Submitted:
02 March 2026
Posted:
05 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. From Standalone Agents to AI Scientist Systems: Progress and Bottlenecks
3. Why Protocol Ecosystems Matter: MCP as an Interoperability Layer
4. Scaling Many-Agent Discovery Needs Human-Inspired Organizational Design
4.1. What Breaks When We Scale Beyond Small Agent Teams
4.2. Organizational Primitives as First-Class Interfaces
4.3. A Minimal Hierarchy is an Information Contract, Not Just a Role Chart
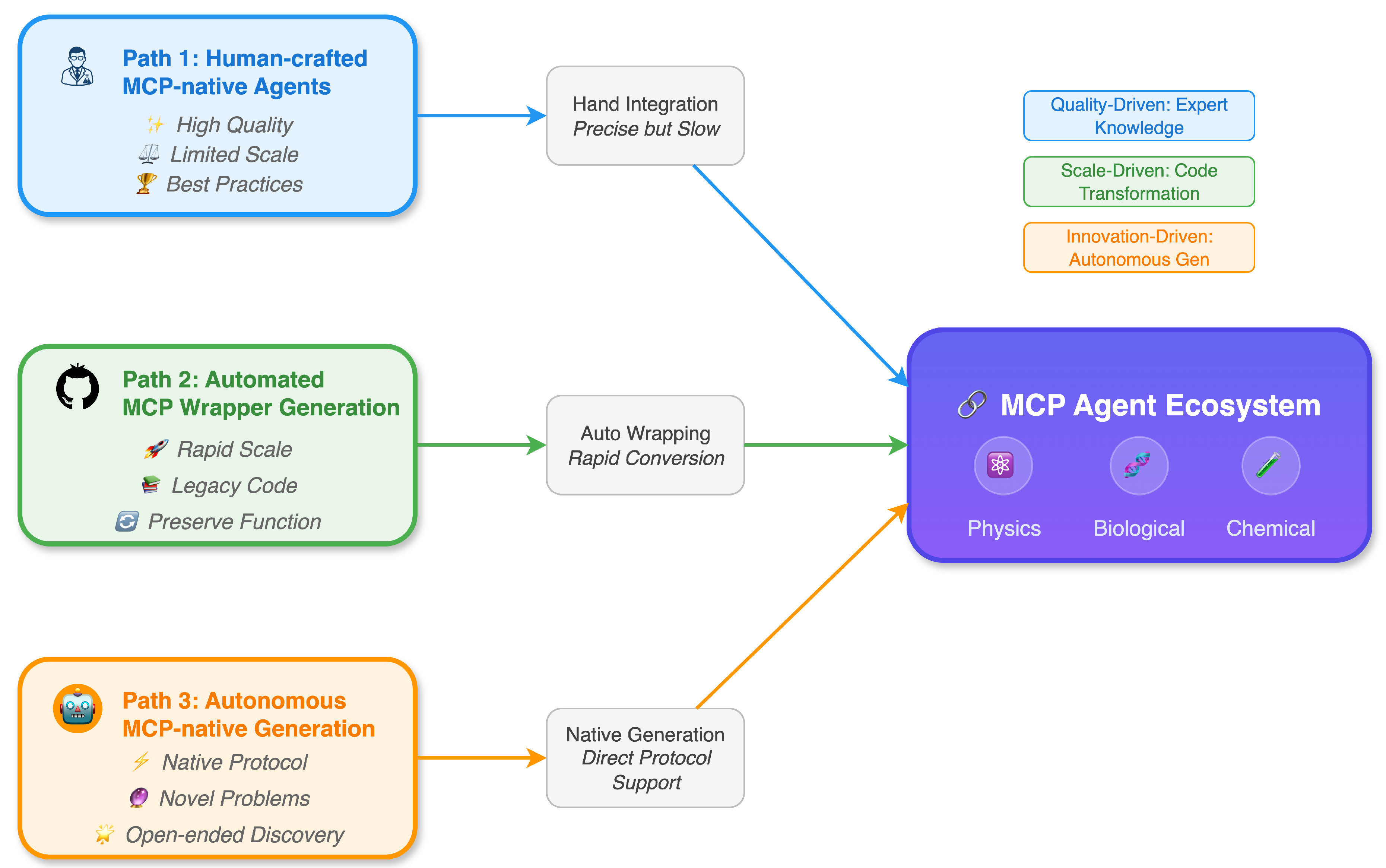
5. Roadmap: Three Pathways to Grow an MCP-Native Scientific Agent Ecosystem
6. Discussion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
| 1 |
N refers to the number of models/agents, and M refers to the number of tools. |
References
- Yao, S.; Zhao, J.; Yu, D.; Du, N.; Shafran, I.; Narasimhan, K.R.; Cao, Y. React: Synergizing reasoning and acting in language models. In Proceedings of the The eleventh international conference on learning representations, 2022. [Google Scholar]
- Schick, T.; Dwivedi-Yu, J.; Dessì, R.; Raileanu, R.; Lomeli, M.; Hambro, E.; Zettlemoyer, L.; Cancedda, N.; Scialom, T. Toolformer: Language models can teach themselves to use tools. Advances in neural information processing systems 2023, 36, 68539–68551. [Google Scholar]
- Wu, Q.; Bansal, G.; Zhang, J.; Wu, Y.; Li, B.; Zhu, E.; Jiang, L.; Zhang, X.; Zhang, S.; Liu, J.; et al. Autogen: Enabling next-gen llm applications via multi-agent conversation. arXiv 2023, arXiv:2308.08155. [Google Scholar]
- Li, G.; Hammoud, H.; Itani, H.; Khizbullin, D.; Ghanem, B. Camel: Communicative agents for" mind" exploration of large language model society. Advances in Neural Information Processing Systems 2023, 36, 51991–52008. [Google Scholar]
- Hong, S.; Zheng, X.; Chen, J.; Cheng, Y.; Wang, J.; Zhang, C.; Wang, Z.; Yau, S.K.S.; Lin, Z.; Zhou, L.; et al. Metagpt: Meta programming for multi-agent collaborative framework. arXiv 2023, arXiv:2308.00352. [Google Scholar]
- Lu, C.; Lu, C.; Lange, R.T.; Foerster, J.; Clune, J.; Ha, D. The ai scientist: Towards fully automated open-ended scientific discovery. arXiv 2024, arXiv:2408.06292. [Google Scholar] [CrossRef]
- Yamada, Y.; Lange, R.T.; Lu, C.; Hu, S.; Lu, C.; Foerster, J.; Clune, J.; Ha, D. The ai scientist-v2: Workshop-level automated scientific discovery via agentic tree search. arXiv 2025, arXiv:2504.08066. [Google Scholar]
- Gottweis, J.; Weng, W.H.; Daryin, A.; Tu, T.; Palepu, A.; Sirkovic, P.; Myaskovsky, A.; Weissenberger, F.; Rong, K.; Tanno, R.; et al. Towards an AI co-scientist. arXiv 2025, arXiv:2502.18864. [Google Scholar] [CrossRef]
- Schmidgall, S.; Su, Y.; Wang, Z.; Sun, X.; Wu, J.; Yu, X.; Liu, J.; Moor, M.; Liu, Z.; Barsoum, E. Agent laboratory: Using llm agents as research assistants. In Findings of the Association for Computational Linguistics: EMNLP; 2025; pp. 5977–6043. [Google Scholar]
