Submitted:
04 April 2025
Posted:
07 April 2025
You are already at the latest version
Abstract
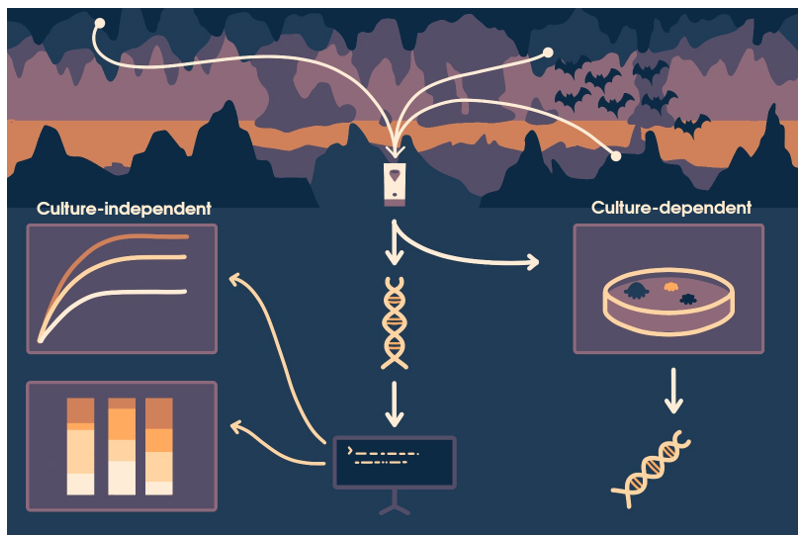
Keywords:
1. Introduction
2. Material and Methods
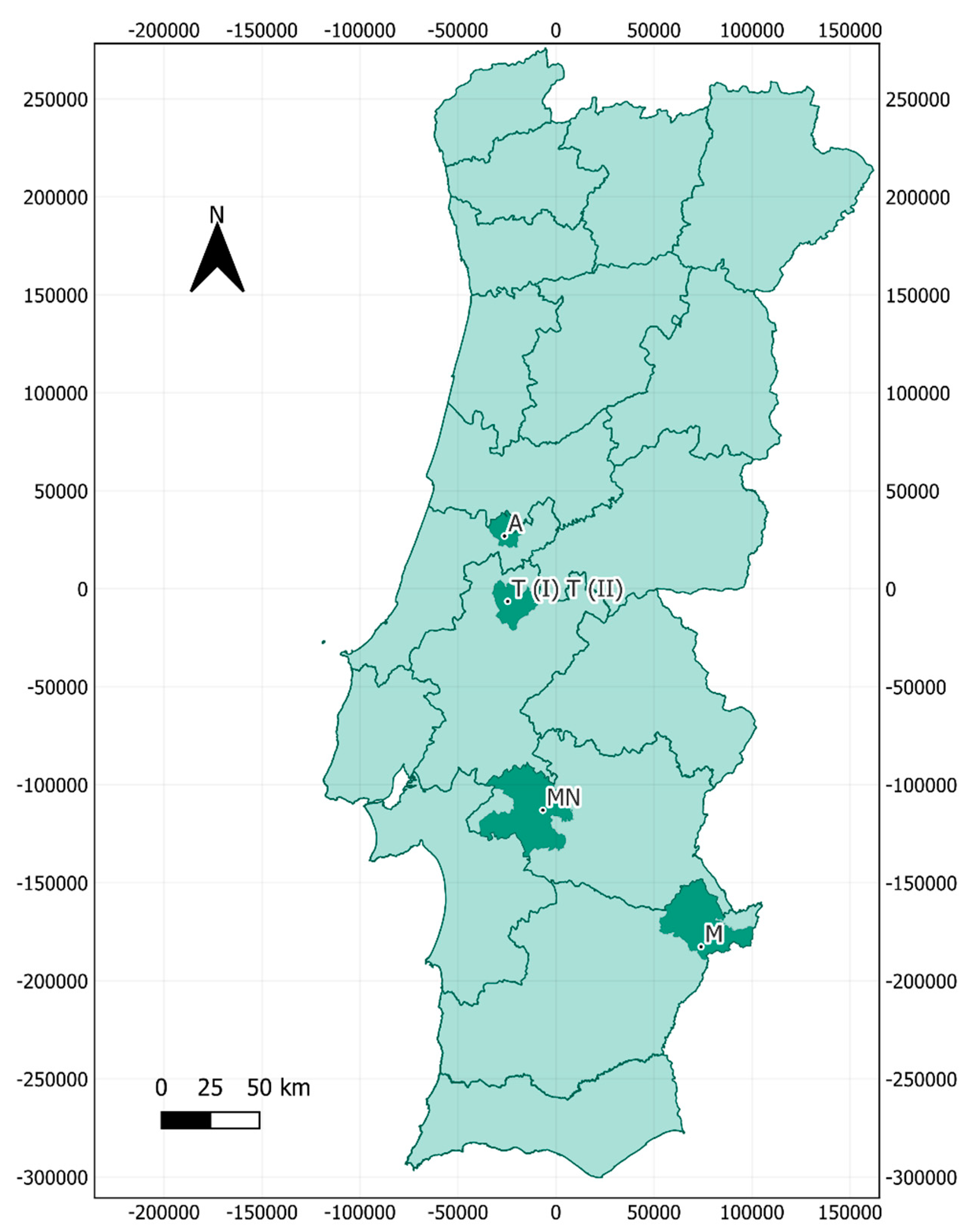
2.1. Sampling Location
2.2. Fungal Culture-Dependent Characterization
2.3. Sequencing of Fungal Isolates
2.4. DNA Extraction and Molecular Identification of the Air Samples
