Submitted:
19 July 2025
Posted:
22 July 2025
Read the latest preprint version here
Abstract
Keywords:
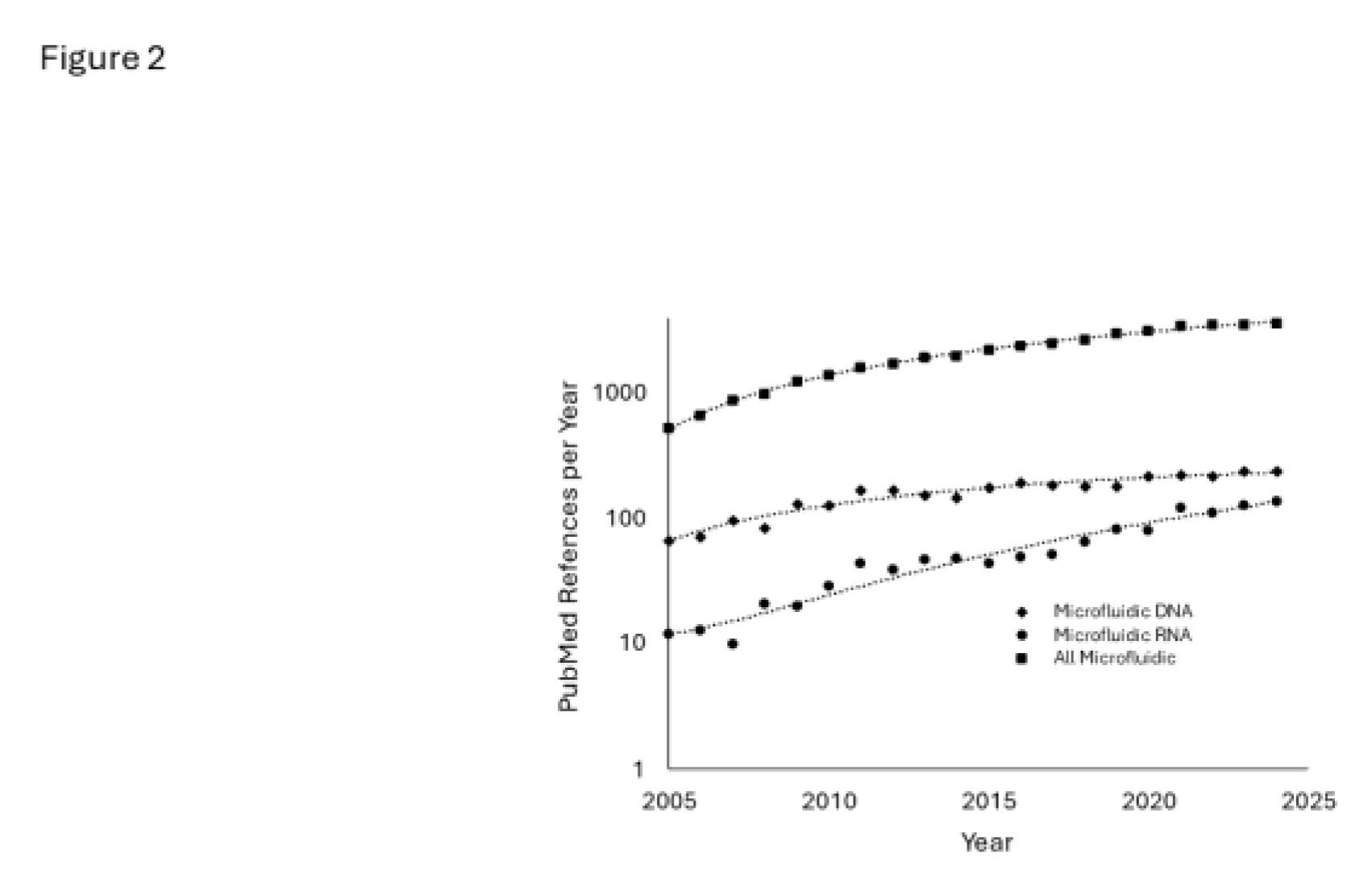
1. Introduction
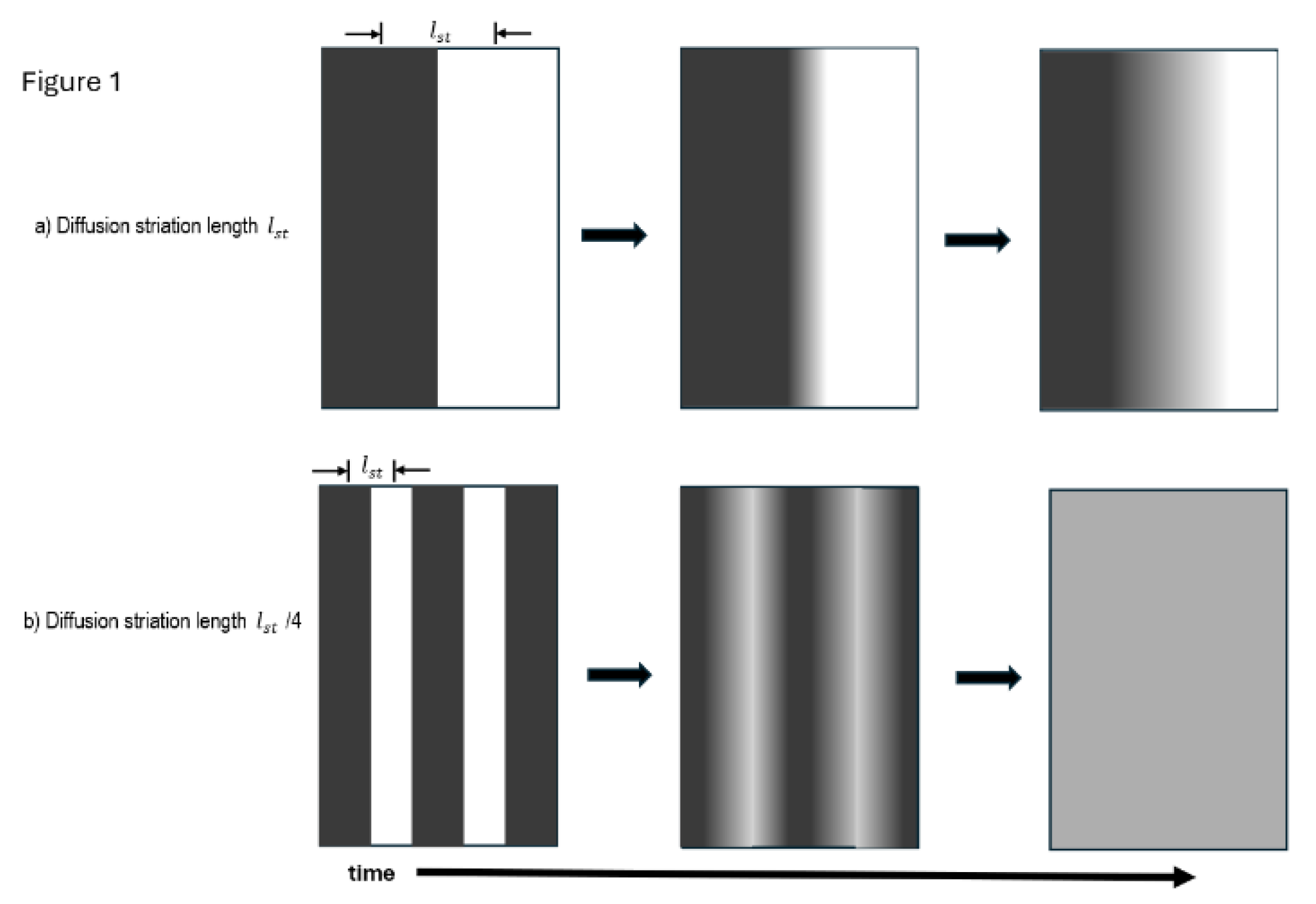
2. Diffusion Mediated Mixing Distances and Time Fundamentals
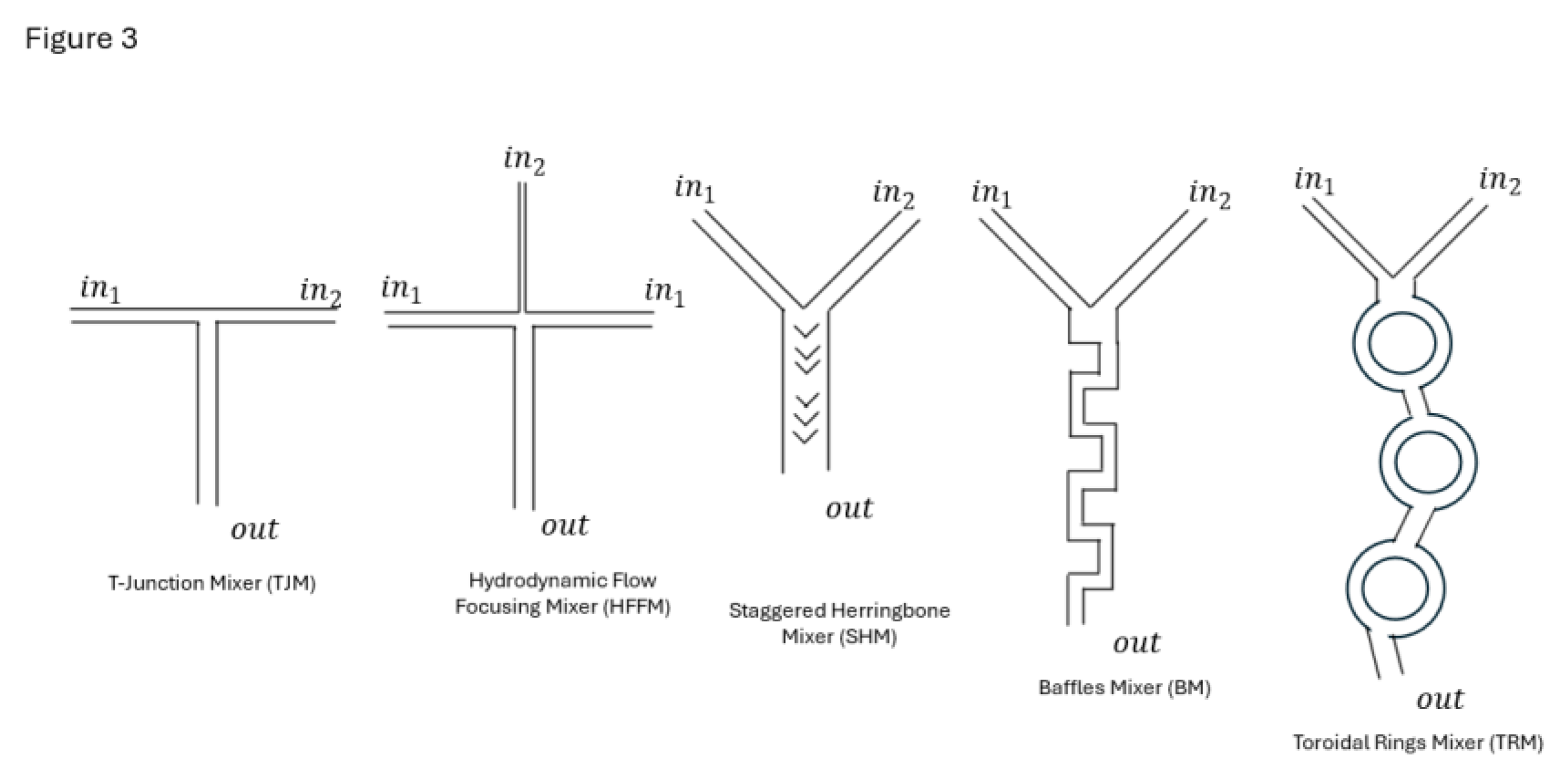
3. Microfluidic Mixer Fundamentals
T-Junction Mixers
Hydrodynamic Flow Focusing Mixers (HFFM)
Staggered Herringbone Mixers (SHM)
Baffles Mixers (BM)
Toroidal Rings Mixers (TRM)
4. Formulation of LNP and mRNA LNP with Microfluidic Mixers
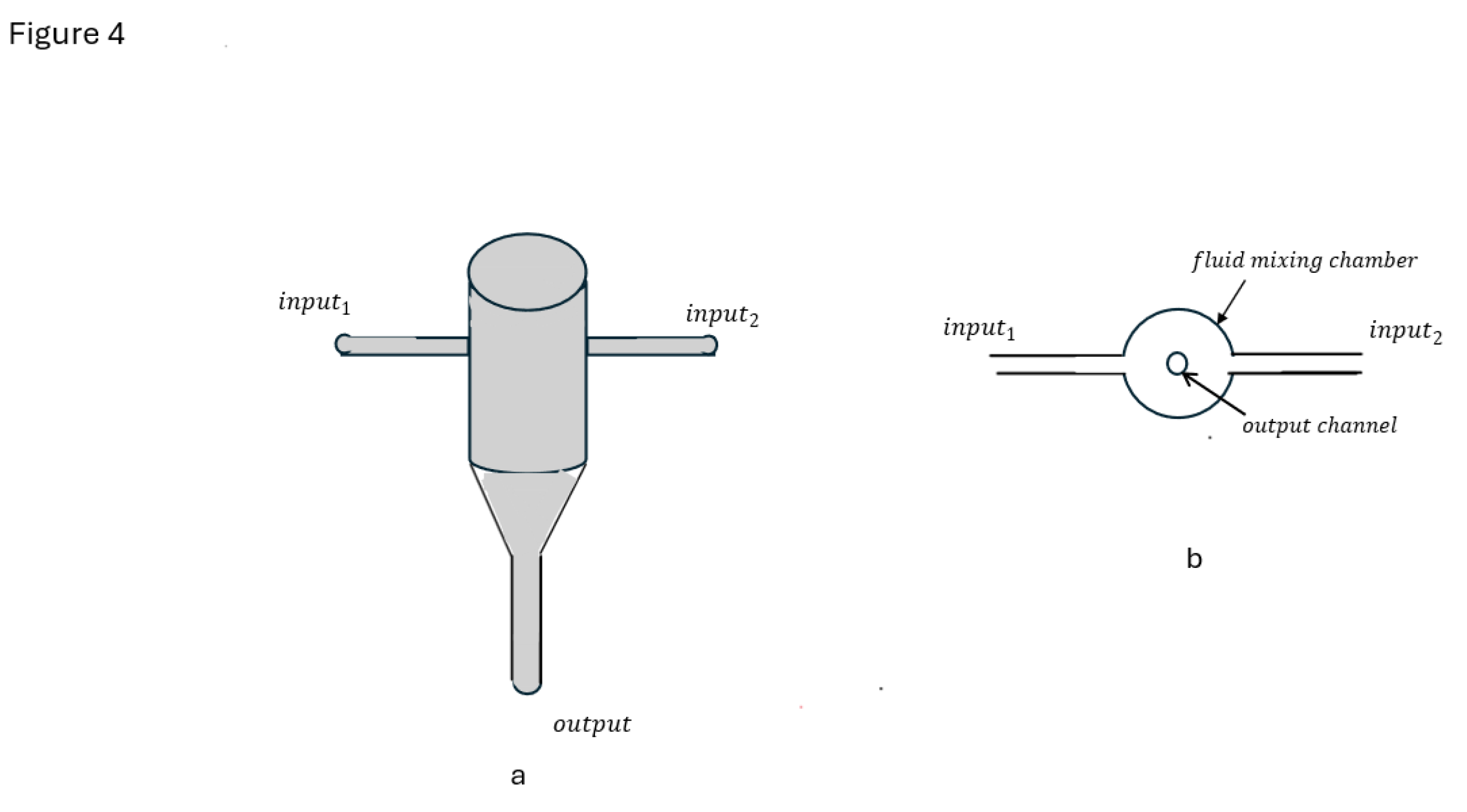
5. Turbulent Flow Mixers Fundamentals
6. Formulation of LNP and mRNA LNP by Turbulent Mixing
7. Flash Nanoprecipitation Self-Assembly and RNA Encapsulation
8. LNP Mixing for Discovery, Scale-up, and Clinical Production
| Product Trade Name | Description | Vendor | Website |
|---|---|---|---|
| NanoAssemblr™ Spark™ | Small-scale rapid formulation screening 25 to 250 µL batch volume |
Cytiva | https://www.cytivalifesciences.com/en/us/shop/lipid-nanoparticle-instruments-and-reagents/nanoparticle-formulation-systems?sort=NameAsc&chunk=1 |
| NanoAssemblr™ Ignite™ | Pre-clinical formulation screening, easy to use TRM, 5 to 60 mL batch |
||
| NanoAssemblr™ Blaze™ | Large-scale for process scale-up , includes TFF 0.2 to 10 L batch |
