Submitted:
02 June 2025
Posted:
03 June 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Molecular Principles of LLPS in the Viral Context
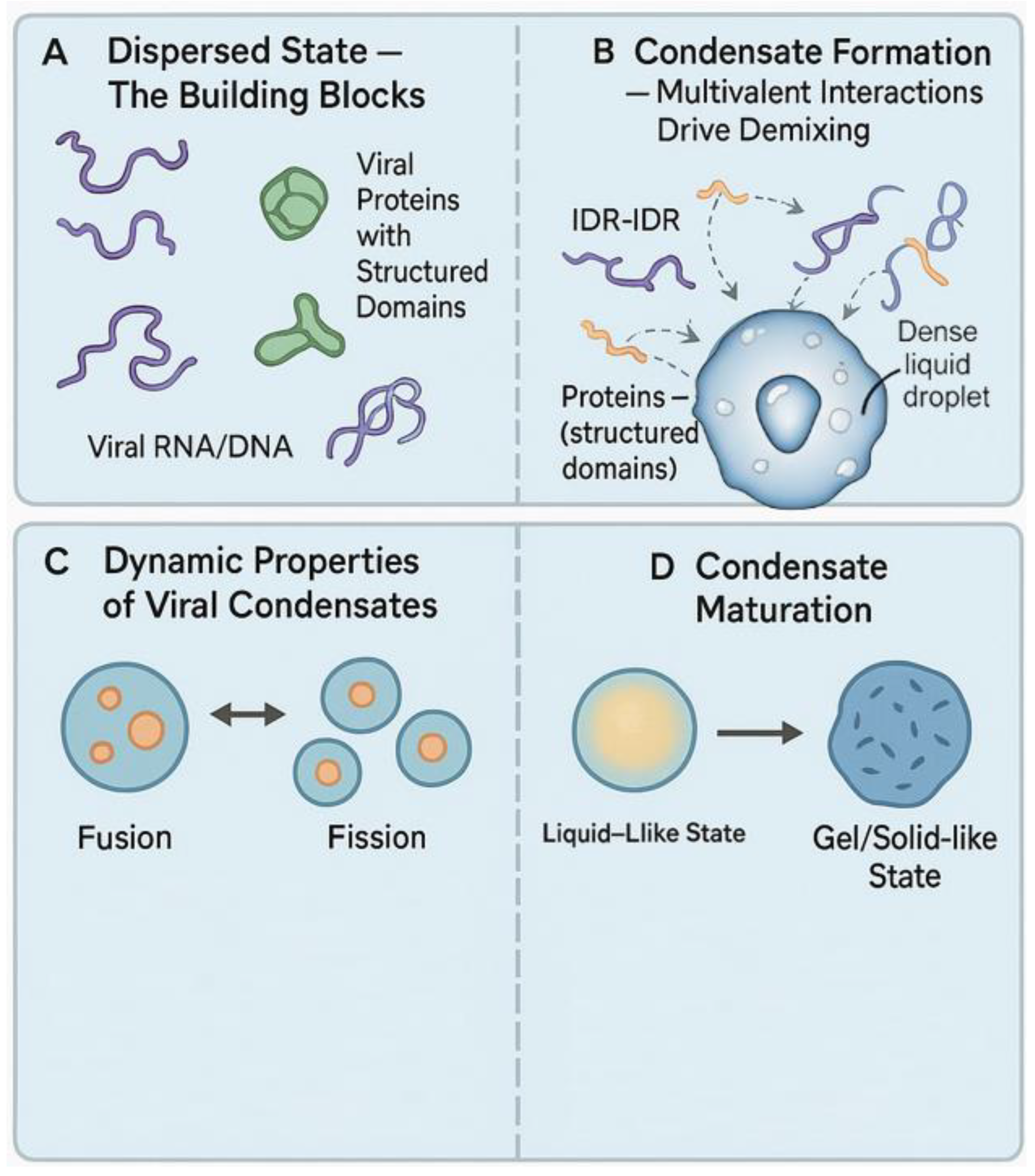
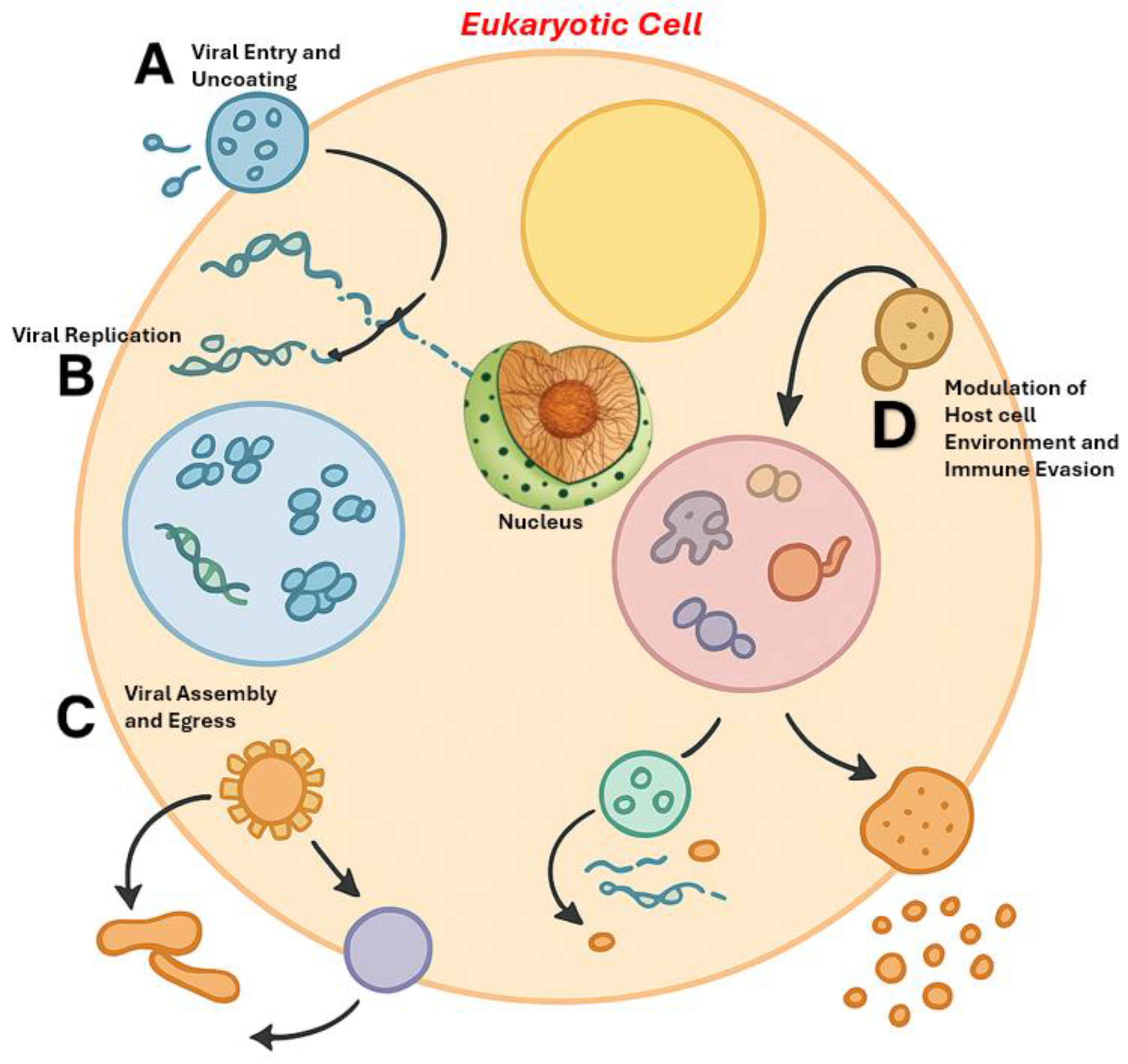
3. LLPS Across the Viral Replication Cycle: New Mechanistic Insights
3.1. Viral Entry and Uncoating
3.2. Genome Replication and Transcription: The Viral Replication Compartment (VRC) as a Biomolecular Condensate
3.2.1. RNA Viruses
3.2.1.1. Paramyxoviruses
3.2.1.2. Coronaviruses
3.2.1.3. Other Negative-Sense RNA Viruses
3.2.1.4. Influenza A Virus (IAV)
3.2.2. DNA Viruses
3.2.2.1. Herpesviruses
3.3. Viral Assembly and Egress
3.3.1. Retroviruses
3.4. Modulation of Host Cell Environment: Immune Evasion and Stress Response Interference
3.4.1. Interference with Stress Granules (SGs)
3.4.2. Sequestration of Immune Factors
3.4.3. Hijacking Host Factors for Viral Benefit
| Virus Family / Specific Virus | Key Viral LLPS Driver(s) | Critical Domains/Motifs | Key Interacting Partners (Viral/Host) | Resulting Condensate / Cellular Process | Key Regulatory Features |
|---|---|---|---|---|---|
| Paramyxoviridae / Measles Virus (MeV) | N and P proteins | P-XD (C-term domain of P), N-IDR (C-term intrinsically disordered region of N) [27] | Viral RNA, host WDR5, host dynein[27] | Inclusion Bodies (IBs) / Viral Replication Factories, Immune Evasion[6] | P protein phosphorylation, Liquid-to-gel transition over time [27] |