- Baek, J.; Jauhar, S.K.; Cucerzan, S.; Hwang, S.J. Researchagent: Iterative research idea generation over scientific literature with large language models. Proceedings of the Proceedings of the 2025 Conference of the Nations of the Americas Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers) 2025, 6709–6738. [Google Scholar]
- Team, N.; Zhang, B.; Feng, S.; Yan, X.; Yuan, J.; Yu, Z.; He, X.; Huang, S.; Hou, S.; Nie, Z.; et al. NovelSeek: When Agent Becomes the Scientist–Building Closed-Loop System from Hypothesis to Verification. arXiv 2025, arXiv:2505.16938. [Google Scholar]
- King, R.D.; Rowland, J.; Aubrey, W.; Liakata, M.; Markham, M.; Soldatova, L.N.; Whelan, K.E.; Clare, A.; Young, M.; Sparkes, A.; et al. The robot scientist Adam. Computer 2009, 42, 46–54. [Google Scholar] [CrossRef]
- Stokols, D.; Hall, K.L.; Taylor, B.K.; Moser, R.P. The science of team science: overview of the field and introduction to the supplement. American journal of preventive medicine 2008, 35, S77–S89. [Google Scholar] [CrossRef] [PubMed]
- Council, N.R.; et al. Enhancing the effectiveness of team science. In Enhancing the Effectiveness of Team Science; National Research Council: Washington, 2015; p. 19007. [Google Scholar]
- Wilson, G.; Aruliah, D.A.; Brown, C.T.; Chue Hong, N.P.; Davis, M.; Guy, R.T.; Haddock, S.H.; Huff, K.D.; Mitchell, I.M.; Plumbley, M.D.; et al. Best practices for scientific computing. PLoS biology 2014, 12, e1001745. [Google Scholar] [CrossRef] [PubMed]
- Simon, H.A. The architecture of complexity. In The Roots of Logistics; Springer, 2012; pp. 335–361. [Google Scholar]
- Parnas, D.L. On the criteria to be used in decomposing systems into modules. Communications of the ACM 1972, 15, 1053–1058. [Google Scholar] [CrossRef]
- Baldwin, C.Y.; Clark, K.B. Design rules . In The power of modularity; MIT press, 2000; Volume 1. [Google Scholar]
- Pan, M.Z.; Arabzadeh, N.; Cogo, R.; Zhu, Y.; Xiong, A.; Agrawal, L.A.; Mao, H.; Shen, E.; Pallerla, S.; Patel, L.; et al. Measuring agents in production. arXiv 2025, arXiv:2512.04123. [Google Scholar] [CrossRef]
- Wang, L.; Ma, C.; Feng, X.; Zhang, Z.; Yang, H.; Zhang, J.; Chen, Z.; Tang, J.; Chen, X.; Lin, Y.; et al. A survey on large language model based autonomous agents. Frontiers of Computer Science 2024, 18, 186345. [Google Scholar] [CrossRef]
- Peng, R.D. Reproducible research in computational science. Science 2011, 334, 1226–1227. [Google Scholar] [CrossRef] [PubMed]
- Sandve, G.K.; Nekrutenko, A.; Taylor, J.; Hovig, E. Ten simple rules for reproducible computational research. PLoS computational biology 2013, 9, e1003285. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, M.D.; Dumontier, M.; Aalbersberg, I.J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.W.; da Silva Santos, L.B.; Bourne, P.E.; et al. The FAIR Guiding Principles for scientific data management and stewardship. Scientific data 2016, 3, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Horling, B.; Lesser, V. A survey of multi-agent organizational paradigms. The Knowledge engineering review 2004, 19, 281–316. [Google Scholar] [CrossRef]