||
| NanoAssemblr™ GMP System | GMP system for clinical supplies 1 to 50 L batch size |
||
| NanoAssemblr™ Commercial Formulation System |
GMP system for large-scale commercial manufacturing |
||
| Tamara | Easy to use, reusable microchips 0.2 to 30 mL batch size | Inside Therapeutics |
https://insidetx.com/product/tamara/ |
| Lipid Nanoparticle Synthesis Pack | Pressure controlled SHM, easy to use. 0.5 ml to 5 L batch size |
Elve Flow | https://www.elveflow.com/microfluidics-application-packs/lipid-nanoparticle-synthesis/ |
| NanoGenerator™ Flex-S | Small-scale discovery screening, multi-sample 1 to 4, 0.1 to 0.5 mL per sample |
PreciGenome LLC | https://www.precigenome.com/ |
| NanoGenerator™ Flex-S Plus | Early discovery, fully automated HT, multi-sample 1 to 96, 0.1 to 0. mL per sample |
||
| NanoGenerator™ Flex-M | Pre-clinical formulation, in-line ethanol dilution 1 to 12 mL batch |
||
| NanoGenerator™ MAX + | cGMP system for clinical manufacturing, product throughput > 10 L /hr. | ||
| Sunscreen | Discovery, microfluid chip options, automated HT, 1 to 96 samples, 0.2 to 2.0 mL per sample |
Unchained Labs | https://www.unchainedlabs.com/lipid-nanoparticles/ |
| Sunshine | Pre-clinical, microfluid chip options, automated 1 to 10 samples, continuous flow up to 30 mL/min |
