Submitted:
08 August 2025
Posted:
14 August 2025
You are already at the latest version
Abstract
Keywords:
Why the Random-Effects Model Is More Natural
Discrepancy Between Subject-Level and Variant-Level Realized Genetic Correlations
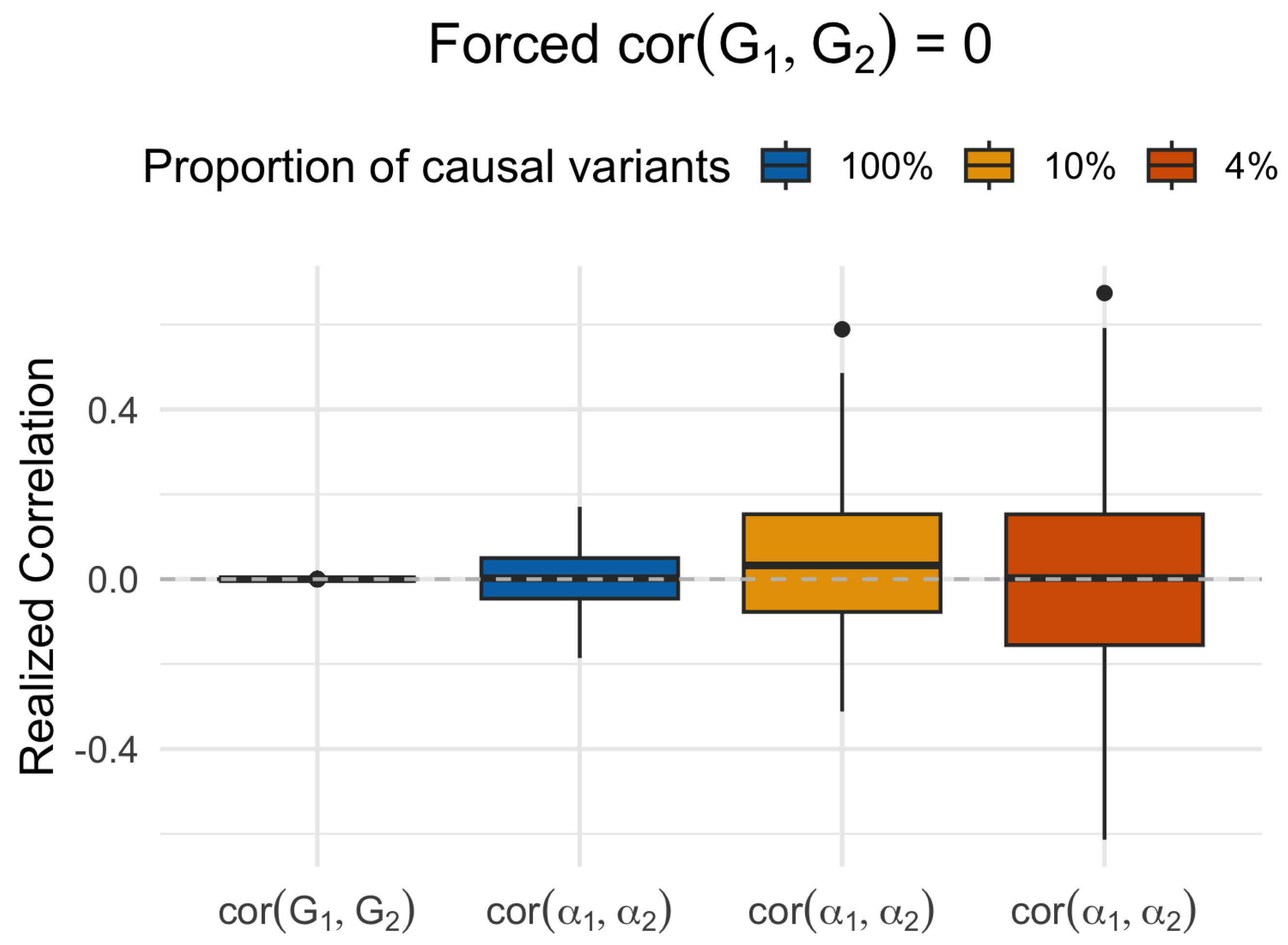
Statistical Artifact in Realized Genetic Correlations
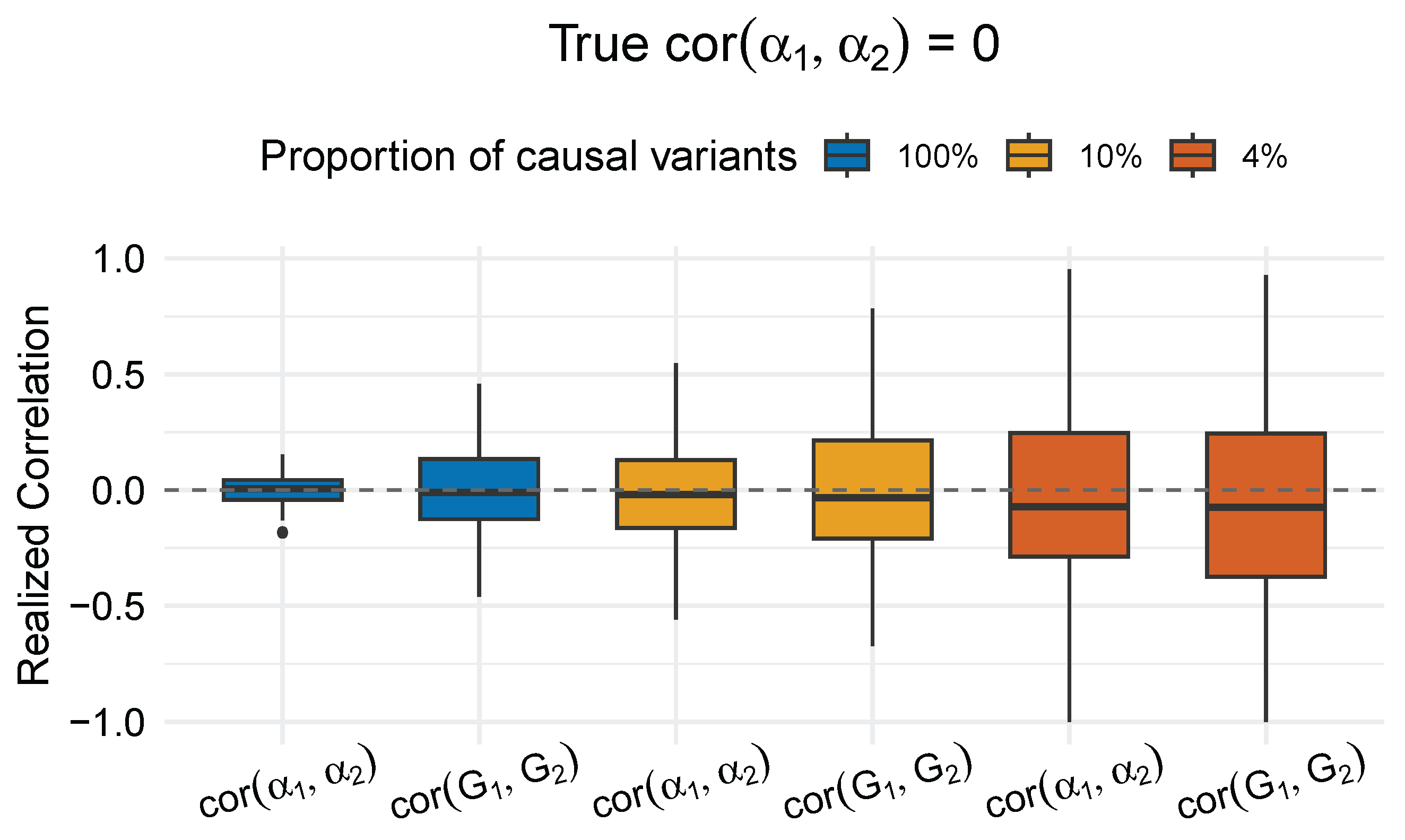
Discussion
- Valid testing requires calibration under a data-generating process. When simulating from a model with true genetic correlation , the realized genetic components and will exhibit non-zero correlation purely due to stochastic variation, especially in small regions or with limited number of causal variants. A valid testing procedure must control type-I error under this null model – that is, it must not declare significance at a rate higher than the nominal level, regardless of fluctuations in observed correlations.
- Inflated type-I error rate is not power. The assertion that the observed inflation in LAVA is due to “power” is misleading. Power refers to the probability of detecting a true effect when it exists. Under the null hypothesis , there is no true effect to detect. Thus, any rejection of the null in this setting is a false positive. Mistaking sensitivity to random noise as “power” indicates a failure to properly model uncertainty in the null distribution.
- Artificial orthogonality does not address the problem. de Leeuw et al. suggest that when subject-level orthogonality is enforced – that is, when is forced in simulations, LAVA’s calibration improves. While true, this is not a valid argument. In real data, the subject-level correlation under the null of no genetic effects will never be zero. A robust method must accommodate such fluctuations and remain conservative. HDL-L achieves this by modeling the variance of the local covariance estimator, thereby avoiding inflation under the null.