- Ehtesham, A.; Singh, A.; Gupta, G.K.; Kumar, S. A survey of agent interoperability protocols: Model context protocol (mcp), agent communication protocol (acp), agent-to-agent protocol (a2a), and agent network protocol (anp). arXiv 2025, arXiv:2505.02279. [Google Scholar]
- Anthropic. Model Context Protocol Specification (Version 2025-11-25) . 2025. [Google Scholar]
- The Linux Foundation (A2A Protocol Project). Agent2Agent (A2A) Protocol Documentation and Specification. Official documentation site. Accessed 2026-02-26. 202. Official documentation site.
- Microsoft. Language Server Protocol Accessed: 2026-02-11. 2021.
- Fielding, R.T. Architectural styles and the design of network-based software architectures; University of California: Irvine, 2000. [Google Scholar]
- Malone, T.W.; Crowston, K. The interdisciplinary study of coordination. ACM Computing Surveys (CSUR) 1994, 26, 87–119. [Google Scholar] [CrossRef]
- Smith, R.G. The contract net protocol: High-level communication and control in a distributed problem solver. In Readings in distributed artificial intelligence; Elsevier, 1988; pp. 357–366. [Google Scholar]
- Grosz, B.J.; Kraus, S. Collaborative plans for complex group action. Artificial Intelligence 1996, 86, 269–357. [Google Scholar] [CrossRef]
- Xu, W.; Huang, C.; Gao, S.; Shang, S. LLM-Based Agents for Tool Learning: A Survey: W. Xu et al. In Data Science and Engineering; 2025; pp. 1–31. [Google Scholar]
- Shah, C.; White, R.W. Agents are not enough. Computer 2025, 58, 87–92. [Google Scholar] [CrossRef]
- Wilson, G.; Bryan, J.; Cranston, K.; Kitzes, J.; Nederbragt, L.; Teal, T.K. Good enough practices in scientific computing. PLoS computational biology 2017, 13, e1005510. [Google Scholar] [CrossRef] [PubMed]
- Kurtzer, G.M.; Sochat, V.; Bauer, M.W. Singularity: Scientific containers for mobility of compute. PloS one 2017, 12, e0177459. [Google Scholar] [CrossRef] [PubMed]
- Köster, J.; Rahmann, S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 2012, 28, 2520–2522. [Google Scholar] [CrossRef] [PubMed]
- Di Tommaso, P.; Chatzou, M.; Floden, E.W.; Barja, P.P.; Palumbo, E.; Notredame, C. Nextflow enables reproducible computational workflows. Nature biotechnology 2017, 35, 316–319. [Google Scholar] [CrossRef] [PubMed]
- Crusoe, M.R.; Abeln, S.; Iosup, A.; Amstutz, P.; Chilton, J.; Tijanić, N.; Ménager, H.; Soiland-Reyes, S.; Gavrilović, B.; Goble, C.; et al. Methods included: standardizing computational reuse and portability with the common workflow language. Communications of the ACM 2022, 65, 54–63. [Google Scholar] [CrossRef]
