Submitted:
23 July 2025
Posted:
24 July 2025
You are already at the latest version
Abstract
Alkaline pectate lyases have great prospects in industry fields such as degumming of papermaking and textile, etc. In this study, a novel pectinase PelA from a Paenibacillus borealis strain was molecular characterized and enzymatically defined. This enzyme represents an important cluster divergent from the well-characterized Bacillus pectinase, shows molecularly active under alkaline condition, and has the optimal pH=9.5. It could be clustered as endo-(1,4)-pectate lyase, and break the α-1,4 glycosidic bonds of the polygalacturonic acid by trans-elimination mode. The extra addition of metal ion Ca2+ could not improve the enzyme activity. In order to achieved its high-level secretory expression and improve its economic competence in bioapplication, the gene copy number of PelA in host genome was improved by constructing the tandem PelA gene expression cassettes. After cultivation condition optimization, cell growth monitoring, the recombinant strain carrying the multi-copy pelA gene reached the expression level of 7520 U/mL culture in a bioreactor. This study has fulfilled the high-level secretory expression of an alkaline pectinase, facilitated its industrial bioapplication and also could be a reference for future work on the heterologous expression of target genes.
Keywords:
1. Introduction
2. Results
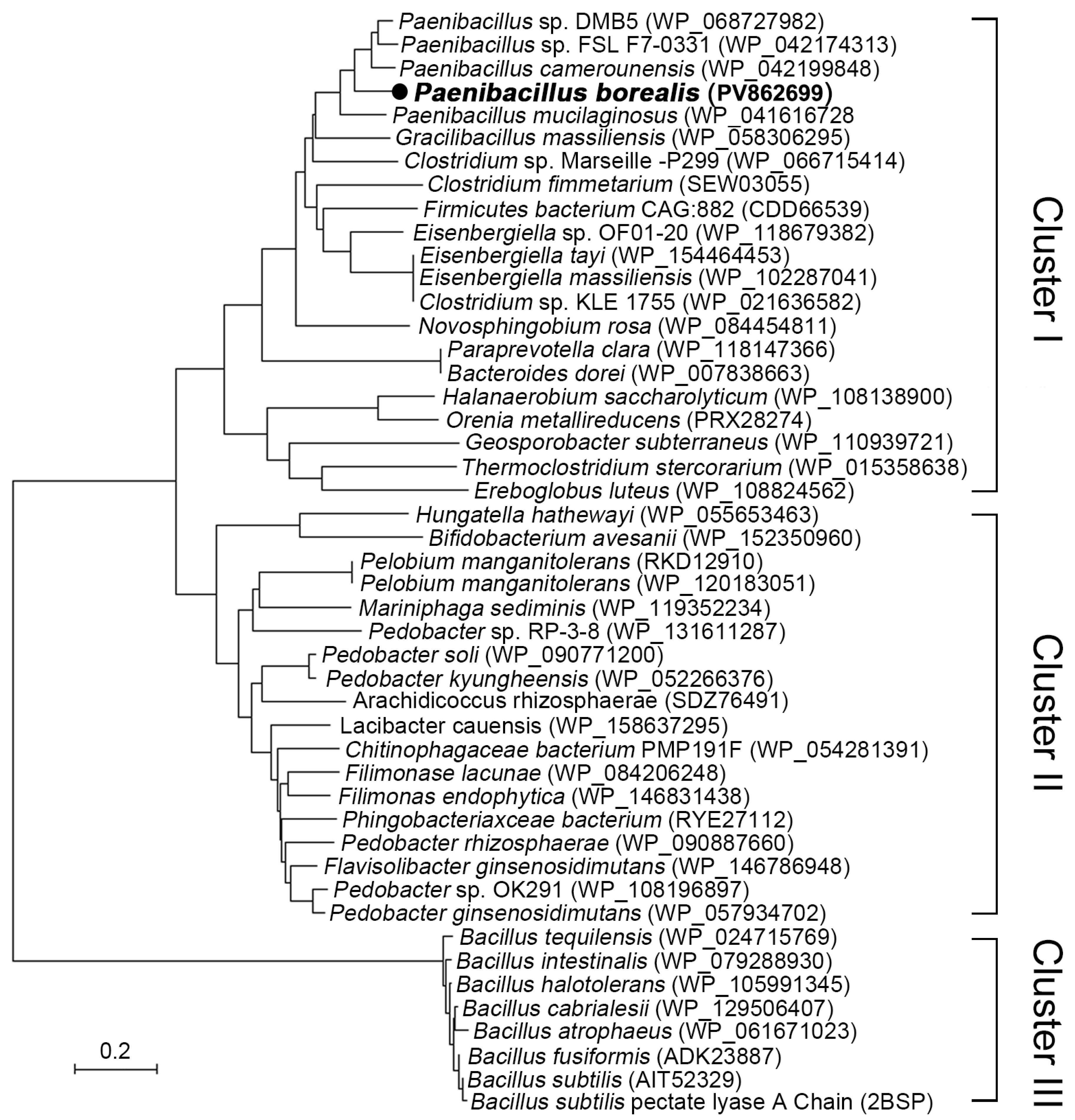
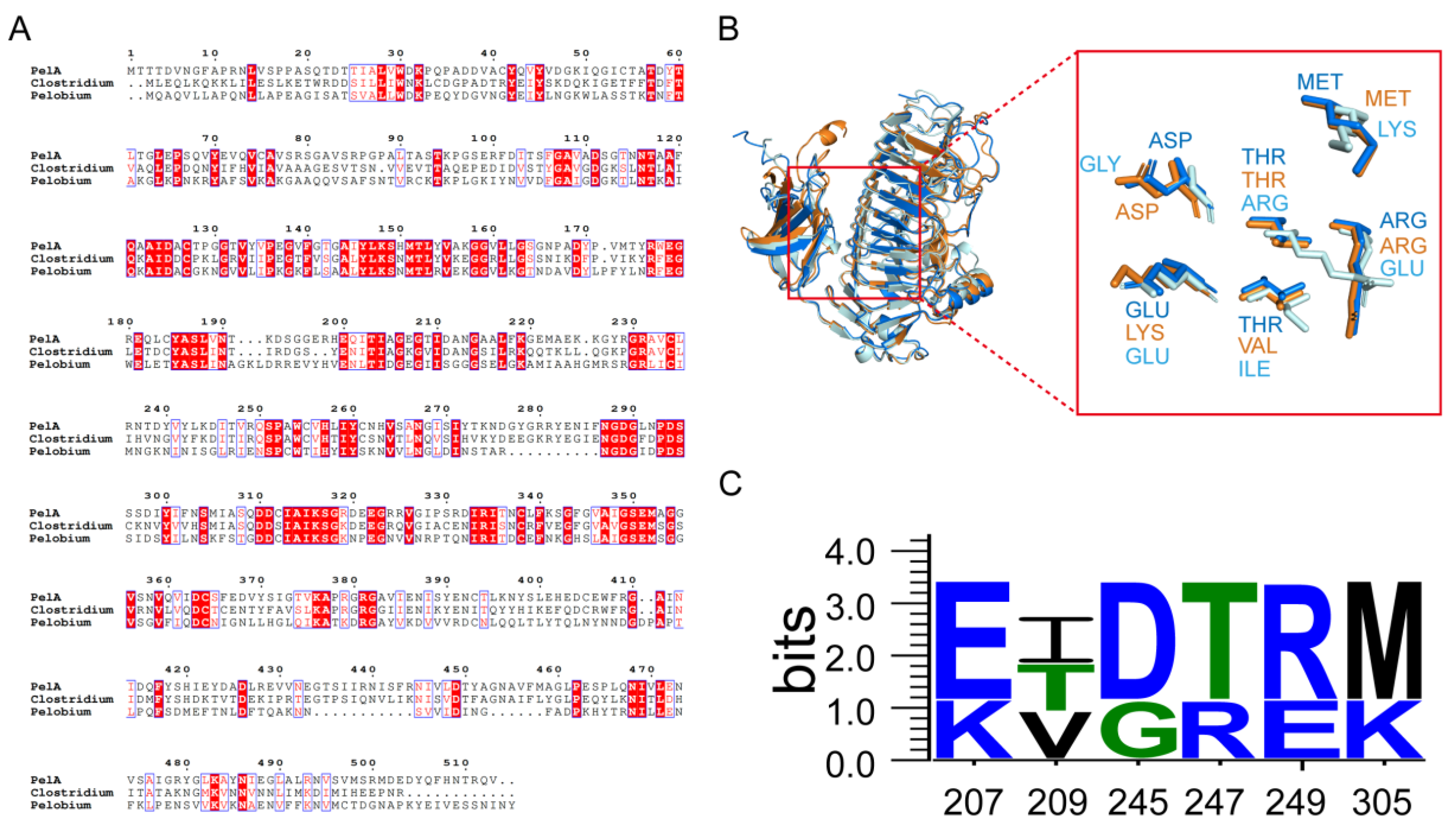
2.1. Phylogeny Analysis, Structure Alignment and Conservation Analysis of PelA from Paenibacillus and Related Genus.
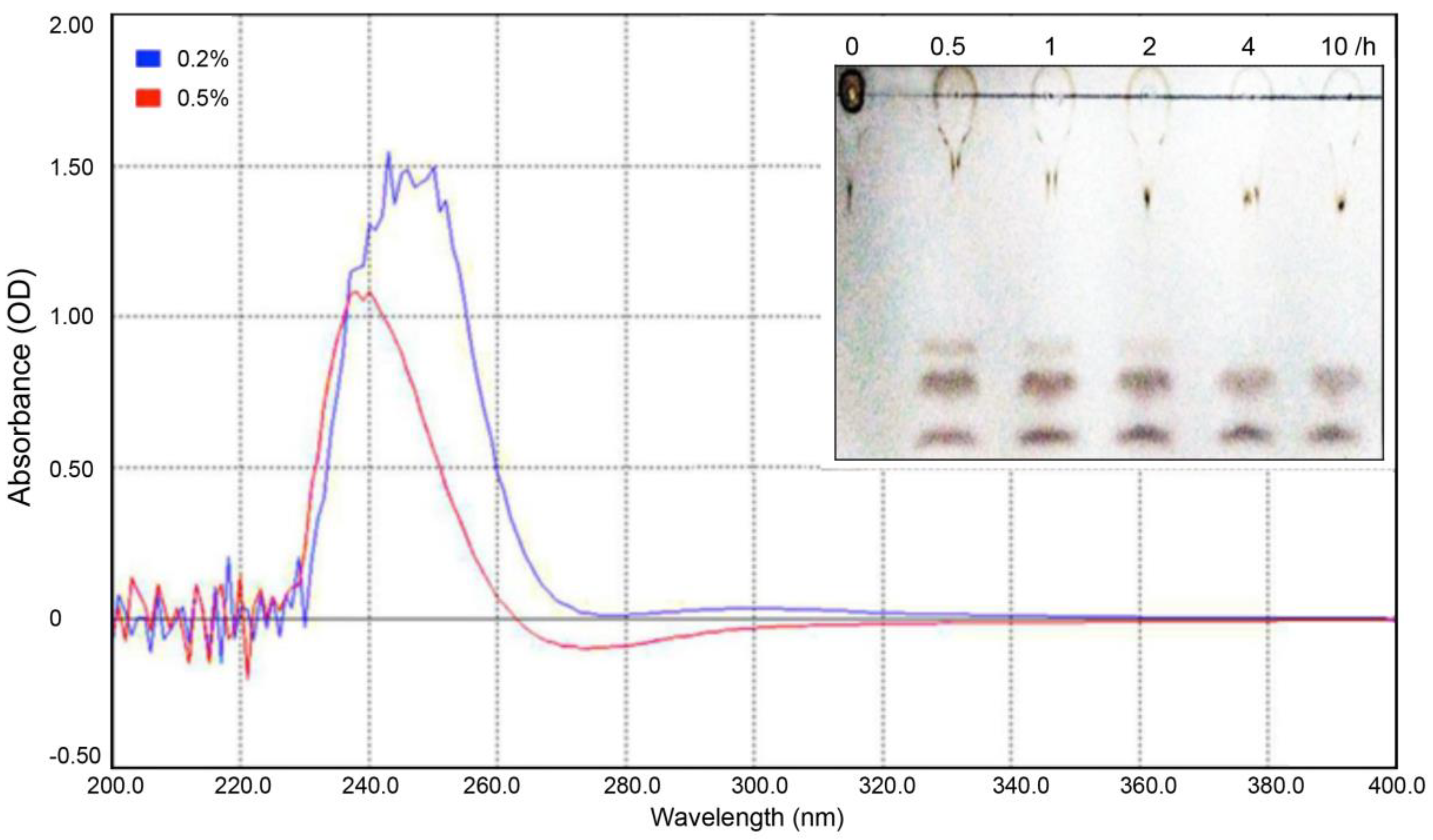
