Submitted:
30 July 2025
Posted:
30 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Work
3. Methodology
- Overlapping triples (): Present in both and .
- Triples with new relations but known entities ().
- Triples with new entities and possibly new relations ().
1. Corpus Partitioning
2. Distant Supervision
3. Model Training
4. Iterative Knowledge Discovery
5. Negative Sampling
6. Algorithm Implementation
4. Experimental Setup
4.1. Datasets
4.2. Model Settings
- BERT (Base, uncased) – general-purpose language model.
- BioClinicalBERT – pre-trained on clinical notes and biomedical articles.
- BiomedRoBERTa – optimized for biomedical scientific literature.
4.3. Training Configuration
- Learning rate:
- Batch size: 20
- Epochs: 4
- Confidence thresholds: , , ,
- Negative sampling ratios: ,
4.4. Evaluation Methods
| Item | Description |
|---|---|
| Coarse KG | 5.2M entities, 7.3M triples |
| Fine Corpus | 240K oncology paragraphs |
| PLMs Used | BERT, BioClinicalBERT, BiomedRoBERTa |
| Test Set Size | 40K sentences |
| Hardware | 4 × NVIDIA A100 GPUs |
5. Results and Discussion
5.1. Held-Out Evaluation
| Model | Precision | Recall | F1 Score |
|---|---|---|---|
| BERT | 0.908 | 0.900 | 0.904 |
| BioClinicalBERT | 0.909 | 0.895 | 0.902 |
| BiomedRoBERTa | 0.908 | 0.901 | 0.905 |
5.2. Manual Evaluation
| Model | |||
|---|---|---|---|
| BERT | 0.90 | 0.58 | 0.70 |
| BioClinicalBERT | 0.90 | 0.66 | 0.62 |
| BiomedRoBERTa | 0.94 | 0.76 | 0.74 |
5.3. Ablation Study
| Model Variant | RE Precision |
|---|---|
| BiomedRoBERTa (Full) | 0.987 |
| w/o Cumulative | 0.983 |
| w/o Iteration | 0.986 |
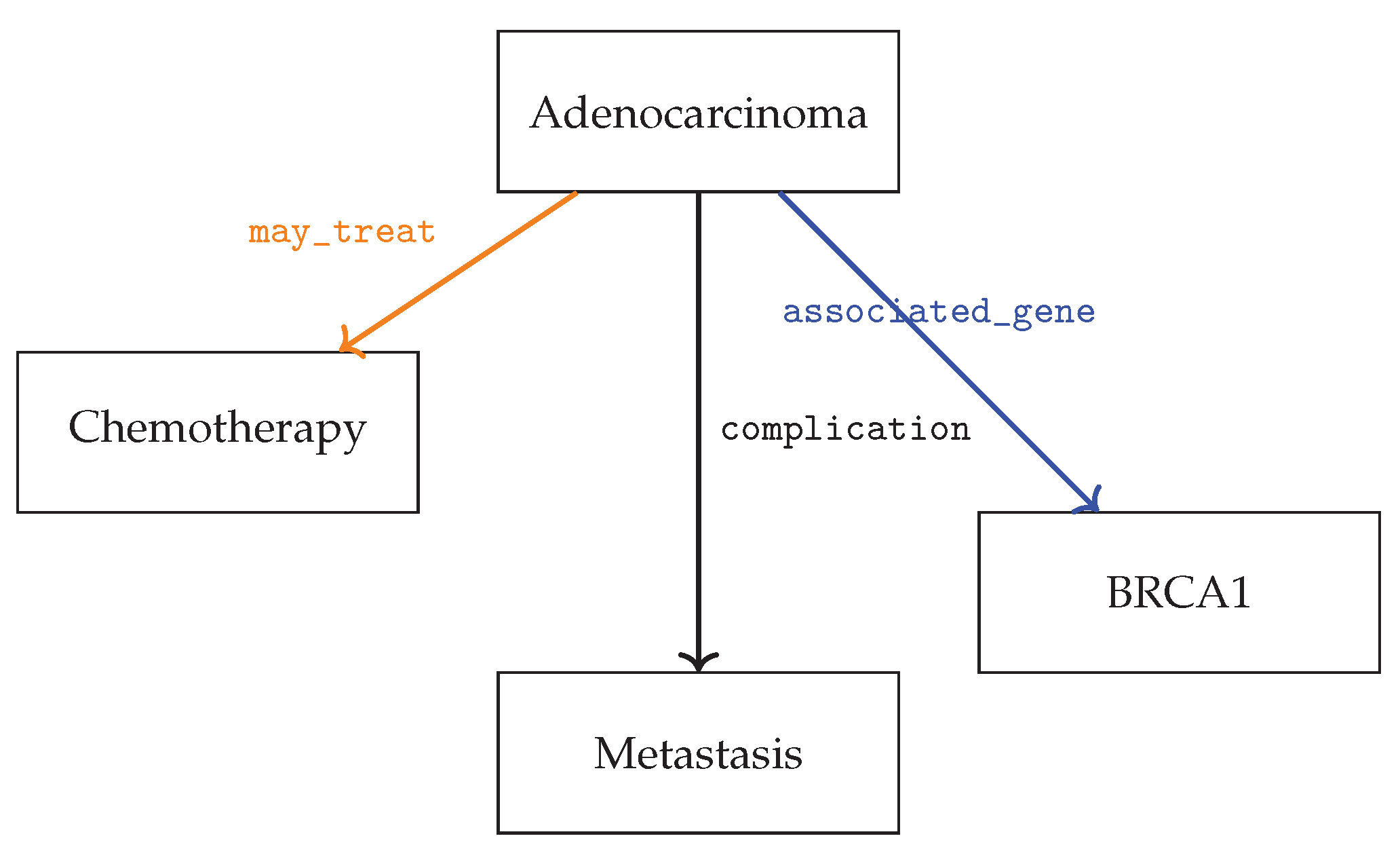
5.4. Case Study
6. Conclusion and Future Work
- Incorporating human-in-the-loop verification for higher-quality extractions.
- Leveraging clinical ontologies and multi-source KGs to enrich supervision.
- Extending the framework to support dynamic entity/relation type discovery.
- Adapting the KGDA approach to integrate with large generative language models like GPT-3.
References
