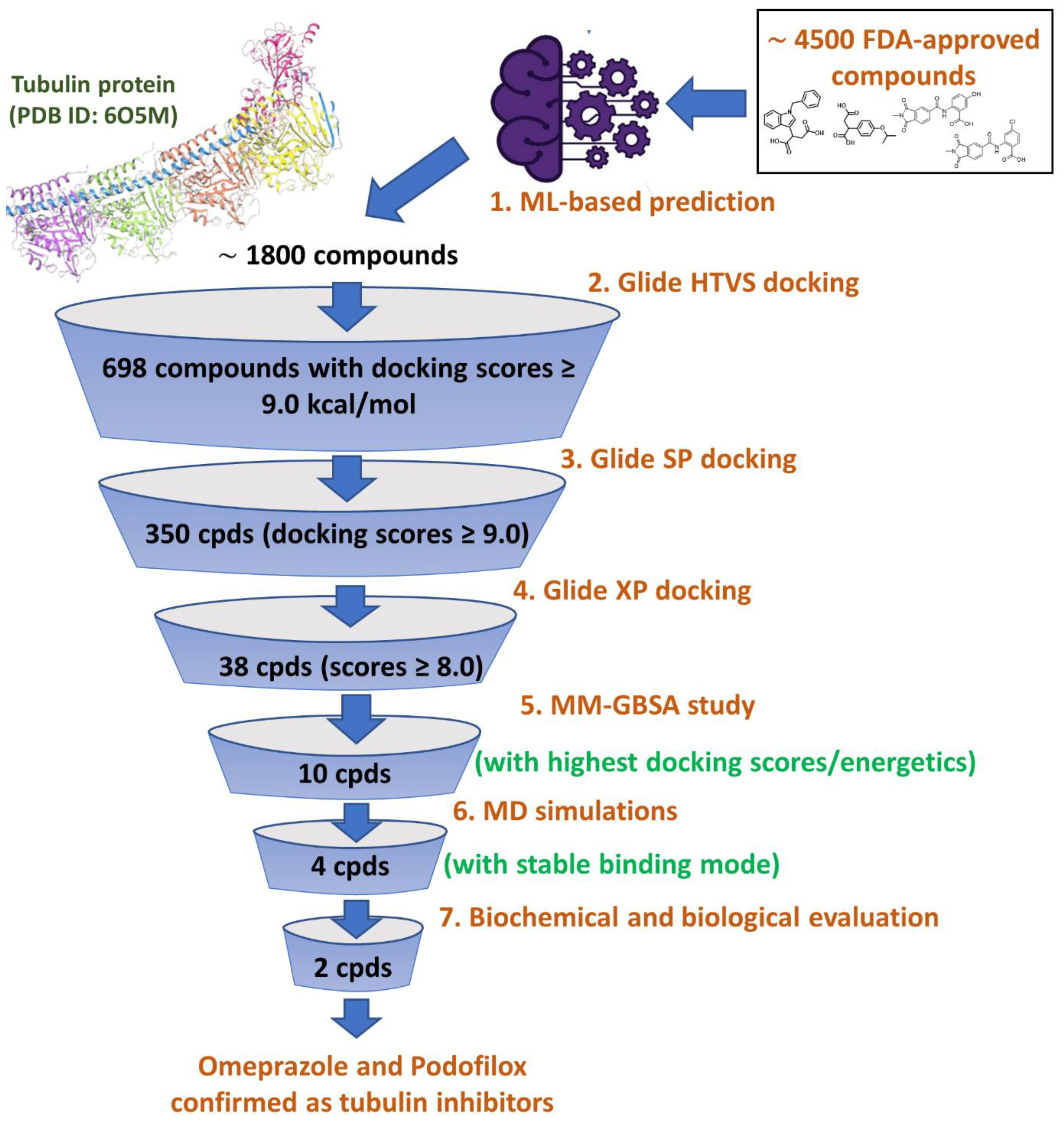
2.1. Machine Learning Workflow
2.1.1. Data Acquisition and Filtration
2.1.2. Decoys Generation
2.1.3. Model Development and Evaluation
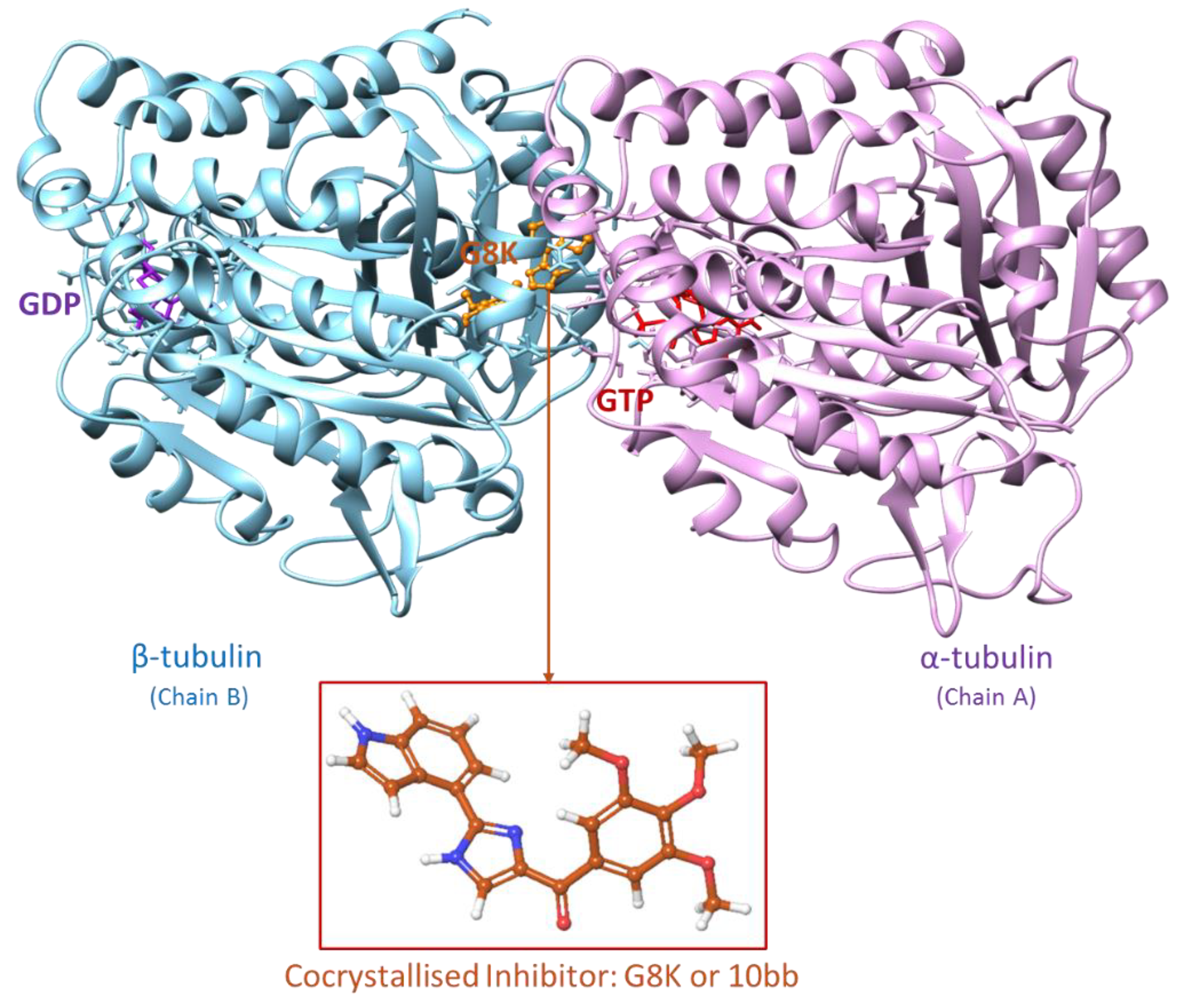
2.2. Protein Preparation and Optimization
2.3. Ligand Preparation
2.4. Receptor Grid Generation
2.5. Molecular Docking
2.6. Binding Free Energy Calculations
2.7. Molecular Dynamics Simulations and Clustering
2.8. Biological Investigations
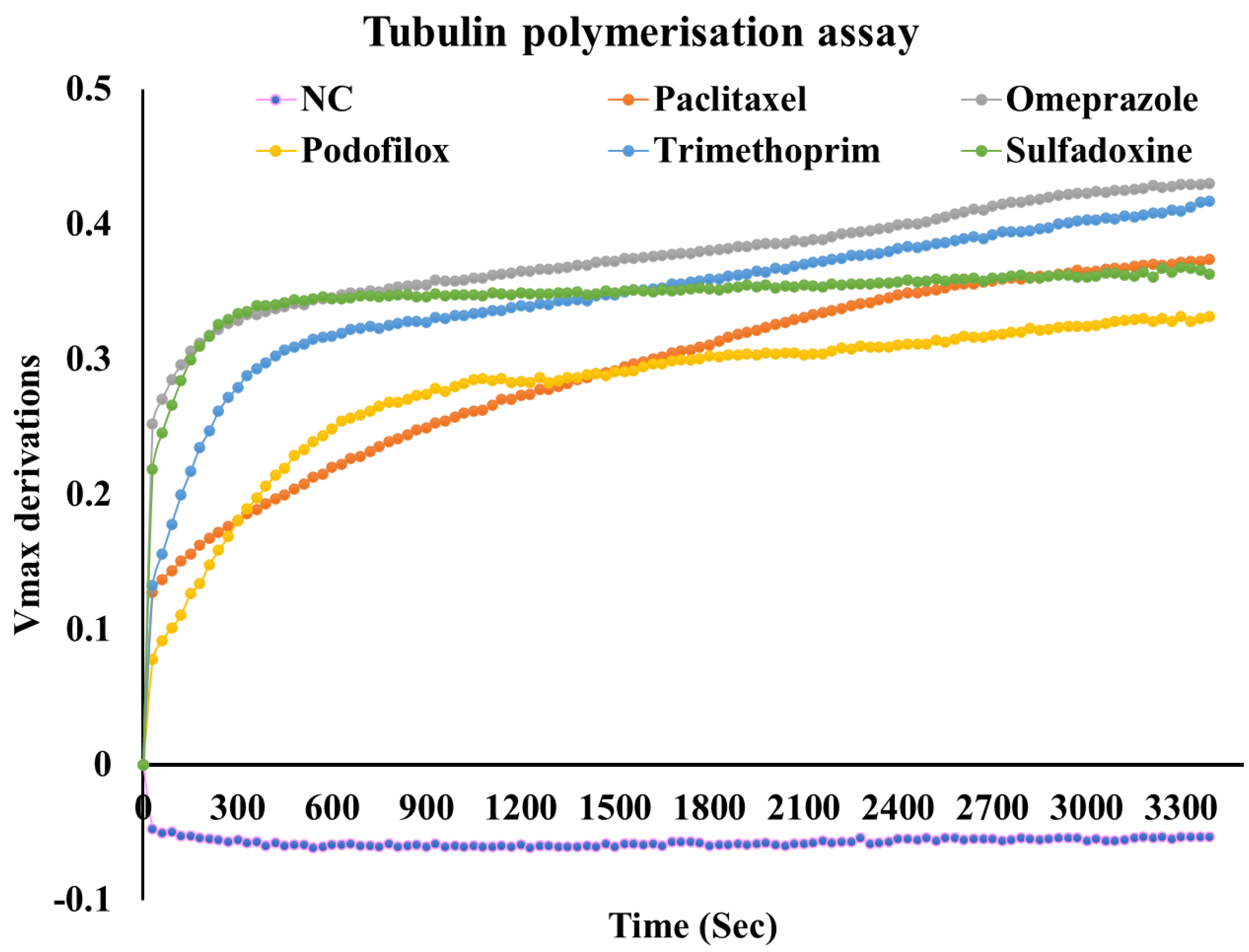
2.8.1. Tubulin Polymerisation Assay
2.8.2. Cell Culture and Cytotoxicity MeasurementsCell Growth and Maintenance
Resazurin Assay
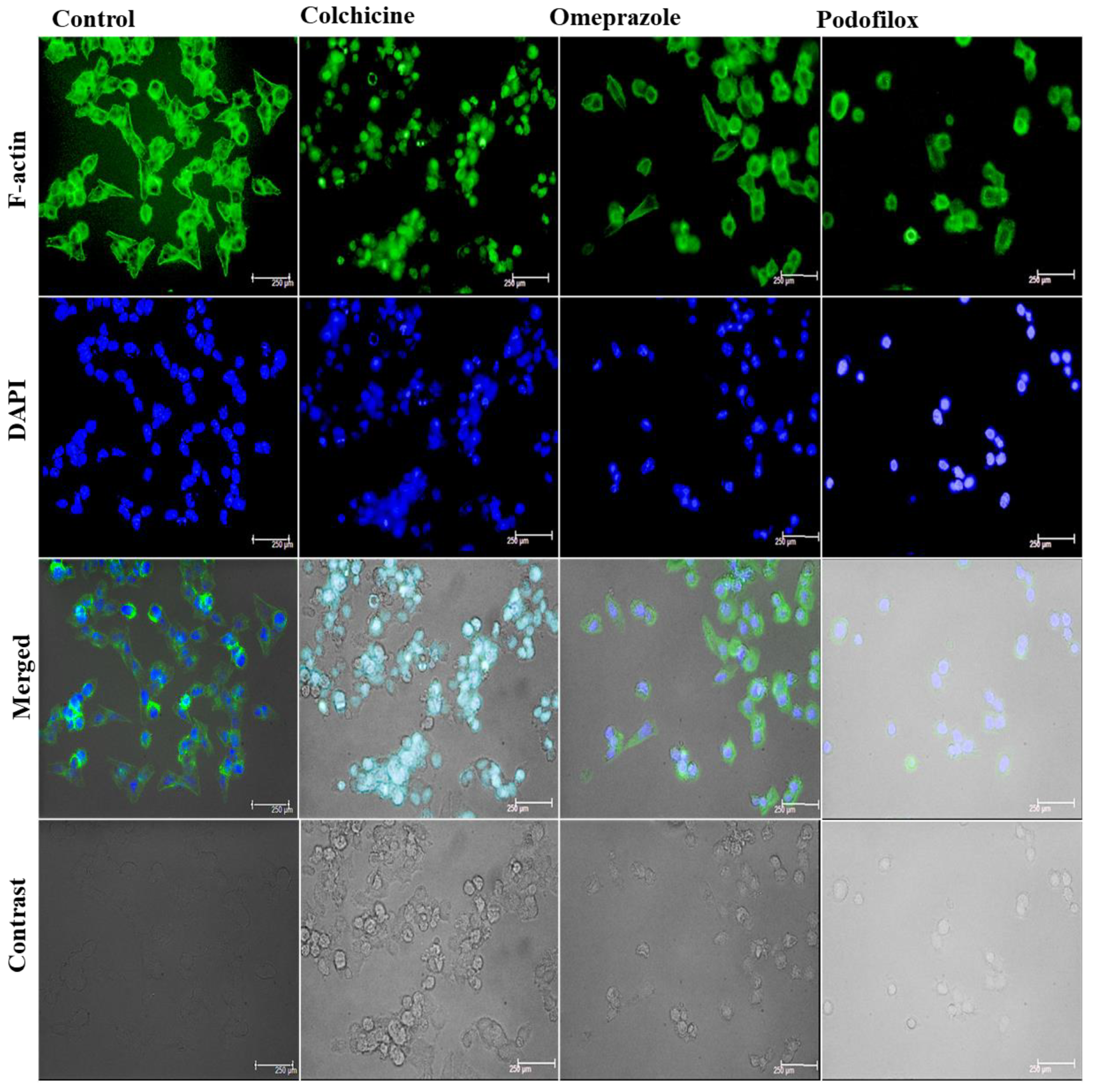
Confocal Analysis to Quantify the Tubulin Disruption
3. Results and Discussion
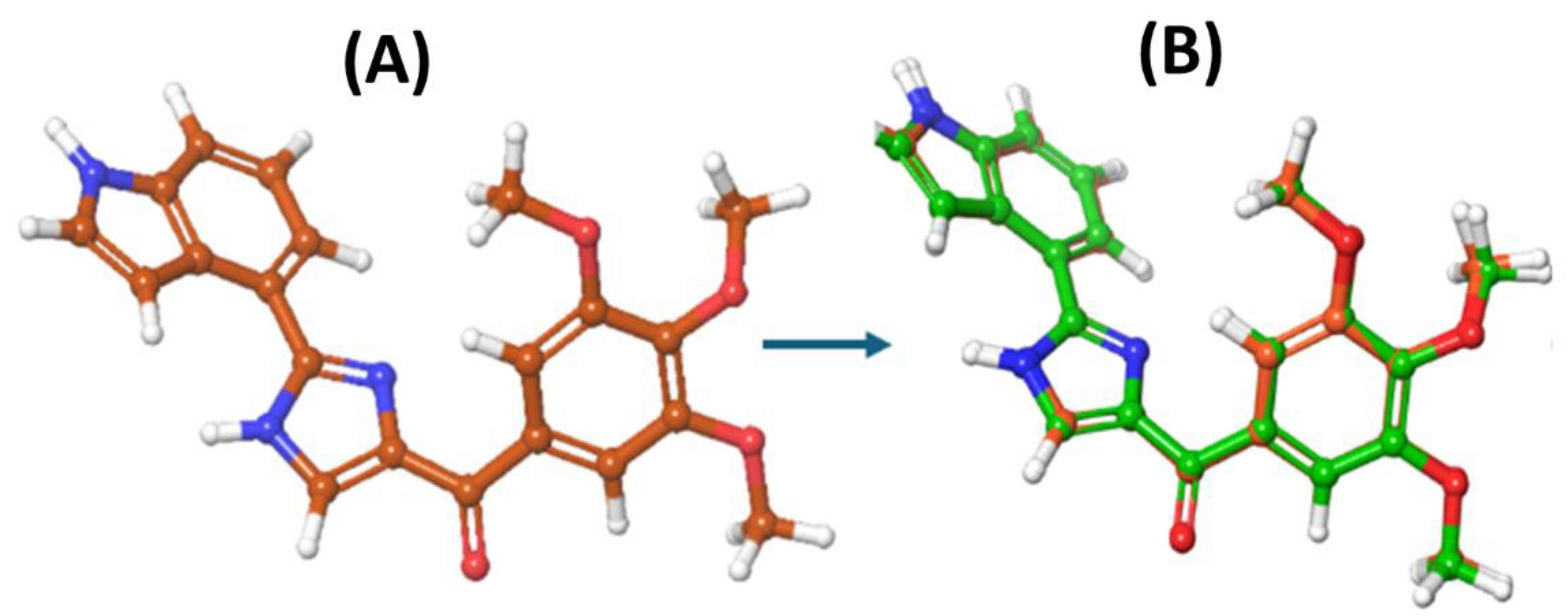
3.1. Machine Learning Model Development
- Bayesian classification (Bayes)
- Recursive partitioning (RP)
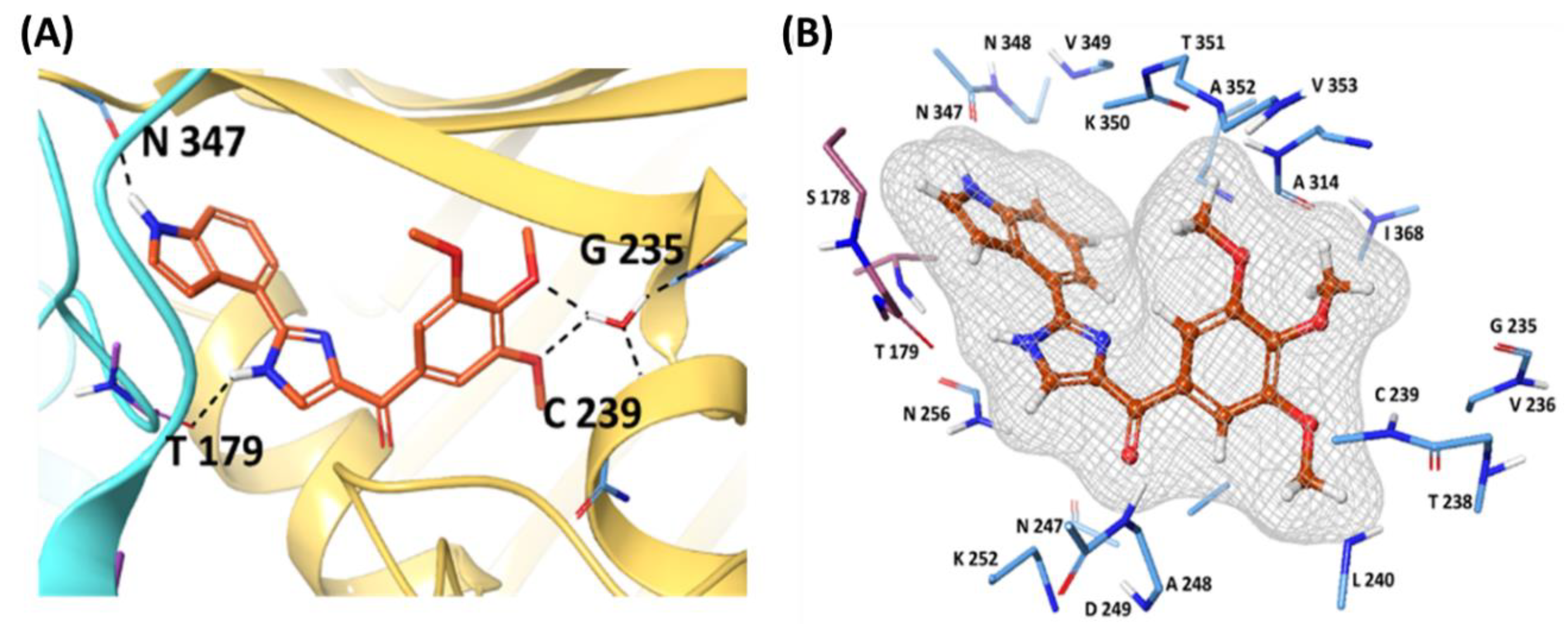
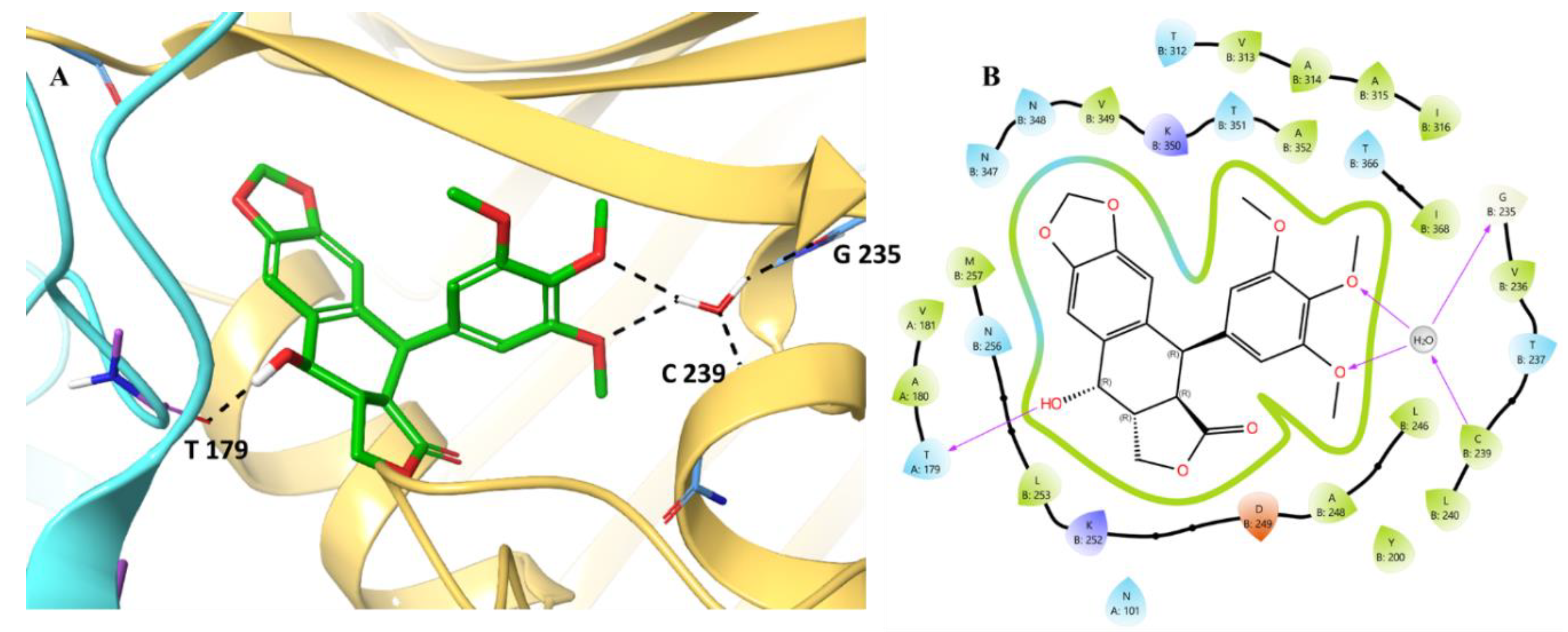
3.2. Molecular Docking
3.3. Molecular Dynamics (MD) Simulations
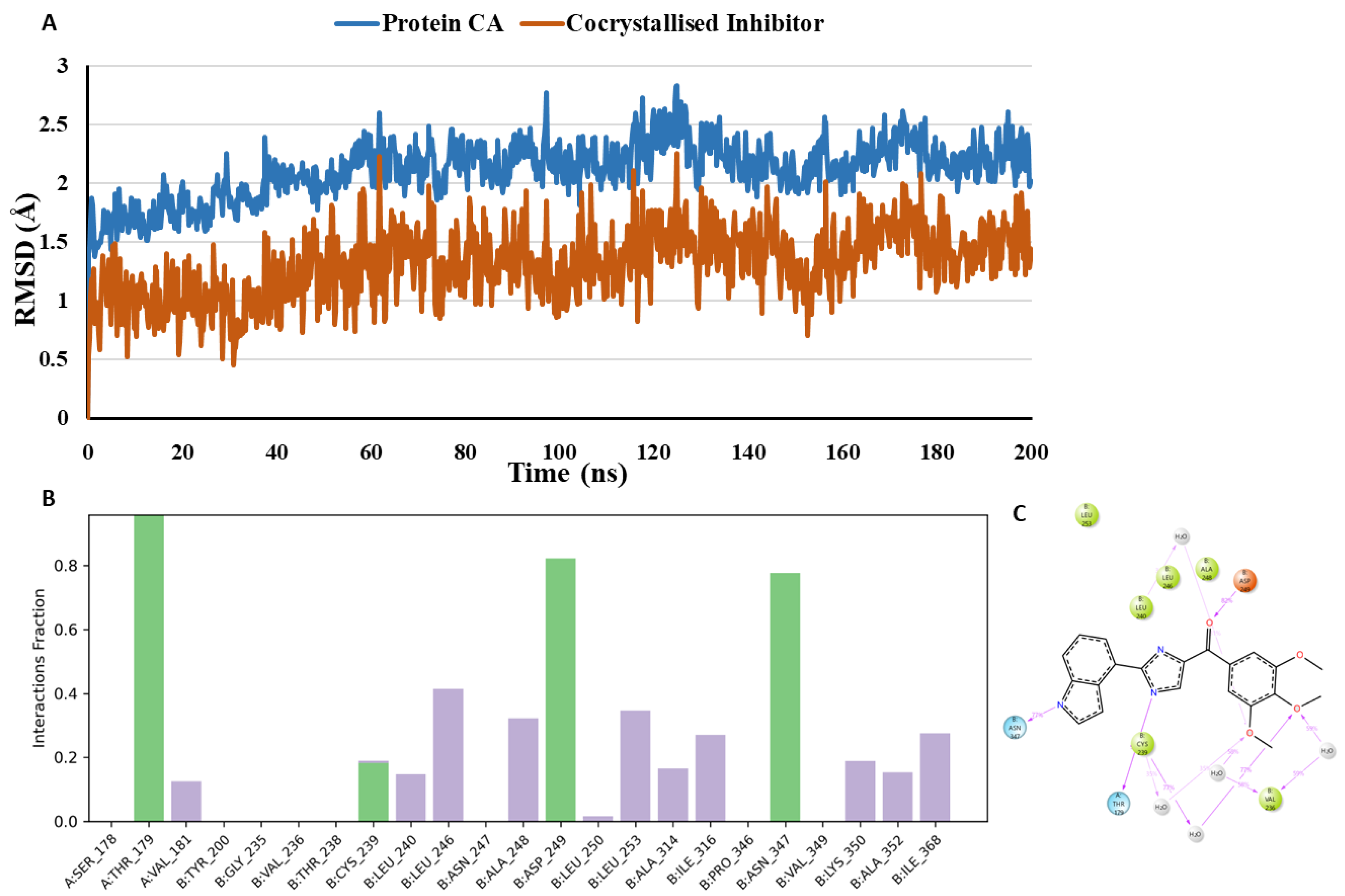
- MD analysis of the co-crystallised inhibitor G8K
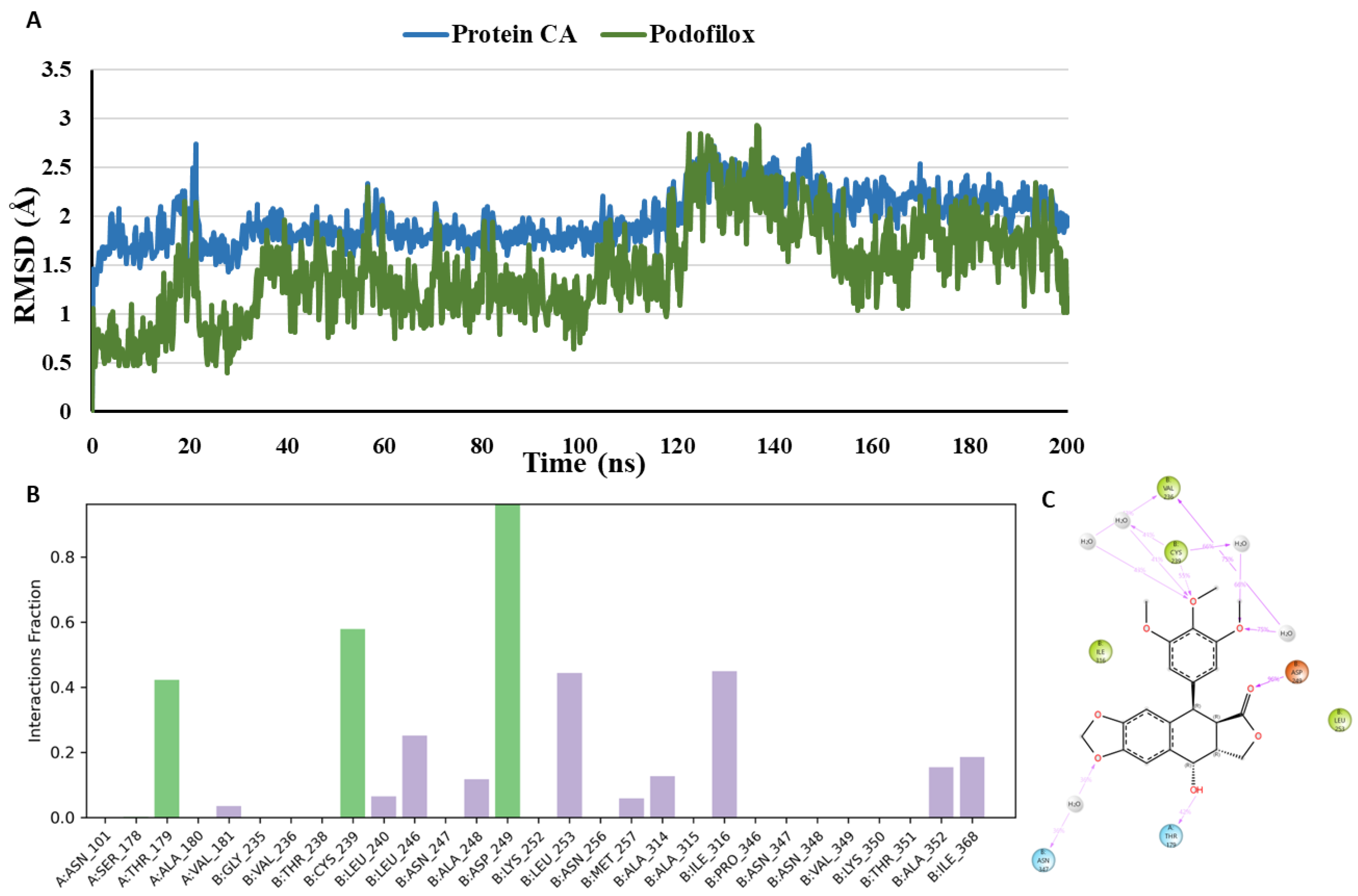
- MD analysis of Podofilox
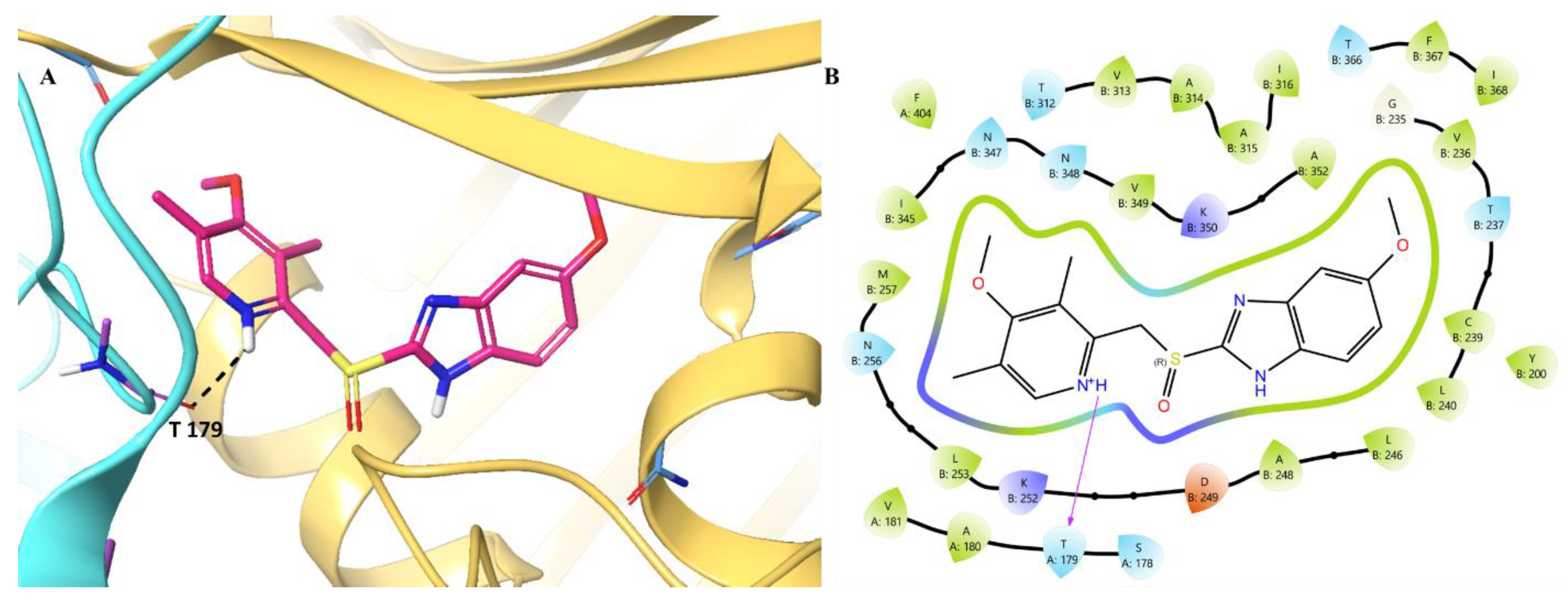
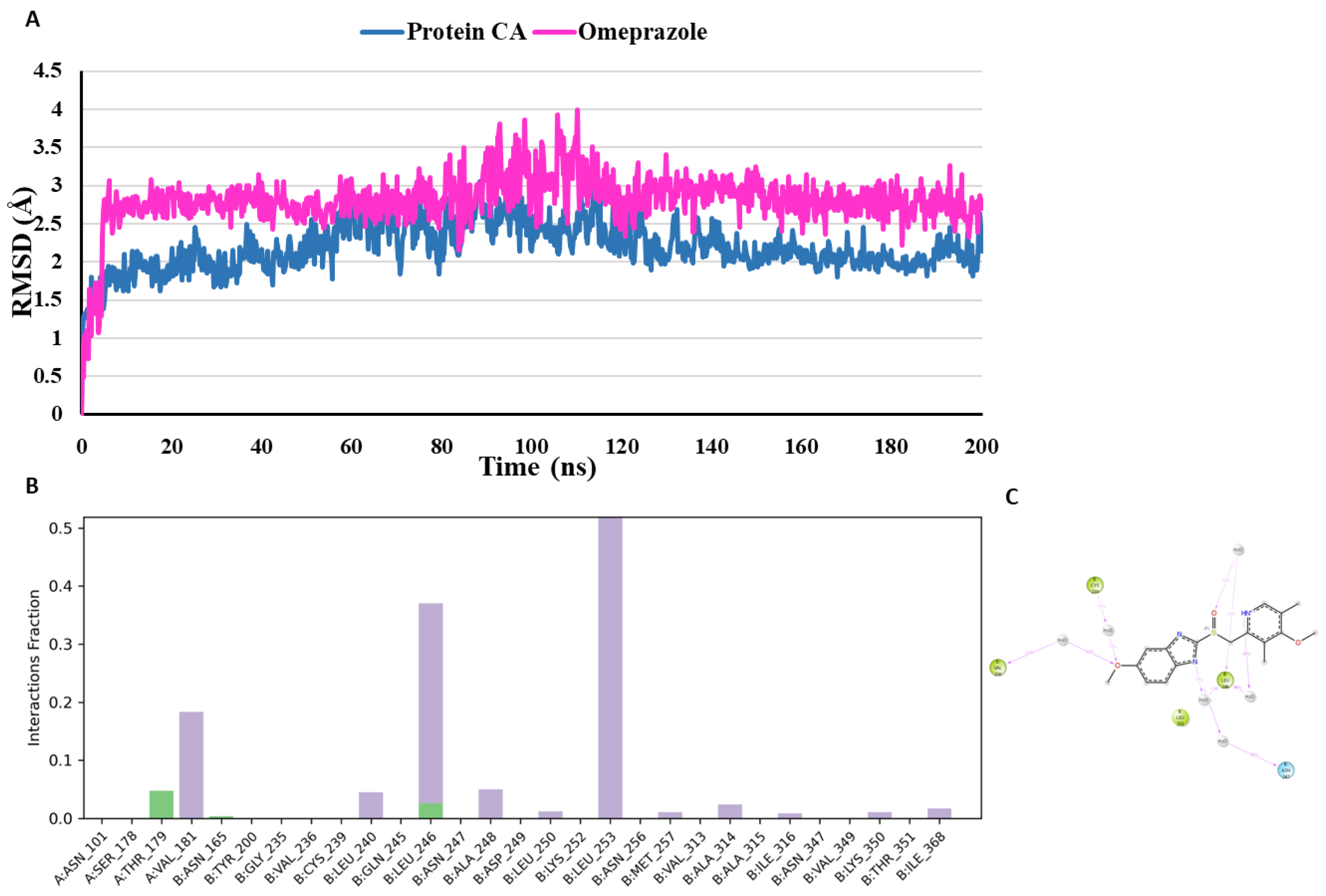
- MD analysis of Omeprazole
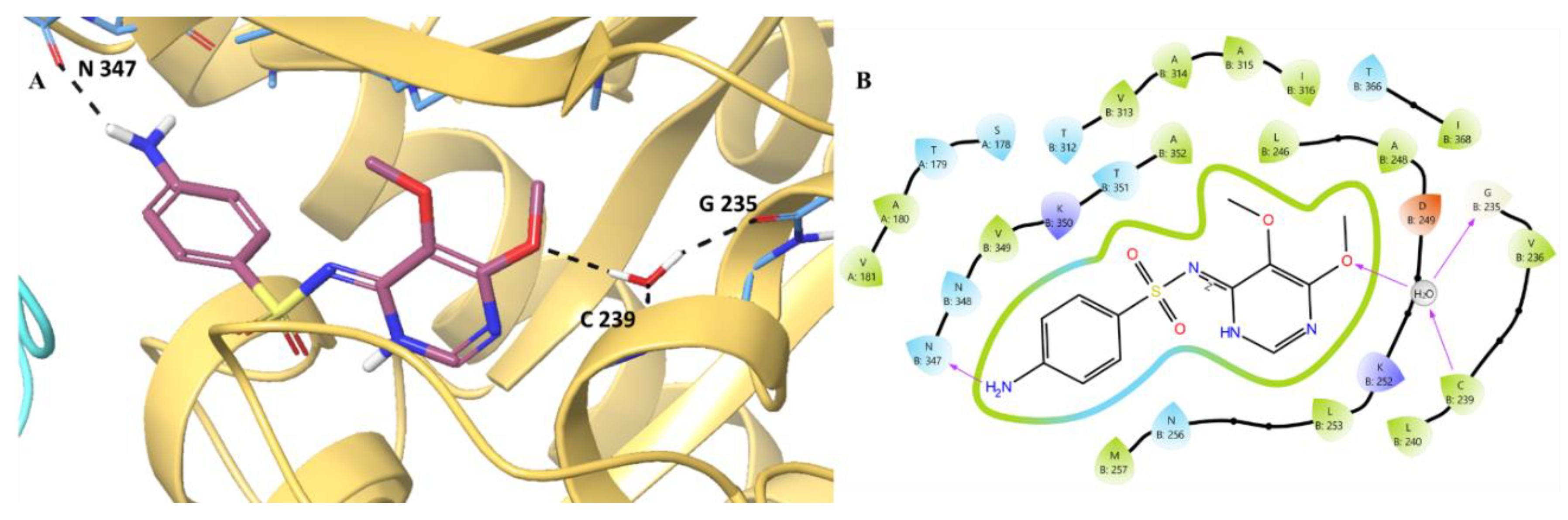
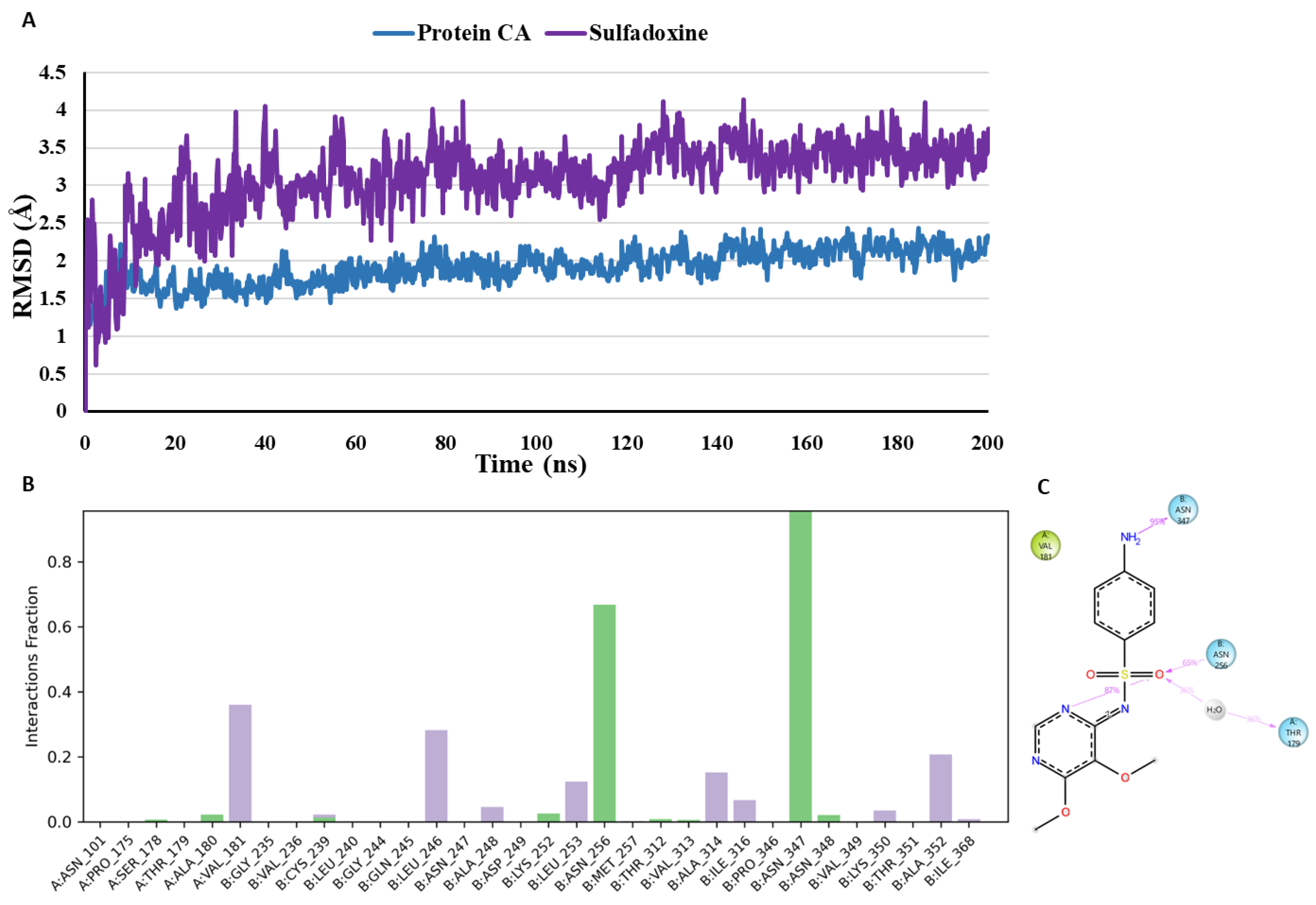
- MD analysis of Sulfadoxine
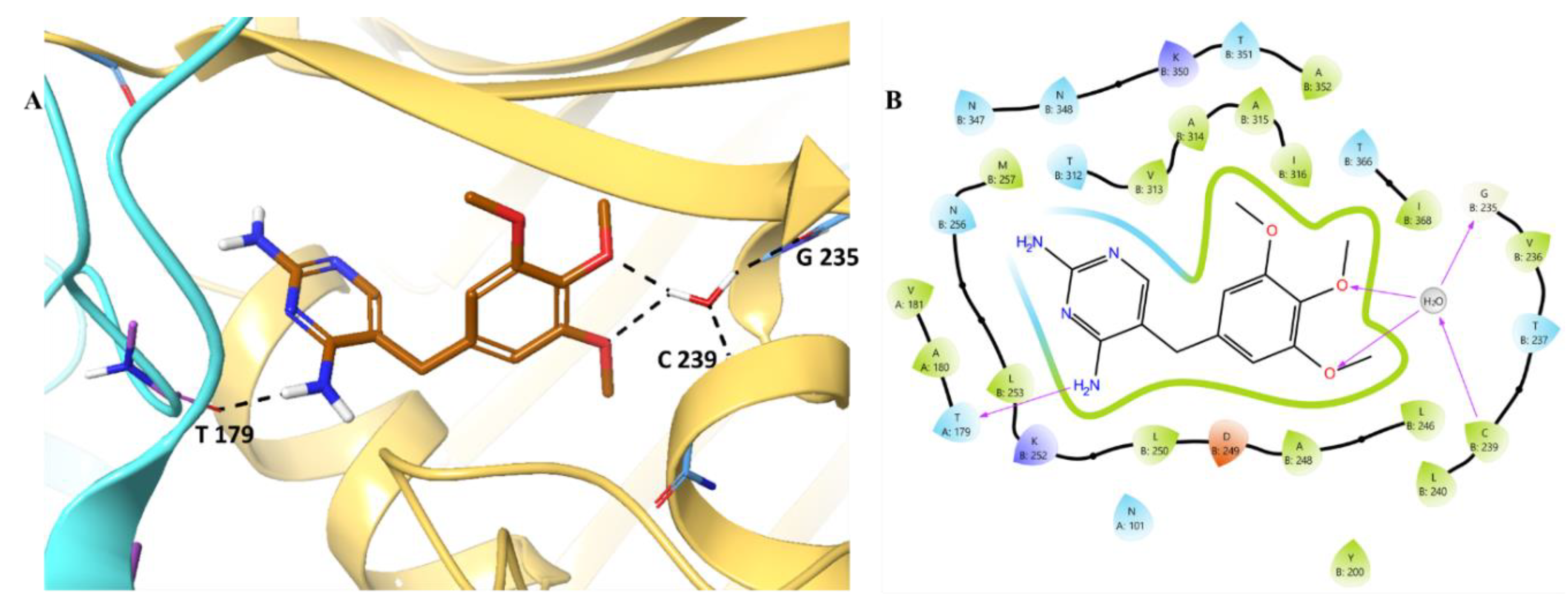