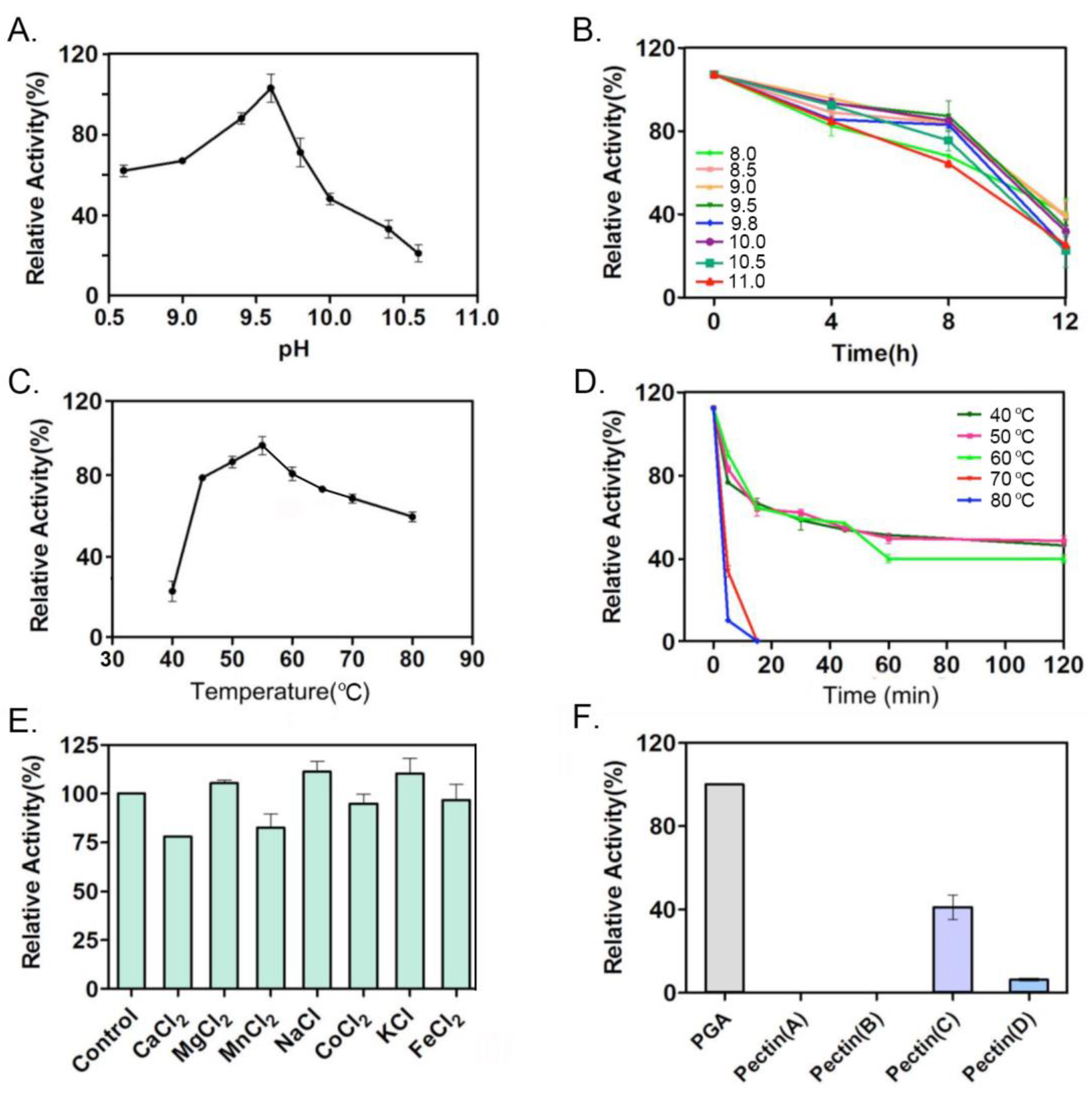
2.2. Enzymatic Definition Reveal PelA Is a Type of Alkaline Endo-Pectate Lyase
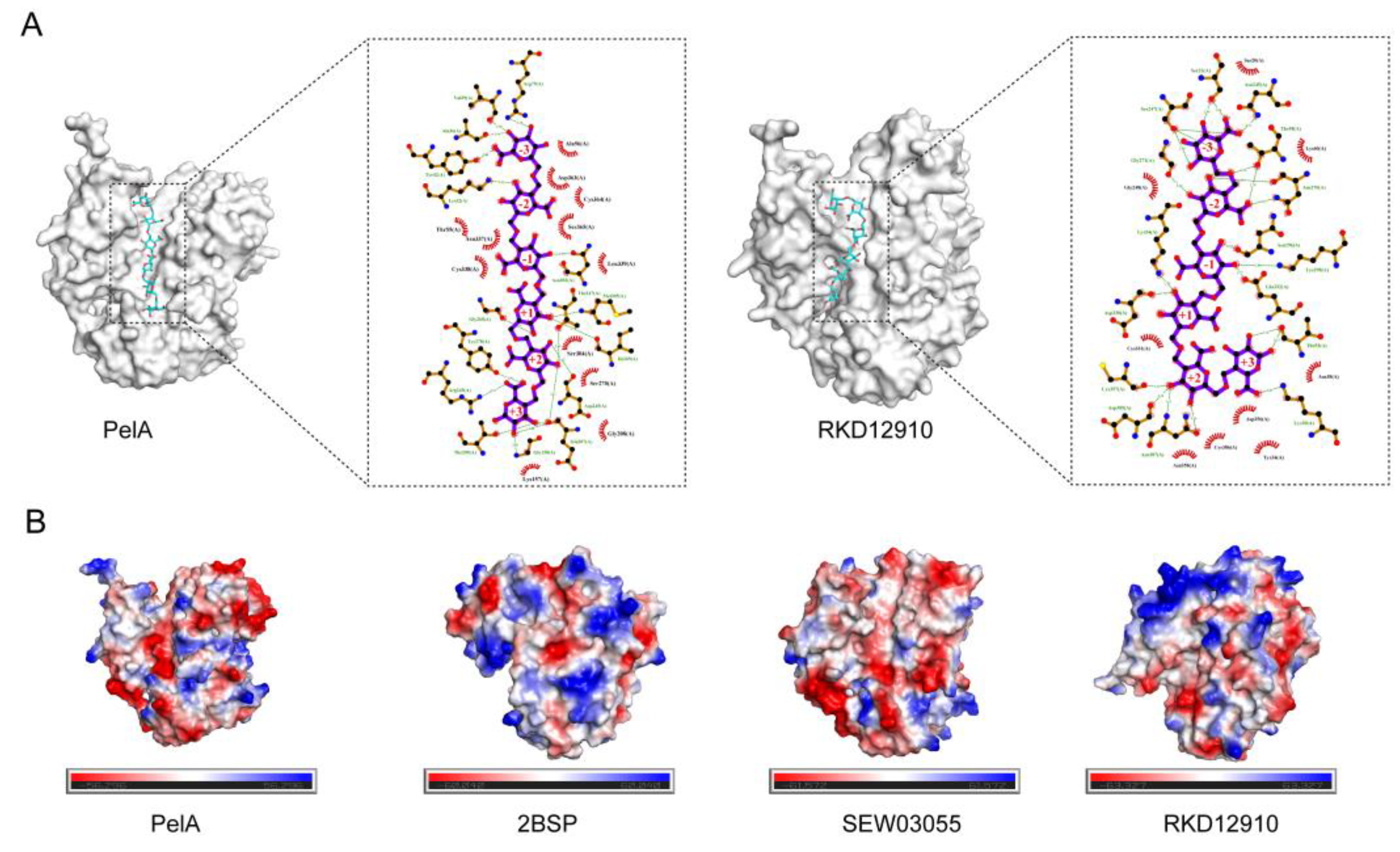
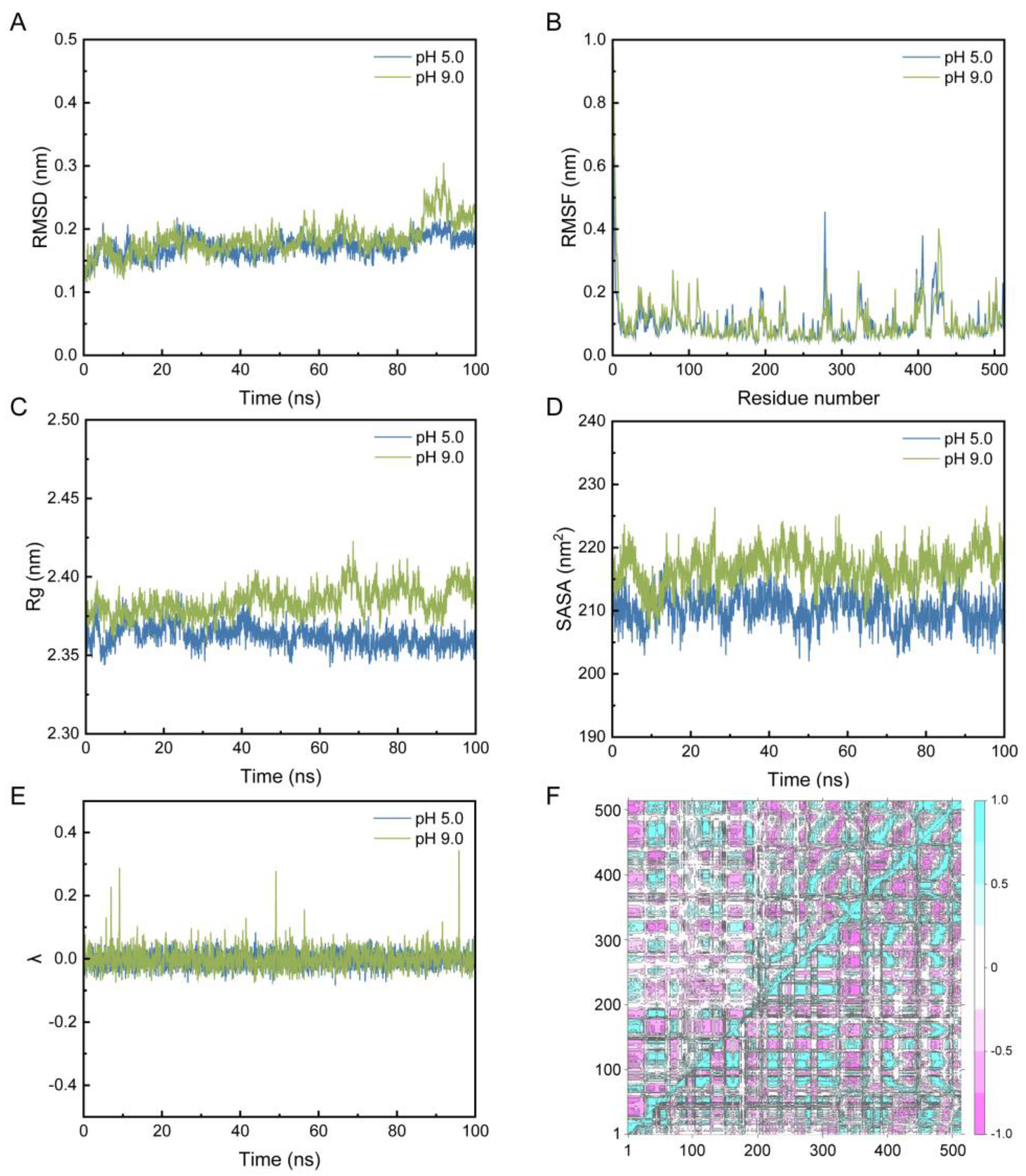
2.3. Molecular Docking and Constant-pH Molecular Dynamics Analysis
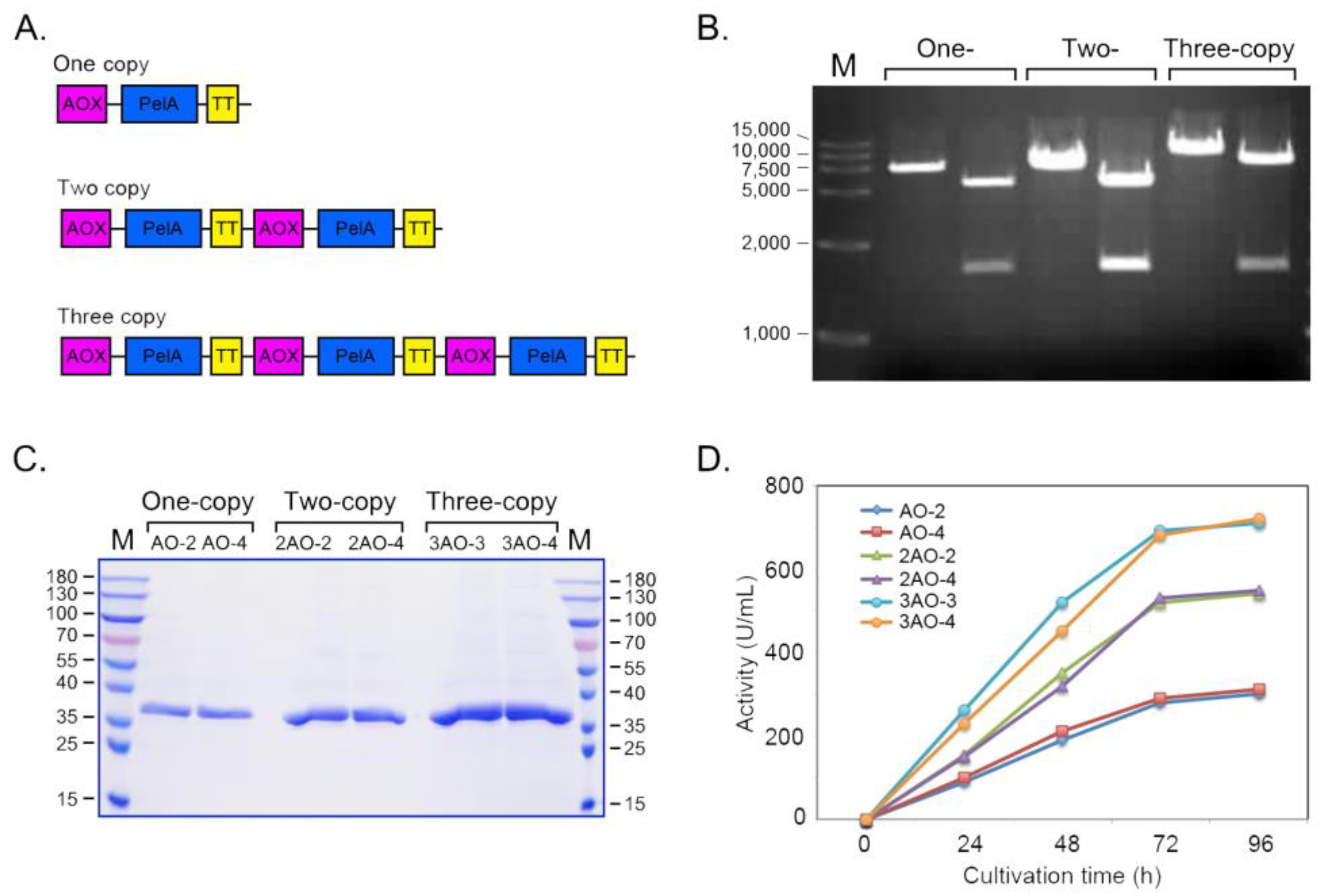
2.4. Improving the Expression Level of PelA by Tandem Expression Cassettes Construction
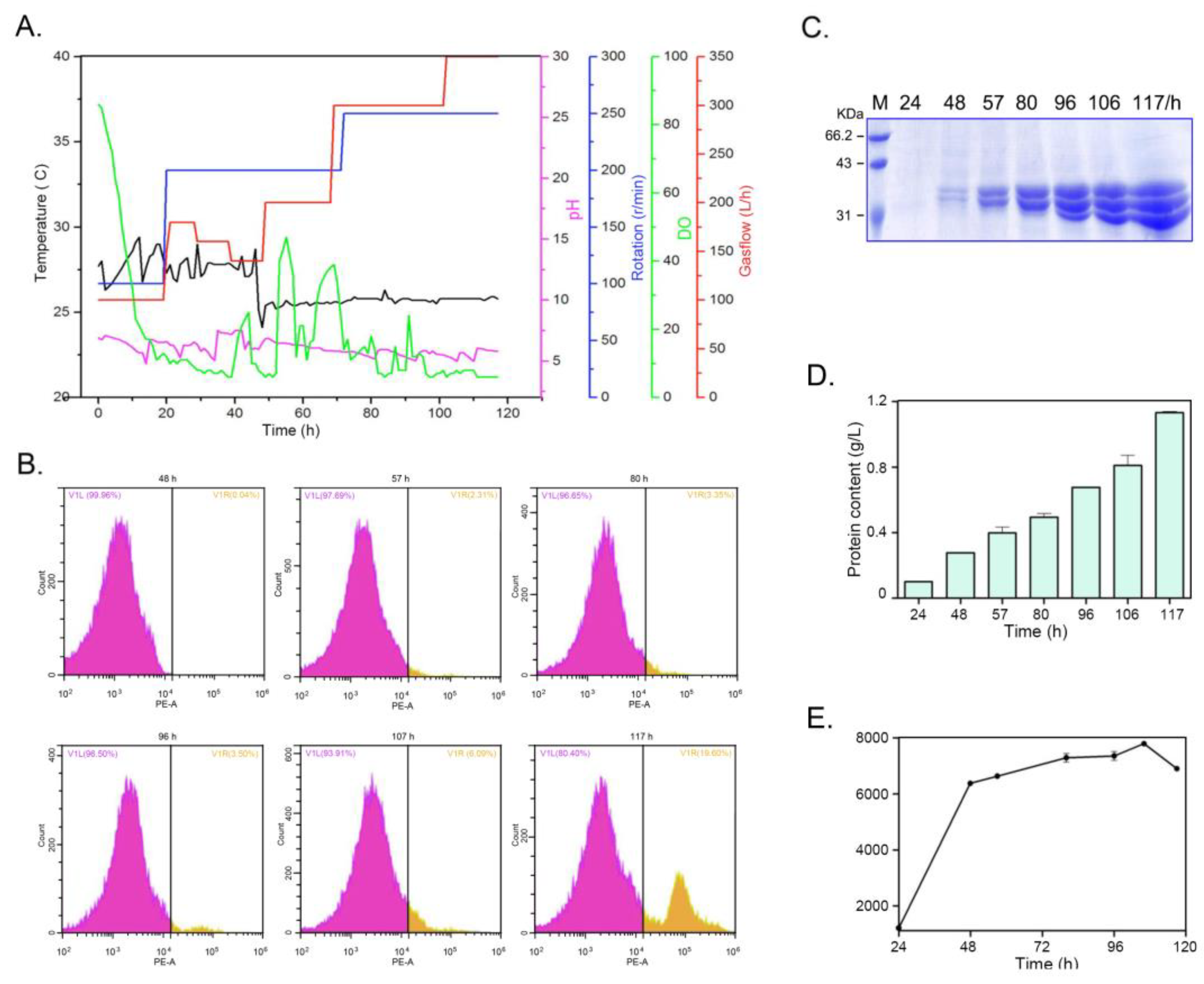
2.5. Expression of PelA in Bioreactor
3. Discussions
3.1. P. borealis Alkaline Pectinase PelA Represent an Important Cluster Divergent from Bacillus pectinase
3.2. Molecular Charactering Revealed PelA Clusters Were Alkaline Adaptation and Structural Divergent from Bacillus pectinase
3.3. Improving the Gene Dosage of PelA and High-Density Cultivation Realized Its High-Level Production
4. Materials and Methods
4.1. Phylogeny Analysis, Structural Alignment and Conservation Analysis of Pectate Lyases
4.2. Gene Cloning, Expression and Enzymatic Characterizing of PelA
4.3. Construction of the Tandem Expression Cassettes of PelA
4.4. Detection of Multiple Copies of the Integration by Quantitative PCR
4.5. Inducible Expression of PelA in Bioreactor and Yeast Cell Viability Counting
4.6. Molecular Docking and Constant-pH Molecular Dynamics Analysis
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Uluisik, S.; Seymour, G.B. Pectate lyases: Their role in plants and importance in fruit ripening [J]. Food Chem. 2020, 309, 125559. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.; P. K. Yadav, D.; Yadav, and K. D. S. Yadav. Pectin lyase: A review. [J]. Process Biochem. 2009, 44, 1–10.