||
| Sunbather | GMP Clinical ready, microfluid chip options, continuous flow up to 1.8 L /hr. |
| Product Trade Name | General Comments | Vendor | Website |
|---|---|---|---|
| DIANT® LARU Discovery | Discovery-scale continuous turbulent jet mixing 2 mL minimum output. volume |
Diant Pharma Inc. | https://diantpharma.com/ |
| DIANT® LARU – Benchtop | Pilot-scale continuous turbulent jet mixing with TFF and PAT max output 0.4 L /min |
||
| DIANT® LiFT – HT | Commercial-scale GMP continuous turbulent jet mixing with TFF and PAT max output 20 L /min |
||
| Nova™ Benchtop | Discovery-scale CIJ mixer system in-line dilution TFR 0.1 to 100 mL /min |
Helix Biotech Inc. | https://www.helixbiotech.com/ |
| Platform for Intracellular Delivery of DNA & RNA | Discovery-scale turbulent mixing technology for intracellular RNA and DNA delivery |
Optimeos Life Sciences Inc. |
https://optimeos.com/ |
| CIJ & MIVM Mixers Design by Dr. Prud'homme's Princeton Lab |
Manufactures CIJ and MIVM turbulent mixer units for lab-scale formulation development |
Holland Applied Technologies |
https://hollandapt.com/products/fittings-components/cij-mivm-mixers/ |
9. Summary and Conclusions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| BM | Microfluidic Baffles Mixer Design |
