Submitted:
01 April 2025
Posted:
01 April 2025
Read the latest preprint version here
Abstract
Keywords:
Why Do We Need to Perform the Normalization for the scRNA-seq Data?
Why Did We Develop a New Method, CLTS, for scRNA-seq Data Normalization?
What Is Hypothesis and the Basic Idea of CLTS Normalization Method?
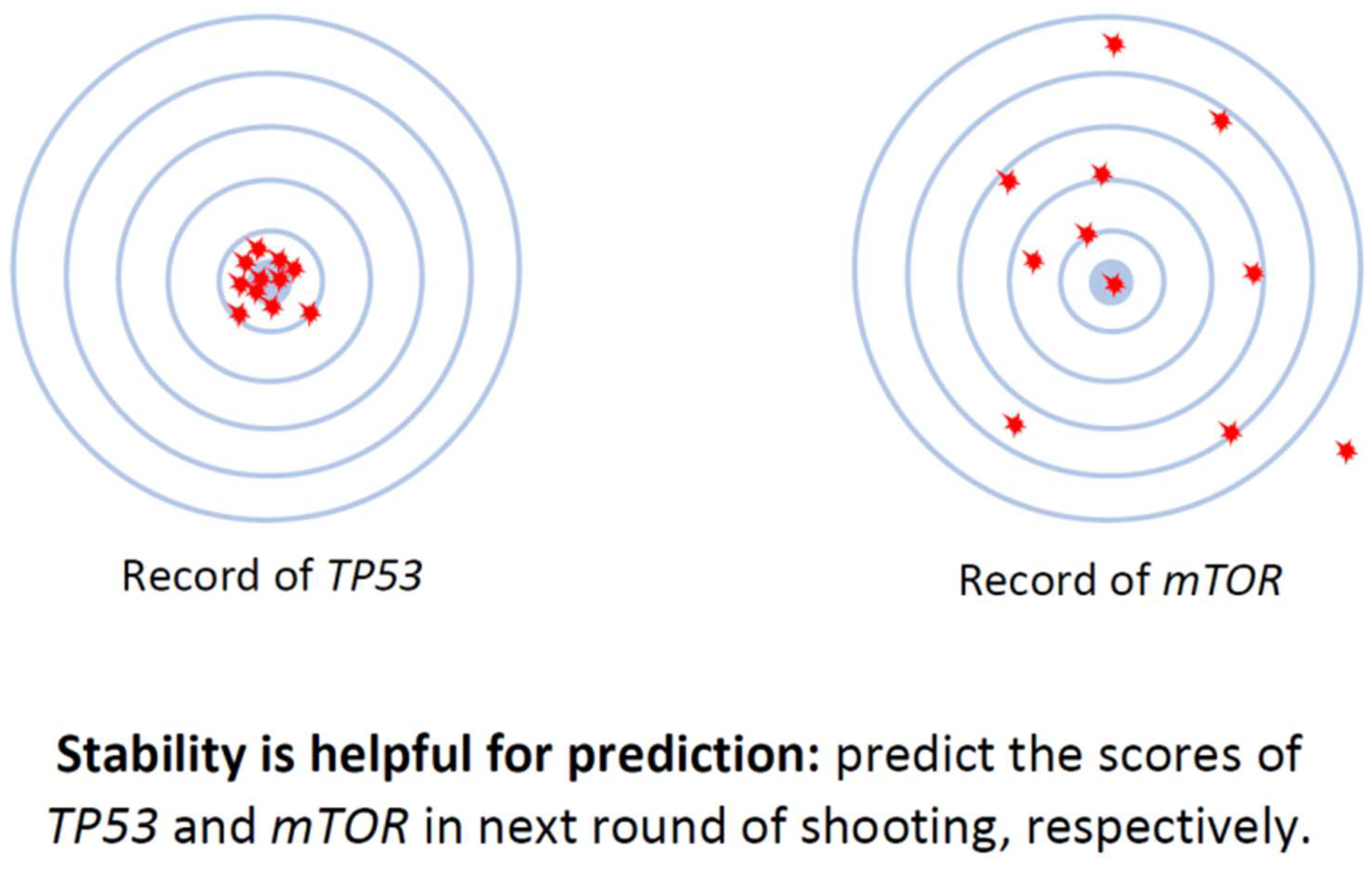
- The value of a cell’s true transcriptome size, or the total amount of RNAs truly expressed by a cell, should remain stable within a narrow range for any type of cell. However, it’s worth noting that the values of true transcriptome sizes may vary significantly among different types of cells.