| Coronaviridae / SARS-CoV-2[6] | Nucleocapsid (N) protein | N-IDRs, RNA-binding domains, Oligomerization domain | Viral RNA, host G3BP1, host kinases (e.g., SRPK1, GSK3) | N-RNA condensates / Replication, Transcription, Genome Packaging, SG modulation | N protein phosphorylation (regulates liquid vs. gel-like state), RNA concentration |
| Retroviridae / HIV-1 | Gag polyprotein (specifically Nucleocapsid - NC domain) | NC Zinc Fingers (ZnF), IDRs in Gag [41] | Viral genomic RNA (gRNA), host factors (e.g., IP6) [23] | Gag assembly sites, NC-RNA condensates / RNA packaging, Virion Assembly, SG modulation [23] | Zn²⁺- [6,27] dependence of NC LLPS, RNA binding[41] |
| Orthomyxoviridae / Influenza A Virus (IAV) [23] | Viral Ribonucleoproteins (vRNPs), Nucleoprotein (NP) | NP oligomerization domains, RNA binding sites[23] | Host Rab11a, other vRNPs | Cytoplasmic vRNP inclusions / Genome segment assembly, Trafficking to budding sites[6] 5 | Interaction with Rab11a, potential modulation by NP PTMs |
| Herpesviridae / HSV-1, KSHV, HCMV [6] | ICP4 (HSV-1), LANA (KSHV), UL112-113 (HCMV), ORF52 (MHV-68) | Various, often IDRs or multivalent domains | Viral DNA, host replication/transcription factors, viral polymerase | Nuclear VRCs (HSV-1, HCMV), LANA-NBs (KSHV), Cytoplasmic VACs (MHV-68) / DNA Replication, Latency, Assembly | Concentration of viral proteins, interaction with viral DNA |
| Rhabdoviridae / Rabies Virus (RABV) [6,27] | N and P proteins | IDRs in N and P, P protein oligomerization domain | Viral RNA, host STAT proteins, host Hsp70 5 | Negri Bodies (NBs) / Viral Replication Factories, Immune Evasion | Protein concentration, interactions between N, P, and L proteins |
4. LLPS: Reshaping Paradigms in Virology
5. Therapeutic Opportunities Targeting Viral LLPS
5.1. Dissolution or Prevention of Condensate Formation
5.2. Aberrant Stabilization or Alteration of Material Properties
5.3. Targeting Specific Components Within Viral Condensates
5.5. Modulating Post-Translational Modifications (PTMs)
- SARS-CoV-2 N protein: Strategies include the use of small molecules like gallic catechin gallate (GCG) from green tea, which can inhibit N protein LLPS. Interference peptides (NIP I-V) have been designed to disrupt N protein interactions essential for LLPS. Furthermore, modulating N protein phosphorylation via kinase activators (e.g., SRPK1) or inhibitors is an active area of investigation, as mutations in N protein IDRs are known to affect LLPS propensity [6].
