Submitted:
29 May 2025
Posted:
29 May 2025
You are already at the latest version
Abstract
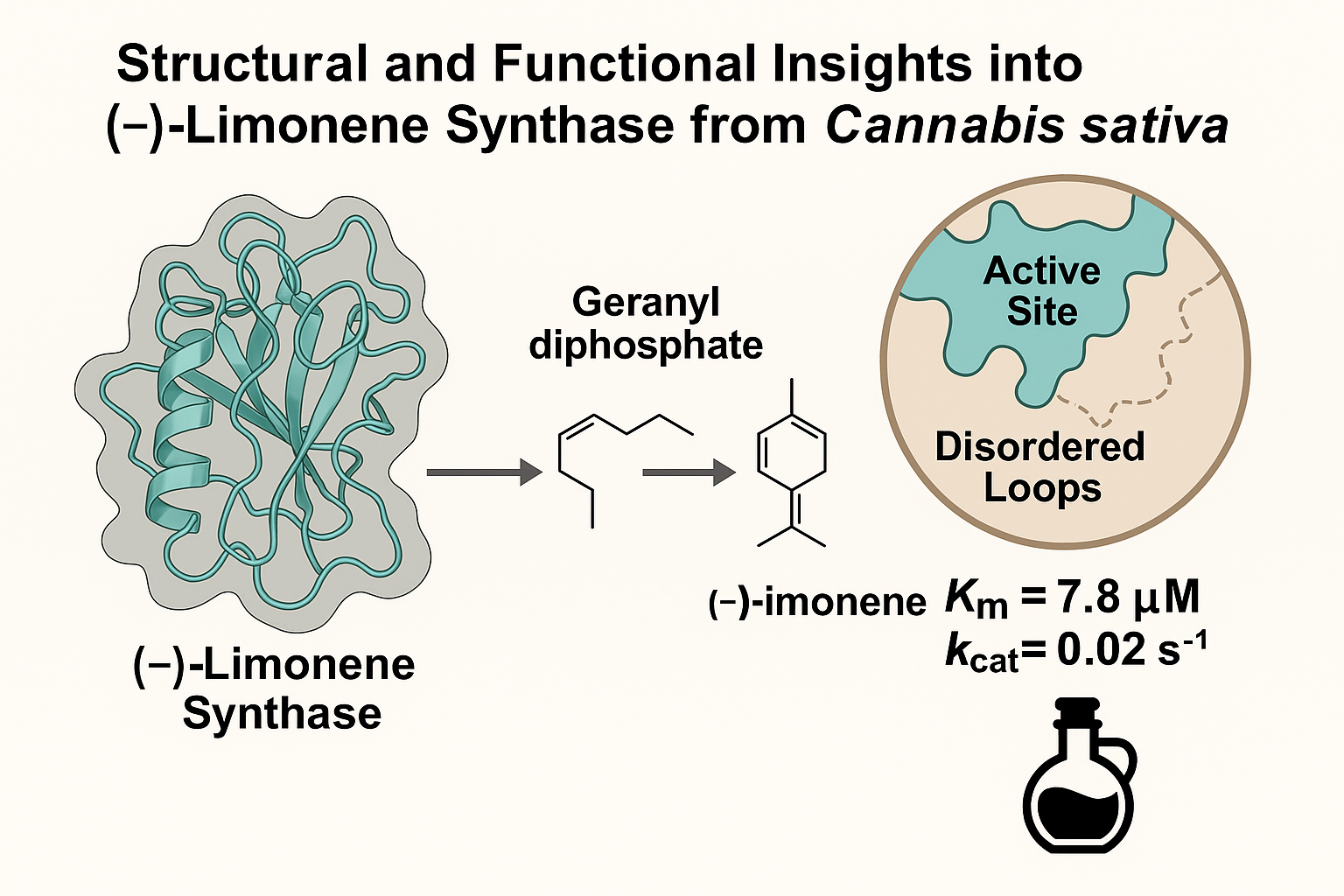
Keywords:
1. Introduction
2. Materials and Methods
2.1. Macromolecule Production
2.2. Crystallisation
2.3. Data Collection, Processing and Structure Solution
2.4. Product Identification by GC-MS
2.5. Steady-State Kinetics
3. Results and Discussion
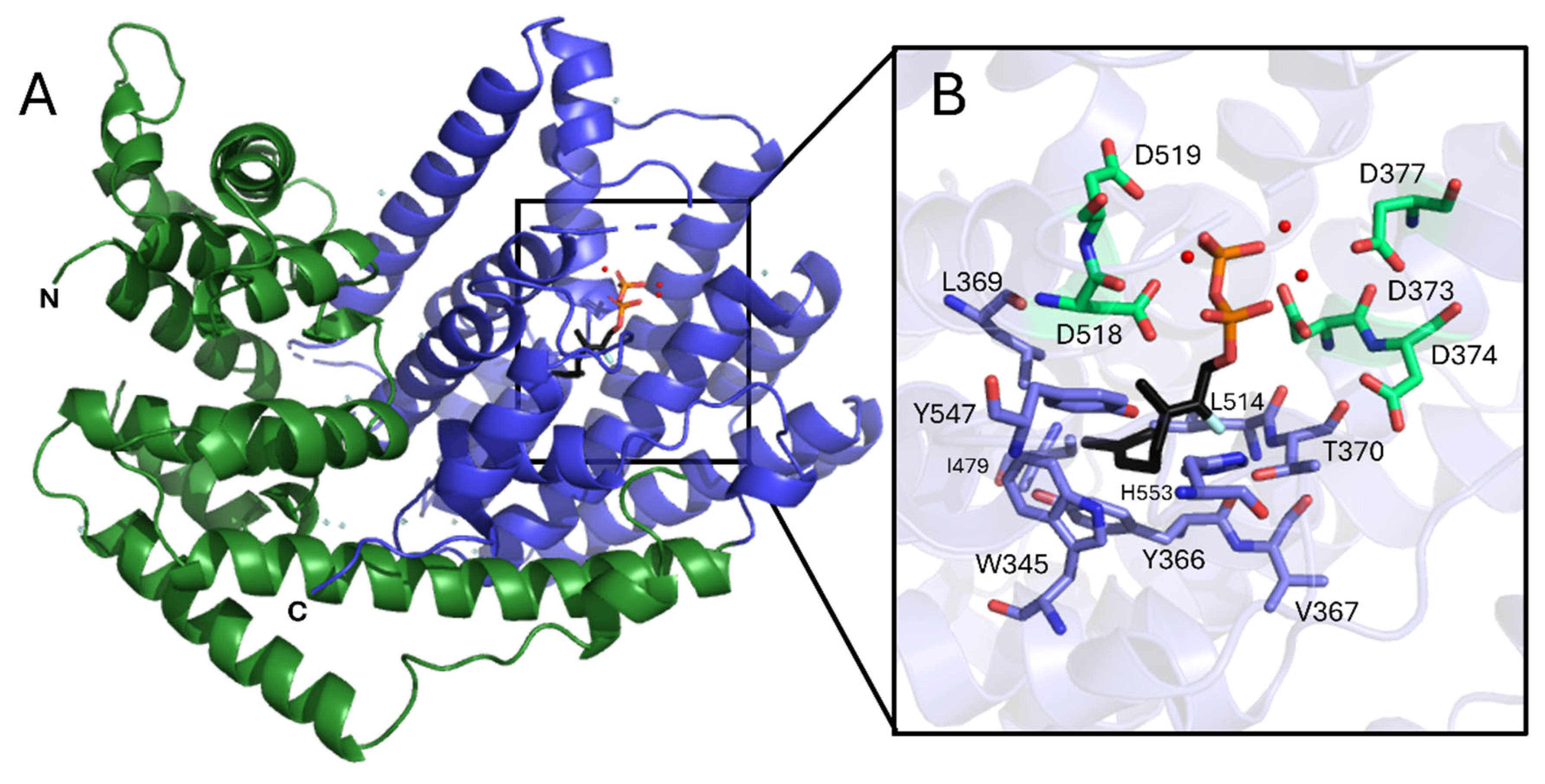
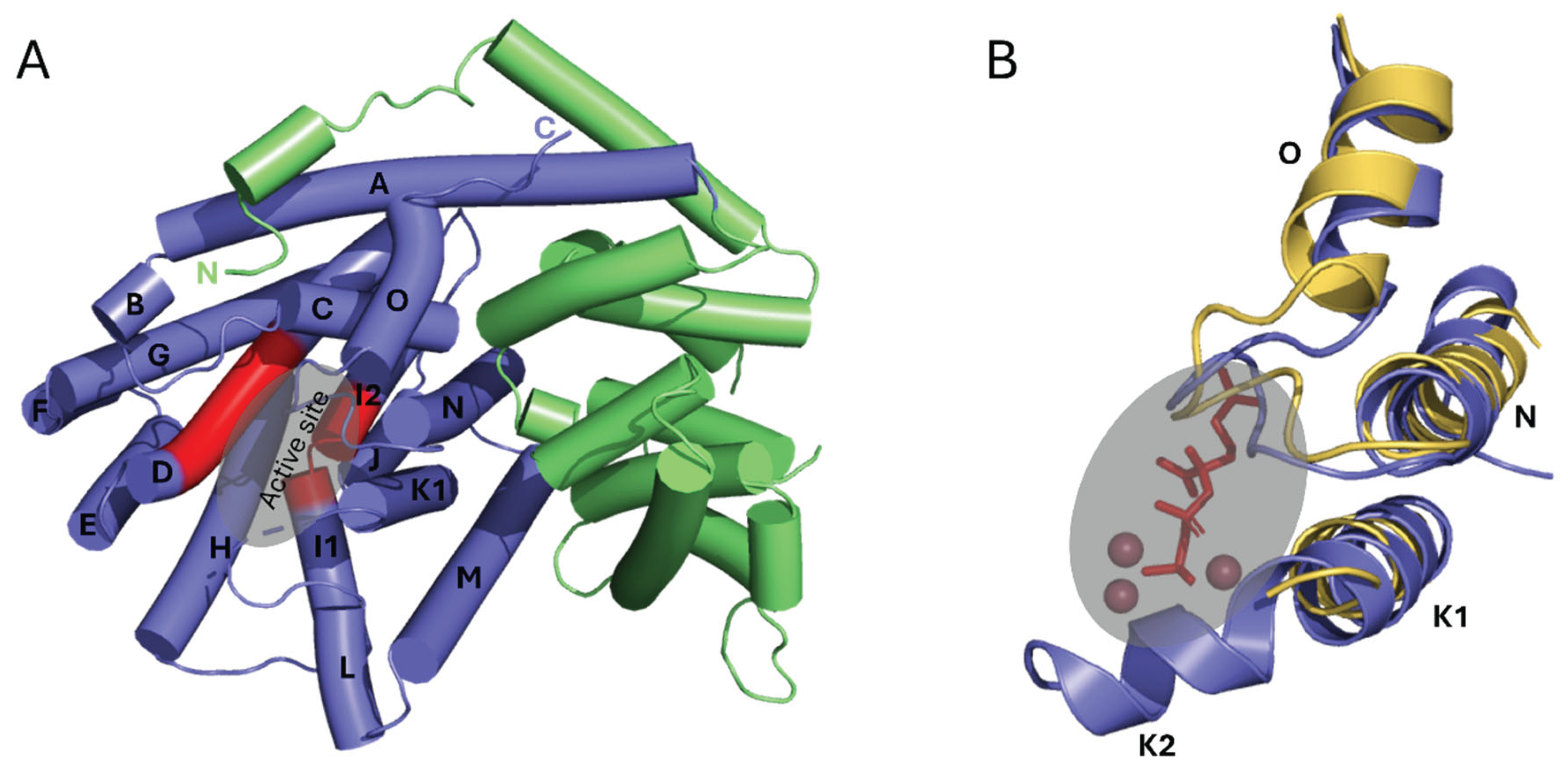
