Submitted:
28 May 2025
Posted:
29 May 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Bacteriophages: Abundance in the Environment
1.2. Bacteriophages: A Vehicle for Resistance Genes
1.3. Antibiotic Resistance Genes Transmission Mechanisms
1.3.1. Conjugation
1.3.2. Transformation
1.3.3. Transduction
2. Resistance Genes Transmission Mechanisms in Bacteriophages
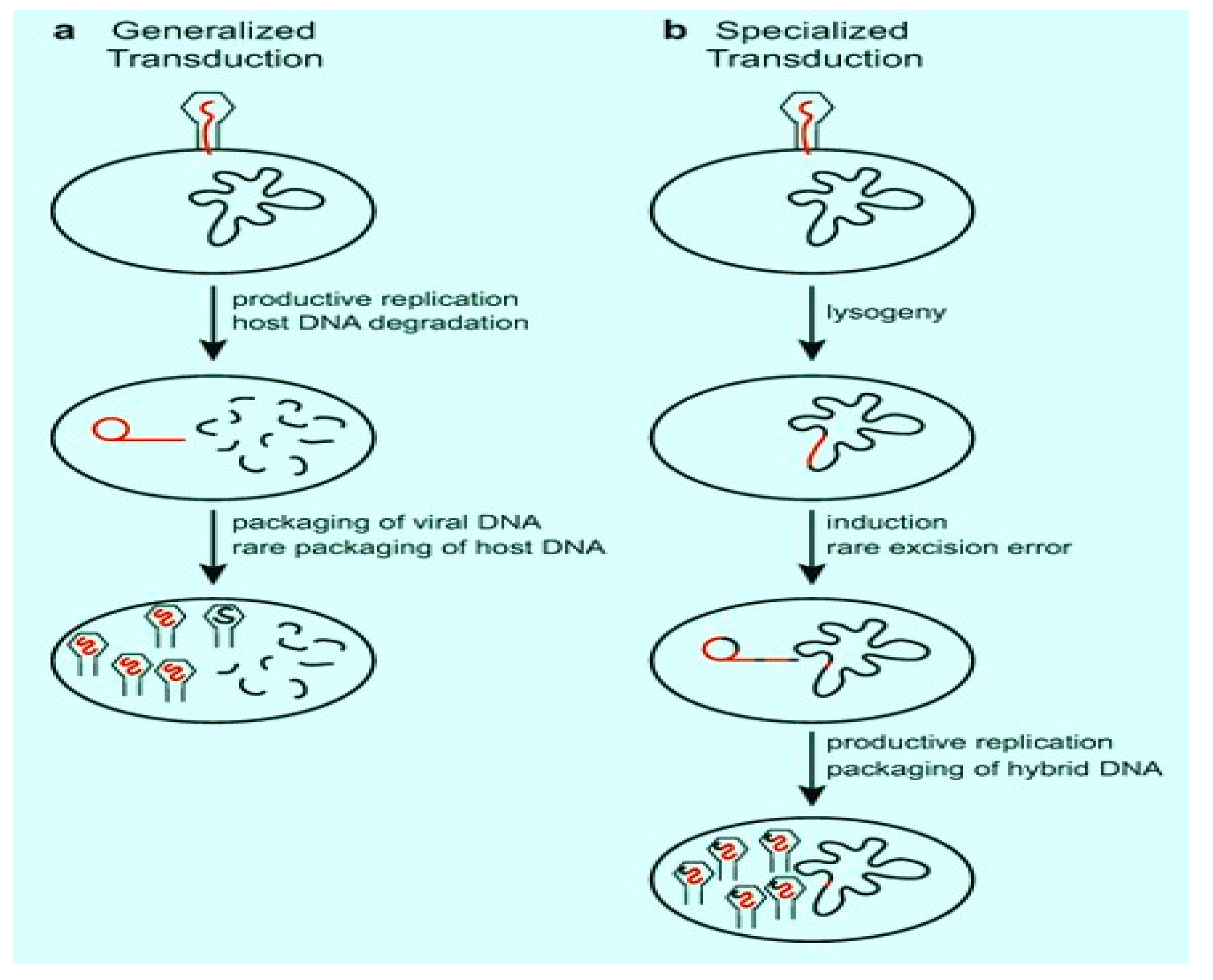
Phages Dissemination of ARGs via Transduction
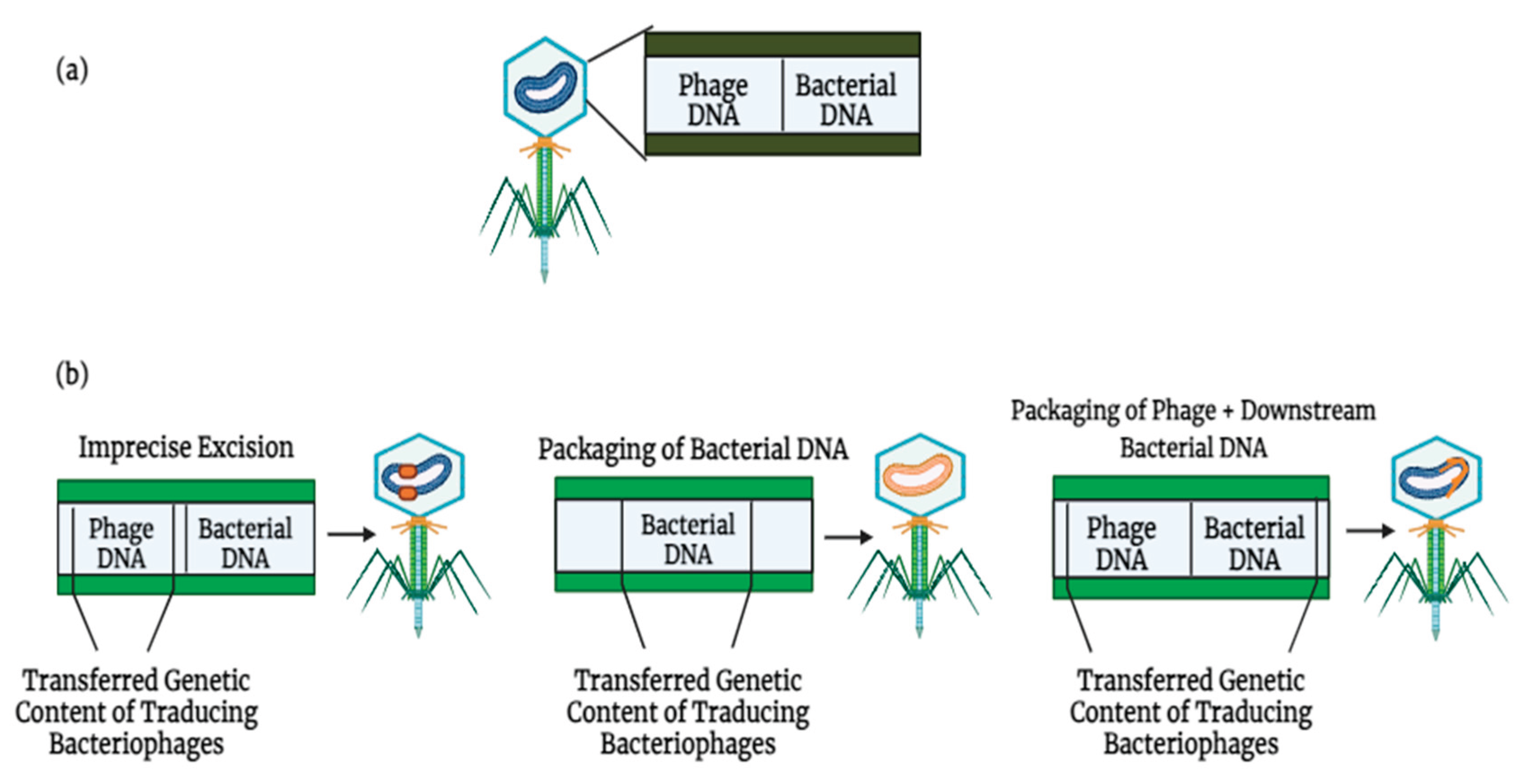
3. Bacterial Genes and ARGs in Viral Communities
ARGs by Bacteriophages
4. Conclusion/Future Perspective
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| ARG | Antibiotic resistance genes |
| HGT | Horizontal gene transfer |
| MGEs | Mobile genetic elements |
| WHO | World Health Organization |
| CDC | Center For Disease Control |
| VBR | Virus-to-bacterium |
| VLPs | Virus-like particles |
References
- Kalyani, A.; Lakshmi, M.M.; Ahalya, N.; Anusha, P.; Seema, S.; Maneesha, T. Classification Of Different Antibiotics And Their Adverse Effects And Uses From Origin To Present. Int. J. Pharm. Sci. 2024, 2, 98–121. [Google Scholar] [CrossRef]
- Halawa, E.M.; Fadel, M.; Al-Rabia, M.W.; Behairy, A.; Nouh, N.A.; Abdo, M.; Olga, R.; Fericean, L.; Atwa, A.M.; El-Nablaway, M. Antibiotic action and resistance: updated review of mechanisms, spread, influencing factors, and alternative approaches for combating resistance. Frontiers in Pharmacology 2024, 14, 1305294. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, G.; Saigal, S.; Elongavan, A. Action and resistance mechanisms of antibiotics: A guide for clinicians. Journal of Anaesthesiology Clinical Pharmacology 2017, 33, 300–305. [Google Scholar] [CrossRef] [PubMed]