- The total raw count obtained from any scRNA-seq data for a cell is essentially a measure of the cell’s true transcriptome size. Moreover, the measured transcriptome size for all cells in the same sample should be proportional to its real value (Although the variance is usually large). Overall, this proportion should be fairly similar for all cells within the same sample. For simplicity, we usually refer to the “measured transcriptome size” as “transcriptome size”.
- The proportion of the measured values to the true transcriptome sizes can vary significantly among cells in different samples. This is what leads to significant differences in gene expression of the same cell type across different samples.
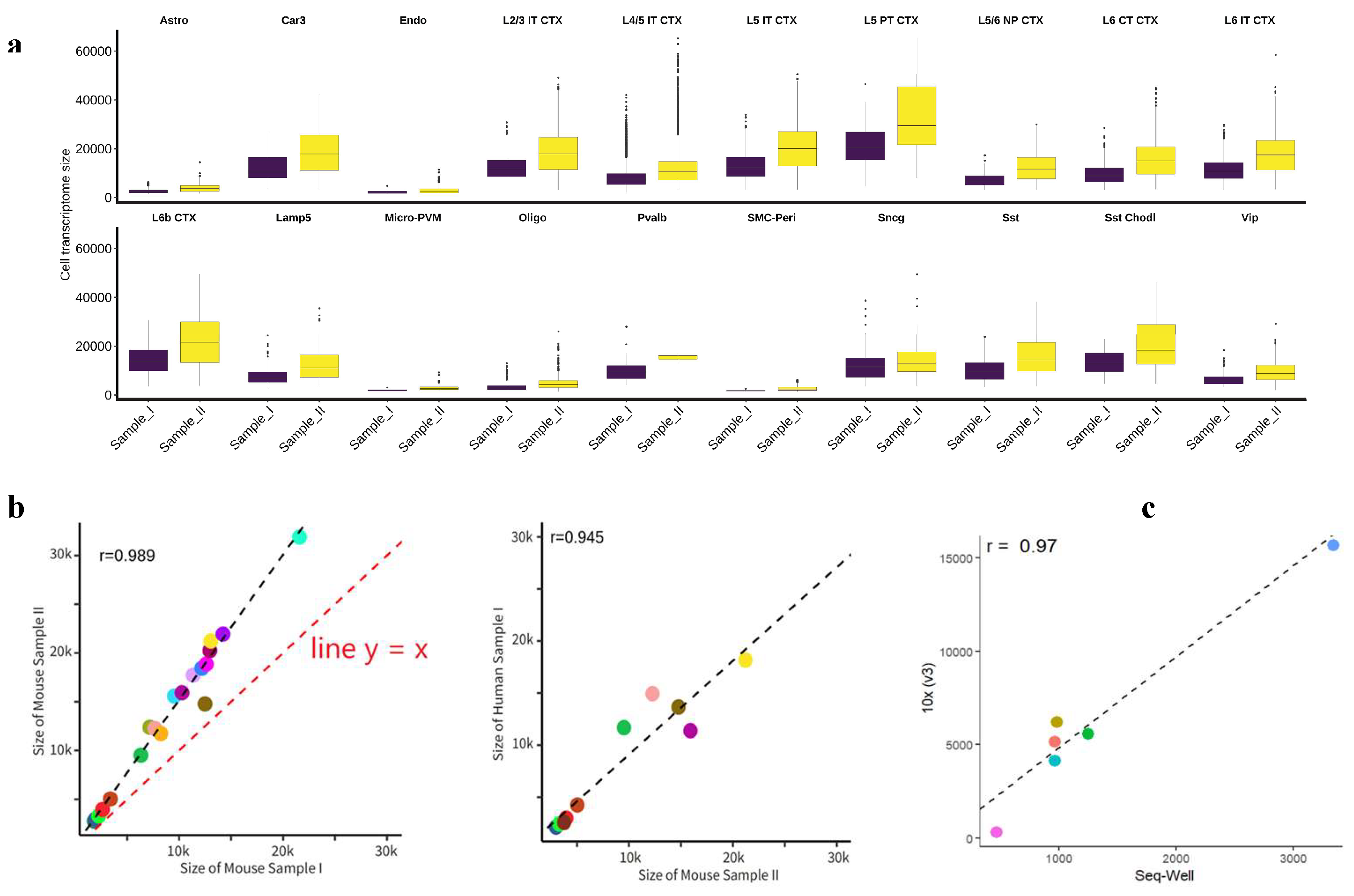
- The transcriptome sizes of different types of cells show considerable variation, while those of the same cell type remain within a narrow range (Figure 2a).