- HIV-1 NC protein: The ZnF domains of NC are critical for its LLPS activity. Zn chelators or mutations in the ZnF domains disrupt NC-LLPS and significantly impair viral replication, identifying the ZnF motif as a druggable target for pan-retroviral therapies [41].
| Therapeutic Strategy | Molecular Target Class | Type of Modulator | Specific Viral Example(s) & Protein(s) | Potential Antiviral Effect | Key Challenges / Current Status |
|---|---|---|---|---|---|
| Disrupt Condensate Formation/Integrity [6,41] | Viral IDRs/LCRs, Viral Oligomerization Domains, Viral-Host Protein Interfaces | Small Molecule (dissolver, binder), Peptide (interface blocker, IDR binder), Antibody/Nanobody | SARS-CoV-2 N (NIP I-V, GCG) ; HIV-1 NC (Zn$^{2+}$ chelators) | Inhibit VRC formation, Block RNA packaging, Disrupt viral assembly | Specificity, Delivery, Off-target effects, Preclinical/Early research |
| Modulate Material Properties[49] | Viral Scaffold Proteins (e.g., NP, N) | Small Molecule (e.g., "glues", stabilizers) | IAV NP (Nucleozin) ; SARS-CoV-2 N | Impair condensate dynamics, Prevent necessary phase transitions (e.g., liquid-to-gel) | Achieving precise modulation, Understanding functional consequence of altered properties |
| Inhibit Scaffold Protein Function[27,51] | Key viral proteins driving LLPS (e.g., N, NC, P) | Small Molecule Inhibitor, Peptide, Nucleic Acid Aptamer | IAV NP (NLS-binding compounds) ; MeV P | Reduce viral replication/transcription, Inhibit viral factory function | Identifying druggable pockets on IDP-rich scaffolds, Specificity |
| Inhibit Client Recruitment [6] | Interaction sites between viral scaffolds and essential viral/host clients | Small Molecule, Peptide | SARS-CoV-2 N and G3BP1 | Prevent concentration of essential factors (e.g., polymerase, RNA) in VRCs, Restore host defenses | High specificity is required to avoid disrupting host protein functions |
| Modulate PTMs Regulating LLPS[27] | Cellular/Viral Enzymes (e.g., kinases, phosphatases) modifying viral LLPS proteins | Small Molecule Enzyme Inhibitor/Activator | SARS-CoV-2 N (SRPK1 activators) ; MeV P (CK2 inhibitors) | Alter condensate properties, Inhibit/promote specific viral processes dependent on PTM state | Specificity for viral targets, Understanding complex PTM codes, Potential host cell toxicity |
6. Challenges and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhang: J.Z., S. Mehta, and J. Zhang, Liquid–liquid phase separation: a principal organizer of the cell’s biochemical activity architecture. Trends in pharmacological sciences, 2021. 42(10): p. 845-856.