- Murray, C.J.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Aguilar, G.R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. The lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- Sher, S.; Richards, G.P.; Parveen, S.; Williams, H.N. Characterization of Antibiotic Resistance in Shewanella Species: An Emerging Pathogen in Clinical and Environmental Settings. Microorganisms 2025, 13, 1115. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhang, Q.; Wang, T.; Xu, N.; Lu, T.; Hong, W.; Penuelas, J.; Gillings, M.; Wang, M.; Gao, W. Assessment of global health risk of antibiotic resistance genes. Nature communications 2022, 13, 1553. [Google Scholar] [CrossRef]
- Zhang, Y.; Guo, Y.; Qiu, T.; Gao, M.; Wang, X. Bacteriophages: Underestimated vehicles of antibiotic resistance genes in the soil. Frontiers in Microbiology 2022, 13, 936267. [Google Scholar] [CrossRef]
- Coque, T.M.; Cantón, R.; Pérez-Cobas, A.E.; Fernández-de-Bobadilla, M.D.; Baquero, F. Antimicrobial resistance in the global health network: known unknowns and challenges for efficient responses in the 21st century. Microorganisms 2023, 11, 1050. [Google Scholar] [CrossRef]
- Salam, M.A.; Al-Amin, M.Y.; Salam, M.T.; Pawar, J.S.; Akhter, N.; Rabaan, A.A.; Alqumber, M.A. Antimicrobial resistance: a growing serious threat for global public health. in Healthcare. 2023. Multidisciplinary Digital Publishing Institute.
- Munita, J.M.; Arias, C.A. Mechanisms of antibiotic resistance. Microbiology spectrum 2016, 4, 15. [Google Scholar] [CrossRef]
- Stokes, H.W.; Gillings, M.R. Gene flow, mobile genetic elements and the recruitment of antibiotic resistance genes into Gram-negative pathogens. FEMS microbiology reviews 2011, 35, 790–819. [Google Scholar] [CrossRef]
- Puxty, R.J.; Millard, A.D. Functional ecology of bacteriophages in the environment. Current Opinion in Microbiology 2023, 71, 102245. [Google Scholar] [CrossRef] [PubMed]
- Batinovic, S.; Wassef, F.; Knowler, S.A.; Rice, D.T.; Stanton, C.R.; Rose, J.; Tucci, J.; Nittami, T.; Vinh, A.; Drummond, G.R. Bacteriophages in natural and artificial environments. Pathogens 2019, 8, 100. [Google Scholar] [CrossRef]