- The average transcriptome sizes of different cell types in different samples show a strong linear relationship. Basically, by multiplying the average transcriptome sizes of all cell types in one of the two samples by a constant, we can make the average transcriptome sizes of all matching cell types in the two samples similar (Figure 2b).
- This linear relationship remains not only between samples of the same species, such as between two mouse brain samples (Figure 2b), but also between samples of different species, such as between a mouse brain sample and a human brain sample (Figure 2c). Additionally, this linear correlation holds across samples with scRNA-seq data from various sequencing platforms, such as 10x (v3) and Seq-well (Figure 2d).
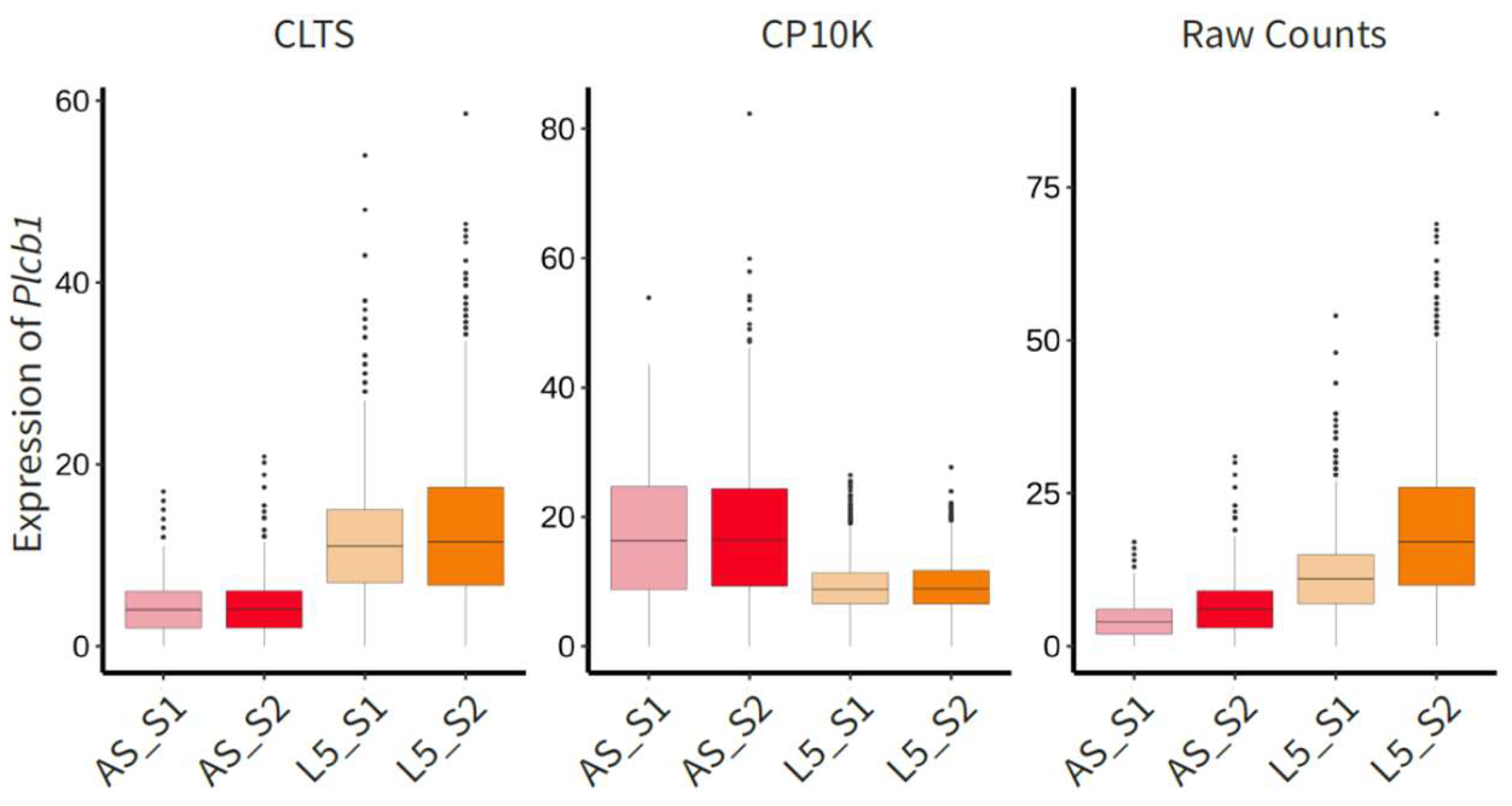
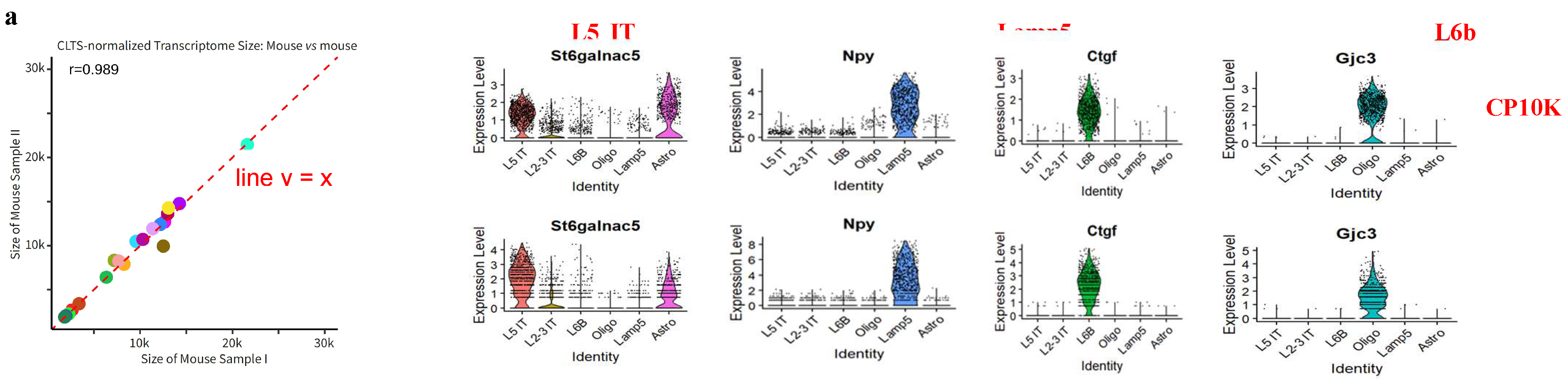
- Our novel model, CLTS, leverages this linear correlation to perform normalization. Consequently, after normalization, the average transcriptome sizes of any given cell type become similar across all samples (Figure 3a). It is obvious that CLTS does not have the scaling issues that CP10K has.
What Should We Notice When Using Seurat for scRNA-seq Data Processing?
- Cell clustering is a process that considers the similarity of cell expression profiles, with CP10K exerting a minimal influence on this step.
- In the process of determining the cell type of each Seurat cluster using cell type markers, typically, the selected type markers, such as Npy, Ctgf and Gjc3 (as shown in Figure 3b), are predominantly expressed in a single cell type or Seurat cluster. Consequently, CP10K does not compromise the precision of cell type annotation. However, if a cell type marker, like St6galnac5 (refer to Figure 3b), demonstrates substantial expression in multiple cell types or Seurat clusters, it becomes imperative to reevaluate the results for potential impacts stemming from CP10K’s scaling issues. Therefore, utilizing the CLTS-normalized data for cell type annotations of Seurat clusters can help prevent annotation errors.