- Zhao, Y.G.; Zhang, H. Phase Separation in Membrane Biology: The Interplay between Membrane-Bound Organelles and Membraneless Condensates. Dev. Cell 2020, 55, 30–44. [CrossRef]
- Lim, J.-Y.; Kim, Y.; Jeon, S.; Jeon, Y.; Kim, W.; Cha, B. Emerging regulatory mechanisms and functions of biomolecular condensates: implications for therapeutic targets. Signal Transduct. Target. Ther. 2025, 10, 1–42. [CrossRef]
- Gomes, E.; Shorter, J. The molecular language of membraneless organelles. J. Biol. Chem. 2019, 294, 7115–7127. [CrossRef]
- Li, J.; Yang, F.; Zahid, A.; Yao, X.; Ismail, H.; Liu, X.; Dou, Z.; Liu, X. Mechanisms and regulation underlying membraneless organelle plasticity control. J. Mol. Cell Biol. 2021, 13, 239–258. [CrossRef]
- Hu, J.; Cui, J.; Jin, S.; Zhang, C.; Yang, S.; Wu, J.; Cai, S.; Guan, X.; Wu, Y.; Shen, W. Molecular mechanisms and cellular functions of liquid-liquid phase separation during antiviral immune responses. Front. Immunol. 2023, 14. [CrossRef]
- Parissi, V.; Li, H.; Ernst, C.; Greb-Markiewicz, B.; Kolonko-Adamska, M.; Man, J.; Ng, B.W.-L. Phase separation in viral infections. Trends Microbiol. 2022, 30, 1217–1231. [CrossRef]
- Ahn, J.-H.; Chung, W.-C.; Song, M.J. Liquid–liquid phase separation drives herpesvirus assembly in the cytoplasm. J. Cell Biol. 2022, 222. [CrossRef]
- Lagaudrière-Gesbert, C.; Gaudin, Y.; Albertini, A.; Glon, D.; Léonardon, B.; Guillemot, A. Biomolecular condensates with liquid properties formed during viral infections. Microbes Infect. 2024, 26, 105402. [CrossRef]
- Li, Z.; Zhang, X.; Zheng, R.; Ma, J. Liquid-liquid Phase Separation in Viral Function. J. Mol. Biol. 2023, 435, 167955. [CrossRef]
- Peng, P.-H., K.-W. Hsu, and K.-J. Wu, Liquid-liquid phase separation (LLPS) in cellular physiology and tumor biology. American journal of cancer research, 2021. 11(8): p. 3766.
- Xue, B.; Habchi, J.; Blocquel, D.; Uversky, A.V.; Uversky, V.N.; Kurgan, L.; Longhi, S. Structural Disorder in Viral Proteins. Chem. Rev. 2014, 114, 6880–6911. [CrossRef]
- Borcherds, W., et al., How do intrinsically disordered protein regions encode a driving force for liquid–liquid phase separation? Current opinion in structural biology, 2021. 67: p. 41-50. [CrossRef]
- Rosen, M.K.; Lin, Y.; Currie, S.L. Intrinsically disordered sequences enable modulation of protein phase separation through distributed tyrosine motifs. J. Biol. Chem. 2017, 292, 19110–19120. [CrossRef]
- Lee, K.S. SARS-CoV-2 immunogenicity models inform novel strategies that can improve mRNA vaccines. 2024. [CrossRef]
- Ke, W., Functional nucleic acid nanoparticles and their delivery. 2020, The University of North Carolina at Charlotte.
- Bergeron, H.C., Blocking the RSV G Protein CX3C Chemokine Motif Protects Against RSV Disease. 2023, University of Georgia.
- Grandori, R.; Brocca, S.; Longhi, S.; Uversky, V. Liquid–Liquid Phase Separation by Intrinsically Disordered Protein Regions of Viruses: Roles in Viral Life Cycle and Control of Virus–Host Interactions. Int. J. Mol. Sci. 2020, 21, 9045. [CrossRef]
- Nosella, M., The Effects Of Post-Translational Modification On Intrinsically Disordered Protein Phase Separation And Nucleosome Dynamics. 2024, University of Toronto (Canada).
- Geiger, F., et al., Rotavirus replication factories are complex ribonucleoprotein condensates. Biorxiv, 2020: p. 2020.12. 18.423429.