- Counterintuitive type-I error inflation for a small proportion of causal variants. We observed that type-I error inflation in LAVA worsens for a smaller proportion of causal variants, even when the true genetic correlation remains zero. Two factors are involved: (i) the mixture of zero and sparse nonzero effects tends to produce higher realized correlations, and, (ii) to maintain a specified heritability, the smaller proportion of causal variants are assigned larger genetic effects. Overall, the fact that LAVA becomes more inflated in these settings suggests that its test statistic is overly sensitive to the presence of a genetic effect, regardless of whether the signal is shared across traits.
Author Contributions
Data Availability Statement
Code Availability
Acknowledgments
Conflicts of Interest
References
- de Leeuw, C., Werme, J. & Posthuma, D. Re-evaluating local genetic correlation analysis in the hdl framework. Preprints (2025). [CrossRef]
- Werme, J., van der Sluis, S., Posthuma, D. & de Leeuw, C. A. An integrated framework for local genetic correlation analysis. Nature Genetics 54, 274–282 (2022). [CrossRef]
- Li, Y., Pawitan, Y. & Shen, X. An enhanced framework for local genetic correlation analysis. Nature Genetics 57, 1053–1058 (2025). [CrossRef]
- Lynch, M., Walsh, B. et al. Genetics and Analysis of Quantitative Traits, vol. 1 (Sinauer Sunderland, MA, 1998). See Chapters 26 and 27 for a detailed description of general mixed models.
- Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nature Genetics 2015, 47, 291–295. [CrossRef] [PubMed]
- Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nature Genetics 2015, 47, 1236–1241. [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