- During downstream analysis, such as identifying genes that are highly expressed in specific cell types, CP10K can significantly influence the results. For instance, several genes that exhibit high expression in Astrocytes may actually have higher expression in L5 cells. Consequently, we strongly advocate for the use of CLTS-normalized data in downstream analysis.
How Do We Integrate Seurat and CLTS for scRNA-seq Data Processing?
- In the first approach, we employ Seurat in a traditional manner for clustering and cell type annotations. Following this, we utilize CLTS for normalization of the scRNA-seq data. We then use this CLTS-normalized data to examine if the cell type markers are impacted by CP10K scaling issues and to conduct other downstream analysis.
- In the second approach, we consider each Seurat cluster as a distinct cell type after the clustering step. Consequently, we use the cluster information and CLTS for normalizing the scRNA-seq data. We then employ CLTS-normalized data for cell type annotation of the Seurat clusters and for performing additional downstream analysis. You can find demonstration codes on how to implement this method on the ReDeconv website.
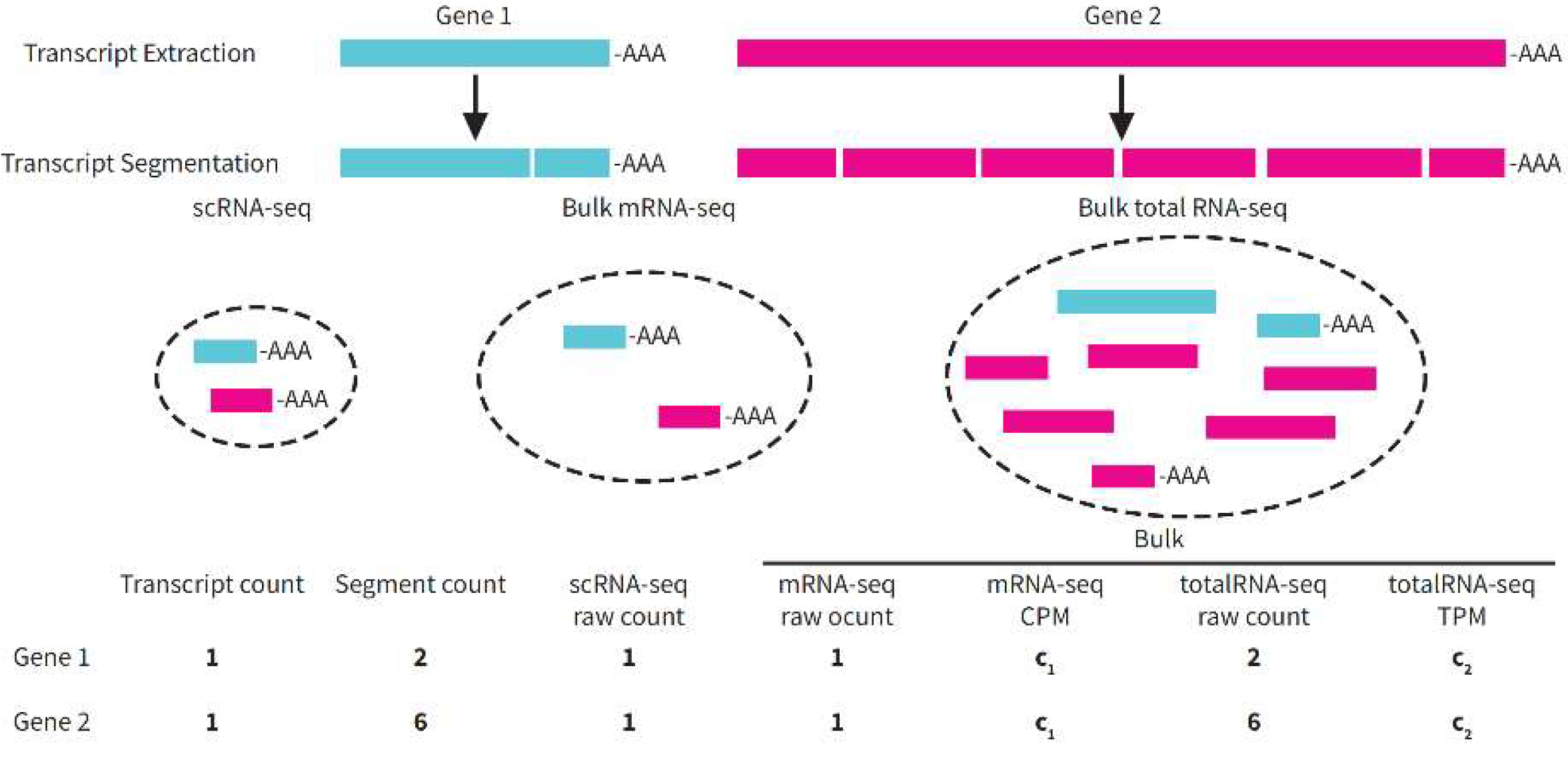
Does the Normalization of Bulk RNA-seq Data Need to Consider the Transcriptome Sizes of Cells?