- Glasner, M.E. Finding enzymes in the gut metagenome. Science 2017, 355, 577–578. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, S.; Baker, K.; Padman, B.S.; Patwa, R.; Dunstan, R.A.; Weston, T.A.; Schlosser, K.; Bailey, B.; Lithgow, T.; Lazarou, M. Bacteriophage transcytosis provides a mechanism to cross epithelial cell layers. MBio 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Naureen, Z.; Dautaj, A.; Anpilogov, K.; Camilleri, G.; Dhuli, K.; Tanzi, B.; Maltese, P.E.; Cristofoli, F.; De Antoni, L.; Beccari, T. Bacteriophages presence in nature and their role in the natural selection of bacterial populations. Acta Bio Medica: Atenei Parmensis 2020, 91, e2020024. [Google Scholar]
- Florent, P.; Cauchie, H.-M.; Herold, M.; Jacquet, S.; Ogorzaly, L. Soil ph, calcium content and bacteria as major factors responsible for the distribution of the known fraction of the DNA bacteriophage populations in soils of Luxembourg. Microorganisms 2022, 10, 1458. [Google Scholar] [CrossRef]
- Simmonds, P.; Aiewsakun, P. Virus classification–where do you draw the line? Archives of virology 2018, 163, 2037–2046. [Google Scholar] [CrossRef]
- Balcazar, J.L. Bacteriophages as vehicles for antibiotic resistance genes in the environment. PLoS pathogens 2014, 10, e1004219. [Google Scholar] [CrossRef]
- Sher, S.; Sultan, S.; Rehman, A. Characterization of multiple metal resistant Bacillus licheniformis and its potential use in arsenic contaminated industrial wastewater. Applied Water Science 2021, 11, 1–7. [Google Scholar] [CrossRef]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile genetic elements associated with antimicrobial resistance. Clinical microbiology reviews 2018, 31. [Google Scholar] [CrossRef]
- Li, B.; Qiu, Y.; Song, Y.; Lin, H.; Yin, H. Dissecting horizontal and vertical gene transfer of antibiotic resistance plasmid in bacterial community using microfluidics. Environment International 2019, 131, 105007. [Google Scholar] [CrossRef] [PubMed]