3.1. Structural Overview of CsLMS
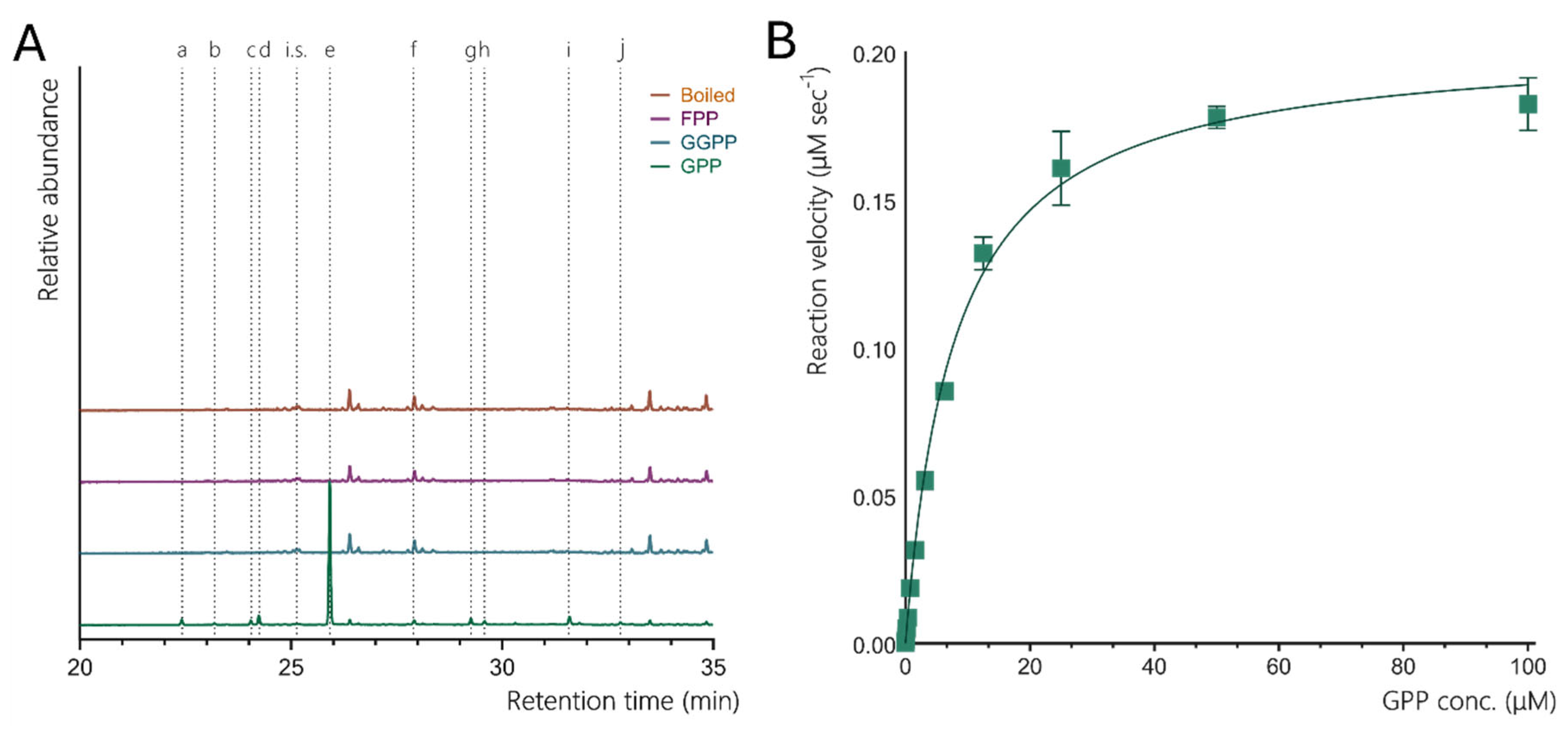
3.2. Enzymatic Activity of CsLMS
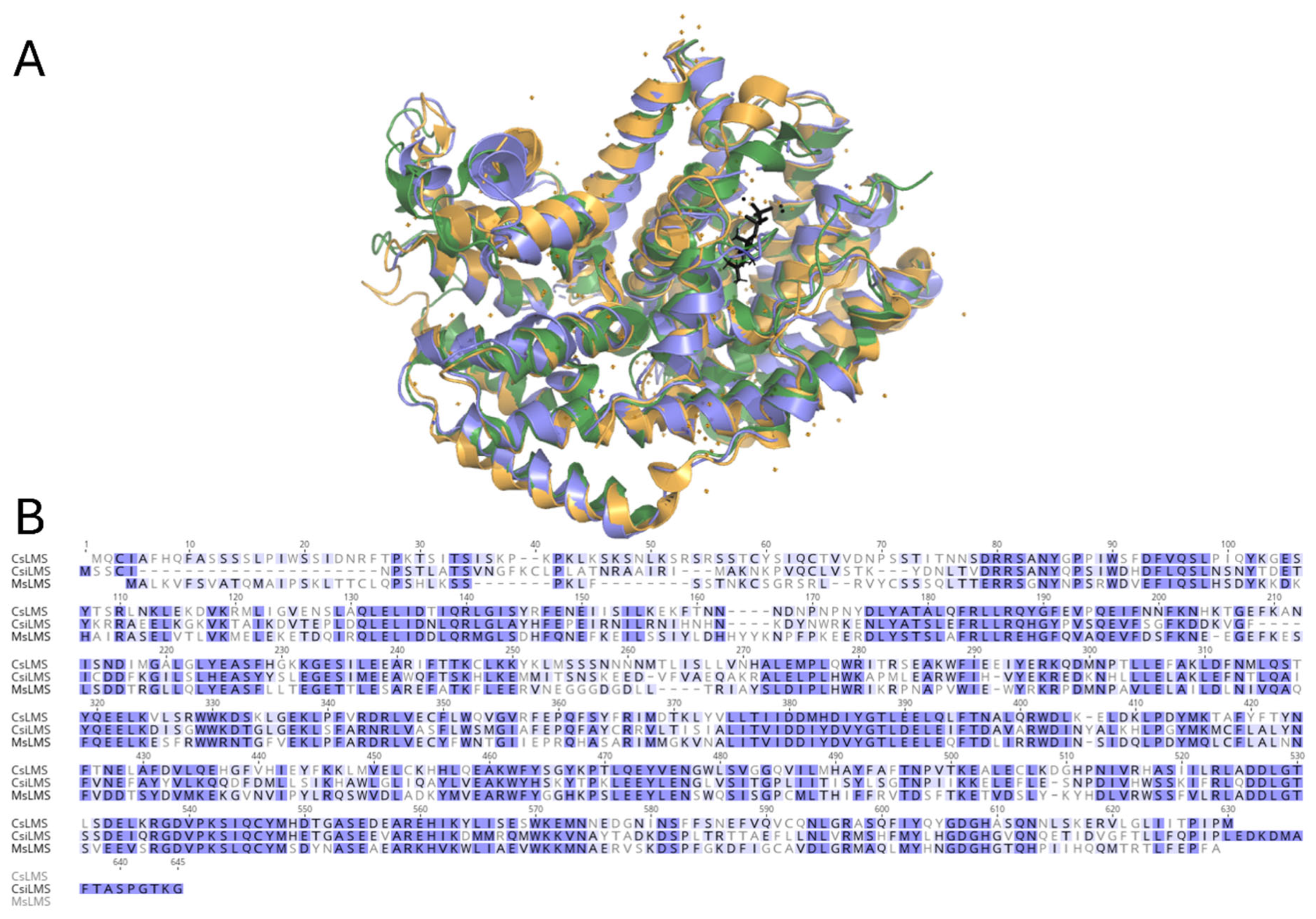
3.3. Comparison of the Overall Structure of CsLMS to That of Other TPS
4. Conclusion
Supplementary Materials
Author Contributions
Acknowledgements
References
- Allen, K. D., McKernan, K., Pauli, C., Roe, J., Torres, A., & Gaudino, R. (2019). Genomic characterization of the complete terpene synthase gene family from Cannabis sativa. PLoS ONE, 14(9), e0222363. [CrossRef]