- Shanker, S.; Estes, M.K.; Ayyar, B.V.; Pollet, J.; Kaur, G.; Prasad, B.V.V.; Stossi, F.; Crawford, S.E.; Kaundal, S.; Anish, R. RNA-dependent RNA polymerase of predominant human norovirus forms liquid-liquid phase condensates as viral replication factories. Sci. Adv. 2024, 10, eadp9333. [CrossRef]
- Sanfeliu-Cerdán, N.; Krieg, M. The mechanobiology of biomolecular condensates. Biophys. Rev. 2025, 6, 011310. [CrossRef]
- Etibor, T.A.; Yamauchi, Y.; Amorim, M.J. Liquid Biomolecular Condensates and Viral Lifecycles: Review and Perspectives. Viruses 2021, 13, 366. [CrossRef]
- Su, J.M.; Wilson, M.Z.; Samuel, C.E.; Ma, D. Formation and Function of Liquid-Like Viral Factories in Negative-Sense Single-Stranded RNA Virus Infections. Viruses 2021, 13, 126. [CrossRef]
- Moncman, C.L.; Leung, D.W.; Creamer, T.P.; El Najjar, F.; Wu, C.; Cifuentes-Munoz, N.; Edmonds, K.; Boggs, K.B.; Ossandón, C.; Roe, M.; et al. Human Metapneumovirus Phosphoprotein Independently Drives Phase Separation and Recruits Nucleoprotein to Liquid-Like Bodies. mBio 2022, 13, e0109922. [CrossRef]
- Gerresheim, G.K.; Dolnik, O.; Biedenkopf, N. New Perspectives on the Biogenesis of Viral Inclusion Bodies in Negative-Sense RNA Virus Infections. Cells 2021, 10, 1460. [CrossRef]
- Zhou, Y.; Su, J.M.; Samuel, C.E.; Ma, D. Measles Virus Forms Inclusion Bodies with Properties of Liquid Organelles. J. Virol. 2019, 93, e00948-19. [CrossRef]
- Yang, Z., et al., Interaction between host G3BP and viral nucleocapsid protein regulates SARS-CoV-2 replication. BioRxiv, 2023.
- Cascarina, S.M. and E.D. Ross, Phase separation by the SARS-CoV-2 nucleocapsid protein: Consensus and open questions. Journal of Biological Chemistry, 2022. 298(3): p. 101677. [CrossRef]
- Nikolic, J.; Le Bars, R.; Lama, Z.; Scrima, N.; Lagaudrière-Gesbert, C.; Gaudin, Y.; Blondel, D. Negri bodies are viral factories with properties of liquid organelles. Nat. Commun. 2017, 8, 58. [CrossRef]
- Nevers, Q.; Albertini, A.A.; Lagaudrière-Gesbert, C.; Gaudin, Y. Negri bodies and other virus membrane-less replication compartments. Biochim. Biophys. Acta 2020, 1867, 118831. [CrossRef]
- Darling, A.L.; Oldfield, C.J.; Liu, Y.; Uversky, V.N. Intrinsically Disordered Proteome of Human Membrane-Less Organelles. Proteomics 2017, 18, e1700193. [CrossRef]
- Bosse, J.B.; Brune, W.; Caragliano, E. Herpesvirus Replication Compartments: Dynamic Biomolecular Condensates?. Viruses 2022, 14, 960. [CrossRef]
- RANGE, H. and I. DOSE, PATHOGEN SAFETY DATA SHEET-INFECTIOUS SUBSTANCES.