- Reid, I.; Ricard, M. Pectinase in papermaking: solving retention problems in mechanical pulps bleached with hydrogen peroxide [J]. Enzyme Microb Technol. 2000, 26, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.S.; Du, Y.M.; Zhang, J.Y. Degumming of ramie fibers by alkalophilic bacteria and their polysaccharide-degrading enzymes [J]. Bioresour Technol. 2001, 78, 89–94. [Google Scholar] [CrossRef] [PubMed]
- Hoondal, G.S.; Tiwari, R.P.; Tewari, R.; et al. Microbial alkaline pectinases and their industrial applications: a review [J]. Appl Microbiol Biotechnol. 2002, 59, 409–418. [Google Scholar] [PubMed]
- Daniell, H.; Ribeiro, T.; Lin, S.; Saha, P.; McMichael, C.; Chowdhary, R.; Agarwal, A. Validation of leaf and microbial pectinases: commercial launching of a new platform technology [J]. Plant Biotechnol J. 2019, 17, 1154–1166. [Google Scholar] [CrossRef] [PubMed]
- Kamijo, J.; Sakai, K.; Suzuki, H.; Suzuki, K.; Kunitake, E.; Shimizu, M.; Kato, M. Identification and characterization of a thermostable pectate lyase from Aspergillus luchuensis var. saitoi [J]. Food Chem. 2019, 276, 503–510. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.; Dubey, A.K.; Anand, G.; Kumar, R.; Yadav, D. Purification and biochemical characterization of an alkaline pectin lyase from Fusarium decemcellulare MTCC 2079 suitable for Crotalaria juncea fiber retting [J]. J Basic Microbiol. 2014, 54, S161–9. [Google Scholar] [CrossRef] [PubMed]
- Kaur, G.; Kumar, S.; Satyanarayana, T. Production, characterization and application of a thermostable polygalacturonase of a thermophilic mould Sporotrichum thermophile Apinis [J]. Bioresour Technol. 2004, 94, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Guan, Y.; Wang, D.; Lv, C.; Zhang, Y.; Gelbic, I.; Ye, X. Archives of microbiology: screening of pectinase-producing bacteria from citrus peel and characterization of a recombinant pectate lyase with applied potential [J]. Arch Microbiol. 2020, 202, 1005–1013. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Li, S.; Xu, Y.; Wang, J.; Wang, F.; Xin, Y.; Shen, Z.; Zhang, H.; Ma, M.; Liu, H. Production of alkaline pectinase: a case study investigating the use of tobacco stalk with the newly isolated strain Bacillus tequilensis CAS-MEI-2-33 [J]. BMC Biotechnol. 2019, 12, 19,45. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Kang, Z.; Ling, Z.; Cao, W.; Liu, L.; Wang, M.; Du, G.; Chen, J. High-level extracellular production of alkaline polygalacturonate lyase in Bacillus subtilis with optimized regulatory elements [J]. Bioresour Technol. 2013, 146, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Kapoor, M.; Sharma, K.K.; Nair, L.M.; Kuhad, R.C. Production and recovery of an alkaline exo-polygalacturonase from Bacillus subtilis RCK under solid-state fermentation using statistical approach [J]. Bioresour Technol. 2008, 99, 937–945. [Google Scholar] [CrossRef] [PubMed]
- Sharma, N.; Rathore, M.; Sharma, M. Microbial pectinase: sources, characterization and applications. [J]. Rev Environ Sci Bio. 2013, 12, 45–60. [Google Scholar] [CrossRef]
- Elo, S.; Suominen, I.; Kämpfer, P.; Juhanoja, J.; Salkinoja-Salonen, M.; Haahtela, K. Paenibacillus borealis sp. nov., a nitrogen-fixing species isolated from spruce forest humus in Finland [J]. Int J Syst Evol Microbiol. 2001, 51, 535–545. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets [J]. Mol Biol Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Mattanovich, D.; Branduardi, P.; Dato, L.; Gasser, B.; Sauer, M.; Porro, D. Recombinant protein production in yeasts [J]. Methods Mol Biol. 2012, 824, 329–538. [Google Scholar] [PubMed]
- Lee, C.; Kim, J.; Shin, S.G.; et al. Absolute and relative QPCR quantification of plasmid copy number in Escherichia coli [J]. J Biotechnol. 2006, 123, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Abad, S.; Kitz, K.; Hörmann, A.; et al. Real-time PCR-based determination of gene copy numbers in Pichia pastoris [J]. Biotechnol J. 2010, 5, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.K.; Chen, Q.C.; Zhou, B.; Wang, XJ.; Liu, SQ. Manno-oligosaccharide preparation by the hydrolysis of konjac flour with a thermostable endo-mannanase from Talaromyces cellulolyticus [J]. J Appl Microbiol. 2019, 127, 520–532. [Google Scholar] [CrossRef] [PubMed]