2.5. Oxford Nanopore Sequencing of ITS
2.6. Taxonomic Classification and Refinement
2.7. Data Analysis
2.8. Custom Scripts for Data Manipulation and Visualization
3. Results
3.1. Culturable Results
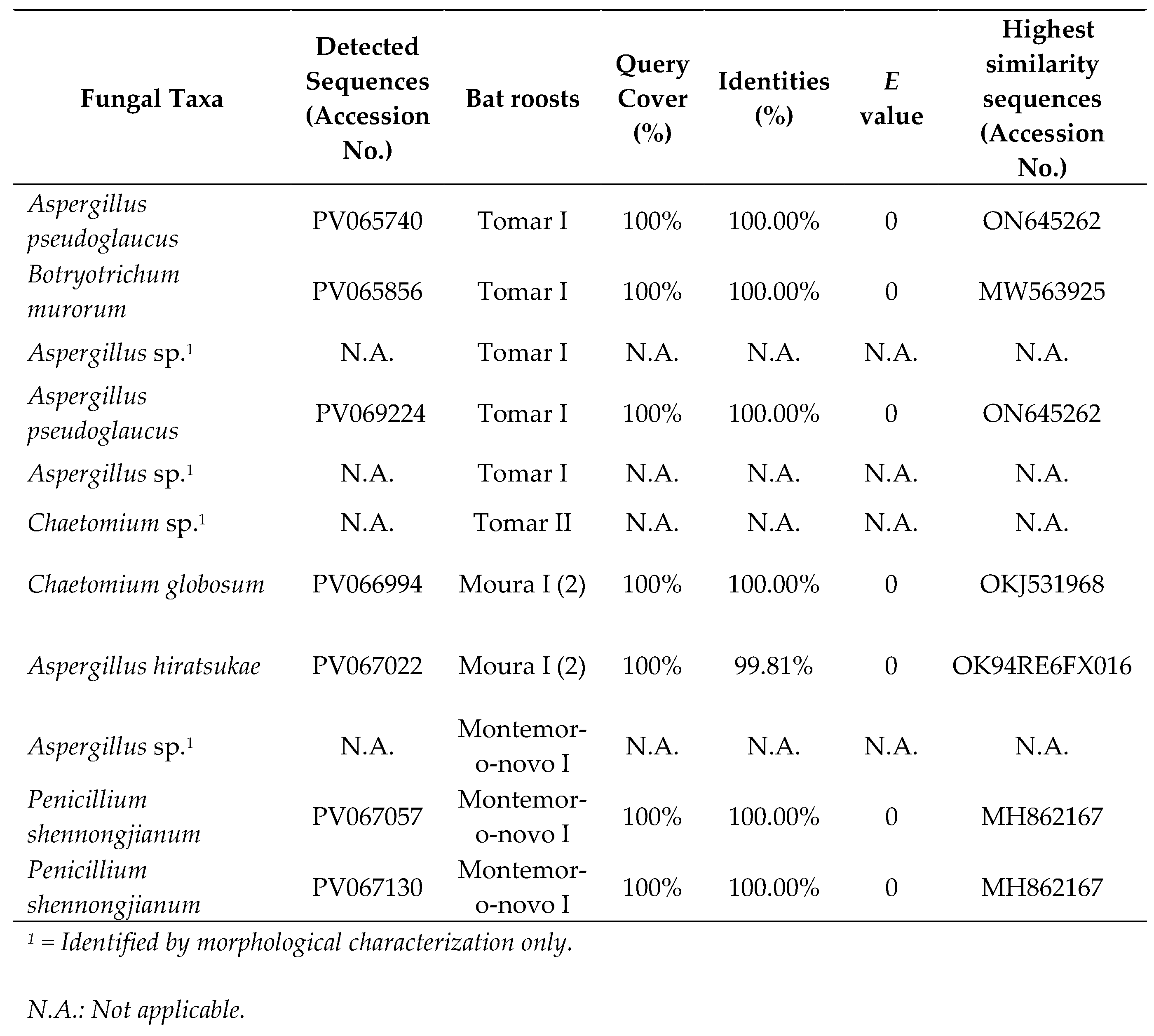 |
3.2. Oxford Nanopore Sequencing
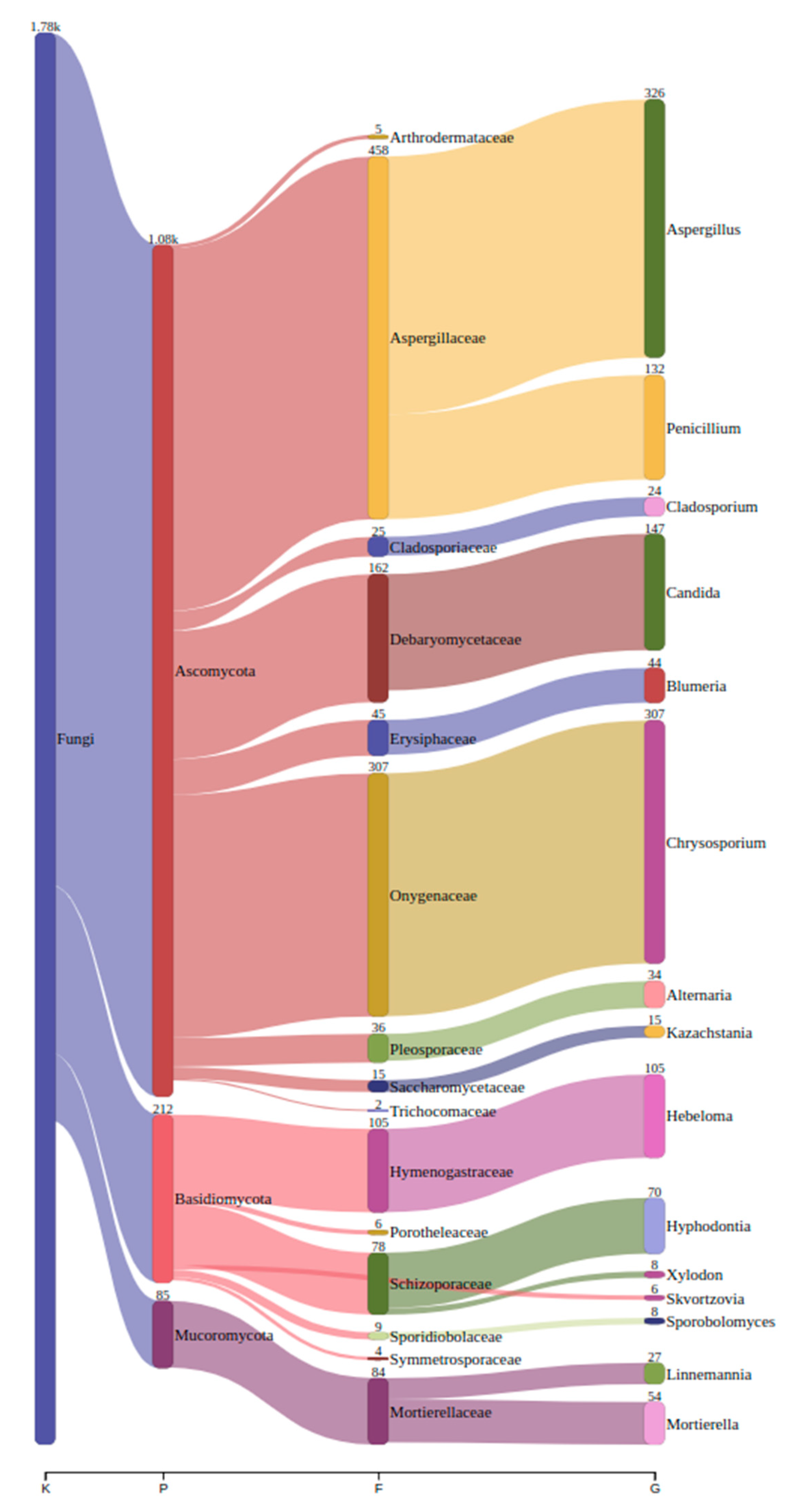
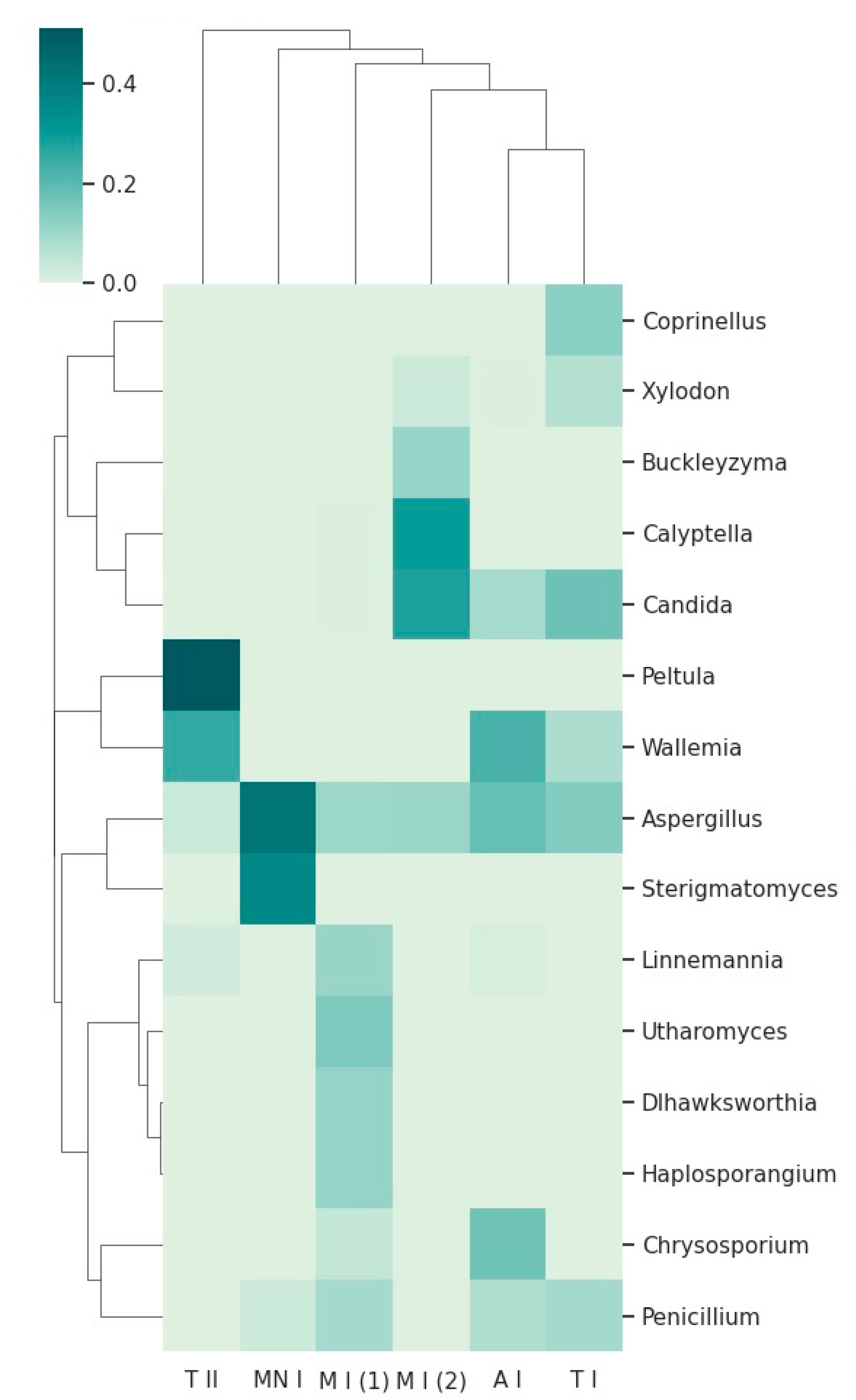