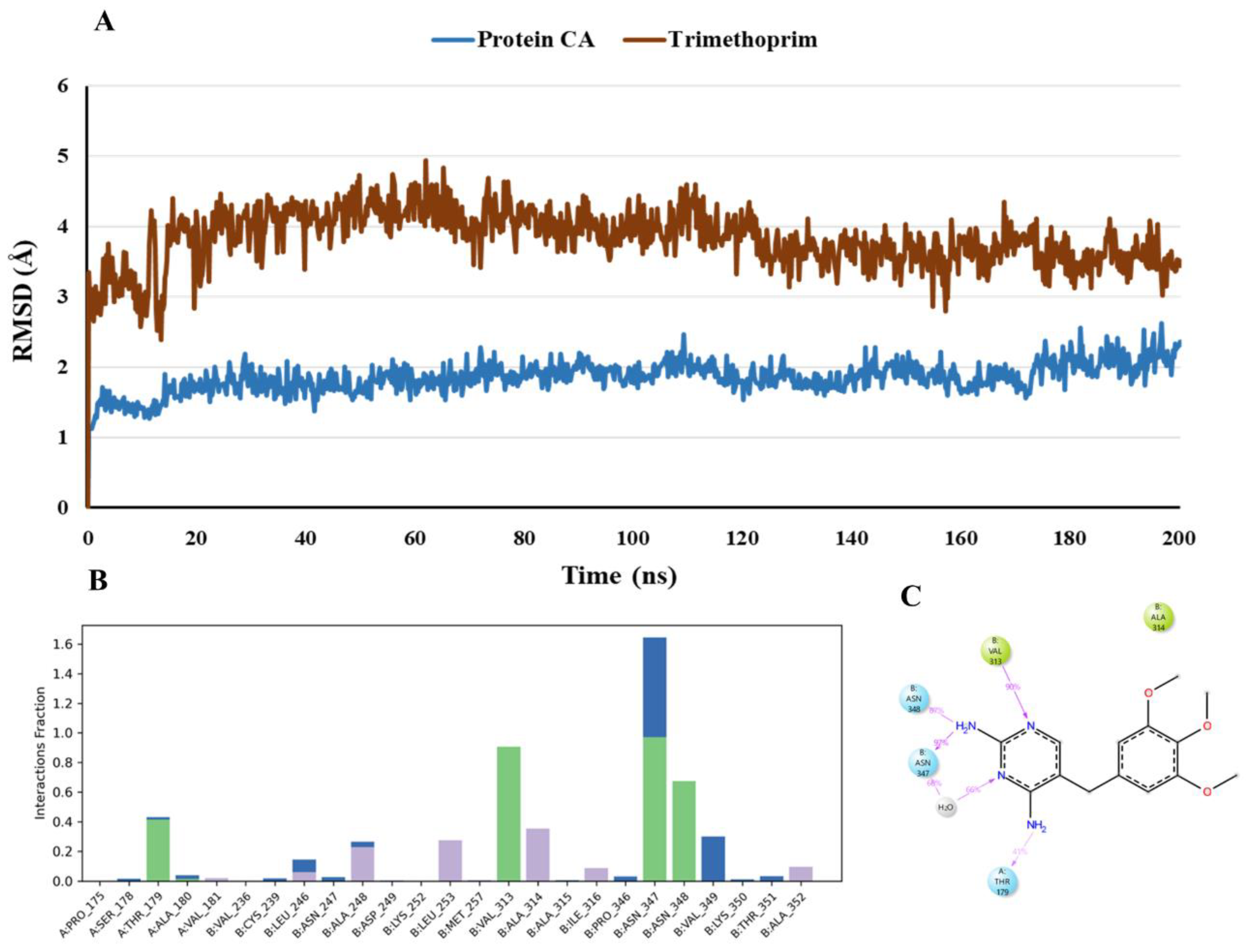
- MD Simulations of Trimethoprim
3.4. Biological Evaluation
4. Conclusion
Data availability
Author contributions
Conflict of interest
References
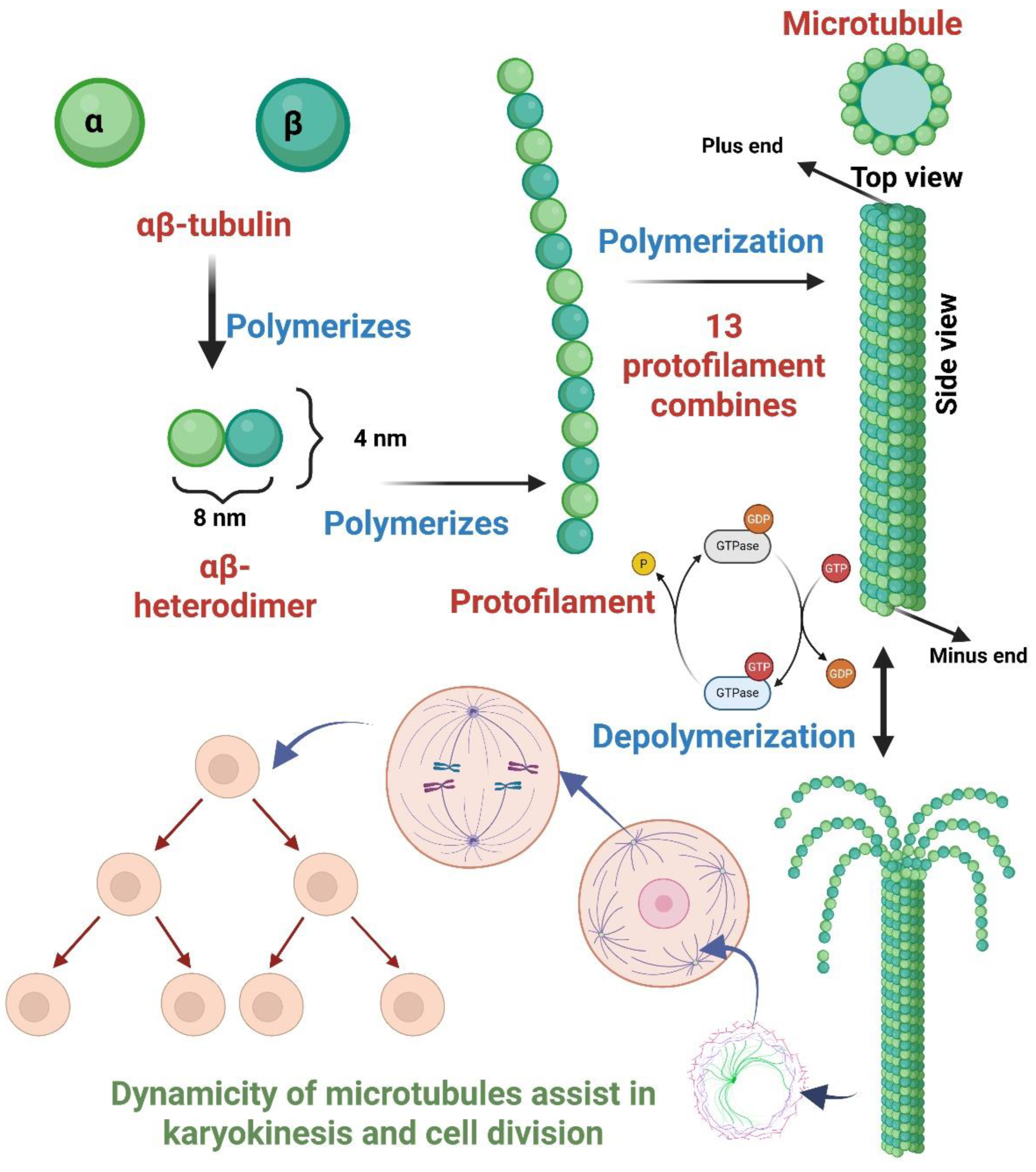
- Nogales, E.; Wolf, S. G.; Downing, K. H. Structure of the Aβ Tubulin Dimer by Electron Crystallography. Nature 1998, 391, 199–203. [Google Scholar] [CrossRef]
- Howard, J.; Hyman, A. A. Dynamics and Mechanics of the Microtubule plus End. Nature 2003, 422, 753–758. [Google Scholar] [CrossRef]
- Brouhard, G. J.; Rice, L. M. The Contribution of Aβ-Tubulin Curvature to Microtubule Dynamics. J. Cell Biol. 2014, 207, 323–334. [Google Scholar] [CrossRef]
- Binarová, P.; Tuszynski, J. Tubulin: Structure, Functions and Roles in Disease. Cells. 2019. [CrossRef] [PubMed]
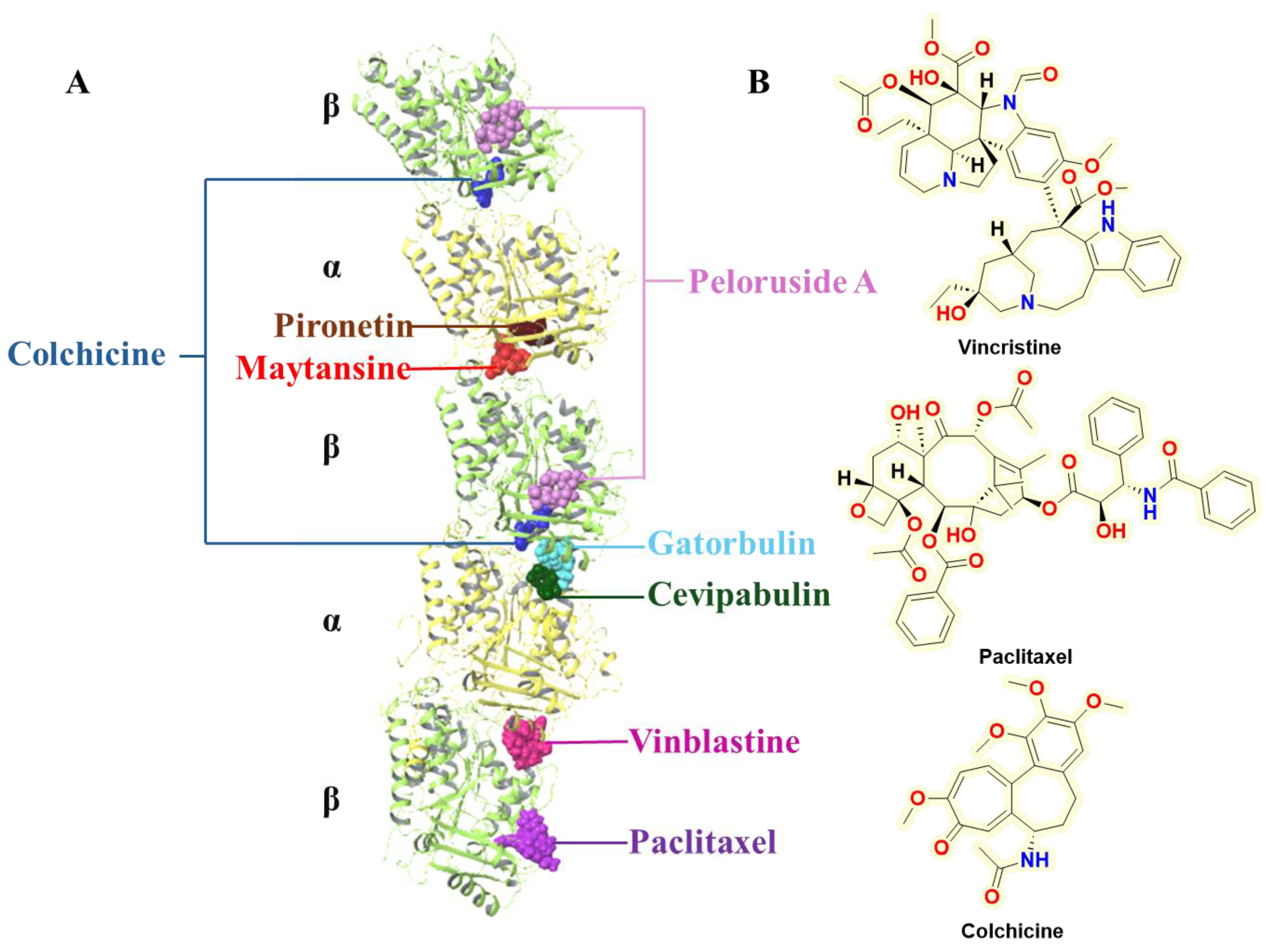
- Wang, X.; Gigant, B.; Zheng, X.; Chen, Q. Microtubule-Targeting Agents for Cancer Treatment: Seven Binding Sites and Three Strategies. MedComm – Oncol. 2023, 2, e46. [Google Scholar] [CrossRef]
- Esterbauer, H.; Schaur, R. J.; Zollner, H. Chemistry and Biochemistry of 4-Hydroxynonenal, Malonaldehyde and Related Aldehydes. Free Radic. Biol. Med. 1991, 11, 81–128. [Google Scholar] [CrossRef] [PubMed]
- Kristensson, M. A. The Game of Tubulins. Cells. 2021. [CrossRef] [PubMed]
