Submitted:
27 April 2025
Posted:
28 April 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Acquisition
2.2. Data Processing
2.3. Phenotype Simulation
2.4. Genomic Selection Methods
2.5. Validation Procedures
2.6. Data Analysis and Visualization
2.7. Software and Code Availability
3. Results
3.1. Summary of Method Performance Across Validations
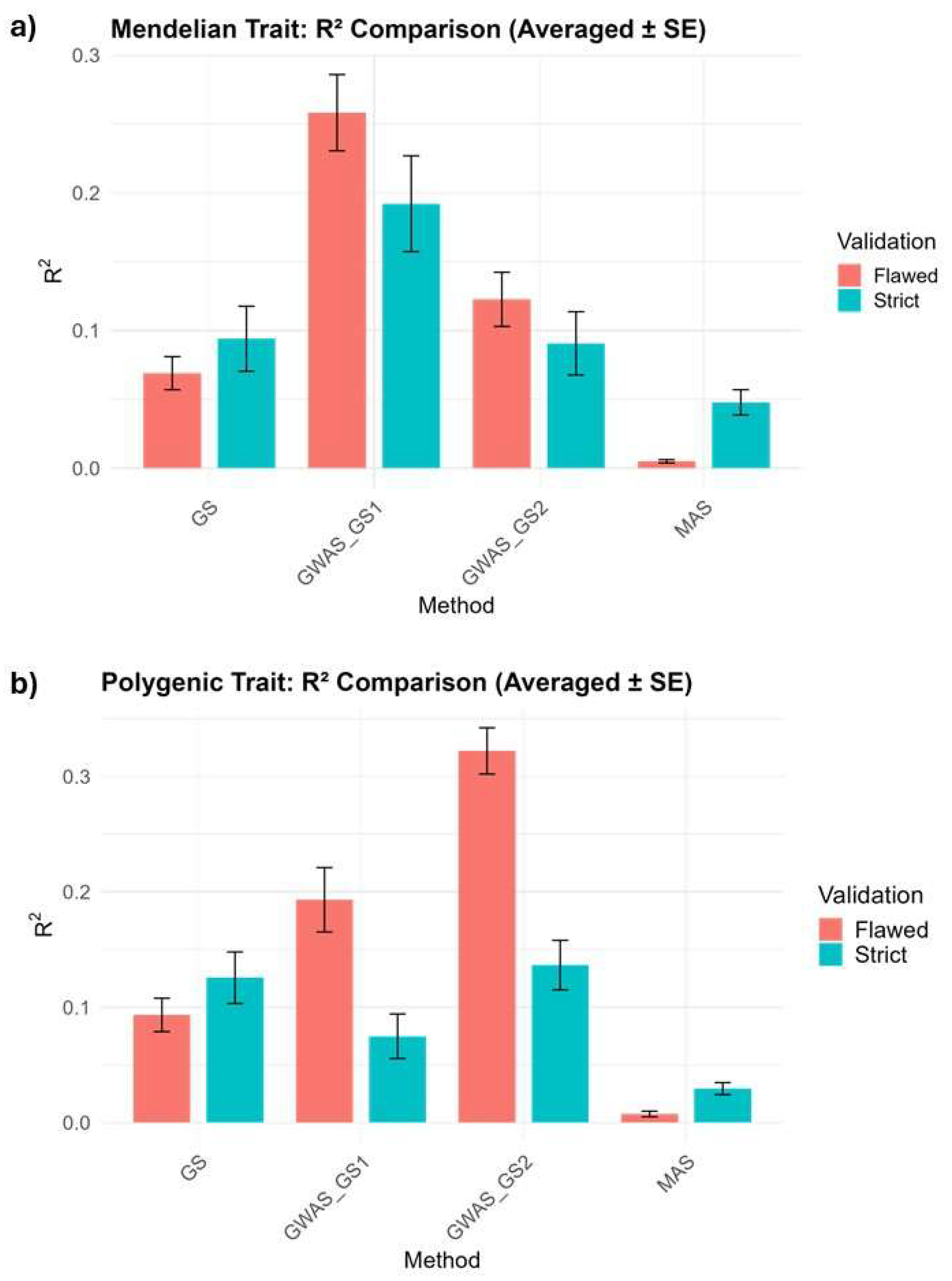
3.2. Mendelian Trait
3.3. Polygenic Trait
3.4. Validation Comparison
4. Discussion
4.1. Validation Bias and Method Performance
4.2. Suitability of Methods for Different Trait Architectures
4.3. Unexpected Patterns in GS and MAS Performance
4.4. Implications for Wheat Breeding
4.5. Limitations and Future Directions
5. Conclusion
Acknowledgement
References
- Bernardo, R. (2014). Genomewide selection when major genes are known. Crop Science, 54(1), 68-75. [CrossRef]
- Budhlakoti, N., Kushwaha, A. K., Rai, A., Chaturvedi, K. K., Kumar, A., Pradhan, A. K.,... & Kumar, S. (2022). Genomic selection: A tool for accelerating the efficiency of molecular breeding for development of climate-resilient crops. Frontiers in Genetics, 13, 832153. [CrossRef]
- Crossa, J., Pérez-Rodríguez, P., Cuevas, J., Montesinos-López, O., Jarquín, D., De Los Campos, G., ….. & Varshney, R. K. (2017). Genomic selection in plant breeding: methods, models, and perspectives. Trends in Plant Science, 22(11), 961-975. [CrossRef]
- Daetwyler, H. D., Calus, M. P. L., Pong-Wong, R., de los Campos, G., & Hickey, J. M. (2010). The impact of genetic architecture on genome-wide evaluation methods. Genetics, 185(3), 1021-1031. [CrossRef]
- Goddard, M. E., & Hayes, B. J. (2009). Mapping genes for complex traits in domestic animals and their use in breeding programmes. Nature Reviews Genetics, 10(6), 381–391. [CrossRef]
- Lande, R., & Thompson, R. (1990). Efficiency of marker-assisted selection in the improvement of quantitative traits. Genetics, 124(3), 743–756. [CrossRef]
- Lipka, A. E., Tian, F., Wang, Q., Peiffer, J., Li, M., Bradbury, P. J.,... & Zhang, Z. (2012). GAPIT: genome association and prediction integrated tool. Bioinformatics, 28(18), 2397-2399. [CrossRef]
- McGowan, M., Wang, J., Dong, H., Liu, X., Jia, Y., Wang, X., Iwata, H., Li, Y., Lipka, A. E., & Zhang, Z. (2021). Ideas in genomic selection with the potential to transform plant molecular breeding: A review. Plant Breeding Reviews, 45, 273–343. [CrossRef]
- Medina, C. A., Kaur, H., Ray, I., & Yu, L. X. (2021). Strategies to increase prediction accuracy in genomic selection of complex traits in alfalfa (Medicago sativa L.). Cells, 10(12), 3372. [CrossRef]