- Guo, Z.; Cheng, S.; Wang, H.; Liang, S.; Qin, Y.; Li, P.; Liu, Z.; Sun, M.; Liu, Y. Stabletoolbench: Towards stable large-scale benchmarking on tool learning of large language models. Proceedings of the Findings of the Association for Computational Linguistics: ACL 2024, 2024, 11143–11156. [Google Scholar]
- Conway, M.E. How do committees invent. Datamation 1968, 14, 28–31. [Google Scholar]
- Park, J.S.; O’Brien, J.; Cai, C.J.; Morris, M.R.; Liang, P.; Bernstein, M.S. Generative agents: Interactive simulacra of human behavior. In Proceedings of the Proceedings of the 36th annual acm symposium on user interface software and technology, 2023; pp. 1–22. [Google Scholar]
- Belhajjame, K.; B’Far, R.; Cheney, J.; Coppens, S.; Cresswell, S.; Gil, Y.; Groth, P.; Klyne, G.; Lebo, T.; McCusker, J.; et al. Prov-dm: The prov data model. W3C Recommendation 2013, 14, 15–16. [Google Scholar]
- Huang, D.; Lapp, H. Software engineering as instrumentation for the long tail of scientific software. arXiv 2013, arXiv:1309.1806. [Google Scholar] [CrossRef]
- Ouyang, C.; Yue, L.; Di, S.; Zheng, L.; Yue, L.; Pan, S.; Yin, J.; Zhang, M.L. Code2MCP: Transforming Code Repositories into MCP Services. arXiv 2025, arXiv:2509.05941. [Google Scholar]
- Wang, G.; Xie, Y.; Jiang, Y.; Mandlekar, A.; Xiao, C.; Zhu, Y.; Fan, L.; Anandkumar, A. Voyager: An open-ended embodied agent with large language models. arXiv 2023, arXiv:2305.16291. [Google Scholar] [CrossRef]
- Tambe, M. Towards flexible teamwork. Journal of artificial intelligence research 1997, 7, 83–124. [Google Scholar] [CrossRef]
- Groth, P.; Moreau, L. PROV-overview. An overview of the PROV family of documents. 2013. [Google Scholar]
- Hu, S.; Lu, C.; Clune, J. Automated design of agentic systems. arXiv 2024, arXiv:2408.08435. [Google Scholar] [CrossRef]
- Zhang, J.; Xiang, J.; Yu, Z.; Teng, F.; Chen, X.; Chen, J.; Zhuge, M.; Cheng, X.; Hong, S.; Wang, J.; et al. Aflow: Automating agentic workflow generation. arXiv 2024, arXiv:2410.10762. [Google Scholar] [CrossRef]
- Zhang, J.; Hu, S.; Lu, C.; Lange, R.; Clune, J. Darwin Godel Machine: Open-Ended Evolution of Self-Improving Agents. arXiv 2025, arXiv:2505.22954. [Google Scholar] [CrossRef]
- Shinn, N.; Cassano, F.; Gopinath, A.; Narasimhan, K.; Yao, S. Reflexion: Language agents with verbal reinforcement learning. Advances in neural information processing systems 2023, 36, 8634–8652. [Google Scholar]
- Ioannidis, J.P. Why most published research findings are false. PLoS medicine 2005, 2, e124. [Google Scholar] [CrossRef] [PubMed]
- Munafò, M.R.; Nosek, B.A.; Bishop, D.V.; Button, K.S.; Chambers, C.D.; Percie du Sert, N.; Simonsohn, U.; Wagenmakers, E.J.; Ware, J.J.; Ioannidis, J.P. A manifesto for reproducible science. Nature human behaviour 2017, 1, 0021. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yu, H.; Zhang, H.; Xu, Y.; Lei, X.; Lai, H.; Gu, Y.; Ding, H.; Men, K.; Yang, K.; et al. Agentbench: Evaluating llms as agents. arXiv 2023, arXiv:2308.03688. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