| CICS | Cylindrical Illumination Confocal Spectroscopy |
| CIJ | Confined Impinging Jet Turbulent Flow Mixer |
| CFD | Computational Fluid Dynamics |
| DMP | Dimethyloxypropane |
| FNC | Flash Nanocomplexation of Nanoparticles |
| FNP | Flash Nanoprecipitation of Nanoparticles |
| FRR | Flow Rate Ratio |
| HFFM | Hydrodynamic Flow Focusing Mixer Design |
| LNP | Lipid Nanoparticle |
| MIVM | Multi-Inlet Vortex Turbulent Flow Mixer |
| SHM | Staggered Herringbone Mixer Design |
| TFR | Total Flow Rate |
| TJM | T-junction Mixer Design |
| TRM | Toroidal Rings Mixer Design |
References
- Chaurasiya, A., et al., A review on multivesicular liposomes for pharmaceutical applications: preparation, characterization, and translational challenges. Drug Deliv Transl Res, 2022. 12(7): p. 1569-1587. [CrossRef]
- Kumari, L., et al., Advancement in Solubilization Approaches: A Step towards Bioavailability Enhancement of Poorly Soluble Drugs. Life (Basel), 2023. 13(5). [CrossRef]
- Wikipedia. Self-assembly. 2025; Available from: https://en.wikipedia.org/wiki/Self-assembly.
- Liu, J., et al., DNA-guided self-assembly in living cells. iScience, 2023. 26(5): p. 106620. [CrossRef]
- Saad, S. and D.F. Jarosz, Protein self-assembly: A new frontier in cell signaling. Curr Opin Cell Biol, 2021. 69: p. 62-69.