- Tao, S.; Chen, H.; Li, N.; Wang, T.; Liang, W. The spread of antibiotic resistance genes in vivo model. Canadian Journal of Infectious Diseases and Medical Microbiology 2022, 2022, 3348695. [Google Scholar] [CrossRef]
- Lopatkin, A.J.; Sysoeva, T.A.; You, L. Dissecting the effects of antibiotics on horizontal gene transfer: Analysis suggests a critical role of selection dynamics. Bioessays 2016, 38, 1283–1292. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Liu, Y.; Zhang, Q.; Jin, L.; Wang, Q.; Zhang, Y.; Wang, X.; Hu, M.; Li, L.; Qi, J. The prevalence of colistin resistance in Escherichia coli and Klebsiella pneumoniae isolated from food animals in China: coexistence of mcr-1 and blaNDM with low fitness cost. International journal of antimicrobial agents 2018, 51, 739–744. [Google Scholar] [CrossRef] [PubMed]
- Evans, B.A.; Kumar, A.; Castillo-Ramírez, S. Genomic basis of antibiotic resistance and virulence in Acinetobacter. Frontiers in Microbiology 2021, 12, 690. [Google Scholar] [CrossRef]
- Yu, Z.; Wang, Y.; Henderson, I.R.; Guo, J. Artificial sweeteners stimulate horizontal transfer of extracellular antibiotic resistance genes through natural transformation. The ISME Journal 2022, 16, 543–554. [Google Scholar] [CrossRef]
- Winter, M.; Buckling, A.; Harms, K.; Johnsen, P.J.; Vos, M. Antimicrobial resistance acquisition via natural transformation: context is everything. Current Opinion in Microbiology 2021, 64, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Voigt, E.; Rall, B.C.; Chatzinotas, A.; Brose, U.; Rosenbaum, B. Phage strategies facilitate bacterial coexistence under environmental variability. PeerJ 2021, 9, e12194. [Google Scholar] [CrossRef]
- Haaber, J.; Penadés, J.R.; Ingmer, H. Transfer of antibiotic resistance in Staphylococcus aureus. Trends in microbiology 2017, 25, 893–905. [Google Scholar] [CrossRef]
- Kasman, L.; Porter, L. Bacteriophages. StatPearls. StatPearls Publishing. 2022.
- Schneider, C.L. Bacteriophage-mediated horizontal gene transfer: transduction. Bacteriophages: biology, technology, therapy 2021, 151-192.