- Bagdadi, N.; Wu, J.; Delaroche, J.; Serre, L.; Delphin, C.; De Andrade, M.; Carcel, M.; Nawabi, H.; Pinson, B.; Vérin, C.; Couté, Y.; Gory-Fauré, S.; Andrieux, A.; Stoppin-Mellet, V.; Arnal, I. Stable GDP-Tubulin Islands Rescue Dynamic Microtubules. J. Cell Biol. 2024, 223, e202307074. [Google Scholar] [CrossRef]
- Manka, S. W.; Moores, C. A. The Role of Tubulin–Tubulin Lattice Contacts in the Mechanism of Microtubule Dynamic Instability. Nat. Struct. Mol. Biol. 2018, 25, 607–615. [Google Scholar] [CrossRef]
- Parker, A. L.; Kavallaris, M.; McCarroll, J. A. Microtubules and Their Role in Cellular Stress in Cancer. Front. Oncol. 2014, 4. [Google Scholar] [CrossRef]
- Dumontet, C.; Jordan, M. A. Microtubule-Binding Agents: A Dynamic Field of Cancer Therapeutics. Nat. Rev. Drug Discov. 2010, 9, 790–803. [Google Scholar] [CrossRef] [PubMed]
- Čermák, V.; Dostál, V.; Jelínek, M.; Libusová, L.; Kovář, J.; Rösel, D.; Brábek, J. Microtubule-Targeting Agents and Their Impact on Cancer Treatment. Eur. J. Cell Biol. 2020, 99, 151075. [Google Scholar] [CrossRef]
- Khrapunovich-Baine, M.; Menon, V.; Yang, C.-P. H.; Northcote, P. T.; Miller, J. H.; Angeletti, R. H.; Fiser, A.; Horwitz, S. B.; Xiao, H. Hallmarks of Molecular Action of Microtubule Stabilizing Agents: EFFECTS OF EPOTHILONE B, IXABEPILONE, PELORUSIDE A, AND LAULIMALIDE ON MICROTUBULE CONFORMATION *. J. Biol. Chem. 2011, 286, 11765–11778. [Google Scholar] [CrossRef] [PubMed]
- Wordeman, L.; Vicente, J. J. Microtubule Targeting Agents in Disease: Classic Drugs, Novel Roles. Cancers. 2021. [CrossRef]