Why Did We Develop a New Model, ReDevonv, for Cell Type Deconvolution?
What Is the Basic Idea of Cell Type Deconvolution?
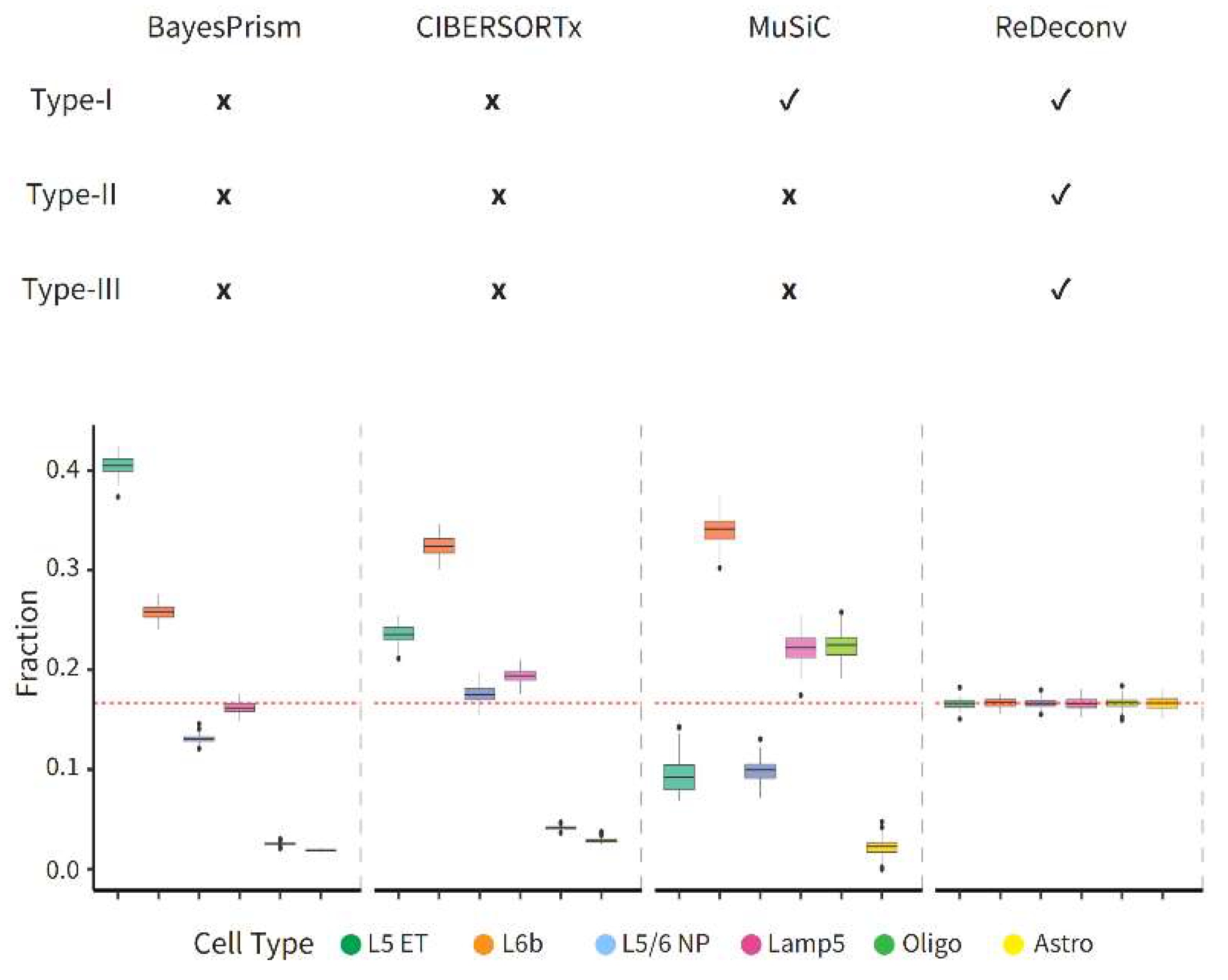
What Is the Type-I Issues?