- Wood, S.J.; Kuzel, T.M.; Shafikhani, S.H. Pseudomonas aeruginosa: infections, animal modeling, and therapeutics. Cells 2023, 12, 199. [Google Scholar] [CrossRef]
- Fišarová, L.; Botka, T.; Du, X.; Mašlaňová, I.; Bárdy, P.; Pantůček, R.; Benešík, M.; Roudnický, P.; Winstel, V.; Larsen, J. Staphylococcus epidermidis phages transduce antimicrobial resistance plasmids and mobilize chromosomal islands. Msphere 2021, 6. [Google Scholar] [CrossRef] [PubMed]
- Frosini, S.M.; Bond, R.; McCarthy, A.J.; Feudi, C.; Schwarz, S.; Lindsay, J.A.; Loeffler, A. Genes on the move: in vitro transduction of antimicrobial resistance genes between human and canine staphylococcal pathogens. Microorganisms 2020, 8, 2031. [Google Scholar] [CrossRef]
- Hussain, H.I.; Aqib, A.I.; Seleem, M.N.; Shabbir, M.A.; Hao, H.; Iqbal, Z.; Kulyar, M.F.-e.-A.; Zaheer, T.; Li, K. Genetic basis of molecular mechanisms in β-lactam resistant gram-negative bacteria. Microbial pathogenesis 2021, 158, 105040. [Google Scholar] [CrossRef]
- Zhang, J.; He, X.; Shen, S.; Shi, M.; Zhou, Q.; Liu, J.; Wang, M.; Sun, Y. Effects of the newly isolated T4-like phage on transmission of plasmid-borne antibiotic resistance genes via generalized transduction. Viruses 2021, 13, 2070. [Google Scholar] [CrossRef]
- Zeman, M.; Mašlaňová, I.; Indráková, A.; Šiborová, M.; Mikulášek, K.; Bendíčková, K.; Plevka, P.; Vrbovská, V.; Zdráhal, Z.; Doškař, J. Staphylococcus sciuri bacteriophages double-convert for staphylokinase and phospholipase, mediate interspecies plasmid transduction, and package mecA gene. Scientific reports 2017, 7, 46319. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Quiles-Puchalt, N.; Chiang, Y.N.; Bacigalupe, R.; Fillol-Salom, A.; Chee, M.S.J.; Fitzgerald, J.R.; Penadés, J.R. Genome hypermobility by lateral transduction. Science 2018, 362, 207–212. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, S.; Fillol-Salom, A.; Quiles-Puchalt, N.; Ibarra-Chávez, R.; Haag, A.F.; Chen, J.; Penadés, J.R. Bacterial chromosomal mobility via lateral transduction exceeds that of classical mobile genetic elements. Nature communications 2021, 12, 6509. [Google Scholar] [CrossRef]
- Chen, Q.; Dharmaraj, T.; Cai, P.C.; Burgener, E.B.; Haddock, N.L.; Spakowitz, A.J.; Bollyky, P.L. Bacteriophage and bacterial susceptibility, resistance, and tolerance to antibiotics. Pharmaceutics 2022, 14, 1425. [Google Scholar] [CrossRef]
- Goh, S.; Hussain, H.; Chang, B.J.; Emmett, W.; Riley, T.V.; Mullany, P. Phage ϕC2 mediates transduction of Tn 6215, encoding erythromycin resistance, between Clostridium difficile strains. MBio 2013, 4. [Google Scholar] [CrossRef]
- Tian, Y.; Yu, H.; Wang, Z. Distribution of acquired antibiotic resistance genes among Enterococcus spp. isolated from a hospital in Baotou, China. BMC research notes 2019, 12, 1–5. [Google Scholar] [CrossRef]
- Colavecchio, A.; Cadieux, B.; Lo, A.; Goodridge, L.D. Bacteriophages contribute to the spread of antibiotic resistance genes among foodborne pathogens of the Enterobacteriaceae family–a review. Frontiers in microbiology 2017, 8, 1108. [Google Scholar] [CrossRef]
- Li, D.; Ge, Y.; Wang, N.; Shi, Y.; Guo, G.; Zou, Q.; Liu, Q. Identification and characterization of a novel major facilitator superfamily efflux pump, SA09310, mediating tetracycline resistance in Staphylococcus aureus. Antimicrobial Agents and Chemotherapy 2023, 67, e01696-22. [Google Scholar] [CrossRef]
- Akindolire, M.A.; Babalola, O.O.; Ateba, C.N. Detection of antibiotic resistant Staphylococcus aureus from milk: A public health implication. International journal of environmental research and public health 2015, 12, 10254–10275. [Google Scholar] [CrossRef]
- Cattoir, V. Mechanisms of Streptococcus pyogenes antibiotic resistance. Streptococcus pyogenes: Basic Biology to Clinical Manifestations [Internet]. 2nd edition, 2022.