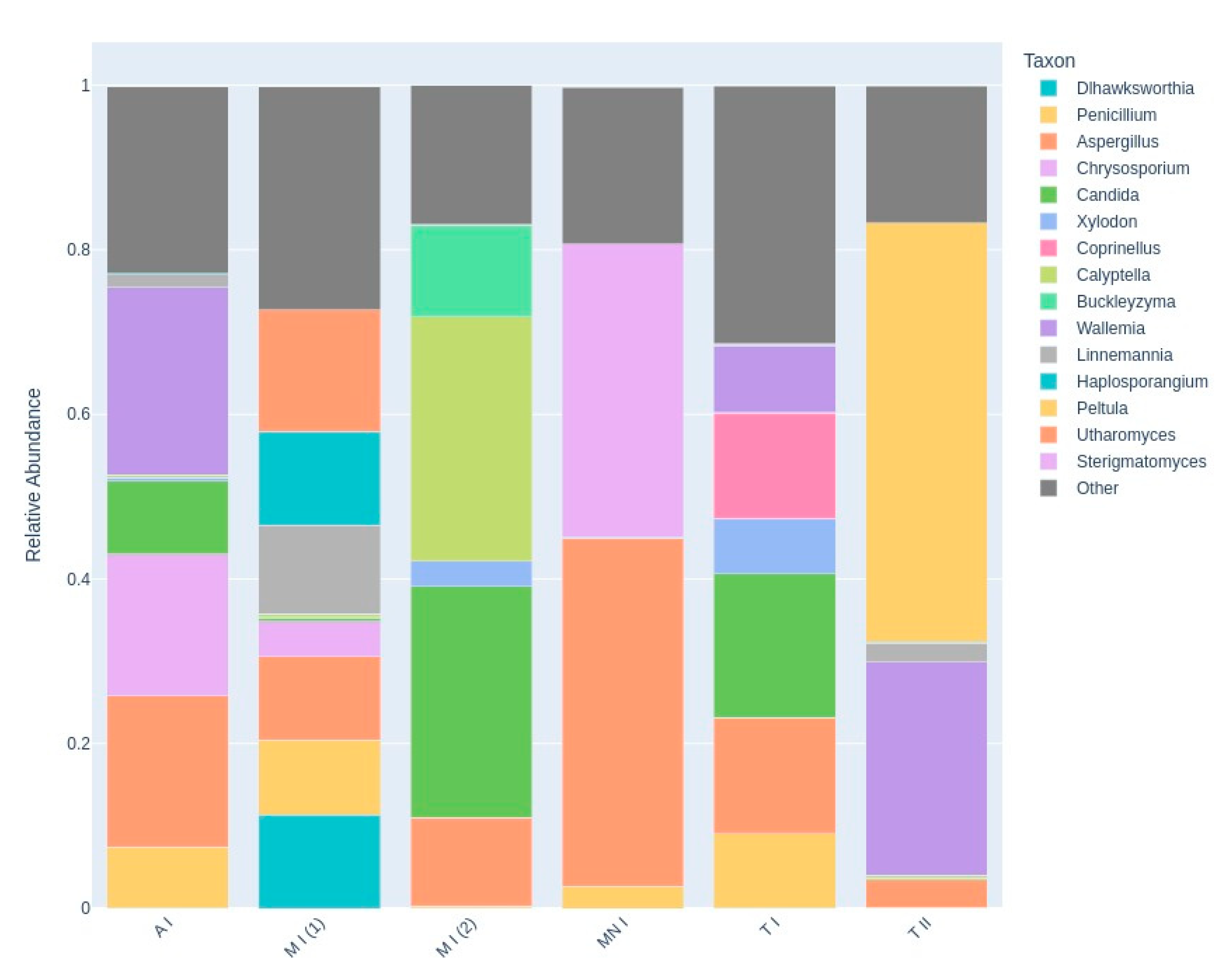
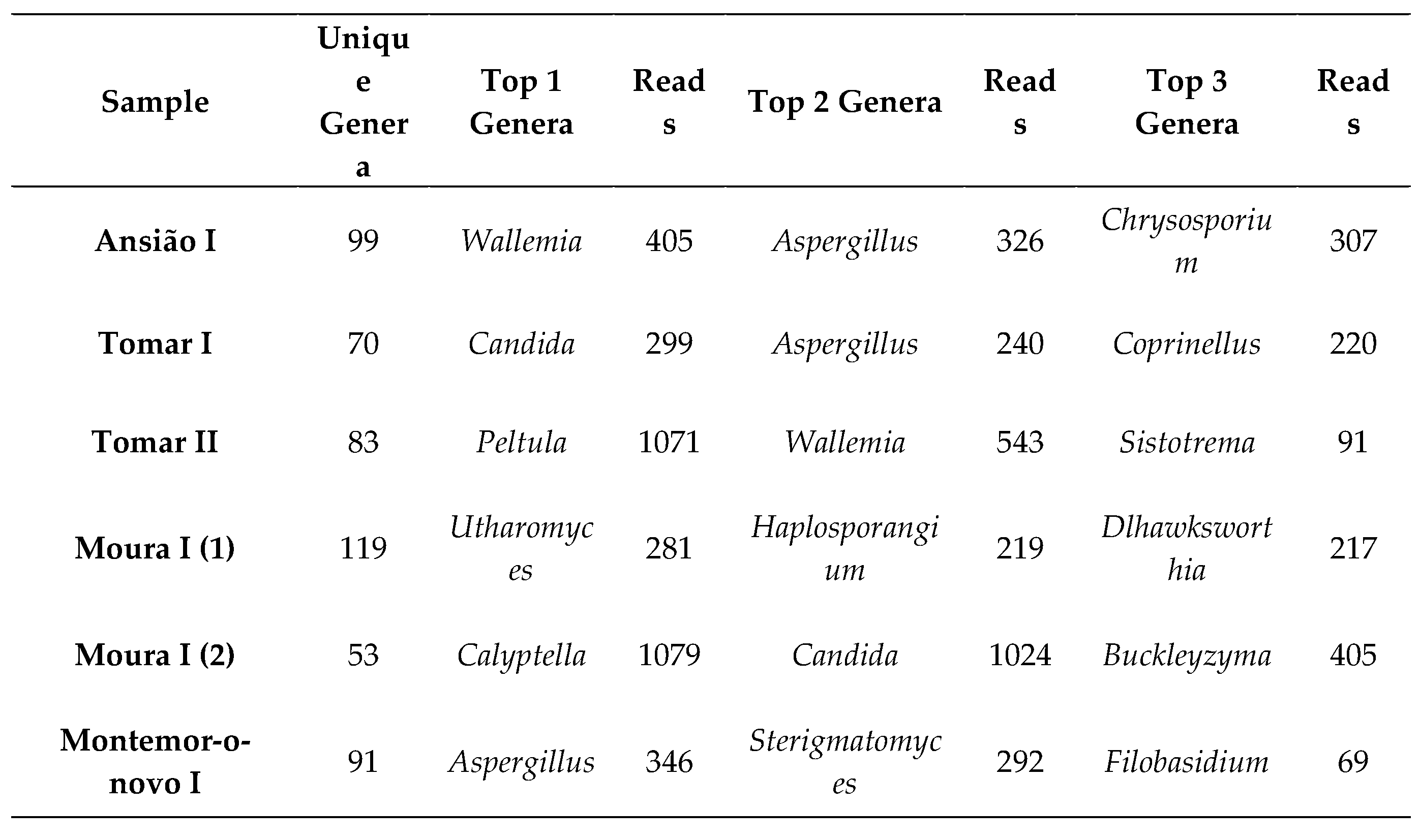
3.2.1. Fungal Taxonomy by Metabarcoding and ONT Sequencing
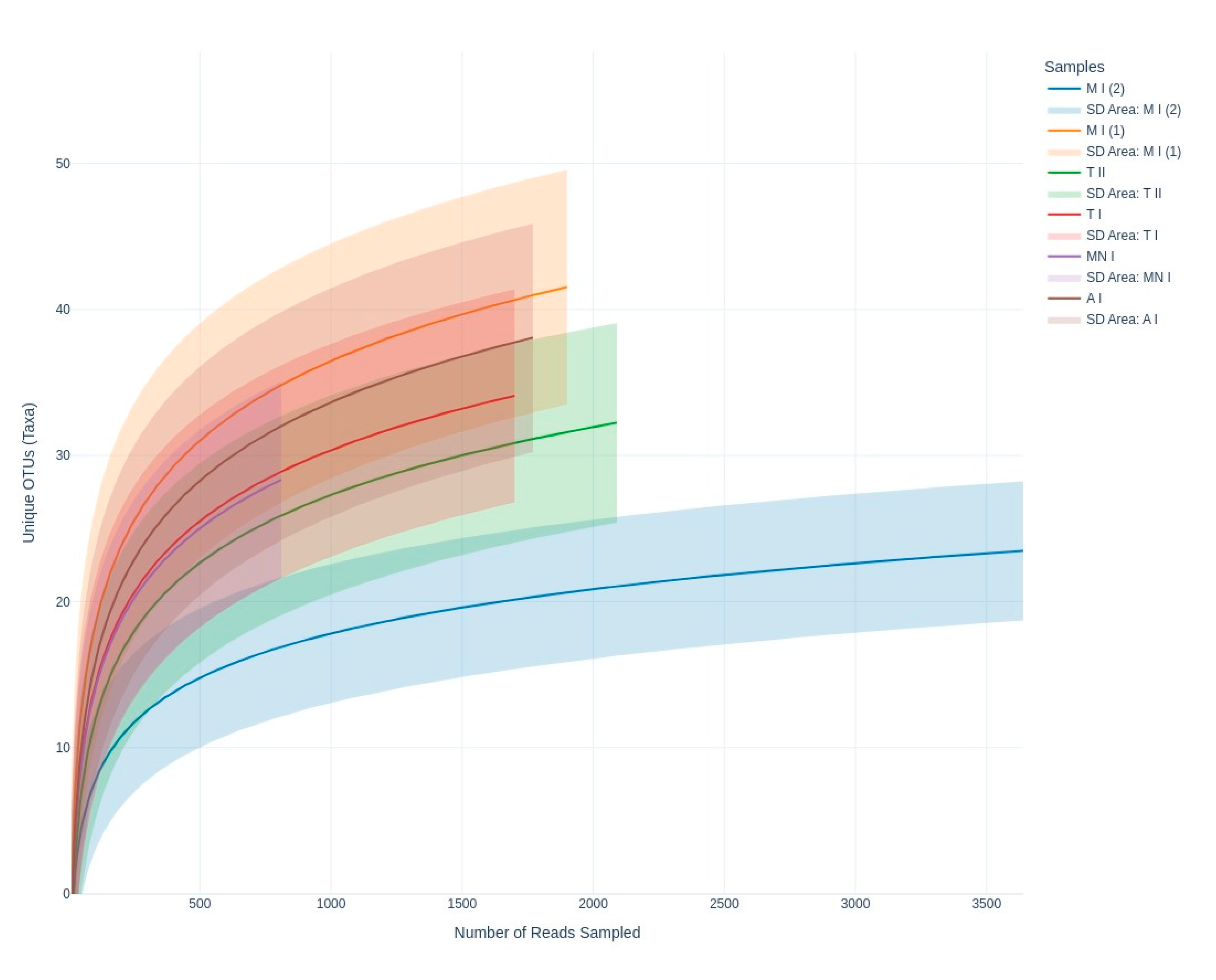
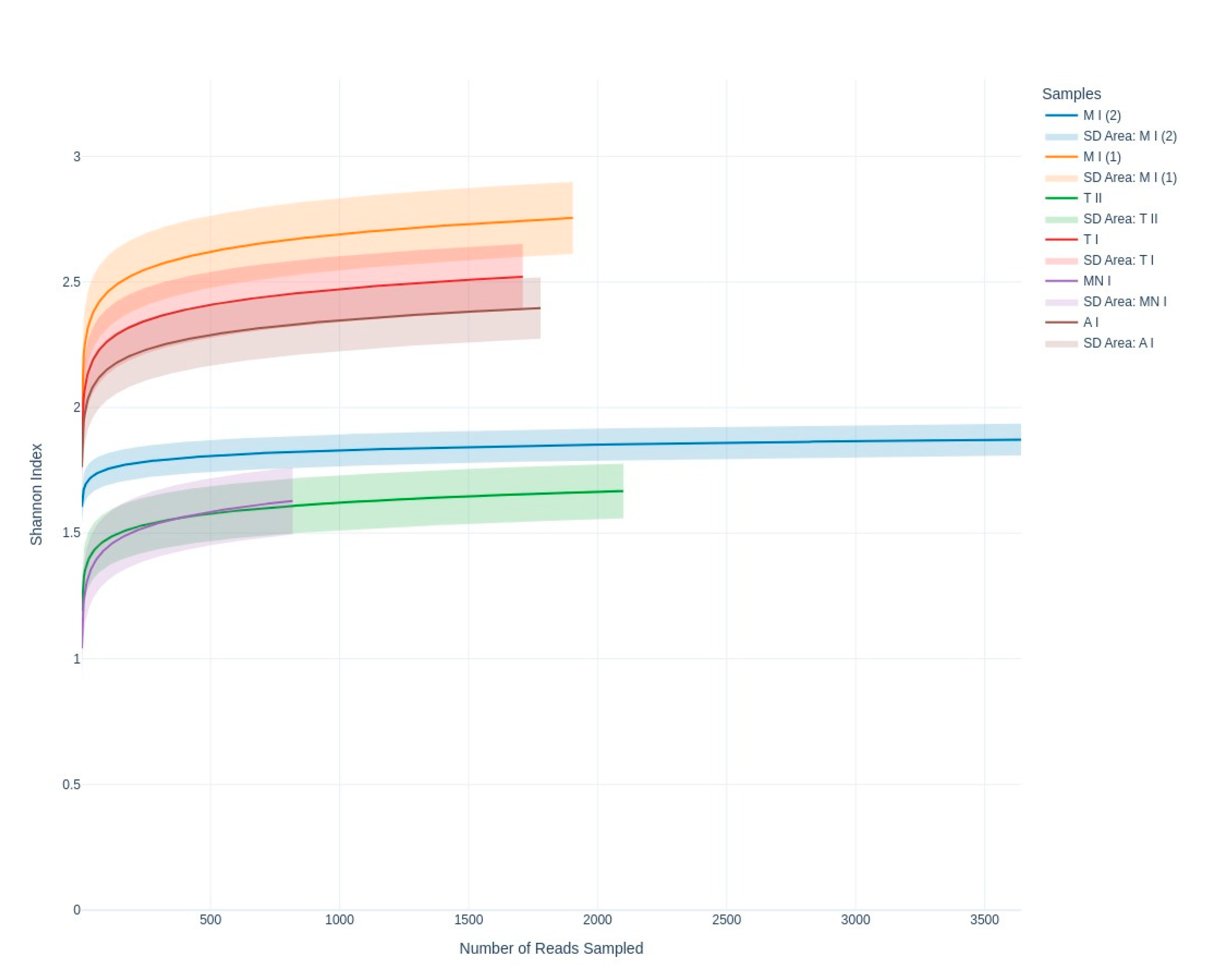
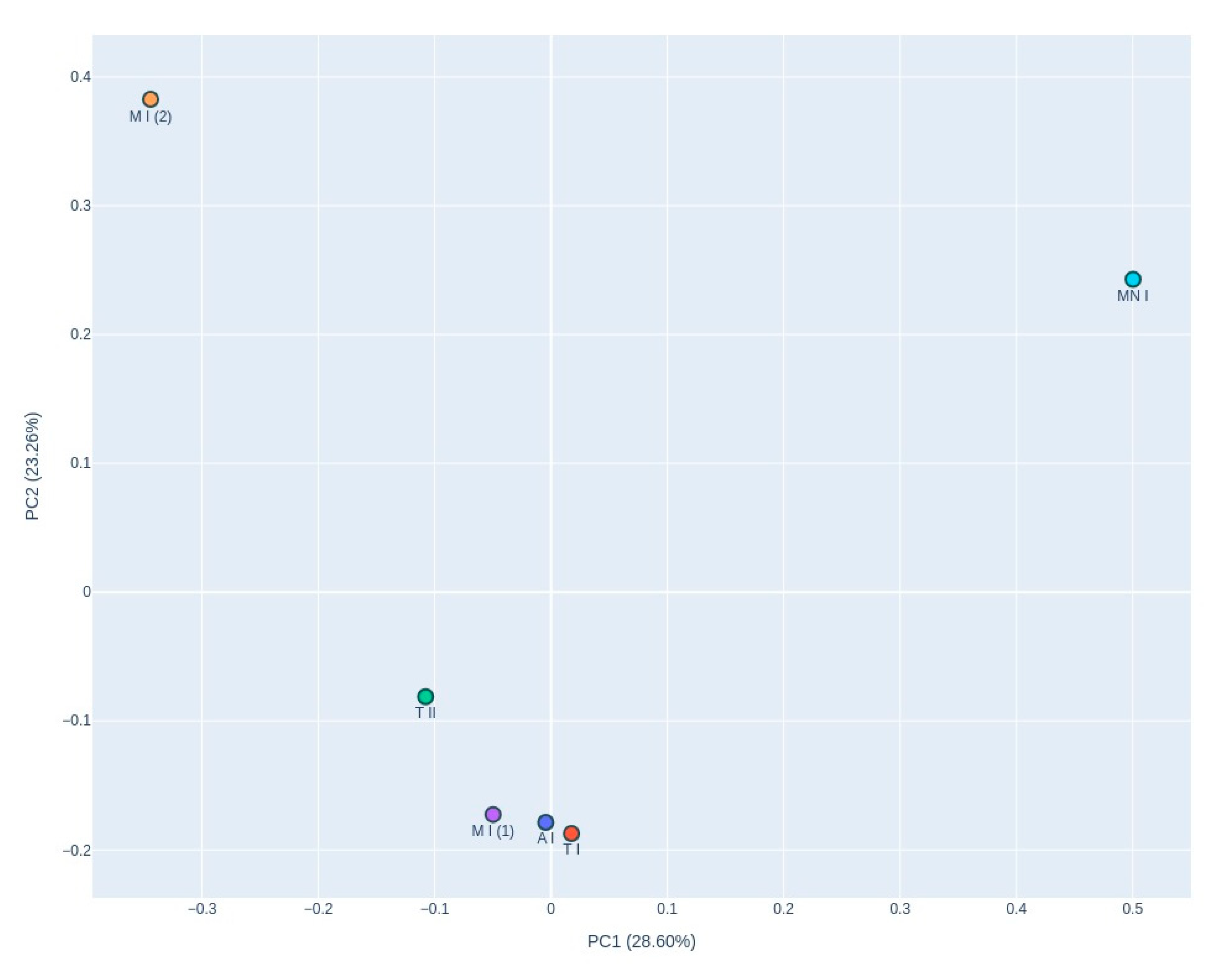
3.2.2. Fungal Diversity
4. Discussion
5. Conclusions
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Abarenkov: K., Zirk, A., Piirmann, T., Pöhönen, R., Ivanov, F., Nilsson, R. H., & Kõljalg, U. (2024). UNITE general FASTA release for Fungi. Version 04.04.2024. UNITE Community. [CrossRef]
- Borzęcka, J., Piecuch, A., Kokurewicz, T., Lavoie, K. H., & Ogórek, R. (2021). Greater mouse-eared bats (Myotis myotis) hibernating in the nietoperek bat reserve (poland) as a vector of airborne culturable fungi. Biology, 10(7). [CrossRef]
- Cantrell, S. A., Dianese, J. C., Fell, J., Gunde-Cimerman, N., & Zalar, P. (2011). Unusual fungal niches. Mycologia, 103(6), 1161–1174. [CrossRef]
- Chen, M., Ding, Z., Zhou, M., Shang, Y., Li, C., Li, Q., Bu, T., Tang, Z., & Chen, H. (2024). The diversity of endophytic fungi in Tartary buckwheat (Fagopyrum tataricum) and its correlation with flavonoids and phenotypic traits. Frontiers in Microbiology, 15(March), 1–16. [CrossRef]
- De Coster, W., & Rademakers, R. (2023). {NanoPack2}: population-scale evaluation of long-read sequencing data. Bioinformatics, 39(5), btad311. [CrossRef]
- De Mandal, S., Zothansanga, Panda, A. K., Bisht, S. S., & Senthil Kumar, N. (2015). First report of bacterial community from a Bat Guano using Illumina next-generation sequencing. Genomics Data, 4, 99–101. [CrossRef]
- Dimkić, I., Fira, D., Janakiev, T., Kabić, J., Stupar, M., Nenadić, M., Unković, N., & Grbić, M. L. (2021). The microbiome of bat guano: for what is this knowledge important? Applied Microbiology and Biotechnology, 105(4), 1407–1419. [CrossRef]
- Domsch, K. H. (1993). Compendium of soil fungi. IHW- Verlag 1, 630–643.