- Alahuhta, M.; Brunecky, R.; Chandrayan, P.; Kataeva, I.; Adams, MW.; Himmel, ME.; Lunin, VV. The structure and mode of action of Caldicellulosiruptor bescii family 3 pectate lyase in biomass deconstruction [J]. Acta Crystallogr D Biol Crystallogr. 2013, 69, 534–539. [Google Scholar] [CrossRef] [PubMed]
- Atanasova, L.; Dubey, M.; Grujić, M.; Gudmundsson, M.; Lorenz, C.; Sandgren, M.; Kubicek, C.P.; Jensen, D.F.; Karlsson, M. Evolution and functional characterization of pectate lyase PEL12, a member of a highly expanded Clonostachys rosea polysaccharide lyase 1 family [J]. BMC Microbiol. 2018, 18, 178. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Wu, P.; Jiang, S.; Selvaraj, J.N.; Yang, S.; Zhang, G. A new cold-active and alkaline pectate lyase from antarctic bacterium with high catalytic efficiency [J]. Appl Microbiol Biotechnol. 2019, 103, 5231–5241. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Rao, L.; Xue, Y.; Zhou, C.; Zhang, Y.; Ma, Y. Cloning, expression, and characterization of a highly active alkaline pectate lyase from alkaliphilic Bacillus sp. N16-5 [J]. J Microbiol Biotechnol. 2010, 20, 670–677. [Google Scholar] [CrossRef] [PubMed]
- Bekli, S.; Aktas, B.; Gencer, D.; Aslim, B. Biochemical and molecular characterizations of a novel ph- and temperature-stable pectate lyase from Bacillus amyloliquefaciens S6 for industrial application [J]. Mol Biotechnol. 2019, 61, 681–693. [Google Scholar] [CrossRef] [PubMed]
- Ko, C.H.; Tsai, C.H.; Tu, J.; Tang, S.H.; Liu, C.C. Expression and thermostability of Paenibacillus campinasensis BL11 pectate lyase and its applications in bast fibre processing. [J]. Ann Appl Biol. 2011, 158, 218–225. [Google Scholar] [CrossRef]
- Yuan, Y.; Zhang, X.Y.; Zhao, Y.; Zhang, H.; Zhou, Y.F.; Gao, J. A novel PL9 pectate lyase from Paenibacillus polymyxa KF-1: cloning, expression, and its application in pectin degradation [J]. Int J Mol Sci. 2019, 22, 20, 3060. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Cao, Y.; Xue, Y.; Liu,W.; Ju, J.; Ma, Y. Structure of an alkaline pectate lyase and rational engineering with improved thermo-alkaline stability for efficient ramie degumming [J]. Int. J. Mol. Sci. 2022, 24, 538. [CrossRef] [PubMed]
- Zhou, C.; Ye, J.; Xue, Y.; Ma, Y. Directed evolution and structural analysis of alkaline pectate lyase from the alkaliphilic bacterium Bacillus sp. strain N16-5 to improve its thermostability for efficient ramie degumming [J]. Appl Environ Microbiol. 2015, 1, 81, 5714–5723. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Huang, C.H.; Liu, W.; Ko, T.P.; Xue, Y.; Zhou, C.; Guo, R.T.; Ma, Y. Crystal structure and substrate-binding mode of a novel pectate lyase from alkaliphilic Bacillus sp. N16-5 [J]. Biochem Biophys Res Commun. 2012, 6, 420, 269–274. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Feulner, P.G.D.; Eizaguirre, C.; Lenz, TL.; Bornberg-Bauer, E.; Milinski, M.; Reusch, TBH.; Chain, FJJ. Genome-wide genotype-expression relationships reveal both copy number and single nucleotide differentiation contribute to differential gene expression between stickleback ecotypes [J]. Genome Biol Evol. 2019, 11, 2344–2359.
- Song, X.; Shao, C.; Guo, Y.; Wang, Y.; Cai, J. Improved the expression level of active transglutaminase by directional increasing copy of mtg gene in Pichia pastoris [J]. BMC Biotechnol. 2019, 30, 19, 54. [Google Scholar] [CrossRef] [PubMed]
- Klug-Santner, B.G.; Schnitzhofer, W.; Vrsanska, M.; et al. Purification and characterization of a new bioscouring pectate lyase from Bacillus pumilus BK2 [J]. J Biotechnol. 2006, 121, 390–401. [Google Scholar] [CrossRef] [PubMed]
| Ct mean of pelA gene | Copy number pelA gene in the reaction | Ct mean of gapdh gene | Copy number of gapdh gene in the reaction | Copy number of pelA in a genome | |
|---|---|---|---|---|---|
| AO-2 | 4.738 | 5.265×10^7 | 16.700 | 4.930×10^7 | 1.068 |
| AO-4 | 5.096 | 5.008×10^7 | 16.501 | 5.090×10^7 | 0.984 |
| 2AO-2 | 3.927 | 9.930×10^7 | 16.503 | 5.010×10^7 | 1.982 |
| 2AO-4 | 3.922 | 9.955×10^7 | 16.495 | 5.079×10^7 | 1.960 |
| 3AO-3 | 3.291 | 1.525×10^8 | 16.498 | 5.095×10^7 | 2.993 |
| 3AO-4 | 3.304 | 1.513×10^8 | 16.501 | 5.058×10^7 | 2.992 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