- Anarat-Cappillino, G., & Sattely, E. S. (2014). The chemical logic of plant natural product biosynthesis. In Current Opinion in Plant Biology (Vol. 19, pp. 51–58). Elsevier Ltd. [CrossRef]
- Booth, J. K., Page, J. E., & Bohlmann, J. (2017). Terpene synthases from Cannabis sativa. PLoS ONE, 12(3). [CrossRef]
- Cao, R., Zhang, Y., Mann, F. M., Huang, C., Mukkamala, D., Hudock, M. P., Mead, M. E., Prisic, S., Wang, K., Lin, F. Y., Chang, T. K., Peters, R. J., & Oldfield, E. (2010). Diterpene cyclases and the nature of the isoprene fold. Proteins: Structure, Function and Bioinformatics, 78(11), 2417–2432. [CrossRef]
- Chen, F., Tholl, D., Bohlmann, J., & Pichersky, E. (2011). The family of terpene synthases in plants: A mid-size family of genes for specialized metabolism that is highly diversified throughout the kingdom. Plant Journal, 66(1), 212–229. [CrossRef]
- Davis, E. M., & Croteau, R. B. (2000). Cyclization enzymes in the biosynthesis of monoterpenes, sesquiterpenes, and diterpenes. In Topics in Current Chemistry (Vol. 301).
- Degenhardt, J., Köllner, T. G., & Gershenzon, J. (2009). Monoterpene and sesquiterpene synthases and the origin of terpene skeletal diversity in plants. In Phytochemistry (Vol. 70, Issues 15–16, pp. 1621–1637). Pergamon. [CrossRef]
- Emsley, P., Lohkamp, B., Scott, W. G., & Cowtan, K. (2010). Features and development of Coot. Acta Crystallographica Section D: Biological Crystallography, 66(4), 486–501. [CrossRef]
- Gao, Y., Honzatko, R. B., & Peters, R. J. (2012). Terpenoid synthase structures: A so far incomplete view of complex catalysis. In Natural Product Reports (Vol. 29, Issue 10, pp. 1153–1175). Royal Society of Chemistry. [CrossRef]
- Gershenzon, J., & Croteau, R. (1990). Biochemistry of the Mevalonic Acid Pathway to Terpenoids.