- Mitchell, M.J., et al., Engineering precision nanoparticles for drug delivery. Nat Rev Drug Discov, 2021. 20(2): p. 101-124.
- Pallares, R.M., et al., Nanoparticle Diagnostics and Theranostics in the Clinic. J Nucl Med, 2022. 63(12): p. 1802-. [CrossRef]
- Mohsen, M.O., et al., Major findings and recent advances in virus-like particle (VLP)-based vaccines. Semin Immunol, 2017. 34: p. 123-132. [CrossRef]
- Chatzikleanthous, D., D.T. O'Hagan, and R. Adamo, Lipid-Based Nanoparticles for Delivery of Vaccine Adjuvants and Antigens: Toward Multicomponent Vaccines. Mol Pharm, 2021. 18(8): p. 2867-2888. [CrossRef]
- Wikipedia. Self-assembly of nanoparticles. 2025; Available from: https://en.wikipedia.org/wiki/Self-assembly_of_nanoparticles.
- FDA, COMIRNATY® (COVID-19 Vaccine, mRNA) Package Insert. U.S. Approval: 2021.
- FDA, SPIKEVAX (COVID-19 Vaccine, mRNA) Package Insert. U.S. Approval: 2022.
- Bauer, A., et al., Current State and Opportunities with Long-acting Injectables: Industry Perspectives from the Innovation and Quality Consortium "Long-Acting Injectables" Working Group. Pharm Res, 2023. 40(7): p. 1601-1631.
- Pardi, N., et al., mRNA vaccines - a new era in vaccinology. Nat Rev Drug Discov, 2018.
- Igyártó, B.Z. and Z. Qin, The mRNA-LNP vaccines - the good, the bad and the ugly? Front Immunol, 2024. 15: p. 1336906.
- Chong, S.H., et al., A review of COVID vaccines: success against a moving target. Br Med Bull, 2022. 144(1): p. 12-44. [CrossRef]
- Friedrichs, S. and D.M. Bowman, COVID-19 may become nanomedicine's finest hour yet. Nat Nanotechnol, 2021. 16(4): p. 362-364.
- Hald Albertsen, C., et al., The role of lipid components in lipid nanoparticles for vaccines and gene therapy. Adv Drug Deliv Rev, 2022. 188: p. 114416. [CrossRef]
- Zhang, Y., et al., Lipids and Lipid Derivatives for RNA Delivery. Chem Rev, 2021. 121(20): p. 12181-12277.
- Wikipedia. Fick's laws of diffusion. 2025; Available from: https://en.wikipedia.org/wiki/Fick%27s_laws_of_diffusion.
- Price, H.C., J. Mattsson, and B.J. Murray, Sucrose diffusion in aqueous solution. Phys Chem Phys, 2016. 18(28): p. 19207-16.
- Johnson, B.K., Prud'homme, R. K., Mechanism for rapid self-assembly of block copolymer nanoparticles. . Phys Rev Lett. , 2003. 91(11): p. 118302.
- Stroock, A.D., et al., Chaotic mixer for microchannels. Science, 2002. 295(5555): p. 647-51. [CrossRef]
- Lee, C.Y., et al., Microfluidic mixing: a review. Int J Mol Sci, 2011. 12(5): p. 3263-87.
- Cheng, Y., et al., Microfluidic technologies for lipid vesicle generation. Lab Chip, 2024. 24(20): p. 4679-4716. [CrossRef]
- Dawson, H., et al., The Rise of the OM-LoC: Opto-Microfluidic Enabled Lab-on-Chip. Micromachines (Basel), 2021. 12(12).
- Narayanamurthy, V., et al., Advances in passively driven microfluidics and lab-on-chip devices: a comprehensive literature review and patent analysis. RSC Adv, 2020. 10(20): p. 11652-11680. [CrossRef]
- Ripoll, M., et al., Optimal self-assembly of lipid nanoparticles (LNP) in a ring micromixer. Sci Rep, 2022. 12(1): p. 9483.
- Cytiva. Nanoparticle formulation systems. 2025; Available from: https://www.cytivalifesciences.com/en/us/shop/lipid-nanoparticle-instruments-and-reagents/nanoparticle-formulation-systems?sort=NameAsc&chunk=1.
- Shen, Y., H. Gwak, and B. Han, Advanced manufacturing of nanoparticle formulations of drugs and biologics using microfluidics. Analyst, 2024. 149(3): p. 614-637. [CrossRef]
- Diant Pharma, I. DIANT’s Nanoparticle Technology. 2025; Available from: https://diantpharma.com/diants-nanoparticle-technology/.
- Liu R.H., S.M.A., Sharp K .V, Olse M,G, Santiag J.G. Adrian R.J., Passive mixing in a three-dimensional serpentine microchannel. Journal of Microelectromechanical Systems, 2000. 9(2): p. 192-197.
- Ying Zheng Liu, B.J.K., Hyung Jin Sun, Two-fluid mixing in a microchannel. International Journal of Heat and Fluid Flow, 2004. 25(6): p. 986-995.
- Maeki, M., et al., Understanding the formation mechanism of lipid nanoparticles in microfluidic devices with chaotic micromixers. PLoS One, 2017. 12(11): p. e0187962.
- Lasic, D.D., The mechanism of vesicle formation. Biochem J, 1988. 256(1): p. 1-11.