- Meuwissen, T. H. E., Hayes, B. J., & Goddard, M. E. (2001). Prediction of total genetic value using genome-wide dense marker maps. Genetics, 157(4), 1819–1829. [CrossRef]
- Morales, N., Ogbonnaya, F. C., & Dreisigacker, S. (2022). T3/Wheat Database: A comprehensive resource for wheat breeding and genomics. Wheat Science Journal, 8(2), 45–56.
- R Core Team. (2023). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/.
- Ravelombola, W. S., Qin, J., Shi, A., Nice, L., Bao, Y., Lorenz, A.,... & Chen, S. (2020). Genome-wide association study and genomic selection for tolerance of soybean biomass to soybean cyst nematode infestation. PLoS One, 15(7), e0235089. [CrossRef]
- Spindel, J. E., Begum, H., Akdemir, D., Collard, B., Redoña, E., Jannink, J. L., & McCouch, S. (2016). Genome-wide prediction models that incorporate de novo GWAS are a powerful new tool for tropical rice improvement. Heredity, 116(4), 395-408.
- Tang, Y., Liu, X., Wang, J., Li, M., Wang, Q., Tian, F.,... & Zhang, Z. (2016). GAPIT version 2: an enhanced integrated tool for genomic association and prediction. The plant genome, 9(2), plantgenome2015-11. [CrossRef]
- Tester, M., & Langridge, P. (2010). Breeding technologies to increase crop production in a changing world. Science, 327(5967), 818–822. [CrossRef]
- Wang, J., & Zhang, Z. (2021). GAPIT version 3: boosting power and accuracy for genomic association and prediction. Genomics, proteomics & bioinformatics, 19(4), 629-640. [CrossRef]
- Zhang, Z., Liu, J., Ding, X., Bijma, P., de Koning, D. J., & Zhang, Q. (2010). Best linear unbiased prediction of genomic breeding values using a trait-specific marker-derived relationship matrix. PloS one, 5(9), e12648. [CrossRef]
- Zhang, Z., Ober, U., Erbe, M., Zhang, H., Gao, N., He, J.,... & Simianer, H. (2014). Improving the accuracy of whole genome prediction for complex traits using the results of genome wide association studies. PloS one, 9(3), e93017. [CrossRef]
| Trait | Method | RMSE | Residual Variance | CV | SE |
|---|---|---|---|---|---|
| Mendelian | GS | 2.7084 | 6.8628 | 1.3765 | 0.0236 |
| MAS | 2.2440 | 5.0418 | 1.0500 | 0.0091 | |
| GWAS_GS1 | 2.6640 | 6.6223 | 0.9919 | 0.0347 | |
| GWAS_GS2 | 2.7124 | 6.8818 | 1.3882 | 0.0229 | |
| Polygenic | GS | 6.2314 | 38.8328 | 0.9684 | 0.0221 |
| MAS | 26.7025 | 712.4898 | 0.9690 | 0.0052 | |
| GWAS_GS1 | 22.1835 | 492.3881 | 1.4096 | 0.0192 | |
| GWAS_GS2 | 6.1794 | 38.2113 | 0.8562 | 0.0213 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