- Gershenzon, J., & Dudareva, N. (2007). The function of terpene natural products in the natural world. In Nature Chemical Biology (Vol. 3, Issue 7, pp. 408–414). [CrossRef]
- Günnewich, N., Page, J. E., Köllner, T. G., Degenhardt, J., & Kutchan, T. M. (2007). Functional expression and characterization of trichome-specific (-)-limonene synthase and (+)-α-pinene synthase from Cannabis sativa. Natural Product Communications, 2(3), 223–232. [CrossRef]
- Holcomb, J., Spellmon, N., Zhang, Y., Doughan, M., Li, C., & Yang, Z. (2017). Protein crystallization: Eluding the bottleneck of X-ray crystallography. In AIMS Biophysics (Vol. 4, Issue 4, pp. 557–575). American Institute of Mathematical Sciences. [CrossRef]
- Hyatt, D. C., Youn, B., Zhao, Y., Santhamma, B., Coates, R. M., Croteau, R. B., & Kang, C. (2007a). Structure of limonene synthase, a simple model for terpenoid cyclase catalysis. Proceedings of the National Academy of Sciences of the United States of America, 104(13), 5360–5365. [CrossRef]
- Hyatt, D. C., Youn, B., Zhao, Y., Santhamma, B., Coates, R. M., Croteau, R. B., & Kang, C. (2007b). Structure of limonene synthase, a simple model for terpenoid cyclase catalysis. Proceedings of the National Academy of Sciences of the United States of America, 104(13), 5360–5365. [CrossRef]
- Jia, Q., Brown, R., K, T. G., Fu, J., Chen, X., Ka-Shu Wong, G., Gershenzon, J., Peters, R. J., & Chen, F. (2022). Origin and early evolution of the plant terpene synthase family. [CrossRef]
- Karunanithi, P. S., & Zerbe, P. (2019). Terpene Synthases as Metabolic Gatekeepers in the Evolution of Plant Terpenoid Chemical Diversity. Frontiers in Plant Science, 10. [CrossRef]
- Kumar, R. P., Morehouse, B. R., Matos, J. O., Malik, K., Lin, H., Krauss, I. J., & Oprian, D. D. (2017). Structural Characterization of Early Michaelis Complexes in the Reaction Catalyzed by (+)-Limonene Synthase from Citrus sinensis Using Fluorinated Substrate Analogues. Biochemistry, 56(12), 1716–1725. [CrossRef]
- Lange, B. M. (2015). The evolution of plant secretory structures and emergence of terpenoid chemical diversity. Annual Review of Plant Biology, 66, 139–159. [CrossRef]
- McCoy, A. J., Grosse-Kunstleve, R. W., Adams, P. D., Winn, M. D., Storoni, L. C., & Read, R. J. (2007). Phaser crystallographic software. Journal of Applied Crystallography, 40(4), 658–674. [CrossRef]
- Morehouse, B. R., Kumar, R. P., Matos, J. O., Olsen, S. N., Entova, S., & Oprian, D. D. (2017). Functional and Structural Characterization of a (+)-Limonene Synthase from Citrus sinensis. Biochemistry, 56(12), 1706–1715. [CrossRef]
- Murshudov, G. N., Vagin, A. A., & Dodson, E. J. (1997). Refinement of macromolecular structures by the maximum-likelihood method. In Acta Crystallographica Section D: Biological Crystallography (Vol. 53, Issue 3, pp. 240–255). [CrossRef]
- Pegan, S., Tian, Y., Sershon, V., & Mesecar, A. (2009). A Universal, Fully Automated High Throughput Screening Assay for Pyrophosphate and Phosphate Release from Enzymatic Reactions. Combinatorial Chemistry & High Throughput Screening, 13(1), 27–38. [CrossRef]
- Pichersky, E., & Gershenzon, J. (2002). The formation and function of plant volatiles: perfumes for pollinator attraction and defense. Current Opinion in Plant Biology, 5(3), 237–243. [CrossRef]
- Potterton, E., Briggs, P., Turkenburg, M., & Dodson, E. (2003). A graphical user interface to the CCP4 program suite. Acta Crystallographica - Section D Biological Crystallography, 59(7), 1131–1137. [CrossRef]
- Rehbein, P., Berz, J., Kreisel, P., & Schwalbe, H. (2019). “CodonWizard” – An intuitive software tool with graphical user interface for customizable codon optimization in protein expression efforts. Protein Expression and Purification, 160, 84–93. [CrossRef]
- Schiff, W. H., & Oprian, D. D. (2023). Mutational Analysis of (+)-Limonene Synthase. Biochemistry, 62(16), 2472–2479. [CrossRef]
- Srividya, N., Davis, E. M., Croteau, R. B., & Lange, B. M. (2015). Functional analysis of (4S)-limonene synthase mutants reveals determinants of catalytic outcome in a model monoterpene synthase. Proceedings of the National Academy of Sciences of the United States of America, 112(11), 3332–3337. [CrossRef]
- Srividya, N., Lange, I., & Lange, B. M. (2020). Determinants of Enantiospecificity in Limonene Synthases. Biochemistry, 59(17), 1661–1664. [CrossRef]
- Tholl, D. (2006). Terpene synthases and the regulation, diversity and biological roles of terpene metabolism. Current Opinion in Plant Biology, 9(3), 297–304. [CrossRef]
- Trapp, S. C., & Croteau, R. B. (2001). Genomic Organization of Plant Terpene Synthases and Molecular Evolutionary Implications.