- Webb, C., et al., Using microfluidics for scalable manufacturing of nanomedicines from bench to GMP: A case study using protein-loaded liposomes. Int J Pharm, 2020. 582: p. 119266.
- Loughney, J.W., et al., Development of an imaged capillary isoelectric focusing method for characterizing the surface charge of mRNA lipid nanoparticle vaccines. Electrophoresis, 2019. 40(18-19): p. 2602-2609. [CrossRef]
- Roces, C.B., et al., Manufacturing Considerations for the Development of Lipid Nanoparticles Using Microfluidics. Pharmaceutics, 2020. 12(11).
- Terada, T., et al., Characterization of Lipid Nanoparticles Containing Ionizable Cationic Lipids Using Design-of-Experiments Approach. Langmuir, 2021. 37(3): p. 1120-1128. [CrossRef]
- Ly, H.H., et al., Optimization of Lipid Nanoparticles for saRNA Expression and Cellular Activation Using a Design-of-Experiment Approach. Mol Pharm, 2022. 19(6): p. 1892-1905.
- Pardi, N., et al., Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J Control Release, 2015. 217: p. 345-51. [CrossRef]
- Maier, M.A., et al., Biodegradable lipids enabling rapidly eliminated lipid nanoparticles for systemic delivery of RNAi therapeutics. Mol Ther, 2013. 21(8): p. 1570-8.
- Leung, A.K., et al., Microfluidic Mixing: A General Method for Encapsulating Macromolecules in Lipid Nanoparticle Systems. J Phys Chem B, 2015. 119(28): p. 8698-706.
- Cullis, P.R. and M.J. Hope, Lipid Nanoparticle Systems for Enabling Gene Therapies. Mol Ther, 2017. 25(7): p. 1467-1475. [CrossRef]
- Cullis, P.R. and P.L. Felgner, The 60-year evolution of lipid nanoparticles for nucleic acid delivery. Nat Rev Drug Discov, 2024. 23(9): p. 709-722. [CrossRef]
- Belliveau, N.M., et al., Microfluidic Synthesis of Highly Potent Limit-size Lipid Nanoparticles for In Vivo Delivery of siRNA. Mol Ther Nucleic Acids, 2012. 1(8): p. e37.
- Yanez Arteta, M., et al., Successful reprogramming of cellular protein production through mRNA delivered by functionalized lipid nanoparticles. Proc Natl Acad Sci U S A, 2018. 115(15): p. E3351-e3360.
- Schoenmaker, L., et al., mRNA-lipid nanoparticle COVID-19 vaccines: Structure and stability. Int J Pharm, 2021. 601: p. 120586. [CrossRef]
- Brader, M.L., et al., Encapsulation state of messenger RNA inside lipid nanoparticles. Biophys J, 2021. 120(14): p. 2766-2770.
- Li, S., et al., Payload distribution and capacity of mRNA lipid nanoparticles. Nat Commun, 2022. 13(1): p. 5561. [CrossRef]
- Tong, X., et al., Correlating Stability-Indicating Biochemical and Biophysical Characteristics with In Vitro Cell Potency in mRNA LNP Vaccine. Vaccines (Basel), 2024. 12(2).
- Abrams, M.T., et al., Evaluation of efficacy, biodistribution, and inflammation for a potent siRNA nanoparticle: effect of dexamethasone co-treatment. Mol Ther, 2010. 18(1): p. 171-80.
- Patel, N., et al., Development and Characterization of an In Vitro Cell-Based Assay to Predict Potency of mRNA-LNP-Based Vaccines. Vaccines (Basel), 2023. 11(7). [CrossRef]
- Offit, P.A., Tell Me When It’s Over. 2024, Washington, DC: National Geographic Partners, LLC.
- Pardi, N., et al., Nucleoside-modified mRNA immunization elicits influenza virus hemagglutinin stalk-specific antibodies. Nat Commun, 2018. 9(1): p. 3361. [CrossRef]
- Liang, F., et al., Efficient Targeting and Activation of Antigen-Presenting Cells In Vivo after Modified mRNA Vaccine Administration in Rhesus Macaques. Mol Ther, 2017. 25(12): p. 2635-2647. [CrossRef]
- Bahl, K., et al., Preclinical and Clinical Demonstration of Immunogenicity by mRNA Vaccines against H10N8 and H7N9 Influenza Viruses. Mol Ther, 2017. 25(6): p. 1316-1327.
- Lederer, K., et al., SARS-CoV-2 mRNA Vaccines Foster Potent Antigen-Specific Germinal Center Responses Associated with Neutralizing Antibody Generation. Immunity, 2020. 53(6): p. 1281-1295.e5.
- Laczkó, D., et al., A Single Immunization with Nucleoside-Modified mRNA Vaccines Elicits Strong Cellular and Humoral Immune Responses against SARS-CoV-2 in Mice. Immunity, 2020. 53(4): p. 724-732.e7. [CrossRef]
- Corbett, K.S., et al., Evaluation of the mRNA-1273 Vaccine against SARS-CoV-2 in Nonhuman Primates. N Engl J Med, 2020. 383(16): p. 1544-1555.
- Lopes, C., et al., Microfluidic production of mRNA-loaded lipid nanoparticles for vaccine applications. Expert Opin Drug Deliv, 2022. 19(10): p. 1381-1395.