- Liu, W.; Jia, H.; Guan, M.; Cui, M.; Lan, Z.; He, Y.; Guo, Z.; Jiang, R.; Dong, G.; Wang, S. Discovery of Novel Tubulin Inhibitors Targeting the Colchicine Binding Site via Virtual Screening, Structural Optimization and Antitumor Evaluation. Bioorg. Chem. 2022, 118, 105486. [Google Scholar] [CrossRef]
- Khwaja, S.; Kumar, K.; Das, R.; Negi, A. S. Microtubule Associated Proteins as Targets for Anticancer Drug Development. Bioorg. Chem. 2021, 116, 105320. [Google Scholar] [CrossRef] [PubMed]
- Calinescu, A.-A.; Castro, M. G. Microtubule Targeting Agents in Glioma. Transl. Cancer Res. 2016, 5. [Google Scholar] [CrossRef]
- Behl, T.; Sharma, A.; Sharma, L.; Sehgal, A.; Zengin, G.; Brata, R.; Fratila, O.; Bungau, S. Exploring the Multifaceted Therapeutic Potential of Withaferin A and Its Derivatives. Biomedicines. 2020. [CrossRef] [PubMed]
- Ikeda, R.; Kurosawa, M.; Okabayashi, T.; Takei, A.; Yoshiwara, M.; Kumakura, T.; Sakai, N.; Funatsu, O.; Morita, A.; Ikekita, M.; Nakaike, Y.; Konakahara, T. 3-(3-Phenoxybenzyl)Amino-β-Carboline: A Novel Antitumor Drug Targeting α-Tubulin. Bioorg. Med. Chem. Lett. 2011, 21, 4784–4787. [Google Scholar] [CrossRef]
- Zhang, D.; Kanakkanthara, A. Beyond the Paclitaxel and Vinca Alkaloids: Next Generation of Plant-Derived Microtubule-Targeting Agents with Potential Anticancer Activity. Cancers. 2020. [CrossRef]