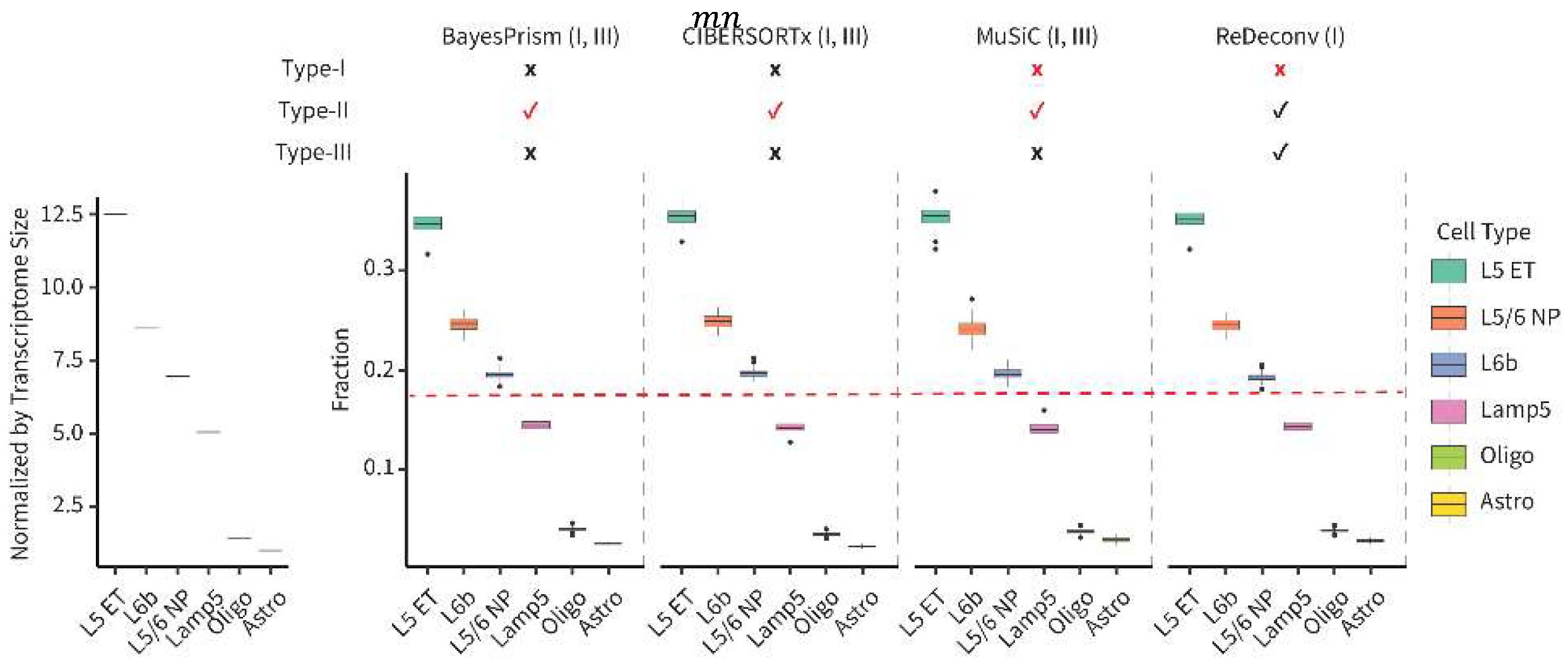
What Is the Type-II Issues?