- Gallego-Clemente, E., Moreno-González, V., Ibáñez, A., Calvo-Peña, C., Ghoreshizadeh, S., Radišek, S., Cobos, R., & Coque, J. J. R. (2023). Changes in the Microbial Composition of the Rhizosphere of Hop Plants Affected by Verticillium Wilt Caused by Verticillium nonalfalfae. Microorganisms, 11(7). [CrossRef]
- Hagiwara, D., Sakamoto, K., Abe, K., & Gomi, K. (2016). Signaling pathways for stress responses and adaptation in Aspergillus species: stress biology in the post-genomic era. Bioscience, Biotechnology, and Biochemistry, 80(9), 1667–1680. [CrossRef]
- Jurado, V., Laiz, L., Rodriguez-Nava, V., Boiron, P., Hermosin, B., Sanchez-Moral, S., & Saiz-Jimenez, C. (2010). Pathogenic and opportunistic microorganisms in caves. International Journal of Speleology, 39(1), 15–24. [CrossRef]
- Keck, F., Blackman, R., Bossart, R., Brantschen, J., Couton, M., Hürlemann, S., Kirschner, D., Locher, N., Zhang, H., & Altermatt, F. (2022). Meta-analysis shows both congruence and complementarity of DNA and eDNA metabarcoding to traditional methods for biological community assessment. Molecular Ecology, 31, 1820–1835. [CrossRef]
- Klich, M. A. (2002). Biogeography of Aspergillus species in soil and litter. Mycologia, 94(1), 21–27. [CrossRef]
- Kokurewicz, T., Ogórek, R., Pusz, W., & Matkowski, K. (2016). Bats Increase the Number of Cultivable Airborne Fungi in the “Nietoperek” Bat Reserve in Western Poland. Microbial Ecology, 72(1), 36–48. [CrossRef]
- Krijgsheld, P., Bleichrodt, R., Veluw, G., Wang, F., Müller, W., Dijksterhuis, J., & Wösten, H. (2012). Development in Aspergillus. Studies in Mycology, 74, 1–29. [CrossRef]
- Lu, J., Breitwieser, F. P., Thielen, P., & Salzberg, S. L. (2017). Bracken: estimating species abundance in metagenomics data. PeerJ Computer Science, 3, e104. [CrossRef]
- Moore, J. A. M., Anthony, M. A., Pec, G. J., Trocha, L. K., Trzebny, A., Geyer, K. M., van Diepen, L. T. A., & Frey, S. D. (2021). Fungal community structure and function shifts with atmospheric nitrogen deposition. Global Change Biology, 27(7), 1349–1364. [CrossRef]
- Moreira, G. (2025). GmoreiraVet/Guitools: v1.1.1. Zenodo. [CrossRef]
- Morschhäuser, J. (2024). Adaptation of Candida albicans to specific host environments by gain-of-function mutations in transcription factors. PLoS Pathogens, 20(11), e1012643. [CrossRef]
- Mühldorfer, K. (2013). Bats and Bacterial Pathogens: A Review. Zoonoses and Public Health, 60(1), 93–103. [CrossRef]
- Ogórek, R., Lejman, A., & Matkowski, K. (2013). Fungi isolated from Niedźwiedzia Cave in Kletno (Lower Silesia, Poland). International Journal of Speleology.
- Rabha, J., & Jha, D. (2018). Metabolic Diversity of Penicillium. 217–234. [CrossRef]
- Raper, K. B. (1965). The Genus Aspergillus. The Williams and Wilkins Co.
- Siqueira, J. P. Z., Sutton, D. A., Gené, J., García, D., Wiederhold, N., & Guarro, J. (2018). Species of Aspergillus section Aspergillus from clinical samples in the United States. Medical Mycology, 56(5), 541–550. [CrossRef]
- Soares, D., & Niemiller, M. L. (2020). Extreme Adaptation in Caves. Anatomical Record, 303(1), 15–23. [CrossRef]
- Voigt, C. C., Phelps, K. L., Aguirre, L. F., Schoeman, M. C., Vanitharani, J., & Zubaid, A. (2016). Bats and buildings: The conservation of synanthropic bats. In Bats in the Anthropocene : Conservation of Bats in a Changing World (pp. 427–453).
- Vu, D., Groenewald, M., de Vries, M., Gehrmann, T., Stielow, B., Eberhardt, U., Al-Hatmi, A., Groenewald, J. Z., Cardinali, G., Houbraken, J., Boekhout, T., Crous, P. W., Robert, V., & Verkley, G. J. M. (2019). Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Studies in Mycology, 92, 135–154. [CrossRef]
- Wasti, I. G., Khan, F. A. A., Bernard, H., Hassan, N. H., Fayle, T., & Sathiya Seelan, J. S. (2021). Fungal communities in bat guano, speleothem surfaces, and cavern water in Madai cave, Northern Borneo (Malaysia). Mycology, 12(3), 188–202. [CrossRef]
- Wick, R. (2024, November). rrwick/{Porechop}. https://github.com/rrwick/Porechop (2017).
- Wood, D. E., Lu, J., & Langmead, B. (2019). Improved metagenomic analysis with {Kraken} 2. Genome Biology, 20(1), 257. [CrossRef]
- Zhang, Z. F., Liu, F., Zhou, X., Liu, X. Z., Liu, S. J., & Cai, L. (2017). Culturable mycobiota from Karst caves in China, with descriptions of 20 new species. In Persoonia (Vol. 39, Issue 1, pp. 1–31). [CrossRef]
| Bat species |
|---|
| Myotis myotis |
| Miniopterus schreibersii |
| Rhinolophus ferrumequinum |
| Rhinolophus mehelyi |
 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