- Assunção, H. C.; Silva, P. M. A.; Bousbaa, H.; Cidade, H. Recent Advances in Microtubule Targeting Agents for Cancer Therapy. Molecules. 2025. [CrossRef]
- Sargsyan, A.; Sahakyan, H.; Nazaryan, K. Effect of Colchicine Binding Site Inhibitors on the Tubulin Intersubunit Interaction. ACS Omega 2023, 8, 29448–29454. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Chen, J.; Xiao, M.; Li, W.; Miller, D. D. An Overview of Tubulin Inhibitors That Interact with the Colchicine Binding Site. Pharm. Res. 2012, 29, 2943–2971. [Google Scholar] [CrossRef]
- Wu, D.; Li, Y.; Zheng, L.; Xiao, H.; Ouyang, L.; Wang, G.; Sun, Q. Small Molecules Targeting Protein–Protein Interactions for Cancer Therapy. Acta Pharm. Sin. B 2023, 13, 4060–4088. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Zhou, Q.; He, J.; Jiang, Z.; Peng, C.; Tong, R.; Shi, J. Recent Advances in the Development of Protein–Protein Interactions Modulators: Mechanisms and Clinical Trials. Signal Transduct. Target. Ther. 2020, 5, 213. [Google Scholar] [CrossRef]
- Field, J. J.; Díaz, J. F.; Miller, J. H. The Binding Sites of Microtubule-Stabilizing Agents. Chem. Biol. 2013, 20, 301–315. [Google Scholar] [CrossRef]
- Horgan, M. J.; Zell, L.; Siewert, B.; Stuppner, H.; Schuster, D.; Temml, V. Identification of Novel β-Tubulin Inhibitors Using a Combined In Silico/In Vitro Approach. J. Chem. Inf. Model. 2023, 63, 6396–6411. [Google Scholar] [CrossRef]