What Is the Type-III Issues?
Why Have These Issues Not Been Noticed Before in Model Evaluations?
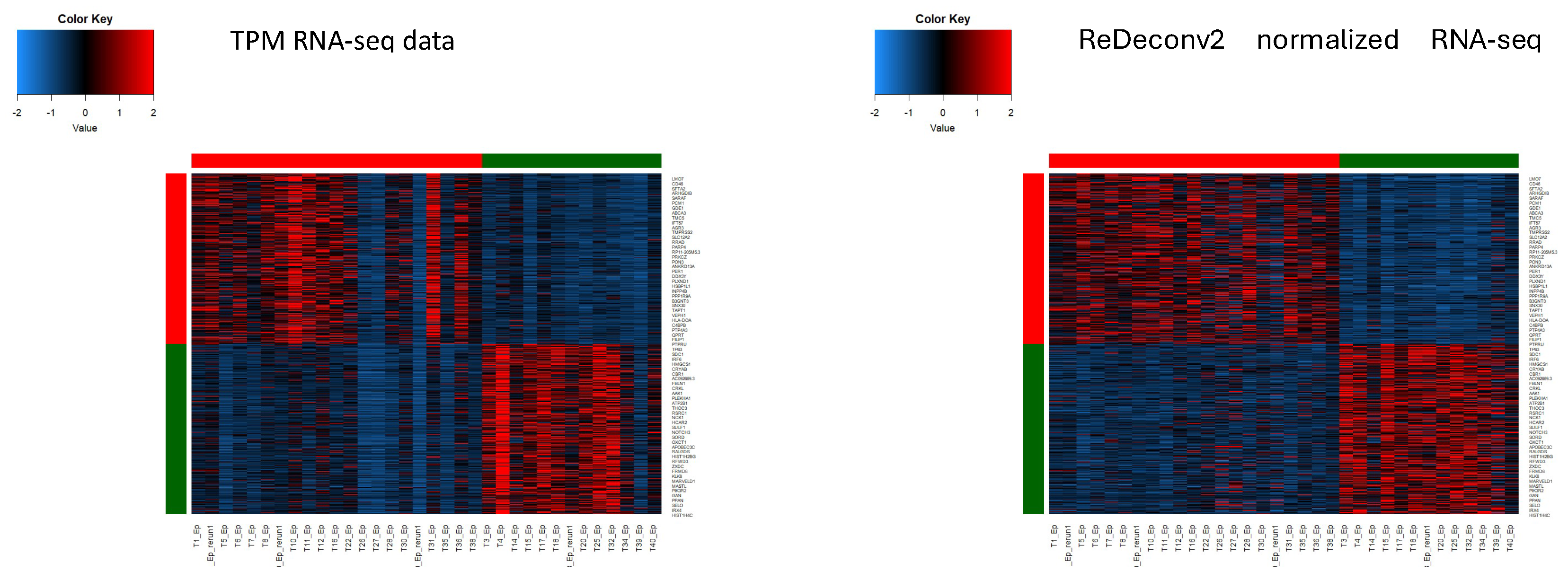
- Notes: Certain models, like BayesPrism and CIBERSORTx, that automatically apply CP10K/CPM to the scRNA-seq data, invariably have Type-I issues, even when utilizing raw-count or CLTS-normalized scRNA-seq data as references. Prior to using any deconvolution models, it’s advisable to verify if the models exhibit any types of issues. If CLTS-normalized scRNA-seq data and TPM/FPKM bulk RNA-seq data are used as inputs for ReDeconv, then all Type-I, II, III issues are effectively addressed. For more detailed information about the ReDeconv model and instructions on how to use ReDeconv, please refer to our paper, “Transcriptome Size Matters for Single-Cell RNA-seq Normalization and Bulk Deconvolution” (Nature Communications, Feb. 2025), and visit the website – https://redeconv.stjude.org/home. Most of the figures in this summary are derived from our paper in Nature Communications.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