- Egorov, E., et al., Robotics, microfluidics, nanotechnology and AI in the synthesis and evaluation of liposomes and polymeric drug delivery systems. Drug Deliv Transl Res, 2021. 11(2): p. 345-352.
- Rapalli, V.K., Khosa A, Singhvi G, Girdhar V, Jain R, Dubey, SK, Application of QbQ Principles in Nanocarrier-Based Drug Delivery Systems, in Pharmaceutical Quality by Design, S. Beg, Hasnain M S, Editor. 2019.
- Amiri, M.M. Tubulance: An Introduction. 2020; Available from: https://www.youtube.com/watch?v=sUqfatUVUPM&list=PLLYH8CQiuCNUS0o1dzgmvPK1mjt5BXVc7.
- Joseph, H. New ‘Superdiffusion’ Proof Probes the Mysterious Math of Turbulence. 2025; Available from: https://www.quantamagazine.org/new-superdiffusion-proof-probes-the-mysterious-math-of-turbulence-20250516/?print=1.
- Johnson, B.K., Prud'homme R. K., Chemical Processing and Micromixing in Confined Impinging Jets. AIcE Journal, 2003. 49(9): p. 2264-2281. [CrossRef]
- Baldyga, J., Bourne, J.R., Walker, B, Non-isothermal Micromixing in Liquids: Theory and Experiment. Can. J. Chem. Eng., 1998. 76: p. 641-649.
- Baldyga, J., Bourne, J.R., Time Constants for Kenics Mixer, in Turbulent Mixing and Chemical Reaction, J. Baldyga, Bourne, J.R., Editor. 1999, John Wiley & Sons: New York. p. 733-763.
- Taylor, R.A., Penney, W.R., Hanh, X. V., Scale-up Methods for Fast Competitive Chemical Reactions in Pipeline Mixers. Ind. Eng. Chem. Res. , 2005. 44: p. 6095-6102. [CrossRef]
- Ying Liu, C.C., Ying Liu, Robert K. Prud’homme, Rodney O. Fox,, Mixing in a multi-inlet vortex mixer (MIVM) for flash nano-precipitation. Chemical Engineering Science 2008. 63: p. 2829-2842.
- Hu, H., et al., Flash Technology-Based Self-Assembly in Nanoformulation: From Fabrication to Biomedical Applications. Mater Today (Kidlington), 2021. 42: p. 99-116.
- Markwalter, C.E. and R.K. Prud'homme, Design of a Small-Scale Multi-Inlet Vortex Mixer for Scalable Nanoparticle Production and Application to the Encapsulation of Biologics by Inverse Flash NanoPrecipitation. J Pharm Sci, 2018. 107(9): p. 2465-2471. [CrossRef]
- Gindy, M.E., et al., Mechanism of macromolecular structure evolution in self-assembled lipid nanoparticles for siRNA delivery. Langmuir, 2014. 30(16): p. 4613-22.
- Gindy, M.E., et al., Stabilization of Ostwald ripening in low molecular weight amino lipid nanoparticles for systemic delivery of siRNA therapeutics. Mol Pharm, 2014. 11(11): p. 4143-53. [CrossRef]
- Espeseth, A.S., et al., Modified mRNA/lipid nanoparticle-based vaccines expressing respiratory syncytial virus F protein variants are immunogenic and protective in rodent models of RSV infection. NPJ Vaccines, 2020. 5(1): p. 16. [CrossRef]
- Shepherd, S.J., et al., Scalable mRNA and siRNA Lipid Nanoparticle Production Using a Parallelized Microfluidic Device. Nano Lett, 2021. 21(13): p. 5671-5680. [CrossRef]
- Warne, N., et al., Delivering 3 billion doses of Comirnaty in 2021. Nat Biotechnol, 2023. 41(2): p. 183-188.
- Gupta, S. CNN Video on Comirnaty Manufacture. 2021; Available from: https://www.cnn.com/2021/03/31/health/pfizer-vaccine-manufacturing/index.html.
- Subraveti, S.N., et al., Synthesizing Lipid Nanoparticles by Turbulent Flow in Confined Impinging Jet Mixers. J Vis Exp, 2024(210). [CrossRef]
- D'Addio, S.M. and R.K. Prud'homme, Controlling drug nanoparticle formation by rapid precipitation. Adv Drug Deliv Rev, 2011. 63(6): p. 417-26.
- Wikipedia. Kelvin Equation. 2025; Available from: https://en.wikipedia.org/wiki/Kelvin_equation.
- Lin, P.J., et al., Influence of cationic lipid composition on uptake and intracellular processing of lipid nanoparticle formulations of siRNA. Nanomedicine, 2013. 9(2): p. 233-46.
- O'Brien Laramy, M.N., et al., Process Robustness in Lipid Nanoparticle Production: A Comparison of Microfluidic and Turbulent Jet Mixing. Mol Pharm, 2023. 20(8): p. 4285-4296.
- Kulkarni, J.A., et al., On the role of helper lipids in lipid nanoparticle formulations of siRNA. Nanoscale, 2019. 11(45): p. 21733-21739. [CrossRef]
- Tran, V., et al., Current and Near-Future Technologies to Quantify Nanoparticle Therapeutic Loading Efficiency and Surface Coating Efficiency with Targeted Moieties. Bioengineering (Basel), 2025. 12(4).
- Semple, S.C., et al., Efficient encapsulation of antisense oligonucleotides in lipid vesicles using ionizable aminolipids: formation of novel small multilamellar vesicle structures. Biochim Biophys Acta, 2001. 1510(1-2): p. 152-66.