- Wang, J.; Miller, D. D.; Li, W. Molecular Interactions at the Colchicine Binding Site in Tubulin: An X-Ray Crystallography Perspective. Drug Discov. Today 2022, 27, 759–776. [Google Scholar] [CrossRef]
- Cheng, Z.; Lu, X.; Feng, B. A Review of Research Progress of Antitumor Drugs Based on Tubulin Targets. Transl. Cancer Res. 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Pushpakom, S.; Iorio, F.; Eyers, P. A.; Escott, K. J.; Hopper, S.; Wells, A.; Doig, A.; Guilliams, T.; Latimer, J.; McNamee, C.; Norris, A.; Sanseau, P.; Cavalla, D.; Pirmohamed, M. Drug Repurposing: Progress, Challenges and Recommendations. Nat. Rev. Drug Discov. 2019, 18, 41–58. [Google Scholar] [CrossRef]
- Greenwood, J. R.; Calkins, D.; Sullivan, A. P.; Shelley, J. C. Towards the Comprehensive, Rapid, and Accurate Prediction of the Favorable Tautomeric States of Drug-like Molecules in Aqueous Solution. J. Comput. Aided. Mol. Des. 2010, 24, 591–604. [Google Scholar] [CrossRef]
- Jha, V.; Galati, S.; Volpi, V.; Ciccone, L.; Minutolo, F.; Rizzolio, F.; Granchi, C.; Poli, G.; Tuccinardi, T. Discovery of a New ATP-Citrate Lyase (ACLY) Inhibitor Identified by a Pharmacophore-Based Virtual Screening Study. J. Biomol. Struct. Dyn. 2020, 1–9. [Google Scholar] [CrossRef]
- Bharadwaj, S.; Deepika, K.; Kumar, A.; Jaiswal, S.; Miglani, S.; Singh, D.; Fartyal, P.; Kumar, R.; Singh, S.; Singh, M. P.; Gaidhane, A. M.; Kumar, B.; Jha, V. Exploring the Artificial Intelligence and Its Impact in Pharmaceutical Sciences: Insights Toward the Horizons Where Technology Meets Tradition. Chem. Biol. Drug Des. 2024, 104, e14639. [Google Scholar] [CrossRef] [PubMed]