- Vardakou, M., Salmon, M., Faraldos, J. A., & O’Maille, P. E. (2014). Comparative analysis and validation of the malachite green assay for the high throughput biochemical characterization of terpene synthases. MethodsX, 1, e187–e196. [CrossRef]
- Wendt, U., & Schulz, G. E. (1998). Isoprenoid biosynthesis: manifold chemistry catalyzed by similar enzymes. Structure, 6, 127–133. http://biomednet.com/elecref/0969212600600127.
- Wiles, D., Roest, J., Shanbhag, B., Vivian, J., & Beddoe, T. (2025). Integrated Platform for Structural and Functional Analysis of Cannabis sativa Terpene Synthases. [CrossRef]
- Xu, J., Ai, Y., Wang, J., Xu, J., Zhang, Y., Phytochemistry, D. Y.-, & 2017, undefined. (n.d.). Converting S-limonene synthase to pinene or phellandrene synthases reveals the plasticity of the active site. Elsevier. Retrieved June 7, 2021, from https://www.sciencedirect.com/science/article/pii/S0031942217300882?casa_token=kPos2qvmfeMAAAAA:lzc4Ji2MyvlW5mrxQpgFeCUgcblqvTv1p2m9RyEMM3JbTbij43FfnnO1FpNqamueTCVtfZan5WUt.
- Xu, J., Xu, J., Ai, Y., Farid, R. A., Tong, L., & Yang, D. (2018). Mutational analysis and dynamic simulation of S-limonene synthase reveal the importance of Y573: Insight into the cyclization mechanism in monoterpene synthases. Archives of Biochemistry and Biophysics, 638(December 2017), 27–34. [CrossRef]
- Yu, Q. (2017). Exploring the Monoterpene Cyclization Mechanism by Studying (+)-Limonene Synthase Using Novel Fluorinated Substrate Analogues. 13(3), 1576–1580. [CrossRef]
| Data collection statistics | |
| Temperature (K) | 100 |
| X-ray source | MX2 Australian synchrotron |
| Space group | P3221 |
| Cell Dimensions (Å) | a = 97.3, b = 97.3, c = 117.7 |
| Resolution (Å) | 60 – 3.2 (3.25 – 3.20) |
| Total no. observations | 81866 (4326) |
| No. unique observations | 10899 (545) |
| Multiplicity | 7.5 (7.9) |
| Data completeness (%) | 99.1 (100) |
| I/sI | 7.5 (5.2) |
| Rmerge (%)1 | 0.29 (0.45) |
| CC (1/2) | 0.98 (0.12) |
| Refinement statistics | |
| Non-hydrogen atoms | |
| Protein | 4290 |
| Rfactor (%)2 | 18 |
| Rfree (%)2 | 23.0 |
| r.m.s.d. from ideality | |
| Bond lengths (Å) | 0.003 |
| Bond angles (°) | 0.591 |
| Dihedrals (°) | 14.5 |
| Ramachandran plot | |
| Favoured regions (%) | 94.9 |
| Allowed regions (%) | 5.1 |
| B-factors (Å2) | |
| Wilson | 41 |
| Average protein | 42 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