- Mutuku, C.; Gazdag, Z.; Melegh, S. Occurrence of antibiotics and bacterial resistance genes in wastewater: resistance mechanisms and antimicrobial resistance control approaches. World Journal of Microbiology and Biotechnology 2022, 38, 152. [Google Scholar] [CrossRef] [PubMed]
- Pailhoriès, H.; Herrmann, J.-L.; Velo-Suarez, L.; Lamoureux, C.; Beauruelle, C.; Burgel, P.-R.; Héry-Arnaud, G. Antibiotic resistance in chronic respiratory diseases: from susceptibility testing to the resistome. European Respiratory Review 2022, 31. [Google Scholar] [CrossRef] [PubMed]
- Debroas, D.; Siguret, C. Viruses as key reservoirs of antibiotic resistance genes in the environment. The ISME journal 2019, 13, 2856–2867. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Rubio, L.; Haarmann, N.; Schwidder, M.; Muniesa, M.; Schmidt, H. Bacteriophages of Shiga toxin-producing Escherichia coli and their contribution to pathogenicity. Pathogens 2021, 10, 404. [Google Scholar] [CrossRef]
- Ballesté, E.; Blanch, A.R.; Muniesa, M.; García-Aljaro, C.; Rodríguez-Rubio, L.; Martín-Díaz, J.; Pascual-Benito, M.; Jofre, J. Bacteriophages in sewage: abundance, roles, and applications. FEMS microbes 2022, 3, xtac009. [Google Scholar] [CrossRef]
- McShan, W.M.; Nguyen, S.V. The bacteriophages of Streptococcus pyogenes. Streptococcus pyogenes: Basic Biology to Clinical Manifestations [Internet], 2016.
- Giovanetti, E.; Brenciani, A.; Vecchi, M.; Manzin, A.; Varaldo, P.E. Prophage association of mef (A) elements encoding efflux-mediated erythromycin resistance in Streptococcus pyogenes. Journal of Antimicrobial Chemotherapy 2005, 55, 445–451. [Google Scholar] [CrossRef]
- Mazaheri Nezhad Fard, R.; Barton, M.; Heuzenroeder, M. Bacteriophage-mediated transduction of antibiotic resistance in enterococci. Letters in applied microbiology 2011, 52, 559–564. [Google Scholar] [CrossRef]
- Colomer-Lluch, M.; Jofre, J.; Muniesa, M. antibiotic resistance genes in environmental bacteriophages.
| Bacteria | Phage | Antibiotic | Transduction frequency (transductants/pfu) |
References |
|---|---|---|---|---|
| Clostridium difficile | ϕC2 | Erythromycin | 10-6 | [42] |
| Enterococcus | EGRM195 | Tetracycline Gentamicin |
10-8-10-9 (tet) 10-7-10-9 (gent) |
[43] |
| Salmonella enterica serovar typhimurium | ES18 | Tetracycline Chloramphenicol |
10-8 (tet) 10-9 (cam) |
[44] |
| Staphylococcal species | Φ80a and ΦJB | Penicillin Tetracycline |
10-5-10-6 | [45] |
| Staphylococcus aureus | 80a | Streptomycin (in SaPi) | 10-1 | [46] |
| Streptococcus pyogenes | nd | Tetracycline Chloramphenicol Macrolides Lincomycin Clindamycin |
10-5-10-6 | [47] |
| Antibiotic | Genes (Resistance) | Natural Source |
|---|---|---|
| Bacitracin | bcrA | Viromes of the human gut, microbiomes in fecal of swine, sputum microbiota of cystic fibrosis |
| B-lactams antibiotics | blaOXA-2, blaPSE-1, blaPSE-4, blaPSE-type genes | Sewage |
| B-lactams antibiotics | blaTEM, blaCTX-M | Water from sewage, river, and animal waste |
| B-lactams antibiotics | blaCTX-M-10 | Enterobacteriaceae |
| Ampicillin | blaCMY-2 | Salmonella enterica |
| Trimethoprim | dfrAa | Microbiomes of Swine fecal |
| Fluoroquinolones |
Fluoroquinolone resistance genes |
sputum microbiota of cystic fibrosis |
| Macrolides | macB | microbiomes in fecal of swine |
| Methicillin | mecA | Water from sewage, river, and animal waste |
| Macrolides | mefA | Streptococcus pyogenes |
| Tetracycline | tetW | Viromes of human gut, microbiomes in the feces of swine |
| Tetracycline | Tet37 | Microbiomes in fecal of swine |
| Tetracycline | tetA, tetB | S. enterica |
| Vancomycin | Vancomycin resistance genes | Viromes of the human gut, microbiomes in the feces of swine |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