- Vemula, D.; Jayasurya, P.; Sushmitha, V.; Kumar, Y. N.; Bhandari, V. CADD, AI and ML in Drug Discovery: A Comprehensive Review. Eur. J. Pharm. Sci. 2023, 181, 106324. [Google Scholar] [CrossRef]
- Wang, K.; Huang, Y.; Wang, Y.; You, Q.; Wang, L. Recent Advances from Computer-Aided Drug Design to Artificial Intelligence Drug Design. RSC Med. Chem. 2024, 15, 3978–4000. [Google Scholar] [CrossRef] [PubMed]
- Goel, K. K.; Chahal, S.; Kumar, D.; Jaiswal, S.; Nainwal, N.; Singh, R.; Mahajan, S.; Rawat, P.; Yadav, S.; Fartyal, P.; Ahmad, G.; Jha, V.; Dwivedi, A. R. Repurposing of USFDA-Approved Drugs to Identify Leads for Inhibition of Acetylcholinesterase Enzyme: A Plausible Utility as an Anti-Alzheimer Agent. RSC Med. Chem. 2024, 15, 4138–4152. [Google Scholar] [CrossRef] [PubMed]
- Mendez, D.; Gaulton, A.; Bento, A. P.; Chambers, J.; De Veij, M.; Félix, E.; Magariños, M. P.; Mosquera, J. F.; Mutowo, P.; Nowotka, M.; Gordillo-Marañón, M.; Hunter, F.; Junco, L.; Mugumbate, G.; Rodriguez-Lopez, M.; Atkinson, F.; Bosc, N.; Radoux, C. J.; Segura-Cabrera, A.; Hersey, A.; Leach, A. R. ChEMBL: Towards Direct Deposition of Bioassay Data. Nucleic Acids Res. 2019, 47, D930–D940. [Google Scholar] [CrossRef]
- Schrödinger Release 2024-2: LigPrep, Schrödinger, LLC, New York, NY, 2024.