- Bodenreider, O. The Unified Medical Language System (UMLS): integrating biomedical terminology. Nucleic Acids Research 2004, 32, D267–D270. [Google Scholar] [CrossRef] [PubMed]
- Angeli, G.; Premkumar, M.J.; Manning, C.D. Leveraging linguistic structure for open domain information extraction. In Proceedings of the Proceedings of the 2015 Conference on Empirical Methods in Natural Language Processing (EMNLP), 2015, pp. 344–354.
- Mintz, M.; Bills, S.; Snow, R.; Jurafsky, D. Distant supervision for relation extraction without labeled data. In Proceedings of the Proceedings of the Joint Conference of the 47th Annual Meeting of the ACL and the 4th International Joint Conference on Natural Language Processing (ACL-IJCNLP), 2009, pp. 1003–1011.
- Zhang, D.; Wang, D. Relation classification via recurrent neural network. arXiv 2015. [Google Scholar] [CrossRef]
- Lee, J.; Yoon, W.; Kim, S.; Kim, D.; So, C.H.; Kang, J. BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 2020, 36, 1234–1240. [Google Scholar] [CrossRef] [PubMed]
- Verandani, A.; Smith, J.; Lee, K. Adaptive Knowledge Graph Construction for Oncology Literature. Journal of Biomedical Informatics In press. 2025. [Google Scholar]
- Wang, H.; Zhang, F.; Xie, X.; Guo, M. DKN: Deep knowledge-aware network for news recommendation. In Proceedings of the Proceedings of the 2018 World Wide Web Conference, 2018, pp. 1835–1844.
- Peng, Y.; Yan, S.; Lu, Z. Transfer learning in biomedical natural language processing: An evaluation of BERT and ELMo on ten benchmarking datasets. Bioinformatics 2019, 35, 3055–3062. [Google Scholar]
- Chen, L.; Zhang, Y.; Wang, H. DistilBERT-KG: Lightweight Knowledge Graph-Aware Language Model for Biomedical Relation Extraction. IEEE Transactions on Computational Biology and Bioinformatics 2023. [Google Scholar] [CrossRef]
- Yan, W.; Hu, J.; Sun, S.; Zhang, L. AutoKG: Towards Automated Construction and Refinement of Domain-Specific Biomedical Knowledge Graphs. In Proceedings of the Proceedings of the 2024 Annual Conference of the Association for Computational Linguistics (ACL); 2024. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