- Maurer, N., et al., Spontaneous entrapment of polynucleotides upon electrostatic interaction with ethanol-destabilized cationic liposomes. Biophys J, 2001. 80(5): p. 2310-26. [CrossRef]
- Pagels, R.F. and R.K. Prud'homme, Polymeric nanoparticles and microparticles for the delivery of peptides, biologics, and soluble therapeutics. J Control Release, 2015. 219: p. 519-535.
- Markwalter, C.E., et al., Polymeric Nanocarrier Formulations of Biologics Using Inverse Flash NanoPrecipitation. Aaps j, 2020. 22(2): p. 18.
- Ristroph, K.D. and R.K. Prud'homme, Hydrophobic ion pairing: encapsulating small molecules, peptides, and proteins into nanocarriers. Nanoscale Adv, 2019. 1(11): p. 4207-4237. [CrossRef]
- Feng, J., et al., Translational formulation of nanoparticle therapeutics from laboratory discovery to clinical scale. J Transl Med, 2019. 17(1): p. 200.
- Gavi E, M.D.L., Barresi A A, CFD modelling and scale-up of Confined Impinging Jet Reactors. Chemical Engineering Science, 2007. 62: p. 2228-2241. [CrossRef]
- Marchisio, D., Large Eddy Simulation of mixing and reaction in a Confined Impinging Jets Reactor. Computers & Chemical Engineering 2009. 33: p. 408-420.
- Dorsey, P.J., et al., Review of machine learning for lipid nanoparticle formulation and process development. J Pharm Sci, 2024. 113(12): p. 3413-3433. [CrossRef]
- Cheng, L., et al., Machine Learning Elucidates Design Features of Plasmid DNA Lipid Nanoparticles for Cell Type-Preferential Transfection. bioRxiv, 2023.
- Carvalho, B.G., et al., Advanced Microfluidic Technologies for Lipid Nano-Microsystems from Synthesis to Biological Application. Pharmaceutics, 2022. 14(1). [CrossRef]
- Zhu, Y., et al., Multi-step screening of DNA/lipid nanoparticles and co-delivery with siRNA to enhance and prolong gene expression. Nat Commun, 2022. 13(1): p. 4282.
- Precigenome. NanoGenerator® Synthesis System Nanoparticle Formulation & Preparation. 2025; Available from: https://www.precigenome.com/.
- Unchained. Lipid Nanoparticles. 2025; Available from: https://www.unchainedlabs.com/lipid-nanoparticles/.
- Ottonelli, I., et al., Microfluidic Technology for the Production of Hybrid Nanomedicines. Pharmaceutics, 2021. 13(9). [CrossRef]
- Holland. The Confined Impinging Jet (CIJ) and the Multi Inlet Vortex Mixer (MIVM). 2019; Available from: https://hollandapt.com/products/fittings-components/cij-mivm-mixers/.
- Hermosilla, J., et al., Analysing the In-Use Stability of mRNA-LNP COVID-19 Vaccines Comirnaty™ (Pfizer) and Spikevax™ (Moderna): A Comparative Study of the Particulate. Vaccines (Basel), 2023. 11(11).
- Fongaro, B., et al., Assessing the physicochemical stability and intracellular trafficking of mRNA-based COVID-19 vaccines. Int J Pharm, 2023. 644: p. 123319. [CrossRef]
| Mixing Method | DLS Diameter, nm |
ζ-Potential, mV | mRNA EE %* | HeLa Cell In vitro Transfection, RLU /1000 cells |
HeLa Cell Viability |
|---|---|---|---|---|---|
| Manual Pipette | ≈ 140 | ≈ 0.5 | >95 % | ≈ 15 | ≈ 100 % |
| Two-Jet CIJ Mixer | ≈ 90 | ≈ 2.0 | >95 % | ≈ 100 | ≈ 100 % |
| Four-Jet MIVM | ≈ 90 | ≈ 1.0 | >95 % | ≈ 100 | ≈ 100 % |
| Process Condition |
Fluorescently Identified Particle | Relative Population by Number | mRNA Copy per Particle | Average mRNA EE % | Average DLS Diameter, nm |
Average ζ -Potential, mV |
|---|---|---|---|---|---|---|
| Initial LNP, Post T-mixing 3:1, Aqueous: Ethanol (v/v) Mixed Solution pH 4.0 |
Lipophilic mRNA Complex (LNC) | 34 ± 8 % | 3.4 ± 0.4 | N/A | 106 ± 13 | + 45 ± 1 |
| Non-Lipophilic mRNA Complex (NLNC) | 25 ± 4 % | 1.3 ± 0.2 | ||||
| mRNA Lipid Nanoparticle (LNP) | 0% | N/A | ||||
| Empty Particle, i.e. No mRNA (ENP) | 41 ± 10 % | 0 | ||||
| Final Process LNP, Post-Dialysis Aqueous pH 7.4 Buffer |
Lipophilic mRNA Complex (LNC) | 0 | N/A | 94.2 ± 0.4 % | 120 ± 6 | - 6.3 ± 1.3 |
| Non-Lipophilic mRNA Complex (NLNC) |
0 | N/A | ||||
| mRNA Lipid Nanoparticle (LNP) | 23 ± 8 % | 2.8 ± 0.4 | ||||
| Empty Particle, i.e. No mRNA (ENP) | 77 ± 8 % | N/A |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