- Schrödinger Release 2024-2: Maestro, Schrödinger, LLC, New York, NY, 2024.
- Mysinger, M. M.; Carchia, M.; Irwin, J. J.; Shoichet, B. K. Directory of Useful Decoys, Enhanced (DUD-E): Better Ligands and Decoys for Better Benchmarking. J. Med. Chem. 2012, 55, 6582–6594. [Google Scholar] [CrossRef]
- Dixon, S. L.; Duan, J.; Smith, E.; Von Bargen, C. D.; Sherman, W.; Repasky, M. P. Autoqsar: An Automated Machine Learning Tool for Best-Practice Quantitative Structure–Activity Relationship Modeling. Future Med. Chem. 2016, 8, 1825–1839. [Google Scholar] [CrossRef]
- Wang, Q.; Arnst, K. E.; Wang, Y.; Kumar, G.; Ma, D.; White, S. W.; Miller, D. D.; Li, W.; Li, W. Structure-Guided Design, Synthesis, and Biological Evaluation of (2-(1H-Indol-3-Yl)-1H-Imidazol-4-Yl)(3,4,5-Trimethoxyphenyl) Methanone (ABI-231) Analogues Targeting the Colchicine Binding Site in Tubulin. J. Med. Chem. 2019, 62, 6734–6750. [Google Scholar] [CrossRef]
- Berman, H. M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T. N.; Weissig, H.; Shindyalov, I. N.; Bourne, P. E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Madhavi Sastry, G.; Adzhigirey, M.; Day, T.; Annabhimoju, R.; Sherman, W. Protein and Ligand Preparation: Parameters, Protocols, and Influence on Virtual Screening Enrichments. J. Comput. Aided. Mol. Des. 2013, 27, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R. A.; Banks, J. L.; Murphy, R. B.; Halgren, T. A.; Klicic, J. J.; Mainz, D. T.; Repasky, M. P.; Knoll, E. H.; Shelley, M.; Perry, J. K.; Shaw, D. E.; Francis, P.; Shenkin, P. S. Glide: A New Approach for Rapid, Accurate Docking and Scoring. 1. Method and Assessment of Docking Accuracy. J. Med. Chem. 2004, 47, 1739–1749. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R. A.; Murphy, R. B.; Repasky, M. P.; Frye, L. L.; Greenwood, J. R.; Halgren, T. A.; Sanschagrin, P. C.; Mainz, D. T. Extra Precision Glide: Docking and Scoring Incorporating a Model of Hydrophobic Enclosure for Protein−Ligand Complexes. J. Med. Chem. 2006, 49, 6177–6196. [Google Scholar] [CrossRef]
- Massova, I.; Kollman, P. A. Combined Molecular Mechanical and Continuum Solvent Approach (MM-PBSA/GBSA) to Predict Ligand Binding. Perspect. Drug Discov. Des. 2000, 18, 113–135. [Google Scholar] [CrossRef]
- Lu, C.; Wu, C.; Ghoreishi, D.; Chen, W.; Wang, L.; Damm, W.; Ross, G. A.; Dahlgren, M. K.; Russell, E.; Von Bargen, C. D.; Abel, R.; Friesner, R. A.; Harder, E. D. OPLS4: Improving Force Field Accuracy on Challenging Regimes of Chemical Space. J. Chem. Theory Comput. 2021, 17, 4291–4300. [Google Scholar] [CrossRef]
- Jorgensen, W. L.; Tirado-Rives, J. The OPLS [Optimized Potentials for Liquid Simulations] Potential Functions for Proteins, Energy Minimizations for Crystals of Cyclic Peptides and Crambin. J. Am. Chem. Soc. 1988, 110, 1657–1666. [Google Scholar] [CrossRef]
- Bowers, K. J.; Chow, D. E.; Xu, H.; Dror, R. O.; Eastwood, M. P.; Gregersen, B. A.; Klepeis, J. L.; Kolossvary, I.; Moraes, M. A.; Sacerdoti, F. D.; Salmon, J. K.; Shan, Y.; Shaw, D. E. Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In SC ’06: Proceedings of the 2006 ACM/IEEE Conference on Supercomputing. [CrossRef]
- Schrödinger Release 2024-2: Desmond Molecular Dynamics System, D. E. Schrödinger Release 2024-2: Desmond Molecular Dynamics System, D. E. Shaw Research, New York, NY, 2021. Maestro-Desmond Interoperability Tools, Schrödinger, New York, NY, 2024.
- Jorgensen, W. L.; Chandrasekhar, J.; Madura, J. D.; Impey, R. W.; Klein, M. L. Comparison of Simple Potential Functions for Simulating Liquid Water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Martyna, G. J.; Klein, M. L.; Tuckerman, M. Nosé–Hoover Chains: The Canonical Ensemble via Continuous Dynamics. J. Chem. Phys. 1992, 97, 2635–2643. [Google Scholar] [CrossRef]
- Wentzcovitch, R. M. Invariant Molecular-Dynamics Approach to Structural Phase Transitions. Phys. Rev. B 1991, 44, 2358–2361. [Google Scholar] [CrossRef]
- Nosé, S. A Unified Formulation of the Constant Temperature Molecular Dynamics Methods. J. Chem. Phys. 1984, 81, 511–519. [Google Scholar] [CrossRef]
| Entry | Model Name | Algorithm | Descriptor Type | Model Score |
| 1. | Bayes_dendritic_1 | Bayesian Classification | Dendritic Fingerprints | 0.9389 |
| 2. | Bayes_linear_20 | Bayesian Classification | Linear Fingerprints | 0.9337 |
| 3. | Bayes_linear_2 | Bayesian Classification | Linear Fingerprints | 0.9257 |
| 4. | Bayes_dendritic_2 | Bayesian Classification | Dendritic Fingerprints | 0.9256 |
| 5. | Bayes_linear_33 | Bayesian Classification | Linear Fingerprints | 0.9237 |
| 6. | Bayes_molprint2D_46 | Bayesian Classification | Mol2D Fingerprints | 0.9200 |
| 7. | Bayes_linear_1 | Bayesian Classification | Linear Fingerprints | 0.9185 |
| 8. | RP_30 | Recursive Partitioning | Mixed Descriptor Set | 0.9158 |
| 9. | Bayes_dendritic_33 | Bayesian Classification | Dendritic Fingerprints | 0.9146 |
| 10. | RP_41 | Recursive Partitioning | Mixed Descriptor Set | 0.9133 |
| Dataset | RMS Err | Correlation (r) | R2/Q2 |
| Training | 0.2246 | 0.9045 | 0.8182 |
| Test | 0.2374 | 0.8988 | 0.8078 |
| Dataset | Class | TP | FP | TN | FN | Recall | Precision | Specificity | F-measure |
| Training set | Active | 125 | 11 | 78 | 4 | 0.969 | 0.919 | 0.876 | 0.943 |
| Inactive | 78 | 4 | 125 | 11 | 0.876 | 0.951 | 0.969 | 0.912 | |
| Test set | Active | 41 | 4 | 25 | 1 | 0.976 | 0.911 | 0.862 | 0.943 |
| Inactive | 25 | 1 | 41 | 4 | 0.862 | 0.962 | 0.976 | 0.909 |
| Entry | Compound | Glide XP Docking Score (Kcal/mol) |
ΔGbind (Kcal/mol) |
ΔGcoulomb (Kcal/mol) |
| 1. | Arformoterol | -10.061 | -18.12 | -21.37 |
| 2. | Podofilox | -9.908 | -38.61 | -22.45 |
| 3. | Tretoquinol | -9.857 | -17.73 | -20.95 |
| 4. | Terizidone | -9.219 | -25.83 | -17.28 |
| 5. | Rucaparib | -9.145 | -19.46 | -15.23 |
| 6. | Aminoglutethimide | -8.99 | -26.59 | -6.77 |
| 7. | Sulfadoxine | -8.763 | -50.56 | 15.89 |
| 8. | Naftifine | -8.531 | -32.33 | -6.58 |
| 9. | Omeprazole | -8.362 | -64.16 | -58.31 |
| 10. | Trimethoprim | -8.117 | -43.89 | -18.9 |
| 11. | Co-crystallized G8K | -12.365 | -74.07 | -29.70 |
| Code | Half Maximal Inhibitory Concentration (IC50) (µM) ± SD | |||||
| SK-MEL-28 (Melanoma) |
A549 (Lung) |
MDAMB-231 (Breast) |
MCF-7 (Breast) |
PC-3 (Prostate) |
HCT-116 (Colon) | |
| Omeprazole | 4.32±0.29 | 17.32±0.19 | 21.65±0.21 | 13.44±0.26 | 12.22±0.18 | 6.22±0.22 |
| Podofilox | 4.98±0.37 | >25 | 19.83±0.24 | 13.61±0.18 | 15.93±0.31 | 5.76±0.18 |
| Trimethoprim | 7.46±0.19 | >25 | >25 | >25 | 18.65±0.28 | 11.32±0.31 |
| Sulfadoxine | 5.22±0.33 | >25 | 16.11±0.19 | 18.22±0.27 | 13.98±0.22 | 8.88±0.27 |
| Paclitaxel | 1.98±0.31 | 2.22±0.35 | 3.89±0.22 | 3.22±0.33 | 4.86±0.19 | NT |
| Colchicine | 2.43±0.19 | 2.11±0.22 | 2.54±0.31 | 2.89±0.24 | 4.02±0.16 | 4.92±0.19 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