- Deng, Z.; De Leo, A.; Vladimirova, O.; Hayden, J.; Lieberman, P.M.; Wiedmer, A. Phase separation and DAXX redistribution contribute to LANA nuclear body and KSHV genome dynamics during latency and reactivation. PLOS Pathog. 2021, 17, e1009231. [CrossRef]
- Uppal, T.; Banerjee, S.; Sun, Z.; Verma, S.C.; Robertson, E.S. KSHV LANA—The Master Regulator of KSHV Latency. Viruses 2014, 6, 4961–4998. [CrossRef]
- Li, S.; Raina, K.; Kazemian, M.; Szymula, A.; George, A.; Ramachandran, A.; Van Sciver, N.; Wang, M.; Zhao, B.; Tumuluri, V.S.; et al. Kaposi’s sarcoma herpesvirus latency-associated nuclear antigen broadly regulates viral gene expression and is essential for lytic infection. PLOS Pathog. 2024, 20, e1011907. [CrossRef]
- Bourqui, L.; Anfossi, M.; Michaelsen, K.; Vogt, B.; Fraefel, C.; Georgi, F.; Seyffert, M.; Tobler, K.; Greber, U.F. The HSV-1 Transcription Factor ICP4 Confers Liquid-Like Properties to Viral Replication Compartments. Int. J. Mol. Sci. 2021, 22, 4447. [CrossRef]
- Hanson, H.M.; Willkomm, N.A.; Yang, H.; Mansky, L.M. Human Retrovirus Genomic RNA Packaging. Viruses 2022, 14, 1094. [CrossRef]
- Moog, C.; Boutant, E.; Réal, E.; Zeiger, M.; Mély, Y.; Anton, H.; Klingler, J. How HIV-1 Gag Manipulates Its Host Cell Proteins: A Focus on Interactors of the Nucleocapsid Domain. Viruses 2020, 12, 888. [CrossRef]
- Niu, M.; Monette, A.; Rao, S.; Chen, L.; Gorelick, R.J.; Mouland, A.J. Pan-retroviral Nucleocapsid-Mediated Phase Separation Regulates Genomic RNA Positioning and Trafficking. Cell Rep. 2020, 31, 107520–107520. [CrossRef]
- Charman, M.; Weitzman, M.D. Replication Compartments of DNA Viruses in the Nucleus: Location, Location, Location. Viruses 2020, 12, 151. [CrossRef]
- Van Royen, T., et al., How RSV proteins join forces to overcome the host innate immune response. Viruses, 2022. 14(2): p. 419. [CrossRef]
- Jobe, F.; Simpson, J.; Hawes, P.; Guzman, E.; Bailey, D. Respiratory Syncytial Virus Sequesters NF-κB Subunit p65 to Cytoplasmic Inclusion Bodies To Inhibit Innate Immune Signaling. J. Virol. 2020, 94, e01380-20. [CrossRef]
- Longhi, S.; Galloux, M. Unraveling Liquid–Liquid Phase Separation (LLPS) in Viral Infections to Understand and Treat Viral Diseases. Int. J. Mol. Sci. 2024, 25, 6981. [CrossRef]
- Bosse, J.B.; Brune, W. Viral dew: Phase separation and the formation of viral replication compartments. PLOS Pathog. 2023, 19, e1011145. [CrossRef]
- Nguyen, A., et al., Modulation of biophysical properties of nucleocapsid protein in the mutant spectrum of SARS-CoV-2. Elife, 2024. 13. [CrossRef]
- Mitrea, D.M.; Gomes, B.F.; Mittasch, M.; Klein, I.A.; Murcko, M.A. Modulating biomolecular condensates: a novel approach to drug discovery. Nat. Rev. Drug Discov. 2022, 21, 841–862. [CrossRef]
- Kong, H.; Amada, J.X.; Ng, B.W.-L.; Scheeff, S. Making it hard to replicate. eLife 2023, 12. [CrossRef]
- Sharma, B., et al., Post-translational modifications (PTMs), from a cancer perspective: an overview. Oncogen, 2019. 2(3): p. 12.
- Yang, F., et al., Discovery of a novel specific inhibitor targeting influenza A virus nucleoprotein with pleiotropic inhibitory effects on various steps of the viral life cycle. Journal of virology, 2021. 95(9): p. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




